Overview of JGI Assembly Alex Copeland Pacific Biosciences

Overview of JGI Assembly Alex Copeland Pacific Biosciences West Coast User Group Meeting 2013 -09 -18 1

Joint Genome Institute 9: 15 – 9: 35 am Overview and Recent Developments Using Pac. Bio 9: 35 – 9: 55 am Large Scale Methylation Study Matthew Blow 9/18/2020 2
Outline Overview • JGI sequencing portfolio and platforms • Programmatic Sequencing goals Microbes • Recent assembly improvements • HGAP results / update Fungi • Genome Improvement / PBJ • HGAP results Latest developments • Metagenomes • 20 kb libraries 3

DOE Joint Genome Institute • Walnut Creek, CA • ~250 employees • $65 M annual budget • Hi. Seq (8), Mi. Seq (5), Pac. Bio (2) Mission: Serving as a genomic user facility in support of the DOE missions in: Bioenergy Carbon cycling Biogeochemistry
Eukaryotic Super Program projects JGI Sequencing JGI Project types: Transcriptome sequencing Plants Fungi Resequencing Prokaryotic Super Program Microbes Metagenomes De-Novo Genome Sequencing Assembly is essential to most JGI Genome projects

Why Sequence ? Plants: • improved drought and pest resistance • conversion to biofuels Fungi: • understanding of taxonomic diversity and relationships • unique enzymatic capabilities Microbes: • discover and catalog phylogenetic diversity • understand plant-microbe interactions Metagenomics: 9/18/2020 • improve understanding of microbial communities • gene- and genome-level comparisons of samples or environments 6

Why Assemble ? Provide context to enable understanding of structure in data Metagenomes: assembly supports understanding community structure and metabolic capabilities. Fungi: reference genomes for understanding taxonomic relationships, and identification of genes involved in carbon cycling. Microbes: reference genomes for clarifying and expanding taxonomic understanding of microbial life. 9/18/2020 7
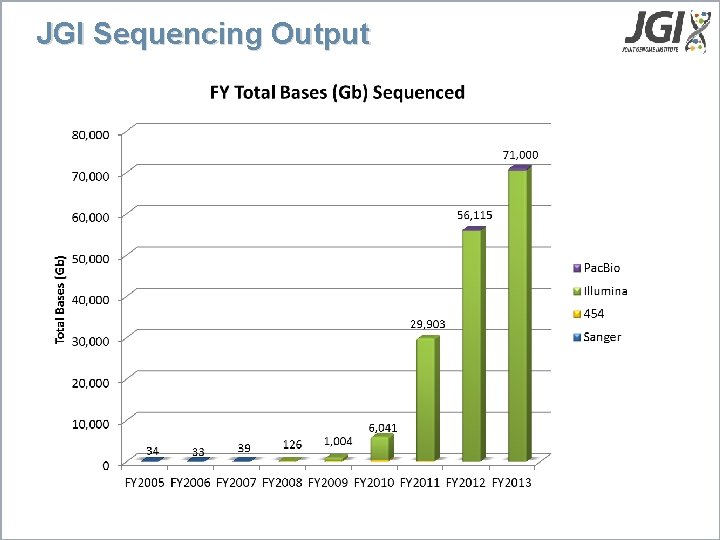
JGI Sequencing Output
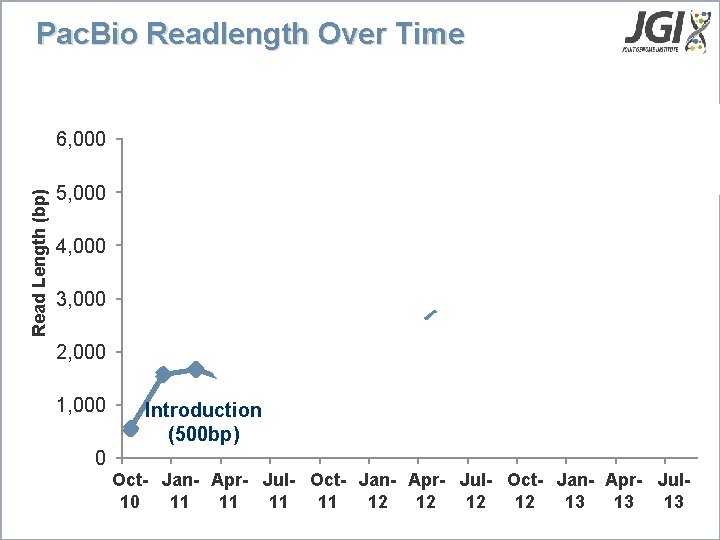
Pac. Bio Readlength Over Time Read Length (bp) 6, 000 XL Upgrade (>4, 000 bp) 5, 000 4, 000 3, 000 First Upgrades (1, 500 bp) P 4 Upgrade (>4, 500 bp) C 2 Upgrade (3, 000 bp) 2, 000 1, 000 Introduction (500 bp) 0 Oct- Jan- Apr- Jul- Oct- Jan- Apr- Jul 10 11 11 12 12 13 13 13
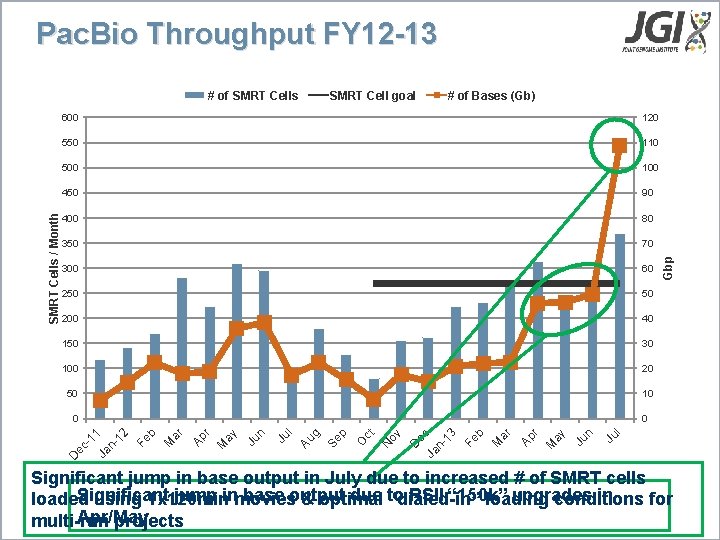
Pac. Bio Throughput FY 12 -13 # of Bases (Gb) 350 70 300 60 250 50 200 40 150 30 100 20 50 10 0 0 D Ju D N O Ju l 80 e Ja c n 13 Fe b M ar Ap r M ay Ju n 400 ov 90 ct 450 Se p 100 Au g 500 l 110 Fe b M ar Ap r M ay Ju n 550 -1 1 Ja n 12 120 Gbp SMRT Cell goal 600 ec SMRT Cells / Month # of SMRT Cells Significant jump in base output in July due to increased # of SMRT cells Significant jump inmovies base output due to RSII “ 150 k” upgrades in loaded using 1 x 120 min & optimal “dialed-in” loading conditions for Apr/May multi-run projects

