Oxidation of NAD by O 2 NADH 2
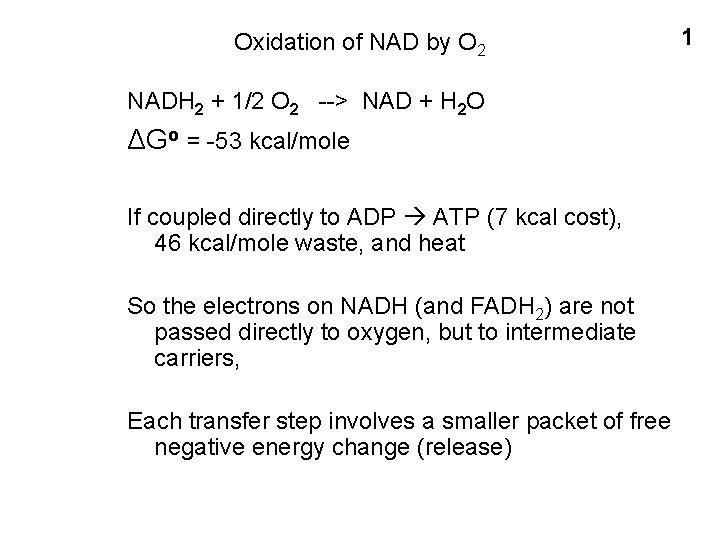
Oxidation of NAD by O 2 NADH 2 + 1/2 O 2 --> NAD + H 2 O ΔGo = -53 kcal/mole If coupled directly to ADP ATP (7 kcal cost), 46 kcal/mole waste, and heat So the electrons on NADH (and FADH 2) are not passed directly to oxygen, but to intermediate carriers, Each transfer step involves a smaller packet of free negative energy change (release) 1
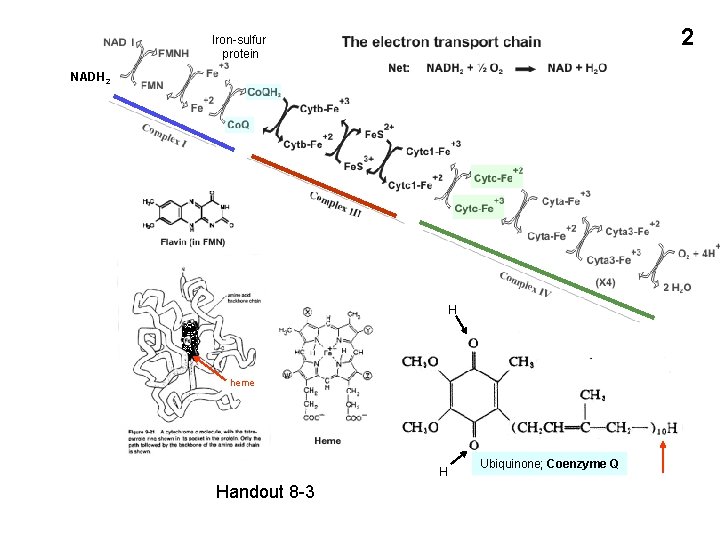
2 Iron-sulfur protein NADH 2 H heme H Handout 8 -3 Ubiquinone; Coenzyme Q
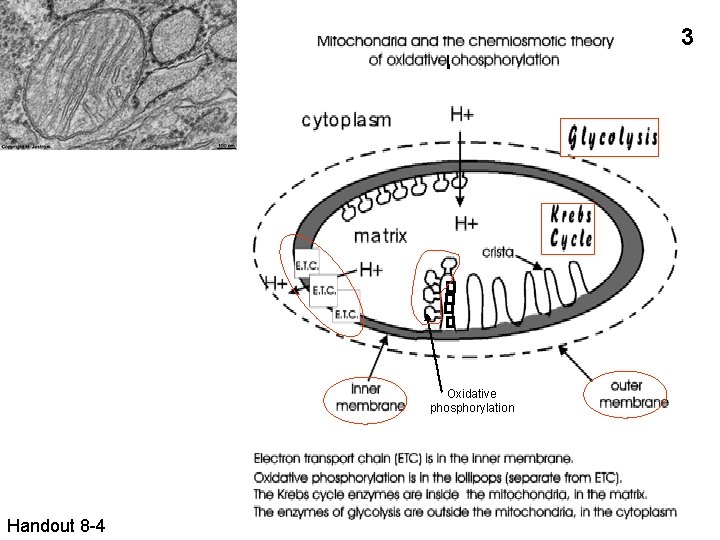
3 Oxidative phosphorylation Handout 8 -4
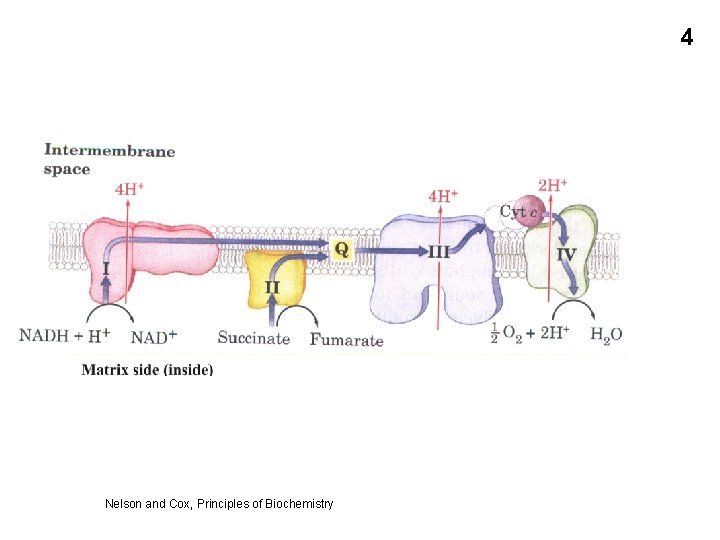
4 Nelson and Cox, Principles of Biochemistry
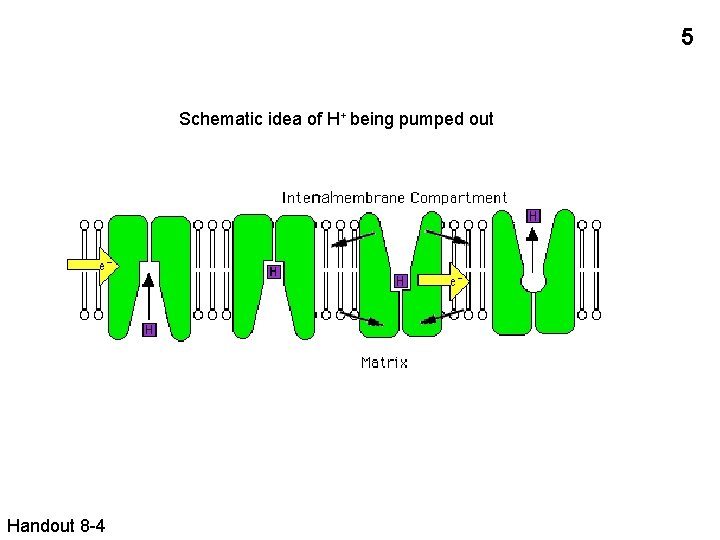
5 Schematic idea of H+ being pumped out nal Handout 8 -4
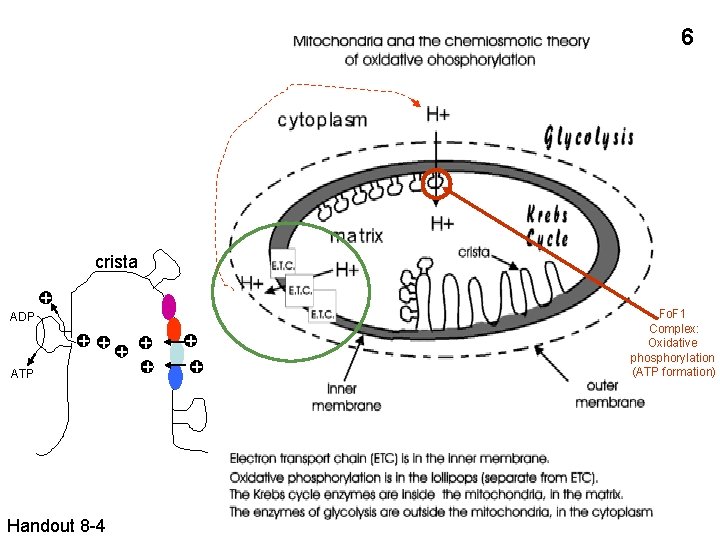
6 crista + ADP ATP + + + Handout 8 -4 + + Fo. F 1 Complex: Oxidative phosphorylation (ATP formation)

