Overview of Shotgun Sequence Analysis Ami S Bhatt

Overview of Shotgun Sequence Analysis Ami S. Bhatt, MD Ph. D | Stanford University | H 3 A Microbiome Workshop University of Witwatersrand | March 29 – 31, 2017 Image courtesy of Fiona Tamburini

Outline • Garbage in, garbage out (Quality filtering, etc) • What is a k-mer • Sequence Taxonomy • k-mer based • marker gene based • Sequence longer sequences/contigs (Assembly) • Gene/ORF prediction from short and long sequences • Gene annotation 3/31/17
TGACTCCTGAATCCCGAGCTTATGCCAAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCTTATGCCAAATCT GTCCCGATCATTGACTCCTAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCTTATG CCAAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCTTATGCCAAATCTGTCCCGAT CATTGACTCCGAATCCCGAGCTTATGCCACCGATCATTGACTCCTAATCTGTCCCGATC ATTGACTCCTGAATCCCGAGCTTATGCCAAATCTGTCCCGATCATTGACTCCTGAATCC CGAGCTTATGCCAAATCTGTCCCGATCATTGACTCCGAATCCCGAGCTTATGCCACTCC TGAATCCCGAGCTTATGCCAAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCCCTG AATCCCGAGCTTATGCCAAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCTAATCT GTCCCGATCATTGACTCCTGAATCCCGAGCTTATGCCAAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCTT ATGCCAAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCTTATGCCAAATCTGTCCC GATCATTGACTCCTAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCTTATGCCAAATCTGTCCCGATCATTGA CTCCGAATCCCGAGCTTATGCCACCGATCATTGACTCCTAATCTGTCCCGATCATTGAC TCCTGAATCCCGAGCTTATGCCAAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCT TATGCCAAATCTGTCCCGATCATTGACTCCGAATCCCGAGCTTATGCCACTCCTGAATC CCGAGCTTATGCCAAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCCCTGAATCCC GAGCTTATGCCAAATCTGTCCCGATCATTGACTCCTGAATCCCGAGCTAATCTGTCCCG

Pick your sequencing technique 16 S sequencing – Gross taxonomic classification

Pick your sequencing technique 16 S sequencing – Gross taxonomic classification Metagenomic sequencing and marker gene analysis – Higher resolution taxonomic classification

Pick your sequencing technique 16 S sequencing – Gross taxonomic classification Metagenomic sequencing and marker gene analysis – Higher resolution taxonomic classification Metagenomic sequencing and full WGS analysis – Species/strain level classification, non-bacterial data, pathways

What is a k-mer? ATTTGCCGGTCTTTCCTGTCCGCAGTATATGTCTCCGGATTTTATGGTG T ATTTGCC, TTTGCCG, CTTTCCT are all k-mers located within the above sequence, where k = 7 (the number of bases) We use k-mers for CLASSIFICATION (taxonomic, functional) and ASSEMBLY 3/31/17
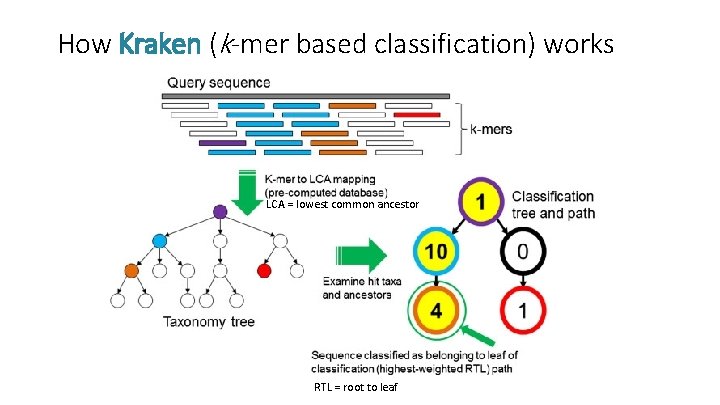
How Kraken (k-mer based classification) works LCA = lowest common ancestor RTL = root to leaf

Why not just use BLASTn? • Large tradeoff between SPEED and ACCURACY • Alignment with BLAST is slow • But the memory footprint for the reference database is pretty small • Kraken is FAST and fairly ACCURATE • But the memory footprint for the reference database is LARGE

