PACKAGING OF DNA DOUBLE HELIX STRUCTURE OF DNA
PACKAGING OF DNA
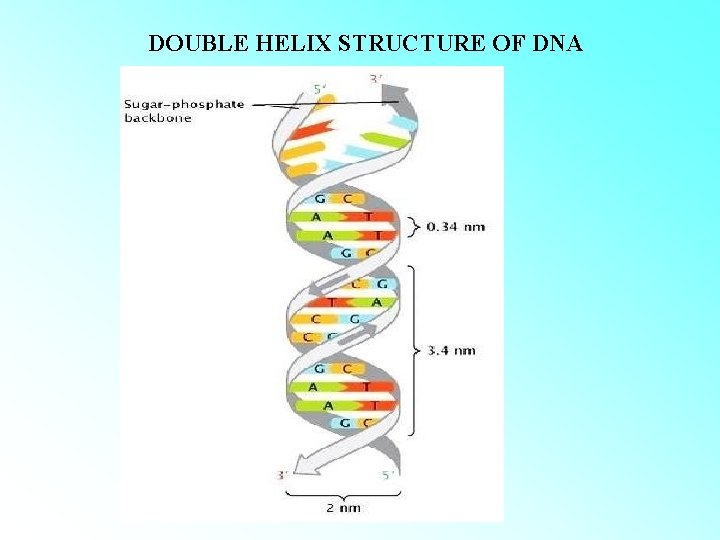
DOUBLE HELIX STRUCTURE OF DNA

CALCULATION OF LENGTH OF DNA • LENGTH OF DNA DOUBLE HELIX= TOTAL NUMBER OF bp X DISTANCE BETWEEN TWO ADJACENT bp. total number of bp : 6. 6 x 109 bp distance between two adjacent bp: 0. 34 nm (0. 34 x 10 -9 m/bp) length of DNA double helix= 6. 6 x 109 bp x 0. 34 x 10 -9 m/bp = 2. 2. meters The dimension of a typical nucleus (approximately 10 -6 m).

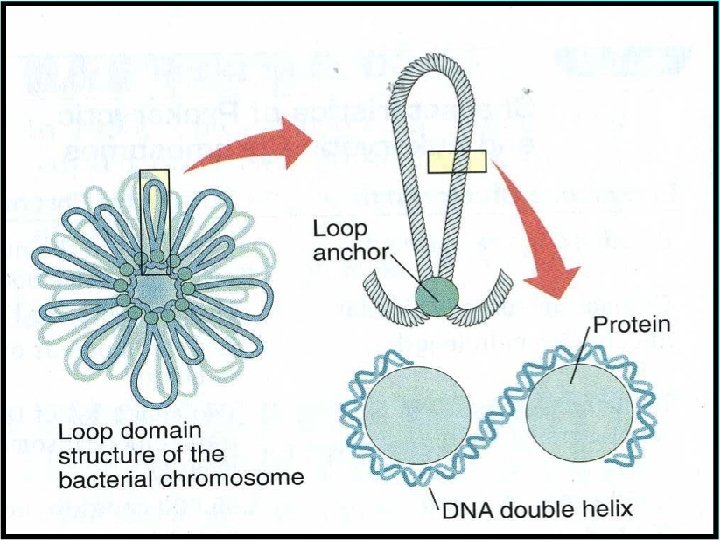
PACKAGING OF DNA IN PROKARYOTES (E. g. E. coli) • Prokaryotes lack a well-defined nucleus. • Genetic material is scattered in the cytoplasm. • Nucleoid: The region where DNA being negatively charged (due to phosphate moiety) is associated with proteins that are positively charged.
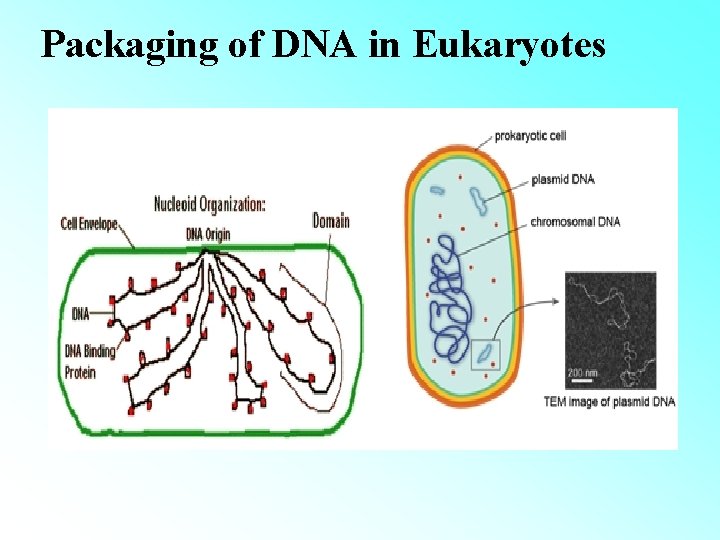
Packaging of DNA in Eukaryotes