7 Chemiosmotic theory (Mitchell hypothesis) Proton motive force (pmf) Chemical gradient Electrochemical gradient Peter Mitchell 1961 (without knowing mechanism) Water-pump-dam analogy 3 exampels of evidence supporting themitchell hypothesis:
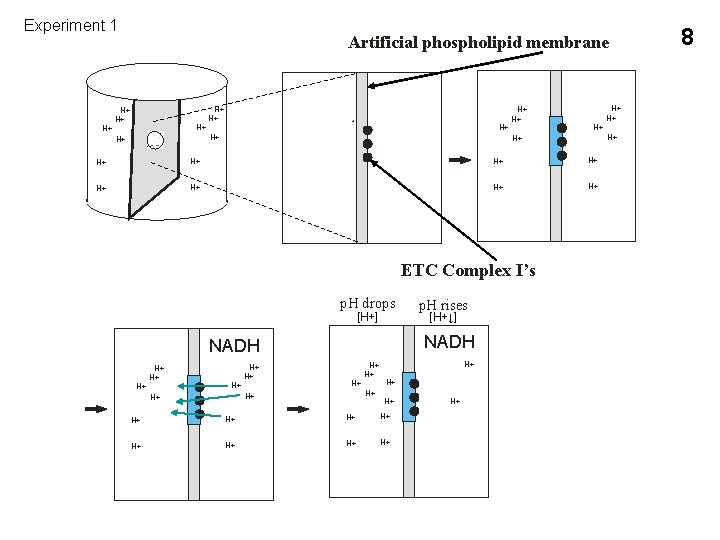
Experiment 1 H+ Artificial phospholipid membrane H+ H+ H+ H+ H+ H+ ETC Complex I’s p. H drops [H+] H+ H+ H+ [H+↓] NADH H+ H+ p. H rises H+ H+ H+ H+ 8
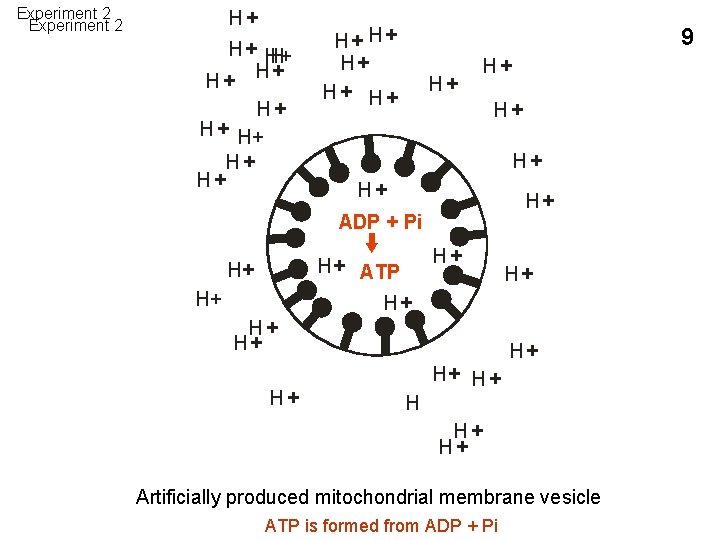
Experiment 2 H+ H + H+ H+ H+ 9 H+ H+ H+ ADP + Pi H+ H+ ATP H+ H+ H+ H+ Artificially produced mitochondrial membrane vesicle ATP is formed from ADP + Pi
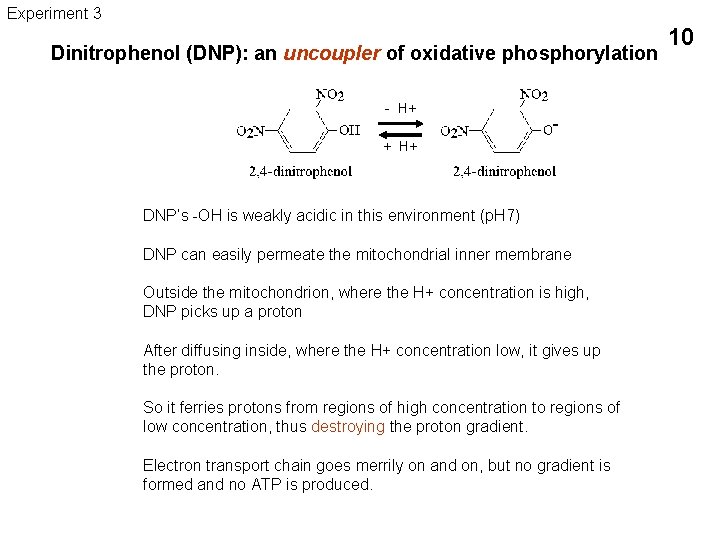
Experiment 3 Dinitrophenol (DNP): an uncoupler of oxidative phosphorylation - H+ - + H+ DNP’s -OH is weakly acidic in this environment (p. H 7) DNP can easily permeate the mitochondrial inner membrane Outside the mitochondrion, where the H+ concentration is high, DNP picks up a proton After diffusing inside, where the H+ concentration low, it gives up the proton. So it ferries protons from regions of high concentration to regions of low concentration, thus destroying the proton gradient. Electron transport chain goes merrily on and on, but no gradient is formed and no ATP is produced. 10