3/31/17
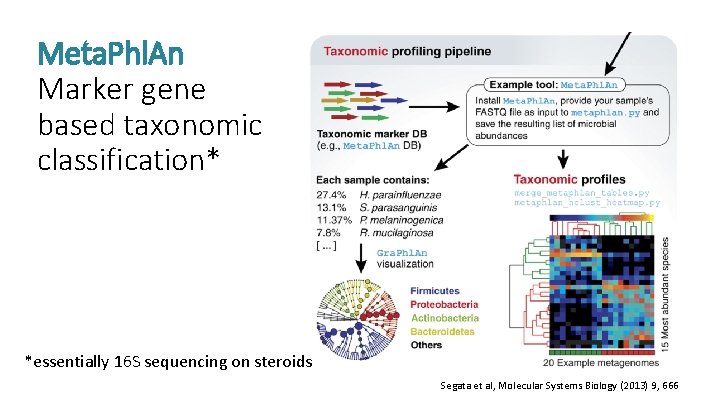
Meta. Phl. An Marker gene based taxonomic classification* *essentially 16 S sequencing on steroids Segata et al, Molecular Systems Biology (2013) 9, 666
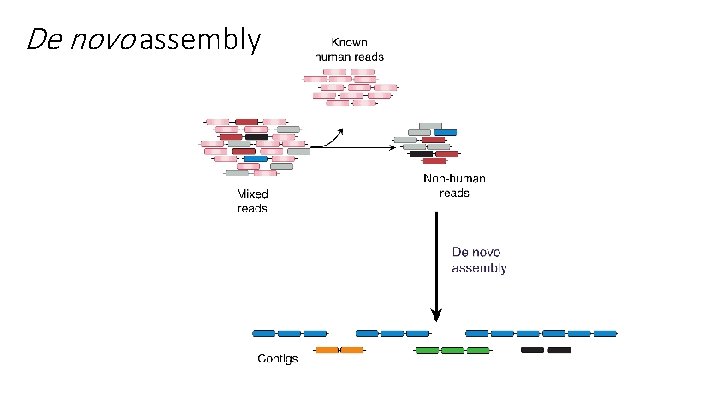
De novo assembly
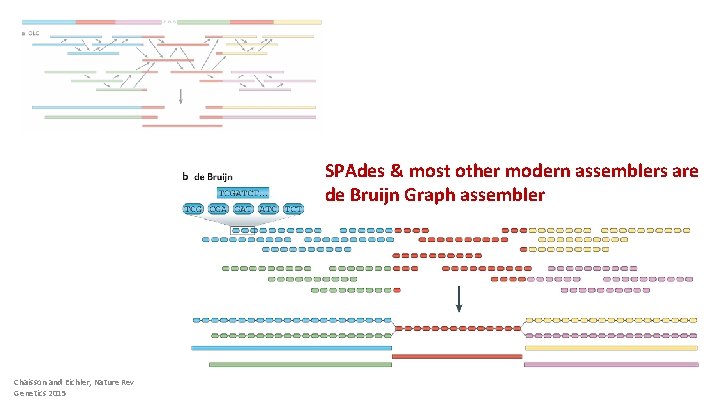
SPAdes & most other modern assemblers are de Bruijn Graph assembler Chaisson and Eichler, Nature Rev Genetics 2015
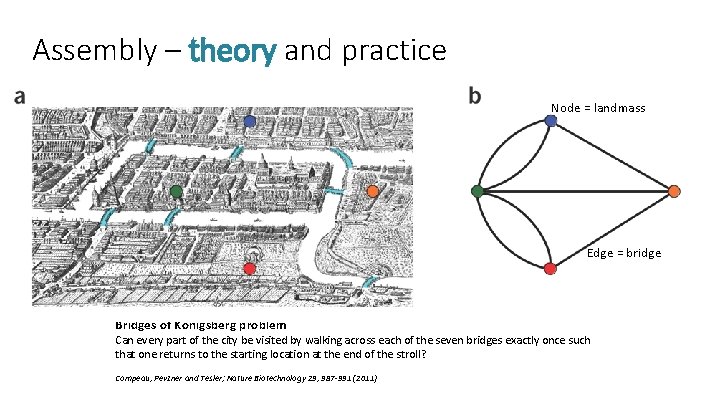
Assembly – theory and practice Node = landmass Edge = bridge Bridges of Königsberg problem Can every part of the city be visited by walking across each of the seven bridges exactly once such that one returns to the starting location at the end of the stroll? Compeau, Pevzner and Tesler; Nature Biotechnology 29, 987 -991 (2011)
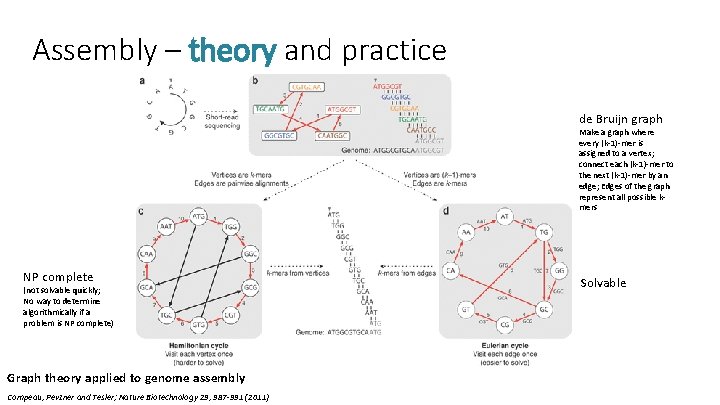
Assembly – theory and practice de Bruijn graph Make a graph where every (k-1)-mer is assigned to a vertex; connect each (k-1)-mer to the next (k-1)-mer by an edge; Edges of the graph represent all possible kmers NP complete (not solvable quickly; No way to determine algorithmically if a problem is NP complete) Graph theory applied to genome assembly Compeau, Pevzner and Tesler; Nature Biotechnology 29, 987 -991 (2011) Solvable