PACKAGING IN EUKARYOTES • In eukaryotes, this organisation is much more complex. • There is a set of positively charged, basic proteins called Histones. • A protein acquires charge depending upon the abundance of amino acids residues with charged side chains. Histones are rich in the basic amino acid residues lysines and arginines. Both the amino acid residues carry positive charges in their side chains.
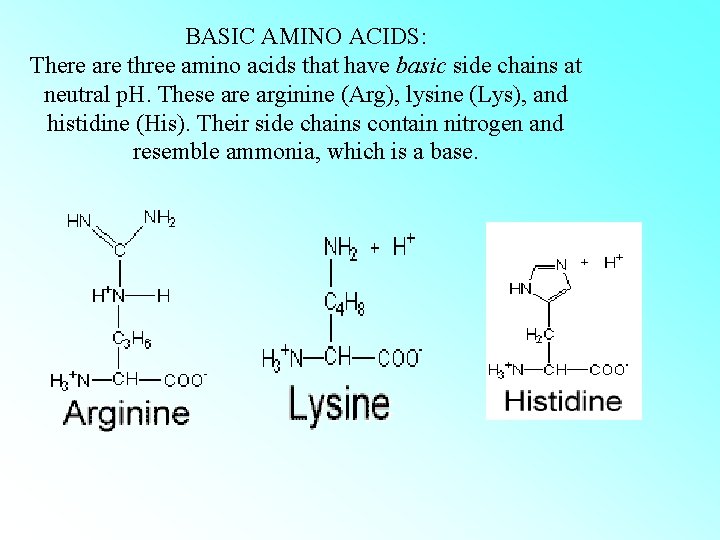
BASIC AMINO ACIDS: There are three amino acids that have basic side chains at neutral p. H. These arginine (Arg), lysine (Lys), and histidine (His). Their side chains contain nitrogen and resemble ammonia, which is a base.