11
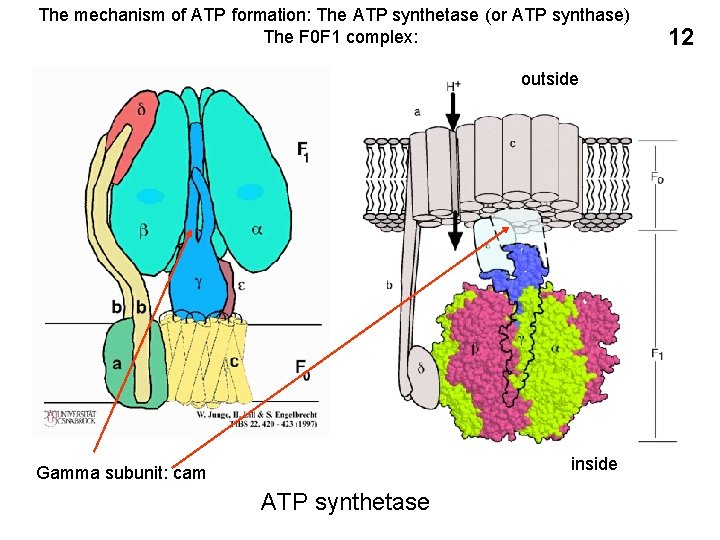
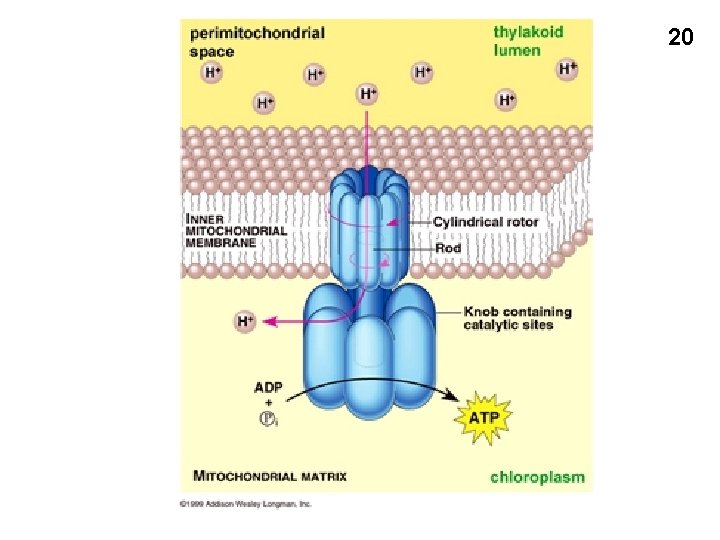
The mechanism of ATP formation: The ATP synthetase (or ATP synthase) The F 0 F 1 complex: outside inside Gamma subunit: cam ATP synthetase 12
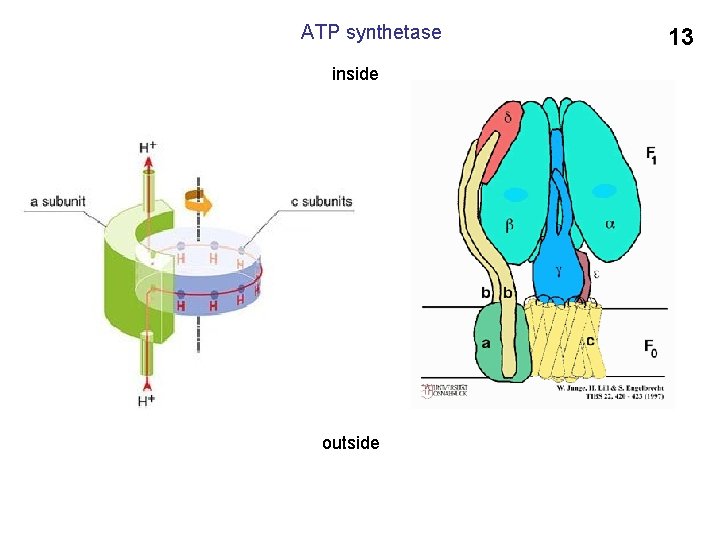
ATP synthetase inside outside 13
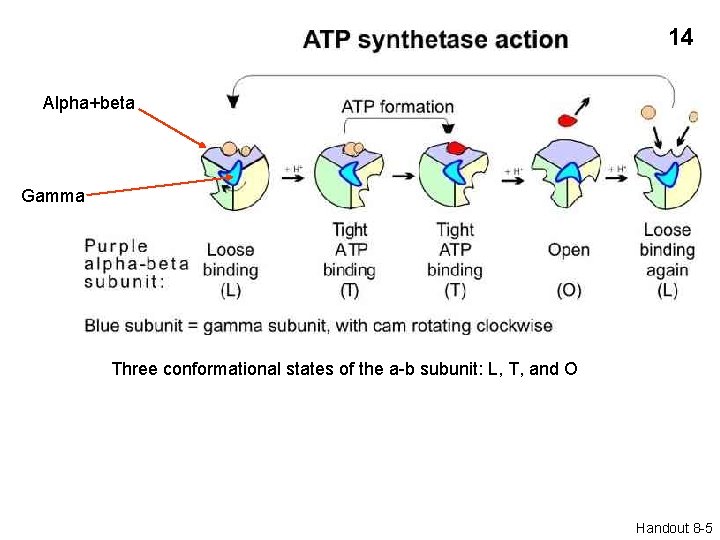
14 Alpha+beta Gamma Three conformational states of the a-b subunit: L, T, and O Handout 8 -5
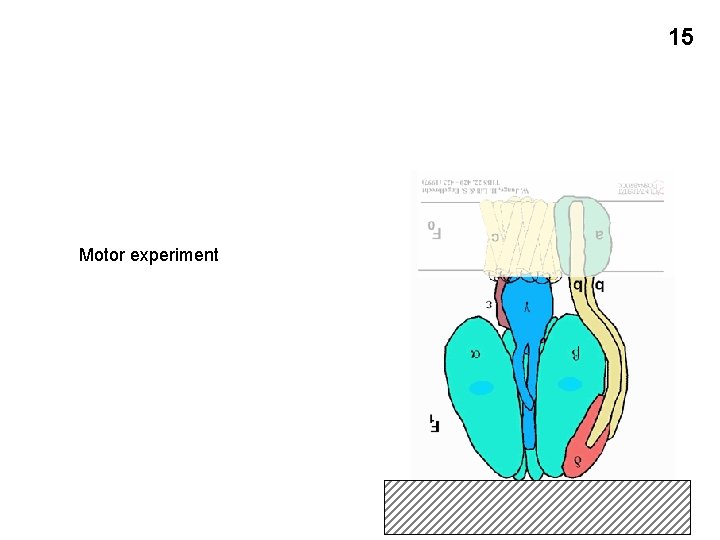
15 Motor experiment
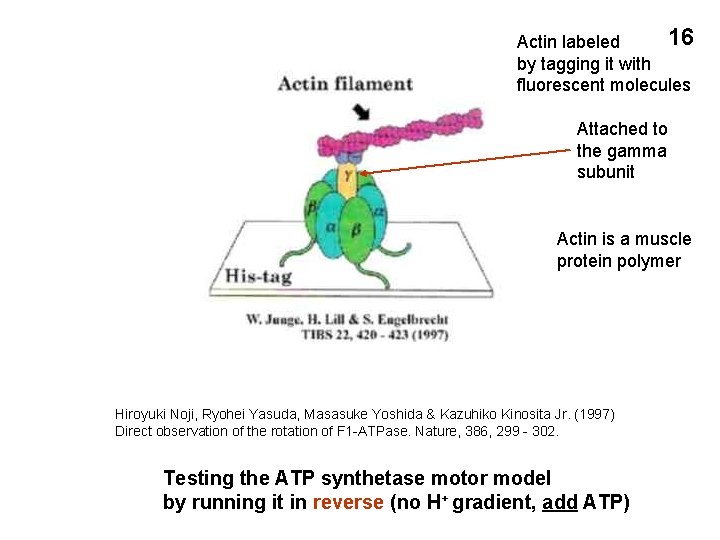
16 Actin labeled by tagging it with fluorescent molecules Attached to the gamma subunit Actin is a muscle protein polymer Hiroyuki Noji, Ryohei Yasuda, Masasuke Yoshida & Kazuhiko Kinosita Jr. (1997) Direct observation of the rotation of F 1 -ATPase. Nature, 386, 299 - 302. Testing the ATP synthetase motor model by running it in reverse (no H+ gradient, add ATP)
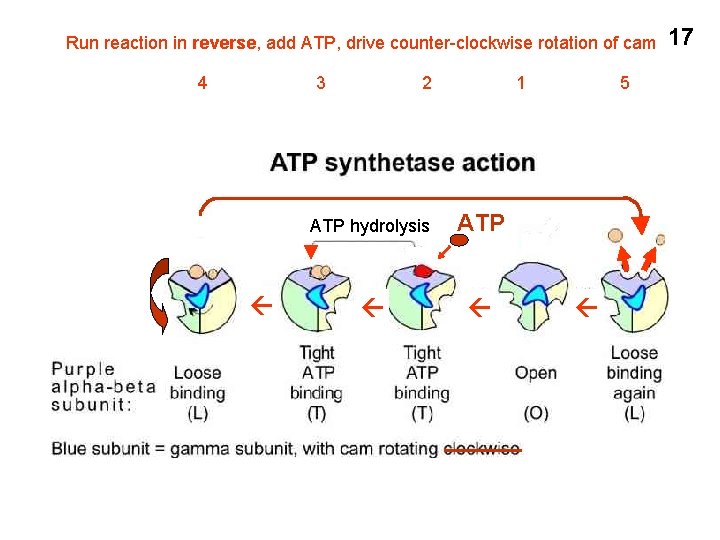
Run reaction in reverse, add ATP, drive counter-clockwise rotation of cam 4 3 2 1 ATP hydrolysis ATP 5 17