Sequencing / Assembly Goals Year 9/18/2020 Standard Improved Metagenomes 2014 350 250 Fungi 2014 100 Microbes 2014 1000 150 11
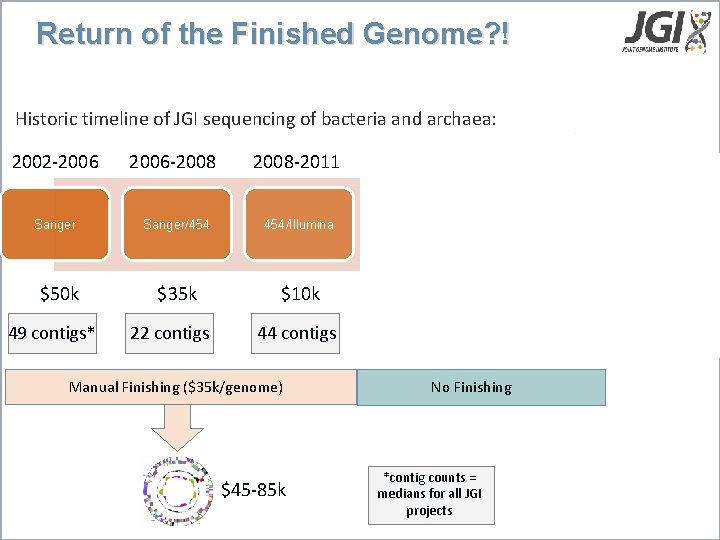
Return of the Finished Genome? ! Historic timeline of JGI sequencing of bacteria and archaea: 2002 -2006 Sanger $50 k 49 contigs* 2006 -2008 -2011 -2013 Pac. Bio Sanger/454 454/Illumina/Pac. Bio $35 k $10 k $1. 5 -3 k $5 k <$2 k 44 contigs 69 contigs 6 contigs 1 contig 22 contigs Manual Finishing ($35 k/genome) $45 -85 k No Finishing *contig counts = medians for all JGI projects
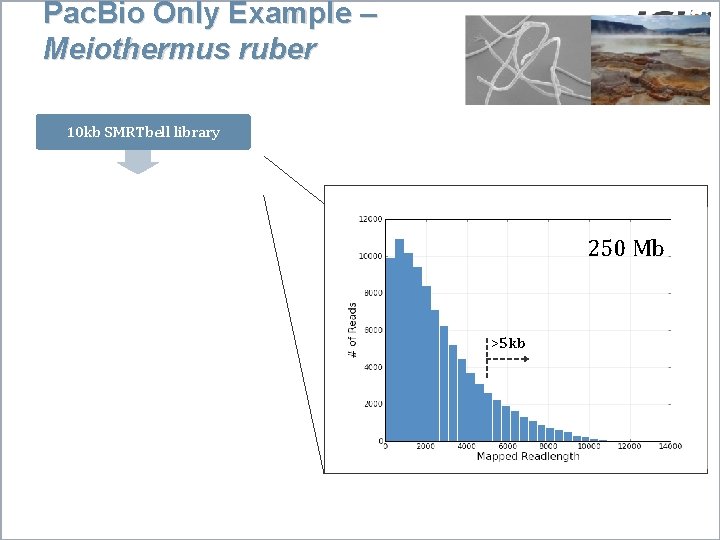
Pac. Bio Only Example – Meiothermus ruber 10 kb SMRTbell library 3 SMRT Cells 250 Mb Long seed reads (>5 kb) pre-assembly Pre-assembled long reads Celera Assembler 5 contigs Polish, Quiver 1 contig >5 kb
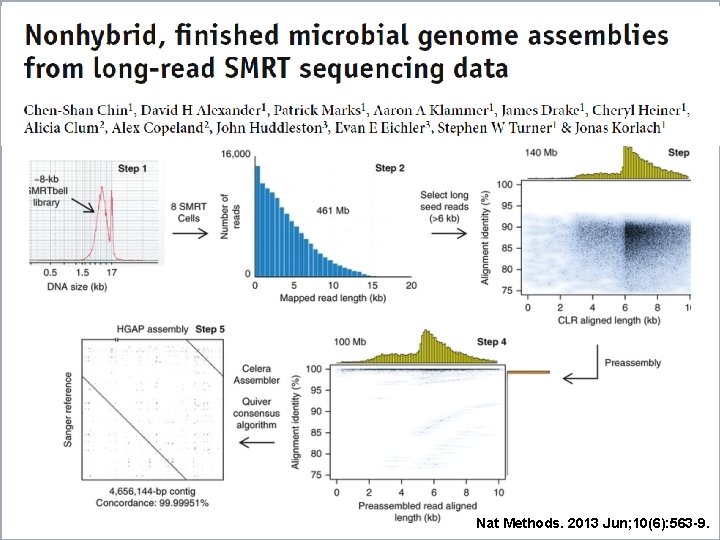
Nat Methods. 2013 Jun; 10(6): 563 -9.
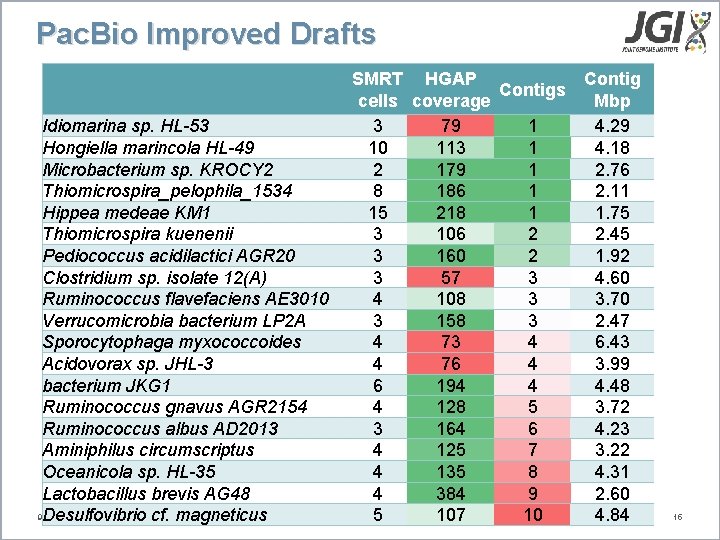
Pac. Bio Improved Drafts Idiomarina sp. HL-53 Hongiella marincola HL-49 Microbacterium sp. KROCY 2 Thiomicrospira_pelophila_1534 Hippea medeae KM 1 Thiomicrospira kuenenii Pediococcus acidilactici AGR 20 Clostridium sp. isolate 12(A) Ruminococcus flavefaciens AE 3010 Verrucomicrobia bacterium LP 2 A Sporocytophaga myxococcoides Acidovorax sp. JHL-3 bacterium JKG 1 Ruminococcus gnavus AGR 2154 Ruminococcus albus AD 2013 Aminiphilus circumscriptus Oceanicola sp. HL-35 Lactobacillus brevis AG 48 9/18/2020 Desulfovibrio cf. magneticus SMRT HGAP Contigs cells coverage Mbp 3 79 1 4. 