Why bother assembling metagenomic data? • Sequence length accuracy of taxonomic classification • Easier to identify full open reading frames for functional predictions • Identify operon structure (related genes located next to one another) • More accurate identification of genomic variations (structural and single nucleotide polymorphisms) 3/31/17
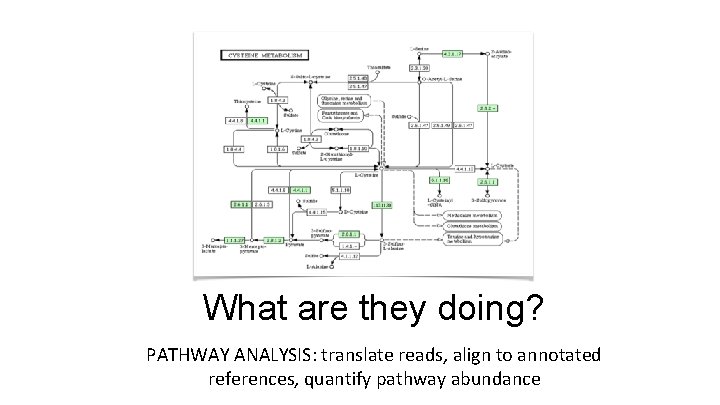
What are they doing? PATHWAY ANALYSIS: translate reads, align to annotated references, quantify pathway abundance
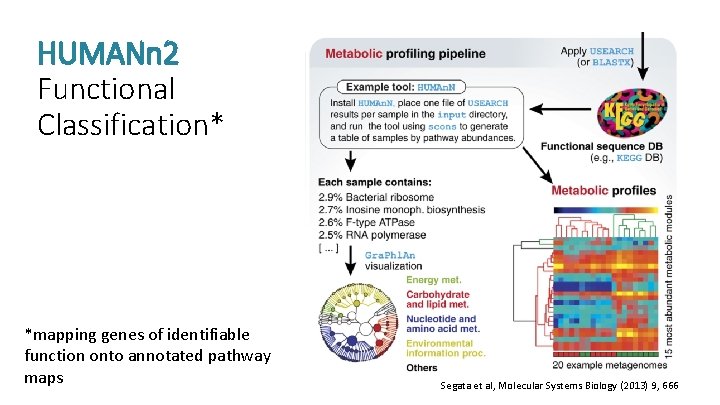
HUMANn 2 Functional Classification* *mapping genes of identifiable function onto annotated pathway maps Segata et al, Molecular Systems Biology (2013) 9, 666
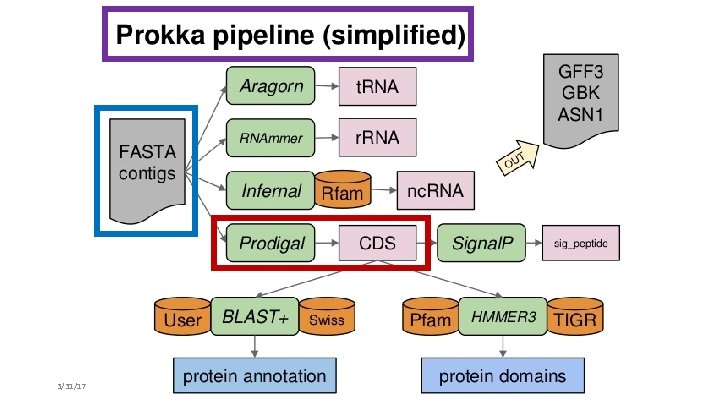
3/31/17
Thank you! bhattlab. com | asbhatt@stanford. edu
- Slides: 20