18 desktop
19
20
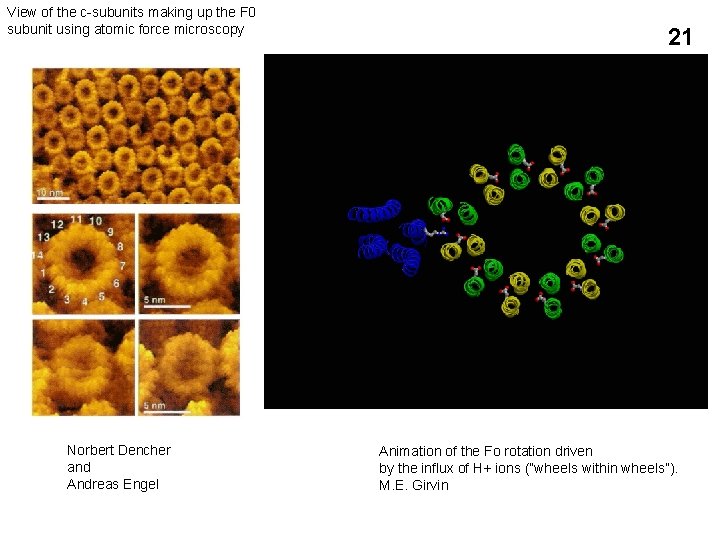
View of the c-subunits making up the F 0 subunit using atomic force microscopy Norbert Dencher and Andreas Engel 21 Animation of the Fo rotation driven by the influx of H+ ions (“wheels within wheels”). M. E. Girvin
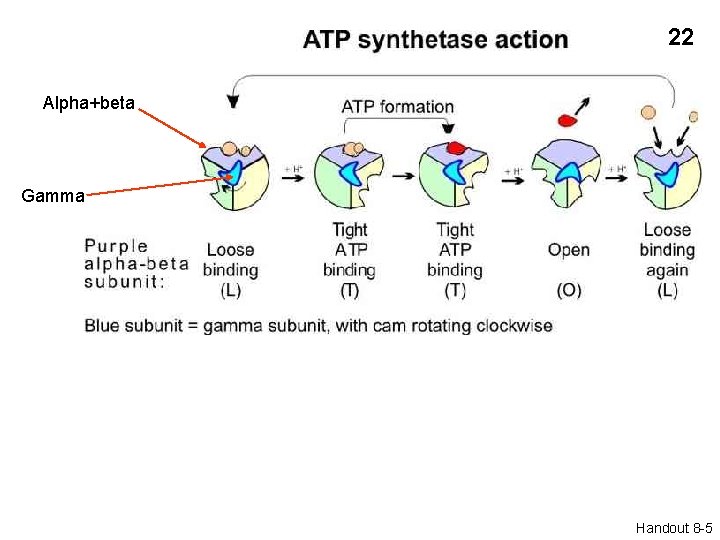
22 Alpha+beta Gamma Handout 8 -5
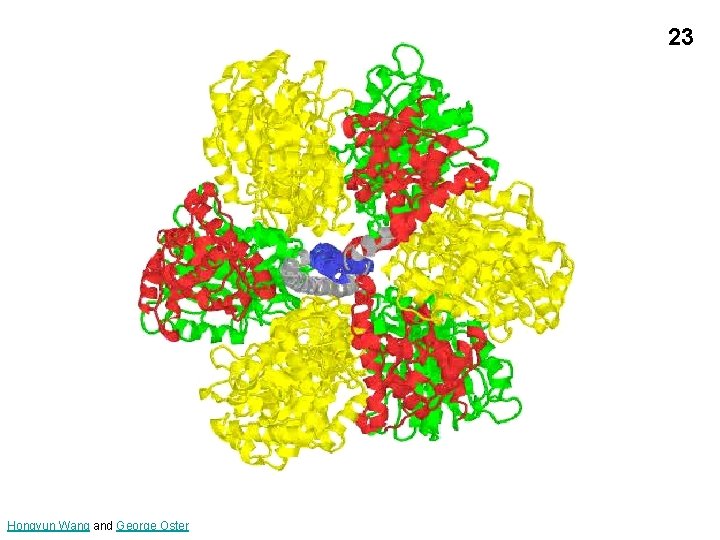
23 Hongyun Wang and George Oster