29 10 113 1 4. 18 2 179 1 2. 76 8 186 1 2. 11 15 218 1 1. 75 3 106 2 2. 45 3 160 2 1. 92 3 57 3 4. 60 4 108 3 3. 70 3 158 3 2. 47 4 73 4 6. 43 4 76 4 3. 99 6 194 4 4. 48 4 128 5 3. 72 3 164 6 4. 23 4 125 7 3. 22 4 135 8 4. 31 4 384 9 2. 60 5 107 10 4. 84 15
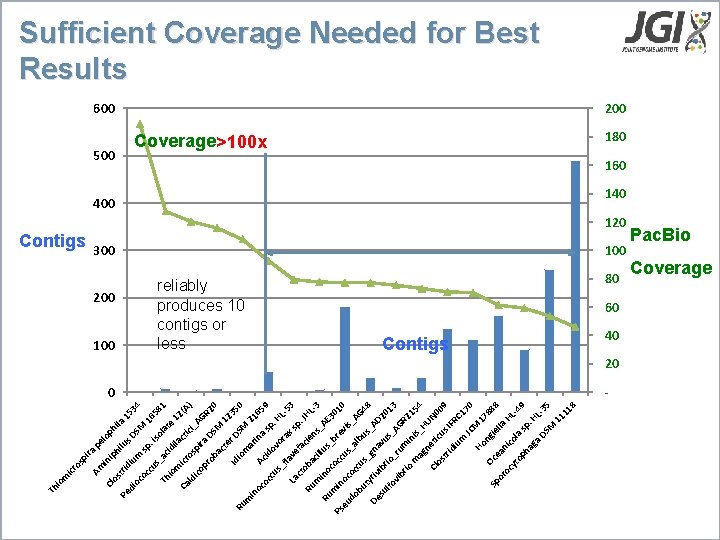
pe lo tri ph ip hi i l Pe us la 1 di 53 di DS um oc 4 M oc sp 16 cu. i 58 so s 1 la Th _ac te id io i 12 m l icr act Ca ici (A) os ld _A ico p pr ira D GR 20 ob ac SM 1 te r D 235 Id 0 Ru io SM m m 2 a in 16 rin oc Ac 59 a oc i sp do cu. vo HL s_ ra fla -5 x 3 ve La ct fa sp. JH cie Ru oba Lns cil 3 m _ l i AE us no Ru _ 3 co br 01 Ps min cc 0 us evis oc eu _a oc _A do lb cu bu us G 48 s_ ty g ri De na _AD vu su vibr 20 lfo io s_ 13 _ A vi ru G br m R 2 io in m is_ 154 ag ne HU N 0 Cl os ticu 09 tri s. I FR di um C 1 70 JC Ho M 1 78 ng Oc 88 Sp ie e l or an la oc HL yt icol -4 a op 9 sp ha. H ga DS L-35 M 11 11 8 in ira Contigs Cl os Am os p icr m Th io Sufficient Coverage Needed for Best Results 600 500 200 100 200 Coverage>100 x reliably produces 10 contigs or less 180 160 400 140 120 300 100 80 60 Contigs 0 40 20 - Pac. Bio Coverage

