Pairwise Sequence Alignment I Lecture for CS 498
























- Slides: 31

Pairwise Sequence Alignment (I) (Lecture for CS 498 -CXZ Algorithms in Bioinformatics) Sept. 22, 2005 Cheng. Xiang Zhai Department of Computer Science University of Illinois, Urbana-Champaign Many slides are taken/adapted from http: //www. bioalgorithms. info/slides. htm

Comparing Genes in Two Genomes • • Small islands of similarity corresponding to similarities between exons Such comparisons are quite common in biology research

Alignment of sequences is one of the most basic and most important problems in bioinformatics…

Outline • Defining the problem of alignment • The longest common subsequence problem • Dynamic programming algorithms for alignment

Aligning Two Strings Given the strings: • v = ATGTTAT • w = ATCGTAC One possible alignment of the strings: AT_GTTAT_ ATCGT_A_C 1 st row – string v with space symbols “-” inserted 2 nd row – string w with space symbols “-” inserted

Aligning Two Strings (cont’d) Another way to represent each row shows the number of symbols of the sequence present up to a given position. For example the above sequences can be represented as: 0 1 2 2 3 4 5 6 7 7 AT_GTTAT_ ATCGT_A_C 0 1 2 3 4 5 5 6 6 7

Alignment Matrix 0 1 2 2 3 4 5 6 7 7 AT_GTTAT_ ATCGT_A_C 0 1 2 3 4 5 5 6 6 7 Both rows of the alignment can be represented in the resulting matrix: 0 1 2 2 3 4 5 6 7 7 0 1 2 3 4 5 5 6 6 7

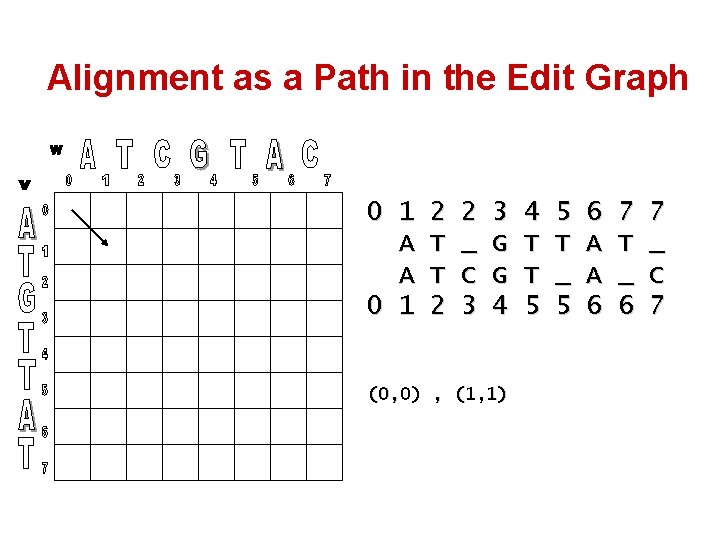
Alignment as a Path in the Edit Graph 0 1 A A 0 1 2 T T 2 2 _ C 3 3 G G 4 (0, 0) , (1, 1) 4 T T 5 5 T _ 5 6 A A 6 7 T _ 6 7 _ C 7