ATP accounting • • • 24 Each of the 3 ETC complex (I, III, IV) pumps enough H+ ions to allow the formation of 1 ATP. So 3 ATPs per pair of electrons passing through the full ETC. So 3 ATPs per 1/2 O 2 So 3 ATPs per NADH 2 But only 2 ATPs per FADH 2 (skips complex 1)
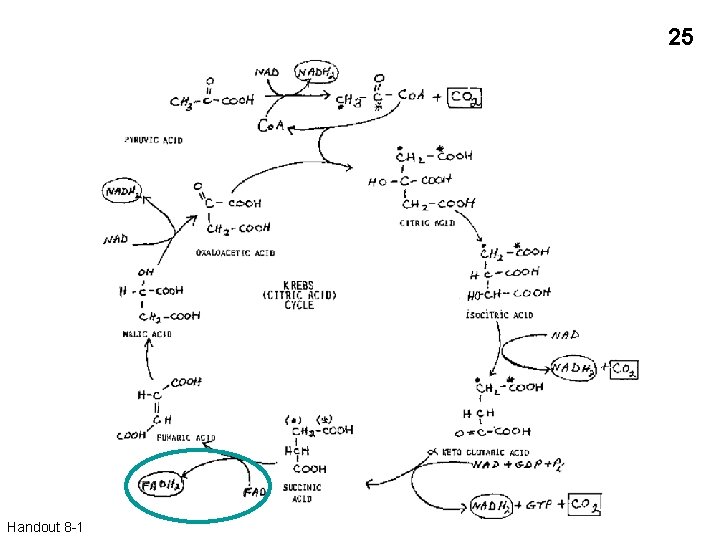
25 Handout 8 -1
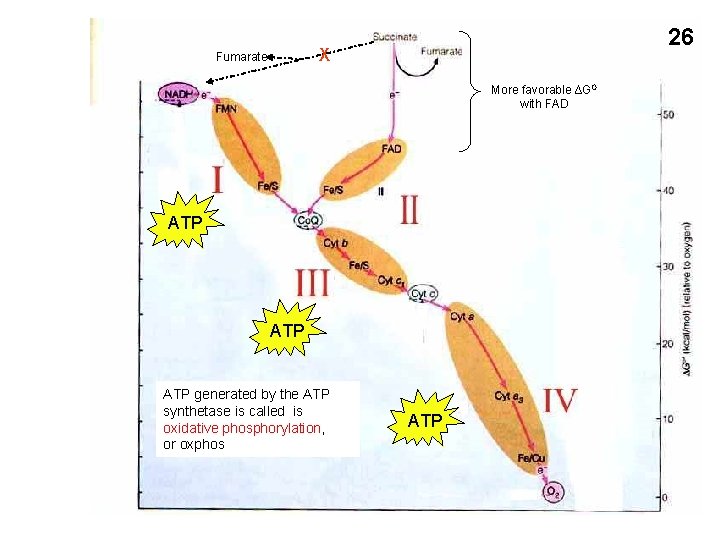
26 X Fumarate More favorable ∆GO with FAD ATP ATP generated by the ATP synthetase is called is oxidative phosphorylation, or oxphos ATP
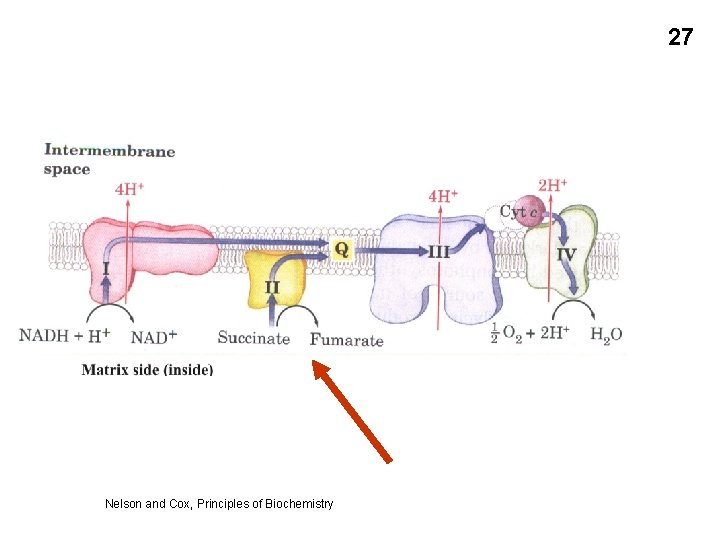
27 Nelson and Cox, Principles of Biochemistry
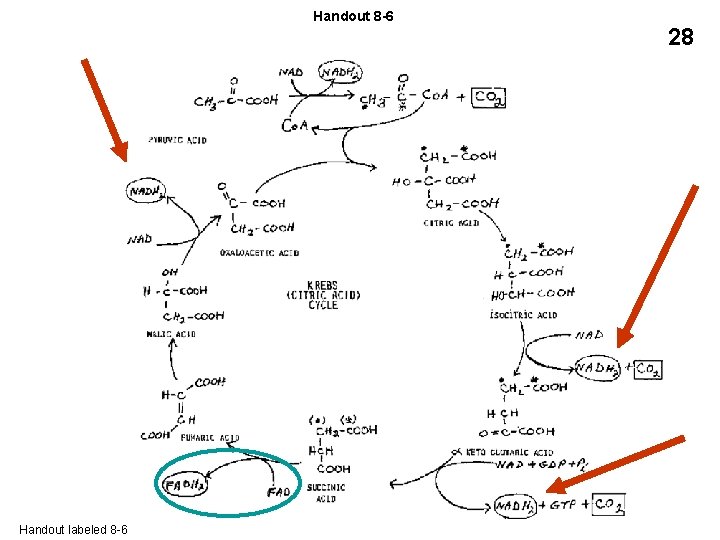
Handout 8 -6 28 OXPHOS: 1 NADH from glycolysis Substrate level phosphorylation (SLP): 2 ATP 1 ATP from Glycolysis 1 ATP (GTP) from Krebs Handout labeled 8 -6 1 NADH from Krebs entry 3 NADH from Krebs 1 FADH 2 from Krebs Total: 17 ATP 5 NADH = 15 ATP 1 FADH 2 = 2 ATP Grand total (E. coli): 17 + 2 = 19 per ½ glucose or 38 per 1 glucose

ATP accounting • 38 ATP/ glucose in E. coli • 36 ATP/glucose in eukaryotes – Cost of bringing in the electrons from NADH from glycolysis into the mitochondrion = 1 ATP per electron pair – So costs 2 ATPs per glucose, subtract from 38 to get 36 net. 29

30 Efficiency • 36 ATP/ glucose, worth 7 X 36 = 252 kcal/mole of glucose • ΔGo for the overall reaction glucose + 6 O 2→ 6 CO 2 + 6 H 2 O: -686 kcal/ mole • Efficiency = 252/686 = 37% • Once again, better than most gasoline engines. • and Energy yield: 36 ATP/ glucose vs. 2 ATP/glucose in fermentation (yet fermentation works) • So with or without oxygen, get energy from glucose
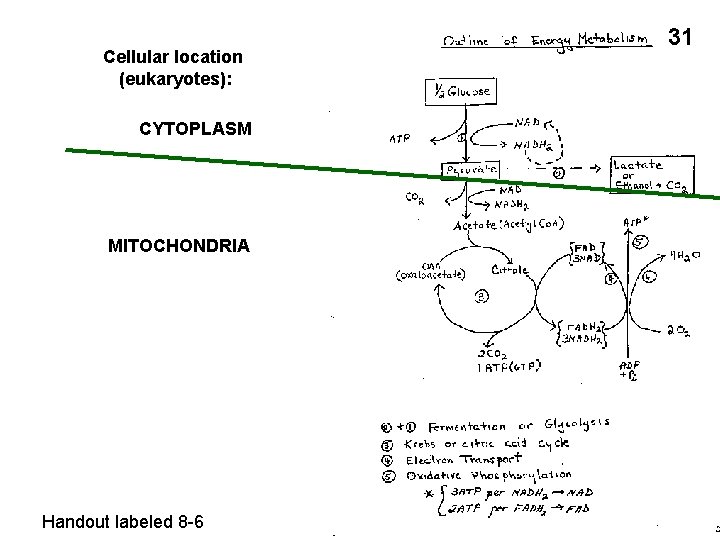
Cellular location (eukaryotes): CYTOPLASM MITOCHONDRIA Handout labeled 8 -6 31

32 Alternative sources of carbon and energy Shake = milk: milk sugar = lactose = disaccharide = glucose – galactose beta-galactosidase +HOH → glucose + galactose glucose → glycolysis, etc. galactose (3 enzymatic steps) glucose

33 Alternative sources of carbon and energy Bun = starch = poly-alpha-glucose G-1 -P → G-6 -P glycolysis

34 Alternative sources of carbon and energy Lettuce = cellulose = polysaccharide Poly-beta glucose →| (stays as the polysaccharide) We have no enzyme for catabolizing cellulose

35 Alternative sources of carbon and energy French fries = fat (oil) = triglyceride
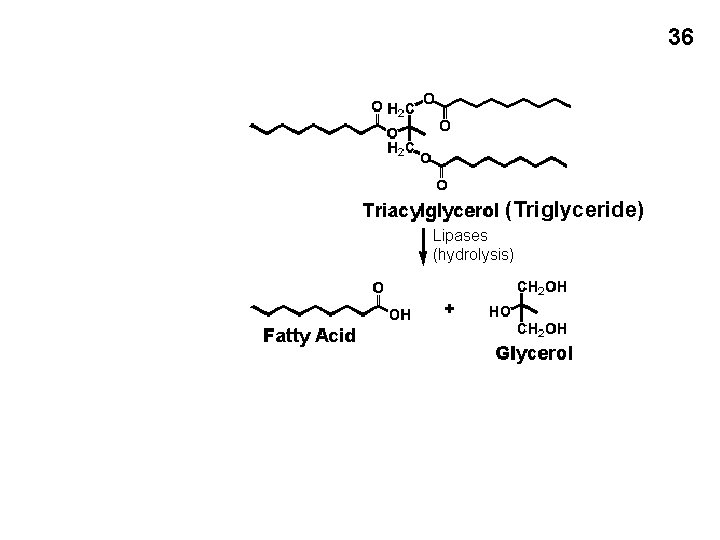
36 (Triglyceride) Lipases (hydrolysis)