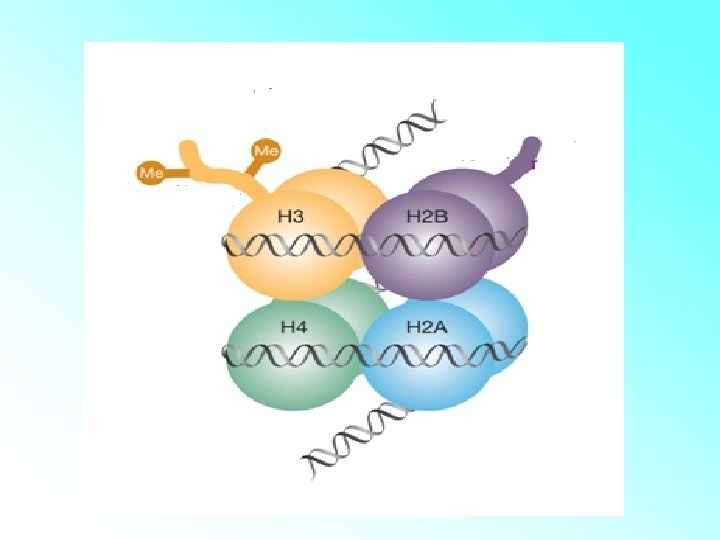
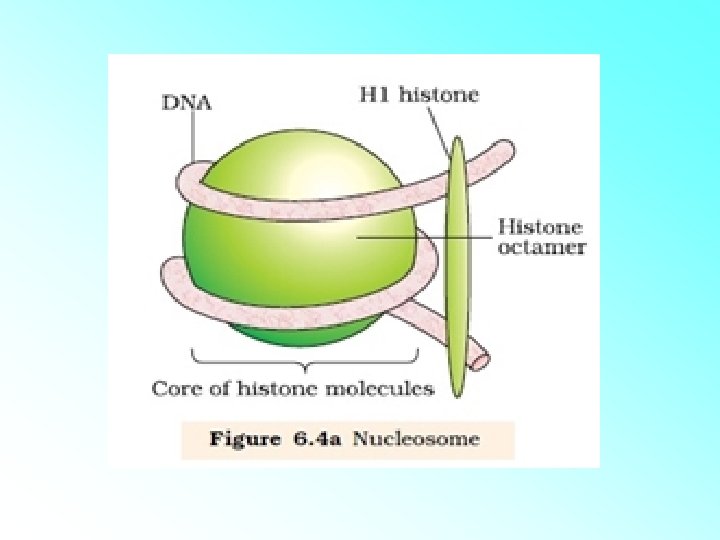
HISTONES: • Histones are organised to form a unit of eight molecules called as histone octamer. The negatively charged DNA is wrapped around the positively charged histone octamer to form a structure called nucleosome. • A typical nucleosome contains 200 bp of DNA helix. Nucleosomes constitute the repeating unit of a structure in nucleus called chromatin, threadlike stained (coloured) bodies seen in nucleus. The nucleosomes in chromatin are seen as ‘beads-onstring

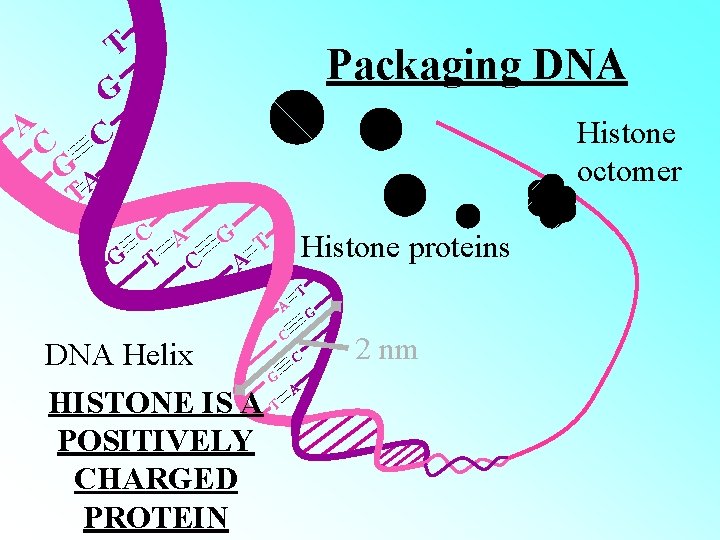
A C T G C Packaging DNA Histone octomer GA T C A G T C A Histone proteins T A DNA Helix G HISTONE IS A T POSITIVELY CHARGED PROTEIN G C C A 2 nm
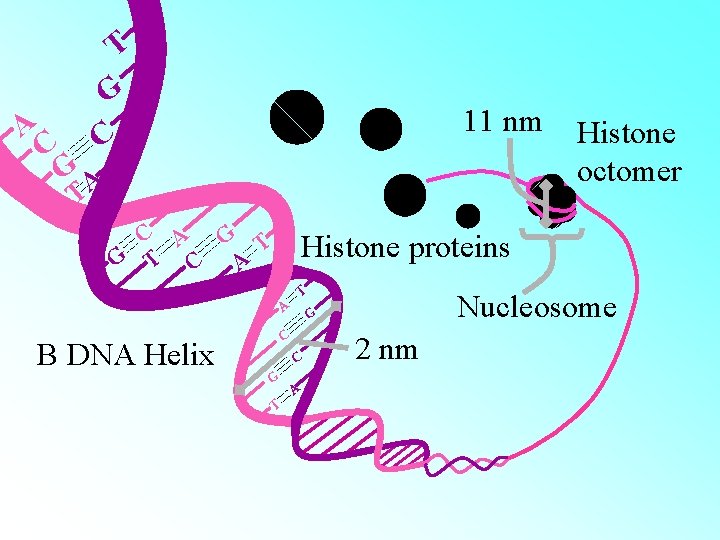
A C T G C 11 nm GA T C A G T C A Histone octomer Histone proteins T A C B DNA Helix C G T Nucleosome G A 2 nm