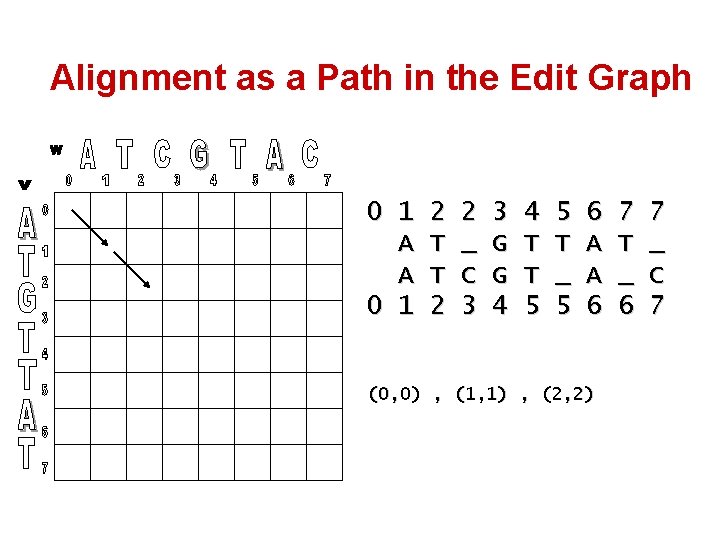
Alignment as a Path in the Edit Graph 0 1 A A 0 1 2 T T 2 2 _ C 3 3 G G 4 4 T T 5 5 T _ 5 6 A A 6 (0, 0) , (1, 1) , (2, 2) 7 T _ 6 7 _ C 7

Alignment as a Path in the Edit Graph 0 1 A A 0 1 2 T T 2 2 _ C 3 3 G G 4 4 T T 5 5 T _ 5 6 A A 6 7 T _ 6 7 _ C 7 (0, 0) , (1, 1) , (2, 2), (2, 3), (3, 4)

Alignment as a Path in the Edit Graph 0 1 A A 0 1 2 T T 2 2 _ C 3 3 G G 4 4 T T 5 5 T _ 5 6 A A 6 7 T _ 6 7 _ C 7 - End Result (0, 0) , (1, 1) , (2, 2), (2, 3), (3, 4), (4, 5), (5, 5), (6, 6), (7, 7)

Alignment as a Path in the Edit Graph Every path in the edit graph corresponds to an alignment:

How to Score an Alignment? • Simplest – Every match scores 1 – Every mismatch scores 0 – An alignment is scored based on the number of common symbols – Lead to the longest common subsequence problem • More sophisticated – ? – To be covered later

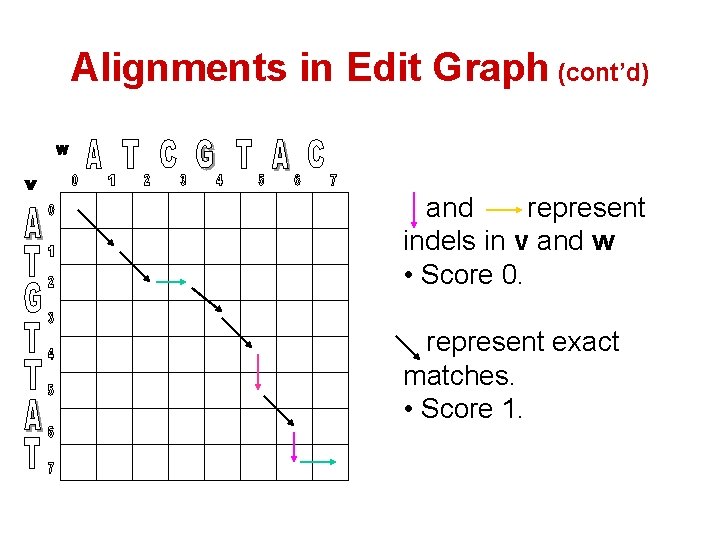
Alignments in Edit Graph (cont’d) and represent indels in v and w • Score 0. represent exact matches. • Score 1.