37 Handout 9 -1 left
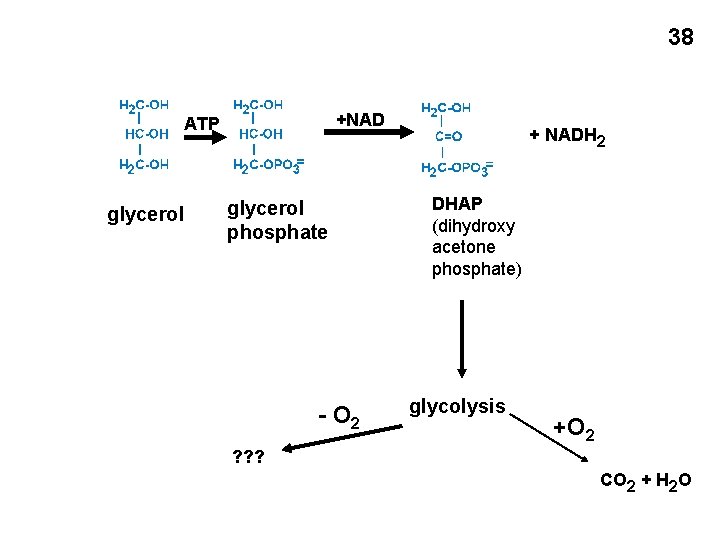
38 +NAD ATP glycerol phosphate - O 2 + NADH 2 DHAP (dihydroxy acetone phosphate) glycolysis +O 2 ? ? ? CO 2 + H 2 O
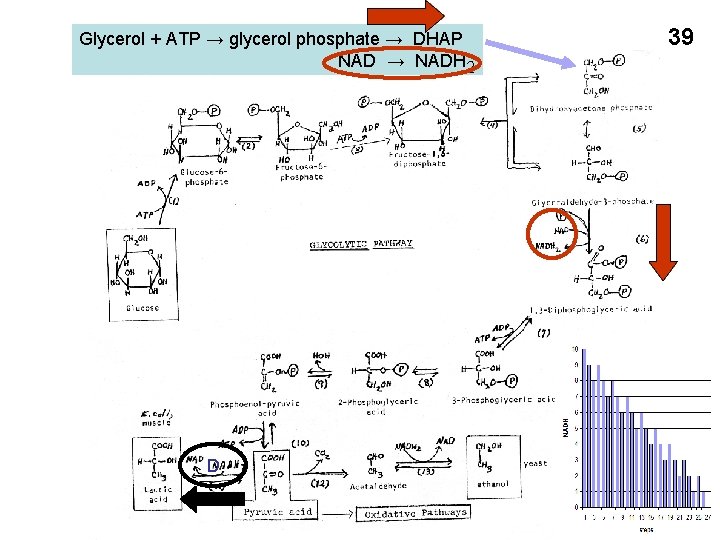
Glycerol + ATP → glycerol phosphate → DHAP NAD → NADH 2 D 39
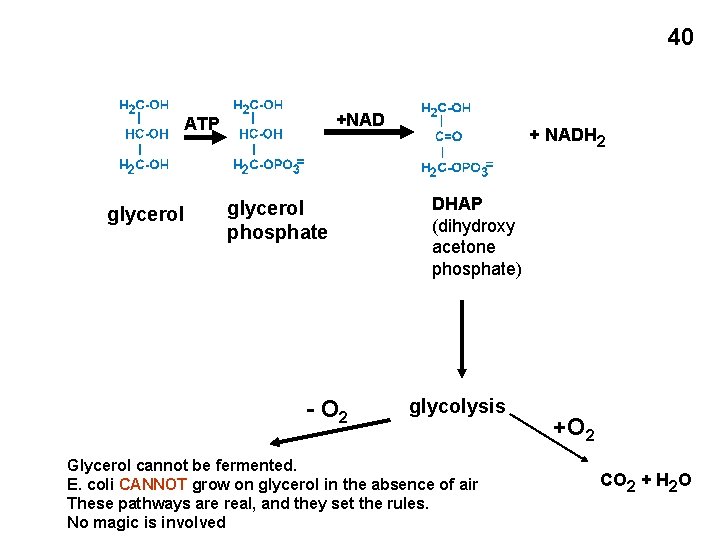
40 +NAD ATP glycerol phosphate - O 2 + NADH 2 DHAP (dihydroxy acetone phosphate) glycolysis Glycerol cannot be fermented. E. coli CANNOT grow on glycerol in the absence of air These pathways are real, and they set the rules. No magic is involved +O 2 CO 2 + H 2 O

