Pairwise Local Alignment and Database Search Csc 487687

Pairwise Local Alignment and Database Search Csc 487/687 Computing for Bioinformatics

Which Program should one use? n Most researchers use methods for determining local similarities: Smith-Waterman (gold standard) Do not find every possible alignment ¡ FASTA of query with database sequence. These ¡ BLAST are used because they run faster than S-W ¡ }

Heuristic Database Search Methods n Smith-Waterman dynamic programming too computer and time intensive for searching big databases ¡ n e. g. , Uni. Prot July 2004 – 1. 5 M sequences Most popular: BLASTx (Altschul et al 1990, 1997) and FASTx (Lipman and Pearson 1985)

BLAST – Basic Local Alignment Search Tool n Basic idea: ¡ n Identify short very similar segment pairs – extend local alignment Critical issues: For every database sequence d significantly similar to q, one should find at least one segment pair ¡ Fewer segment pairs means faster computation ¡

Definitions n n Maximal Segment Pair (MSPqd) – pair of identical length segments having the highest score of all ungapped local alignments between q and d. High-Scoring Segment Pair (HSP) – segment pair for which the score cannot be increased by shortening or extension Word – segment of fixed length w Word pair – pair of segments of length w

Reformulating the Problem n n n Identify those database sequences d such that MSPqd is over a threshold V. A segment pair scoring at least V has with a high probability a word pair scoring at least T. Identify word pairs with score at least T, extend to high-scoring segment pairs – check if score over V

Finding Hits and HSPs n n n Hit – word pair scoring at least T Preprocess q ¡ Find all words o. T (length w) that can score at least T against a word in q ¡ Save in easy-to-use data structure Find the hits ¡ Search in d for all occurrences (od) of the words o. T Extend (heuristically) to high-scoring segment pairs Perform dynamic programming around HSPs scoring over a certain threshold – allows introduction of gaps

Pre-processing q n n Aim: ¡ Allow rapid identification of all words o. T in d – and the location of corresponding words in q to allow extension into HSPs Possibility: table of 20 w entries

Pre-processing q

Finding HSPs n n n For each word in d (starting in position j) hitting a word in q (starting in position i), record the hit indexed by its diagonal (j-i ). Hits close together on the same diagonal are joined before extension to HSPs Extending to HSP: ¡ Ideally – move to the end of the sequences in both directions ¡ Heuristic – if score falls “far below” best seen so far, stop extension
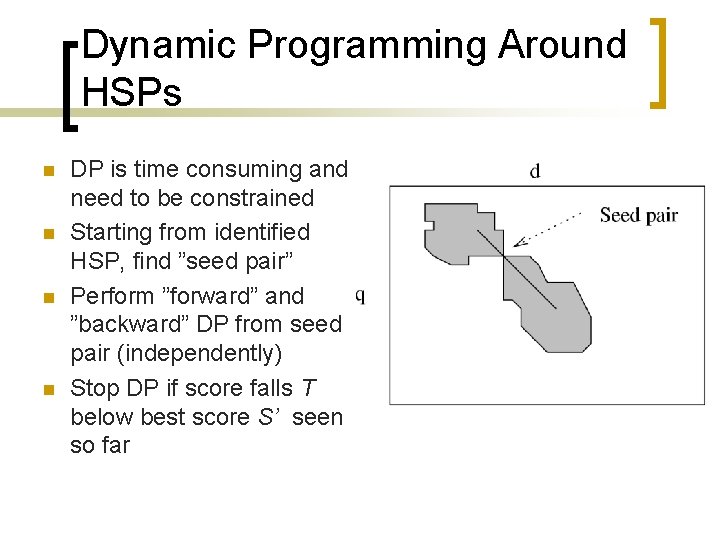
Dynamic Programming Around HSPs n n DP is time consuming and need to be constrained Starting from identified HSP, find ”seed pair” Perform ”forward” and ”backward” DP from seed pair (independently) Stop DP if score falls T below best score S’ seen so far

Significance of alignments n n n Suppose alignment reveals an intriguing similarity between two sequences. Is the similarity significant ? Or could it have arisen by chance?

Significance of alignment n If the score of the alignment observed is no better than might be expected from a random permutation of the sequence, then it is likely to have arisen by chance.

How to Generate the Random Sequences? n Global alignment ¡ n Randomize one of the sequences, many times, realign each result to the second sequence (fixed), and collect the distribution of resulting scores. Local alignment ¡ Uses the population of results returned from the entire database as the population with which to measure the statistics.

Statistical parameters n Z-score ¡ A measure of how unusual our original match is A z-score of 0 means the observed similarity is no better than the average of the control population. The higher the Z-score, the greater the probability. Z-score 5

Statistical parameters n P = the probability that the alignment is better than random ¡ ¡ ¡ P ≤ 10 -100 exact match P in range 10 -100 - 10 -50 sequences very nearly identical P in range 10 -50 - 10 -10 closely-related sequences, homology certain P in range 10 -5 - 10 -1 distant relatives, usually P > 10 -1 match probably insignificant

Statistical parameters n E-value ¡ ¡ ¡ The expected number of sequences that give the same Z-score or better if the database is probed with a random sequence. found by multiplying the value of P by the size of the database probed. Note that E but not P depends on the size of the database.

Statistical parameters n Interpreting E values ¡ ¡ ¡ E ≤ 0. 02 sequences probably homologous E between 0. 02 and 1 homologous cannot be ruled out E >1 you’d have to expect this good a match just by chance

Rules and thinking. . n Percent of identical residues in the optimal alignment ¡ ¡ ¡ over 45%, very similar structures, common or at least a related function. Over 25%, a similar general folding pattern. A lower degree of sequence similarity cannot rule out homology

Rules and thinking. . n n 18%-25% twilight zone, the suggestion of homology is tantalizing but dangerous Absence of significant similarity does not imply that the sequences are not homologous – could be distantly related (twilight zone or beyond)
- Slides: 20