DNA IS WRAPPED AROUND HISTONE OCTOMER INTO A STRUCTURE CALLED NUCLEOSOME C A G T C A T A G C C G T A
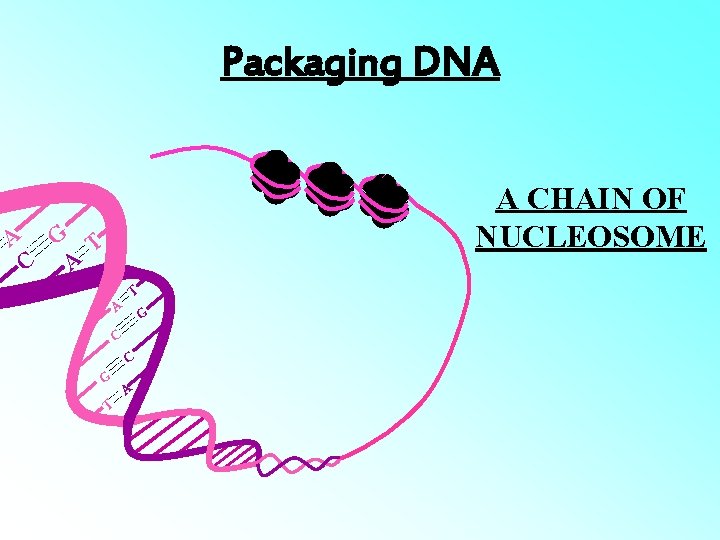
Packaging DNA A A CHAIN OF NUCLEOSOME G T C A T A G C C G T A
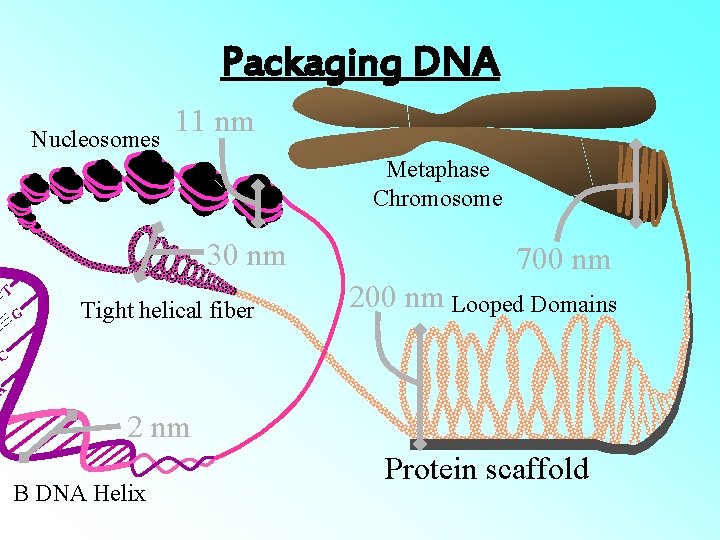
Packaging DNA Nucleosomes 11 nm Metaphase Chromosome 30 nm T G Tight helical fiber 700 nm 200 nm Looped Domains C A 2 nm B DNA Helix Protein scaffold
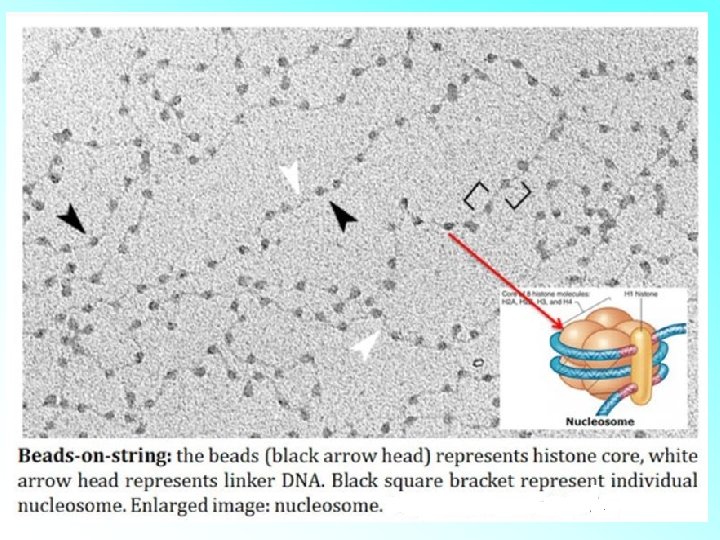
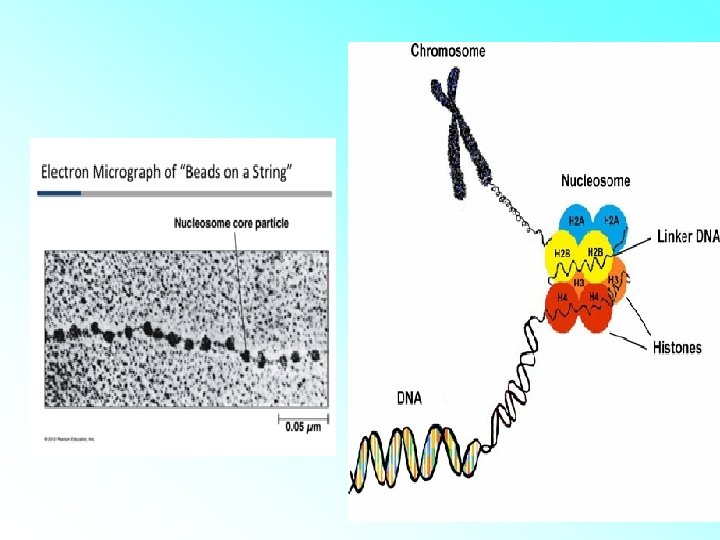
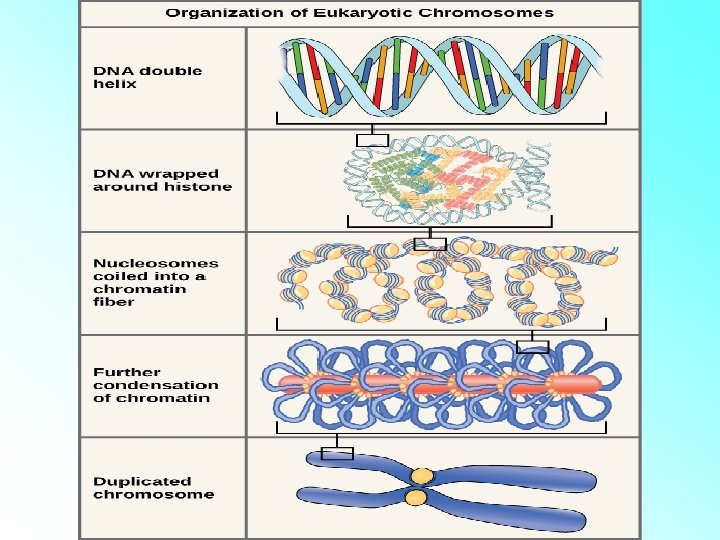

