Pac Bio Meets the Microbiome George Weinstock Pac

Pac. Bio Meets the Microbiome George Weinstock Pac. Bio Users Group Meeting September 18, 2013

Diverse interest in medical metagenomics • • Acne Antibiotics, gut microbiome, and obesity Antibiotic resistance Asthma, allergies • • Bacterial vaginosis Cancer microbiomes Conjunctiva – trachoma microbiome Crohn's disease Cystic fibrosis Diabetes • • • C. difficile VRE MRSA E. coli O 157 H 7 NICU bacteremia Intestinal fat uptake Necrotizing enterocolitis Non-Alcoholic Fatty Liver Disease Oral microbiome • • • Influenza infection Pre-term babies Childhood vaccination Sepsis • • • Maternal microbiome Vitamin D Respiratory microbiome • • Periodontitis Caries Parasitic infection and the microbiome Post-transplant Lymphoproliferative Disorders Pre-term birth • • Oral microbiome Skin microbiome Dietary effects on gut microbiome Fecal transplant HIV and lung microbiome Infection control • • • Acute RSV infection Vitamin D • • • ICU NICU Short-bowel syndromes Urethritis Virus discovery • • Kawasaki Disease Fever of unknown origin in children Transplantation: CMV, BK Immuno-suppression/-compromised
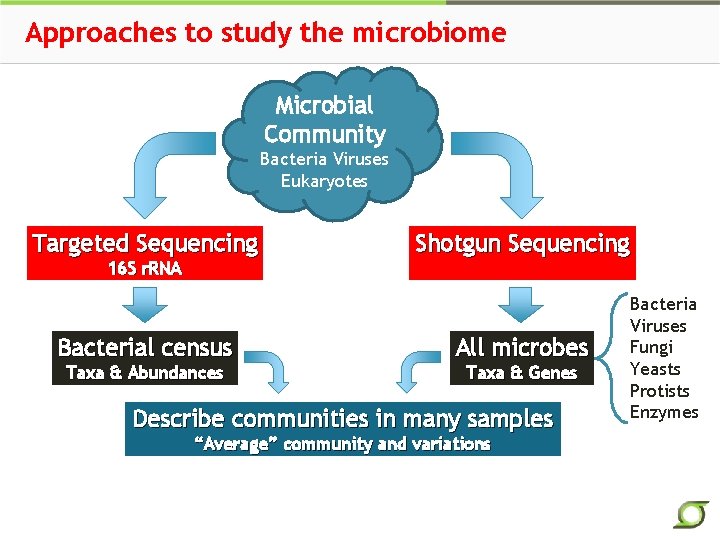
Approaches to study the microbiome Microbial Community Bacteria Viruses Eukaryotes Targeted Sequencing 16 S r. RNA Bacterial census Taxa & Abundances Shotgun Sequencing All microbes Taxa & Genes Describe communities in many samples “Average” community and variations Bacteria Viruses Fungi Yeasts Protists Enzymes

Studying communities - 16 S r. RNA genes Major “enterotypes” of the stool biomes Histograms of genera in each sample Ruminococcus women St. Louis Houston not hispanic/ latino/spanish Hispanic /latino/spanish BMI>=30 25 <= BMI <30 BMI<25 NA Each row a Prevotella different sample Bacteroides

Some Metagenomic Effects Community Structure e. g. content; ecological parameters (biodiversity) Multiple Specific Organisms beneficial ↓ detrimental↑ Specific Organism e. g. C. difficile Genes or Pathways e. g. lactic acid
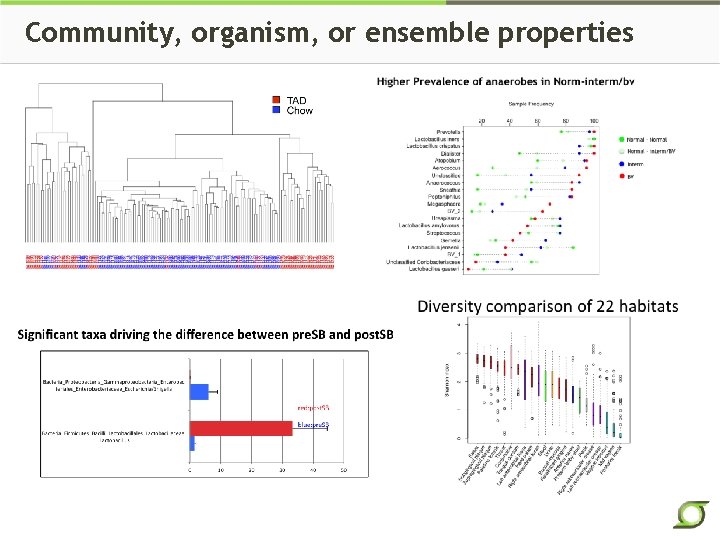
Community, organism, or ensemble properties