41 Alternative sources of carbon and energy Hamburger = protein Proteases (e. g. , trypsin) → → 20 AAs Stomach acid (p. H 1) also helps by denaturing protein making it accessable to proteolytic attack Each of the 20 AA’s has its own catabolic pathway, and ends up in the glycolytic or Krebs cycle pathways But first, the N must be removed:
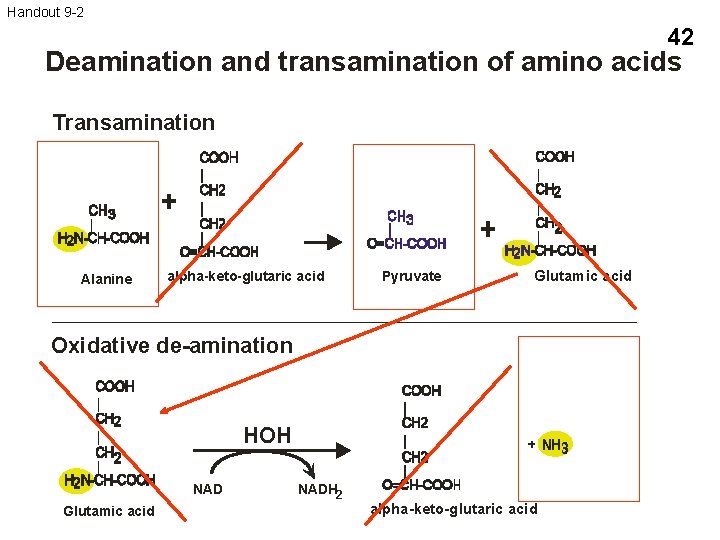
Handout 9 -2 42 Deamination and transamination of amino acids Transamination Alanine alpha-keto-glutaric acid Pyruvate Glutamic acid Oxidative de-amination HOH NAD Glutamic acid NADH 2 alpha-keto-glutaric acid
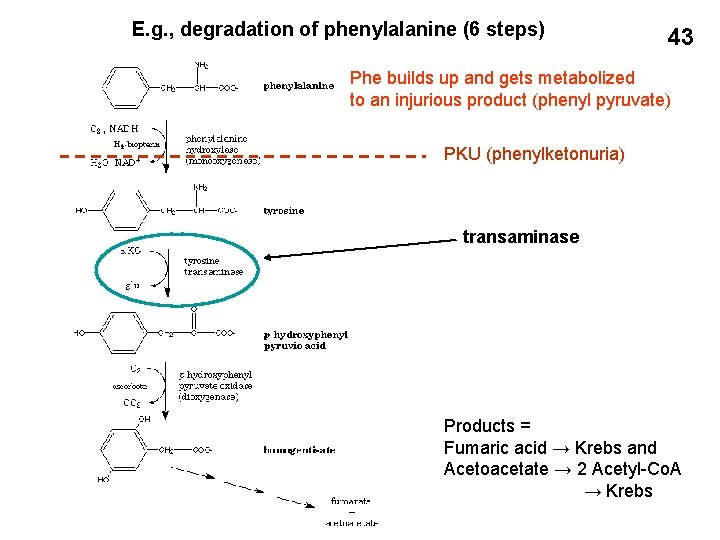
E. g. , degradation of phenylalanine (6 steps) 43 Phe builds up and gets metabolized to an injurious product (phenyl pyruvate) PKU (phenylketonuria) transaminase Products = Fumaric acid → Krebs and Acetoacetate → 2 Acetyl-Co. A → Krebs
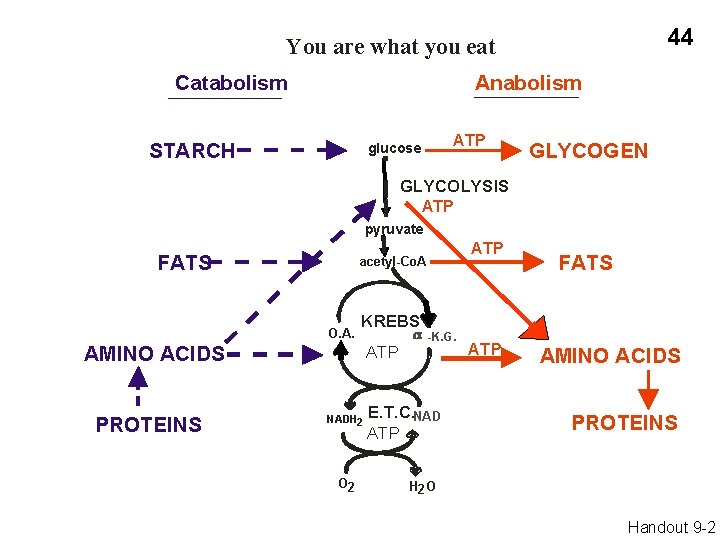
44 You are what you eat Catabolism Anabolism STARCH ATP glucose GLYCOGEN GLYCOLYSIS ATP pyruvate FATS O. A. KREBS AMINO ACIDS PROTEINS ATP acetyl-Co. A ATP NADH 2 O 2 -K. G. E. T. C. NAD ATP FATS AMINO ACIDS PROTEINS H 2 O Handout 9 -2
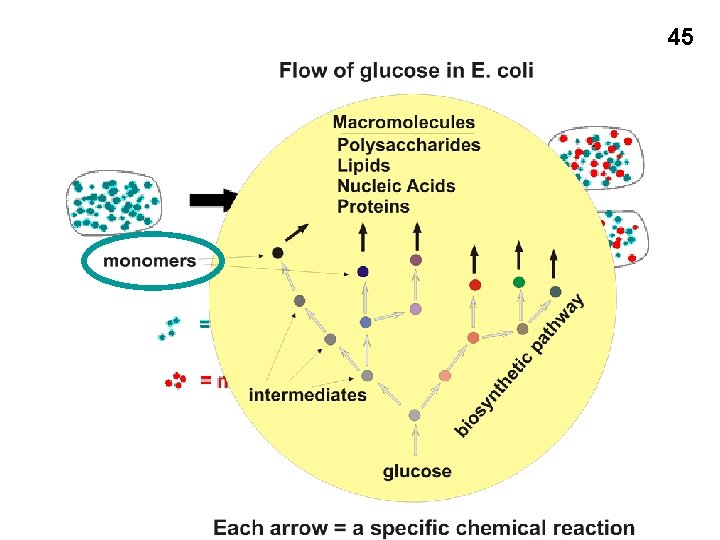
45
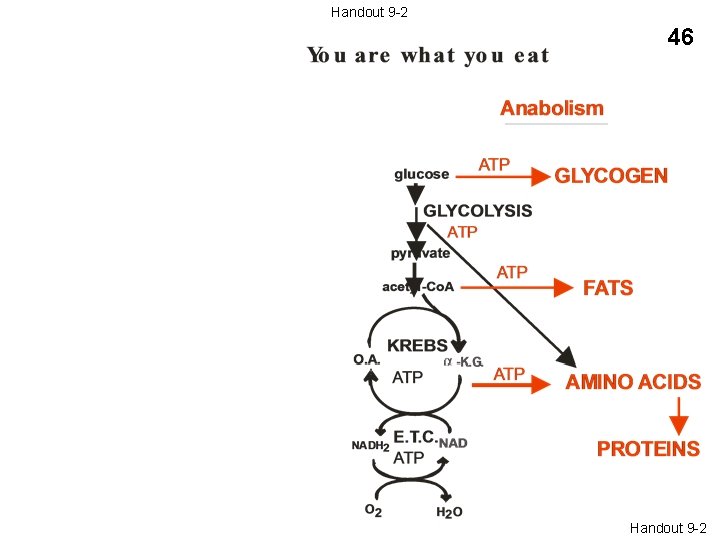
Handout 9 -2 46 Handout 9 -2

Biosynthesis of monomers E. g. , • Fatty acids (acetyl Co. A from Krebs cycle) • Amino acids (Serine: 3 -phospho-glyceric acid from glycolysis) 47
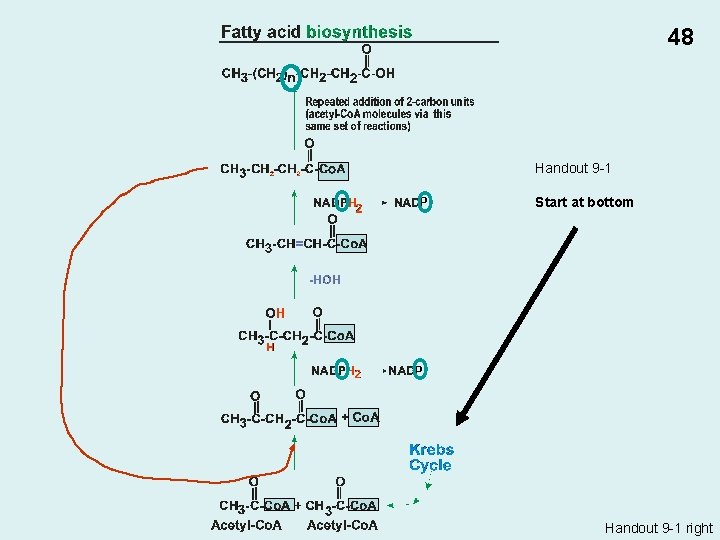
48 Handout 9 -1 P Start at bottom P Handout 9 -1 right
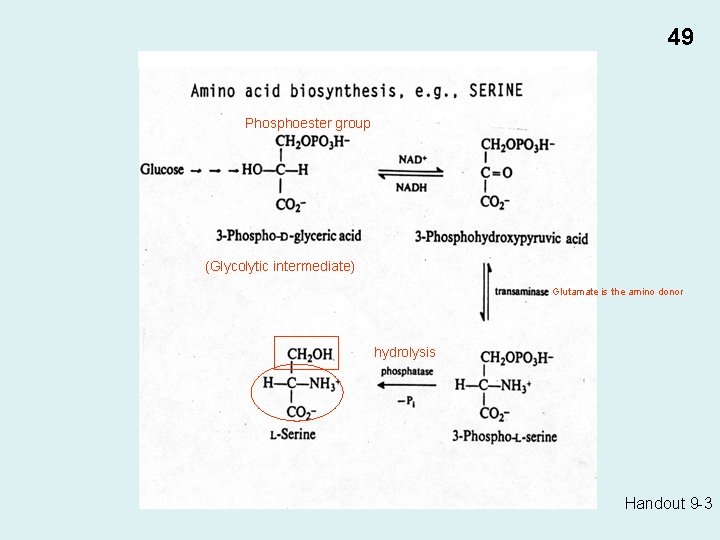
Handout 9 -3 49 Phosphoester group (Glycolytic intermediate) Glutamate is the amino donor hydrolysis Handout 9 -3