• The beads-on-string structure in chromatin is packaged to form chromatin fibers that are further coiled and condensed at metaphase stage of cell division to form chromosomes. • The packaging of chromatin at higher level requires additional set of proteins that collectively are referred to as Non-histone Chromosomal (NHC) proteins
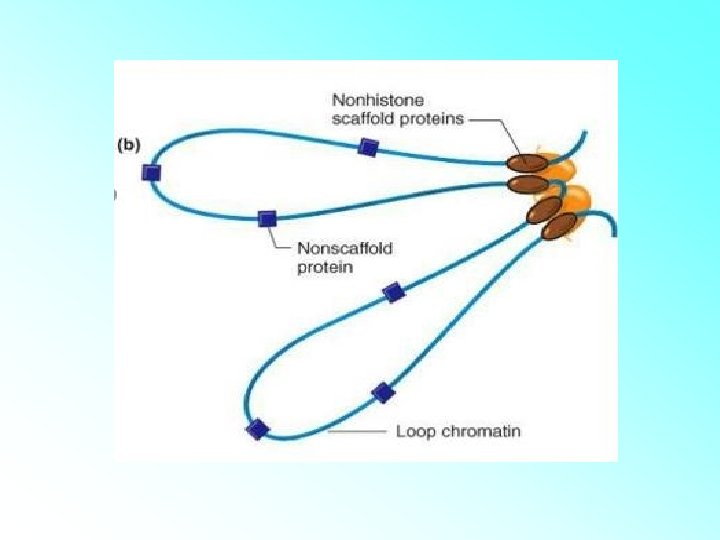