Metagenomic pathogen detection in clinical samples Hospital microbiology lab Patient samples Metagenomic sequencing Compare results Patients with/without hospital acquired diarrhea Alexis Elward David Haslam Greg Storch Rana Elfeghaly Yanjiao Zhou Kristine Wylie
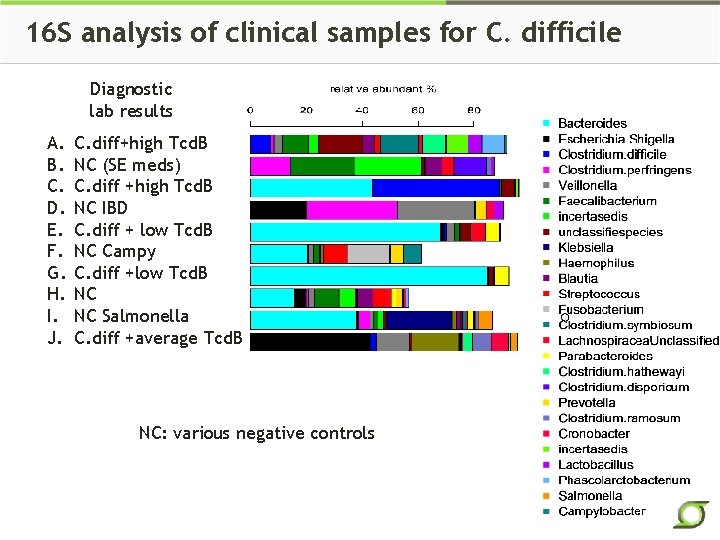
16 S analysis of clinical samples for C. difficile Diagnostic lab results A. B. C. D. E. F. G. H. I. J. C. diff+high Tcd. B NC (SE meds) C. diff +high Tcd. B NC IBD C. diff + low Tcd. B NC Campy C. diff +low Tcd. B NC NC Salmonella C. diff +average Tcd. B NC: various negative controls
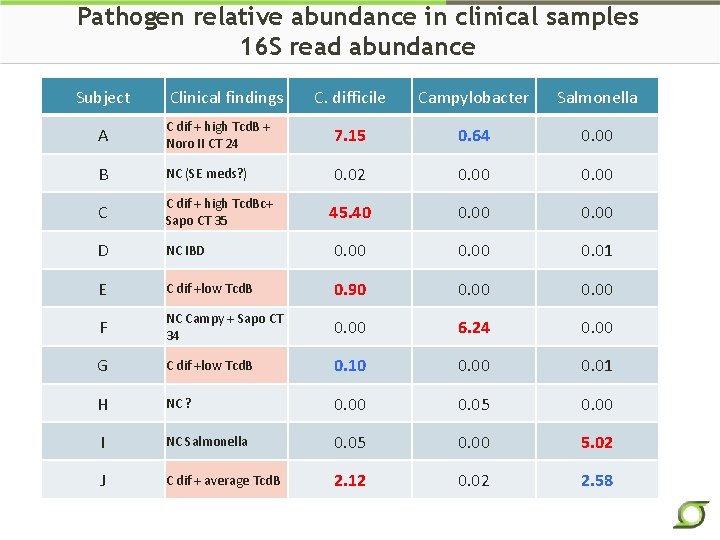
Pathogen relative abundance in clinical samples 16 S read abundance Subject Clinical findings C. difficile Campylobacter Salmonella A C dif + high Tcd. B + Noro II CT 24 7. 15 0. 64 0. 00 B NC (SE meds? ) 0. 02 0. 00 C C dif + high Tcd. Bc+ Sapo CT 35 45. 40 0. 00 D NC IBD 0. 00 0. 01 E C dif +low Tcd. B 0. 90 0. 00 F NC Campy + Sapo CT 34 0. 00 6. 24 0. 00 G C dif +low Tcd. B 0. 10 0. 01 H NC ? 0. 00 0. 05 0. 00 I NC Salmonella 0. 05 0. 00 5. 02 J C dif + average Tcd. B 2. 12 0. 02 2. 58