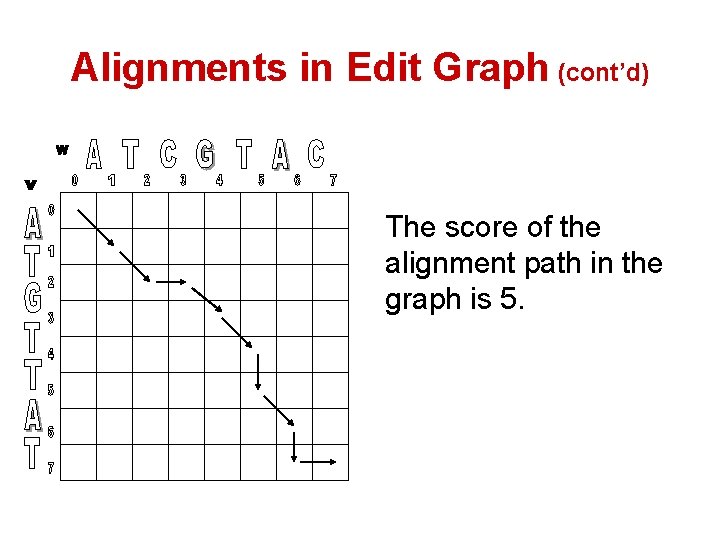
Alignments in Edit Graph (cont’d) The score of the alignment path in the graph is 5.

The Longest Common Subsequence (LCS) Problem • Find the longest subsequence common to two strings. Input: Two strings, v and w. v = ATGTTAT w = ATCGTAC Output: The longest common subsequence of v and w. v = AT GTTAT | | | w = ATCGT AC “ATGTA” A subsequence is not necessarily consecutive Longest common subsequence Best alignment

How to solve the LCS problem efficiently?

Brute Force Approach • Enumerate all the sequences up to length min(|v|, |w|) • For each one, check to see if it is a subsequence of v and w • Very expensive…. (How many sequences do we have to enumerate? )

The Idea of Dynamic Programming • Think of an alignment as a path in an edit graph • We only need to keep track of the best alignment (i. e. , the longest common subsequence) • Score a longer alignment based on shorter alignments

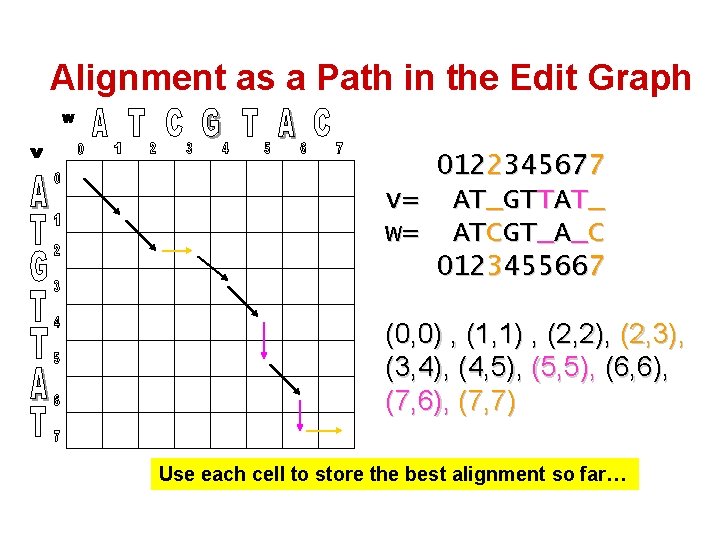
Alignment as a Path in the Edit Graph 0122345677 v= AT_GTTAT_ w= ATCGT_A_C 0123455667 (0, 0) , (1, 1) , (2, 2), (2, 3), (3, 4), (4, 5), (5, 5), (6, 6), (7, 7) Use each cell to store the best alignment so far…

Alignment: Dynamic Programming Use this scoring algorithm si, j = max si-1, j-1+1 if vi = wj si-1, j si, j-1

Dynamic Programming Example • There are no matches in the beginning of the sequence • Label column i=1 to be all zero, and row j=1 to be all zero