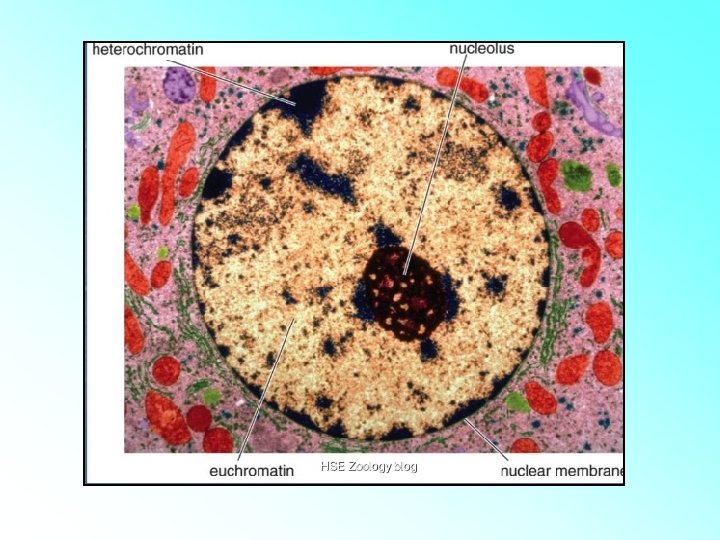
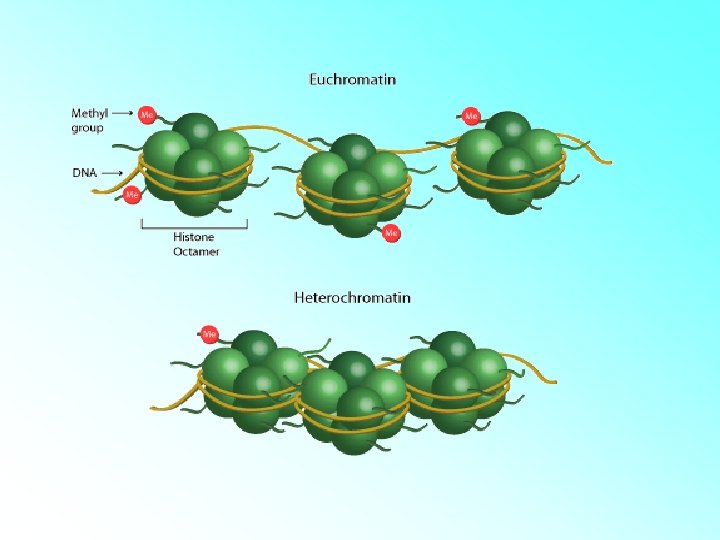
Chromatin Types • Euchromatin: – Lightly Stained. – The loosely coiled region of chromatin. – Transcriptionally active. • Heterochromatin: – Darkly Stained. – The tightly coiled region of chromatin. – Transcriptionally inactive.

ACTIVITY 1. Calculate the length of the DNA of bacteriophage lambda that has 48502 base pairs.

• Distance between two pairs=0. 34 nm=0. 34× 10 -9 m/bp Therefore, • total number of base pairs given are 48502 Total length of DNA=0. 34× 109 m/bp× 48502 =16490. 68× 10 -9 or 16. 49× 10 -6 m

2. A virus has a double stranded genome containing 48500 base pairs, calculate length in micro meters.

0. 34 nm ( the accepted length of one base pair). =485000 x 0. 34 = 164. 9 micrometres.

- Slides: 30