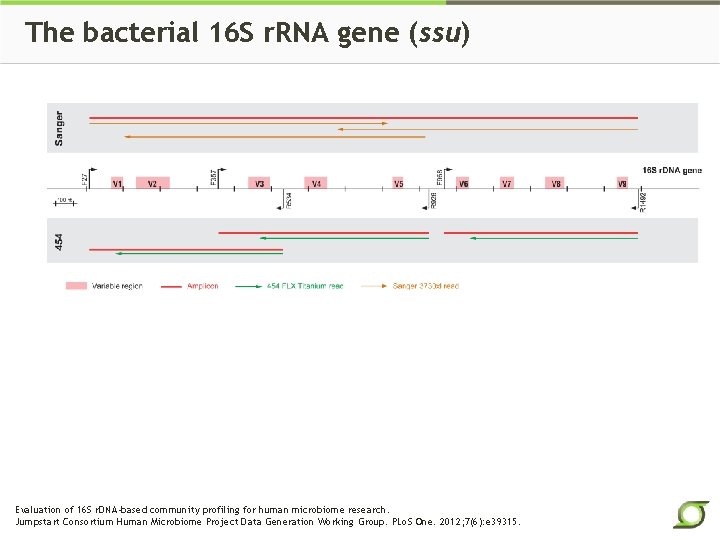
The bacterial 16 S r. RNA gene (ssu) Evaluation of 16 S r. DNA-based community profiling for human microbiome research. Jumpstart Consortium Human Microbiome Project Data Generation Working Group. PLo. S One. 2012; 7(6): e 39315.
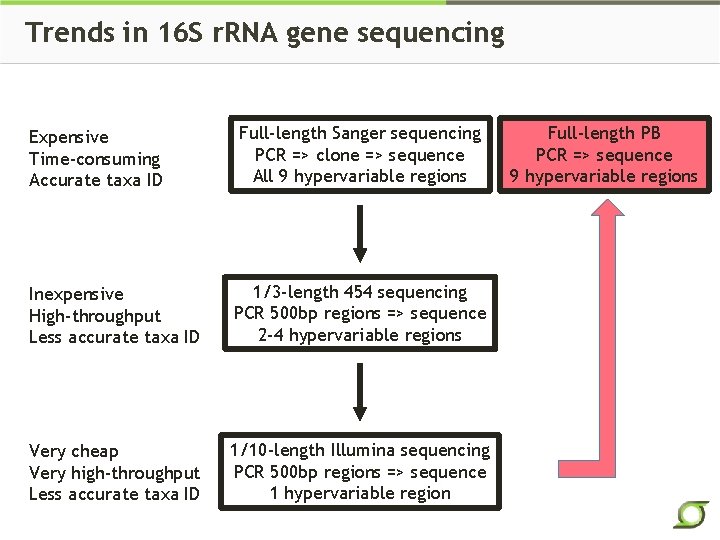
Trends in 16 S r. RNA gene sequencing Expensive Time-consuming Accurate taxa ID Full-length Sanger sequencing PCR => clone => sequence All 9 hypervariable regions Inexpensive High-throughput Less accurate taxa ID 1/3 -length 454 sequencing PCR 500 bp regions => sequence 2 -4 hypervariable regions Very cheap Very high-throughput Less accurate taxa ID 1/10 -length Illumina sequencing PCR 500 bp regions => sequence 1 hypervariable region Full-length PB PCR => sequence 9 hypervariable regions
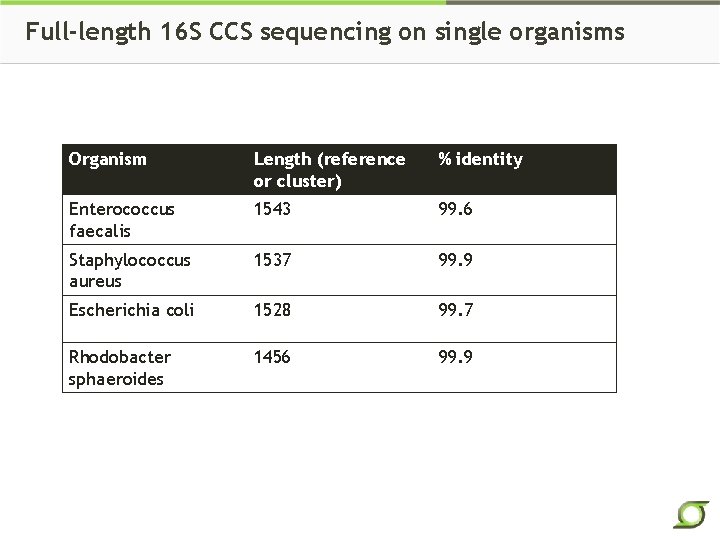
Full-length 16 S CCS sequencing on single organisms Organism Length (reference or cluster) % identity Enterococcus faecalis 1543 99. 6 Staphylococcus aureus 1537 99. 9 Escherichia coli 1528 99. 7 Rhodobacter sphaeroides 1456 99. 9