50
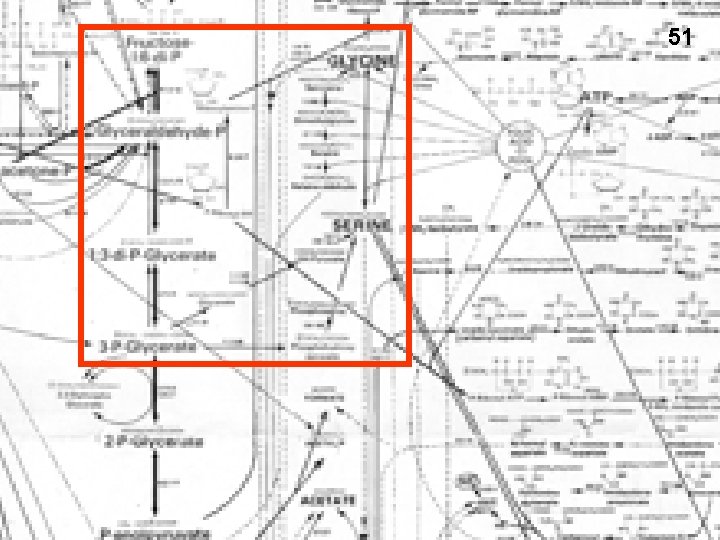
51
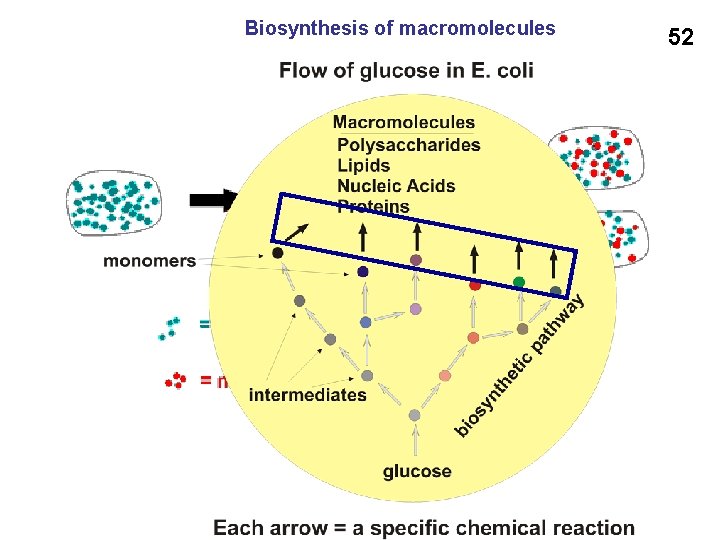
Biosynthesis of macromolecules 52
Biosynthesis of macromolecules 1) Lipids 2) Polysaccharides 3) Proteins 53
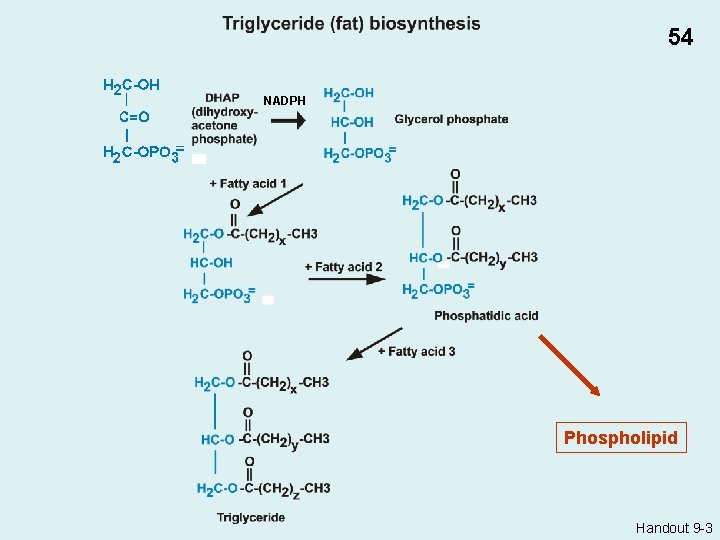
54 NADPH Phospholipid Handout 9 -3
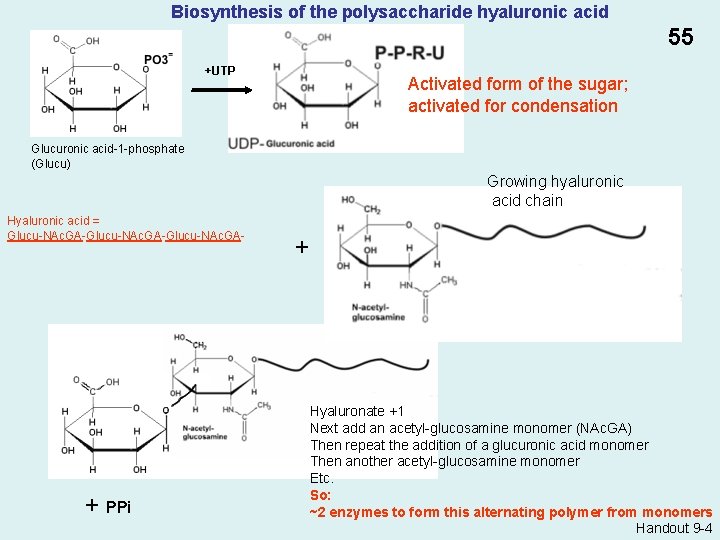
Biosynthesis of the polysaccharide hyaluronic acid 55 +UTP Activated form of the sugar; activated for condensation Glucuronic acid-1 -phosphate (Glucu) Growing hyaluronic acid chain Hyaluronic acid = Glucu-NAc. GA-Glucu-NAc. GA- + PPi + Hyaluronate +1 Next add an acetyl-glucosamine monomer (NAc. GA) Then repeat the addition of a glucuronic acid monomer Then another acetyl-glucosamine monomer Etc. So: ~2 enzymes to form this alternating polymer from monomers Handout 9 -4

Biosynthesis of proteins e. g. , an enzyme like hexokinase: met-val-his-leu-gly …. . If this done like lipids and polysaccharides, we need an enzyme for each linkage First an enzyme that will condense val to met to make met-val. Then an enzyme with a different substrate specificity, which adds his to metval to make met-val-his. Since there are 500 AAs in hexokinase, we need 500 enzymes to do the job. If there are 3000 proteins in E. coli, then we need ~500 X 3000 = 1. 5 million enzymes to make all the different primary structure of all the proteins. But even then, it won’t work, as each of these million enzymes is also a protein that needs to be synthesized. We need a better plan to polymerize the amino acids in the right order. 56
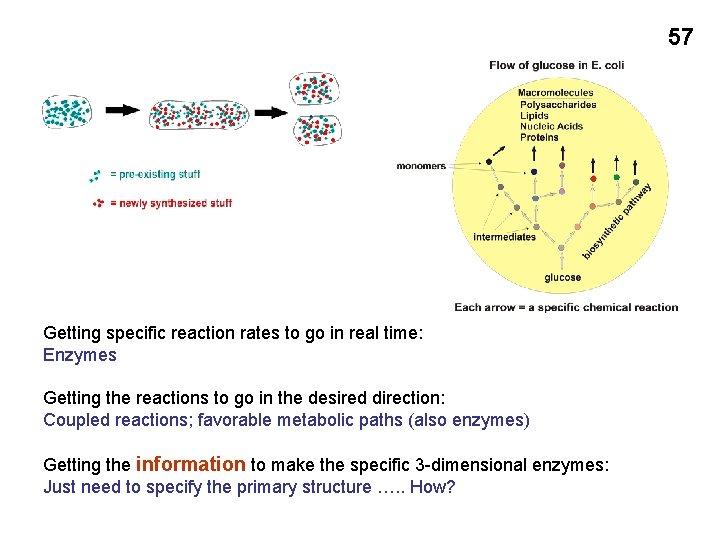
57 Getting specific reaction rates to go in real time: Enzymes Getting the reactions to go in the desired direction: Coupled reactions; favorable metabolic paths (also enzymes) Getting the information to make the specific 3 -dimensional enzymes: Just need to specify the primary structure …. . How?

Nucleic acids Prof. Mowshowitz will continue with this next chapter in the story, leading to the biosynthesis of the all-important proteins. 58
- Slides: 58