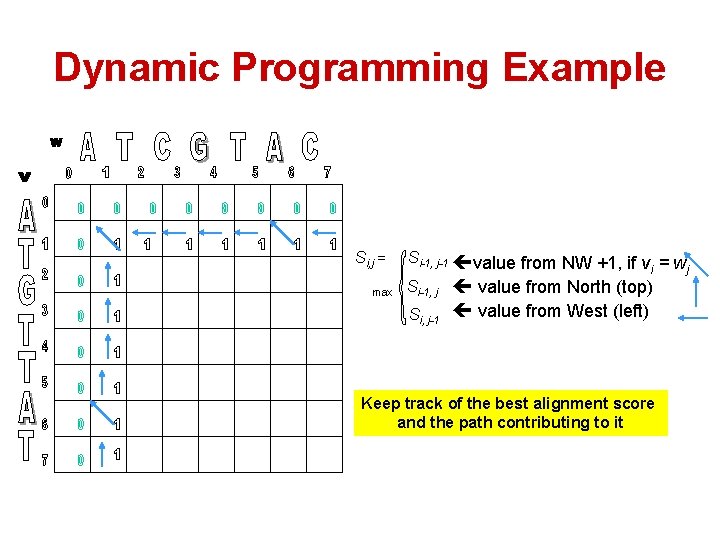
Dynamic Programming Example Si, j = max Si-1, j-1 value from NW +1, if v = w i j Si-1, j value from North (top) Si, j-1 value from West (left) Keep track of the best alignment score and the path contributing to it

Alignment: Backtracking Arrows from. show where the score originated if from the top if from the left if vi = wj

Dynamic Programming Example Continuing with the scoring algorithm gives this result.

LCS Algorithm 1. LCS(v, w) 2. for i 1 to n 3. Si, 0 0 4. for j 1 to m 5. S 0, j 0 6. for i 1 to n 7. for j 1 to m 8. si-1, j 9. si, j max si, j-1 10. si-1, j-1 11. “ “ • • • bi, j + 1, if vi = wj if si, j = si-1, j “ “ if si, j = si, j-1 “ “ if si, j = si-1, j-1 + 1 return (sn, m, b)

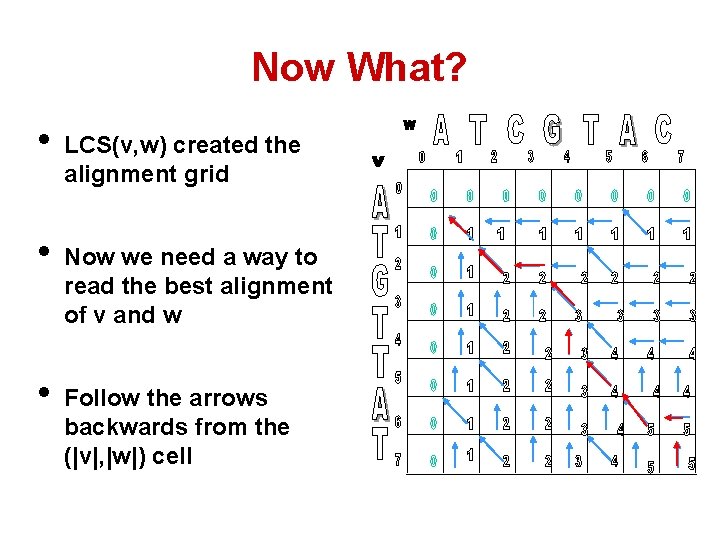
Now What? • • • LCS(v, w) created the alignment grid Now we need a way to read the best alignment of v and w Follow the arrows backwards from the (|v|, |w|) cell

LCS Runtime • To create the nxm matrix of best scores from vertex (0, 0) to all other vertices, it takes O(nm) time. • Why O(nm)? The pseudocode consists of a nested “for” loop inside of another “for” loop to set up a nxm matrix.

How do we improve the scoring of alignments? Can we still find an alignment efficiently? We’ll talk about these later…

The LCS Recurrence Revisited • The formula can be rewritten by adding zero to the edges that come from an indel, since the penalty of indels are 0: si-1, j-1+1 if vi = wj si, j = max Matching score si-1, j + 0 si, j-1 + 0 Insertion/deletion score

What You Should Know • How an alignment corresponds to a path in an edit graph • How the LCS problem corresponds to alignment with a simple scoring method • How the dynamic programming algorithm solves the LCS problem (= simple alignment)