Large-scale single isolate typing • Have ~8000 isolates (microtiter plates) from hospital • Looking for unsequenced species from humans • Need FL 16 S in order to make a species call for typing • 400 base reads from 454 do not give enough specificity • Each well has one strain • Sanger seq’ing of FL PCR products => single sequence w/o cloning • Can Pac. Bio compete: cheaper, higher throughput? • Goal: • Find what species these isolates are • Choose novel isolates • Perform WG sequencing

Large-scale single isolate typing with Pac. Bio • Sanger: do not see alleles of multiple 16 S genes/strain Pac. Bio: can see different alleles since single molecule • Hospital isolates (82): • • 70 samples agree between Sanger and Pac. Bio 4 samples have minor species seen with both platforms 5 samples have strain differences seen with Pac. Bio, not w Sanger 2 samples failed with Sanger, not w Pac. Bio 1 sample disagreement 7 DNA sample controls agree between platforms 4 known culture sample controls agree between platforms • Pac. Bio: can see low level contaminants • 99% agreement between Sanger and Pac. Bio (90/91) • Only 1 disagreement between the platforms • More information from Pac. Bio

Cost is an issue • With 96 samples/1 SMRT cell, the fully loaded cost of Pac. Bio is about 2 x Sanger. • • • SMRT cell Sequencing reagents Library kit and labor Instrument Computation (storage, labor, cpu) • Would need to pool more samples/SMRT cell • Need more bar codes

Sequencing communities of microbes en masse • 16 S r. RNA gene sequencing for community profiling • Full-length gives species-level definition • 454 500 bp reads give genus-level definition • Shotgun sequencing • Longer reads give better assembly (of unknown uncultured) • Bacteria, viruses, fungi and other eukaryotes described
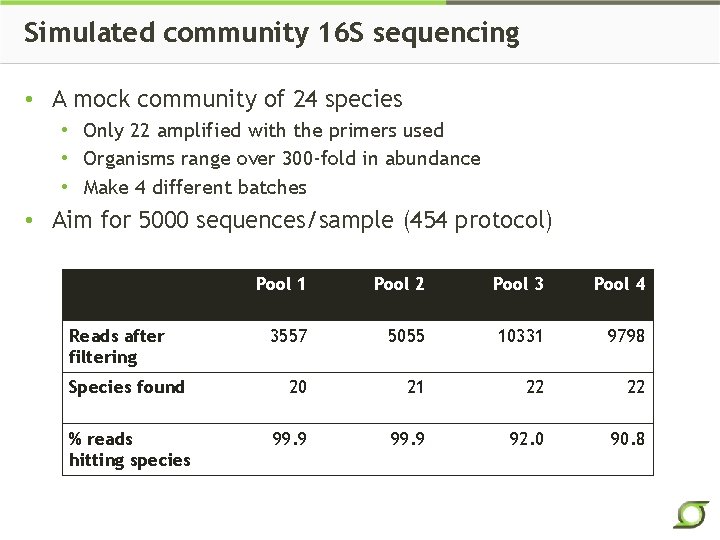
Simulated community 16 S sequencing • A mock community of 24 species • Only 22 amplified with the primers used • Organisms range over 300 -fold in abundance • Make 4 different batches • Aim for 5000 sequences/sample (454 protocol) Pool 1 Pool 2 Pool 3 Pool 4 3557 5055 10331 9798 Species found 20 21 22 22 % reads hitting species 99. 9 92. 0 90. 8 Reads after filtering
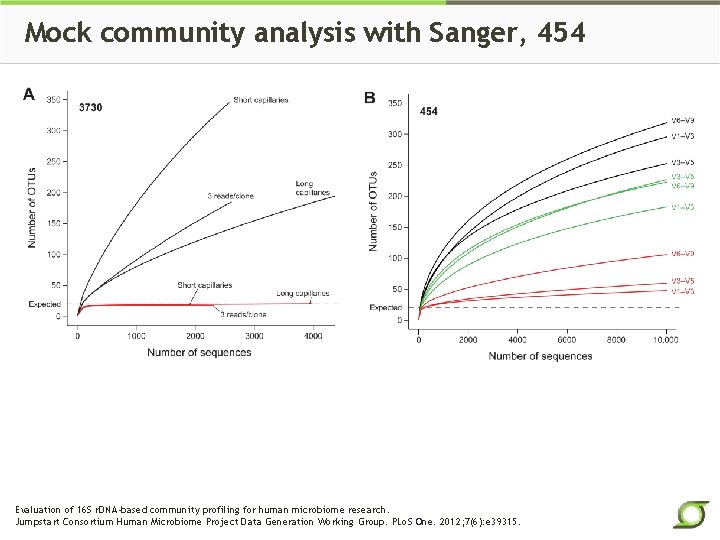
Mock community analysis with Sanger, 454 Evaluation of 16 S r. DNA-based community profiling for human microbiome research. Jumpstart Consortium Human Microbiome Project Data Generation Working Group. PLo. S One. 2012; 7(6): e 39315.

Consistent recognition of an organism in the pool for 4 replicate 16 S amplifications: Shuttleworthia_satelles Selenomonas_sputigena Ruminococcus_gnavus Providencia_alcalifaciens Prevotella_tannerae Parabacteroides_merdae Parabacteroides_johnsonii Neisseria_elongata Mitsuokella_multacida Methanobrevibacter_smithii Kingella_oralis Holdemania_filiformis Gemella_haemolysans Fusobacterium_periodonticum Eubacterium_sireaum Enterobacter_cancerogenus Eikenella_corrodens Dialister_invisus Collinsella_stercoris Clostridium_nexile Citrobacter_youngae Bifidobacterium_dentium Bacteroides_plebeius Actinomyces_odontolyticus Pool 1 Pool 2 Pool 3 Pool 4 0% 10% 20% 30% 40% 50% 60% 70% 80% 90% 100% 300 -fold difference in prevalence of 16 S genes for separate organisms in the pool Methanobrevibacter (an archaea) and Collinsella do not amplify with 16 S primers utilized.

INFECTION Traditional culture-based analysis Sample Culture single species Metagenomic analysis (culture-independent) WGS Assembly, Annotation Strains/Subspecies based on gene content Strains/Subspecies based on SNP/indel content Replace culture-based analysis with metagenomic analysis Alignment 16 S Assembly Annotation Species present Genes of interest Variants of a species Strains/Subs pecies

Acknowledgments Washington University Genome Institute: • Makedonka Mitreva • Erica Sodergren • Sahar Abubucker • Karthik Kota • John Martin • Bruce Rosa Thank you to the • Yanjiao Zhou subjects and • Kristine Wylie their families • Kathie Mihindukulasuriya • Hongyu Gao • Bill Shannon • Patricio La Rosa • Great Production & Informatics Teams Clinical • Greg Storch, WU • Phil Tarr, WU • Barb Warner, WU • J. Dennis Fortenberry, Indiana U • Ellen Li, SUNY-Stony Brook • Huiying Li, UCLA • Brad Warner, WU • Many others • • Funding: NIH Gates Foundation Peer Bork Group • • Siegfried Schloissnig Manimozhiyan Arumugam Shinichi Sunagawa Julien Tap Ana Zhu Alison S. Waller Daniel R. Mende Shamil R. Sunyaev Susan Haake, UCLA Martin Blaser, NYU Richard Hotchkiss, WU Scott Weiss, Harvard Katherine Gregory, Harvard Catherine O’Brien, Toronto Homer Twigg, Indiana U
- Slides: 21