Fungi: Genome Improvement Assembly improvement consists of gap closing and error correction Needed because even the best assemblies contain gaps and imperfections due to • • Genome complexity Sequencing biases Repeats and polymorphism Data and algorithm limitations Traditional finishing methods too costly/time consuming
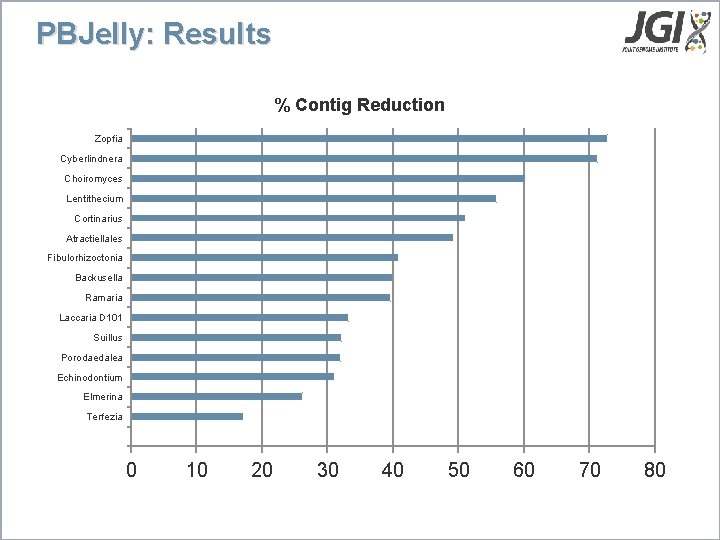
PBJelly: Results % Contig Reduction Zopfia Cyberlindnera Choiromyces Lentithecium Cortinarius Atractiellales Fibulorhizoctonia Backusella Ramaria Laccaria D 101 Suillus Porodaedalea Echinodontium Elmerina Terfezia 0 10 20 30 40 50 60 70 80
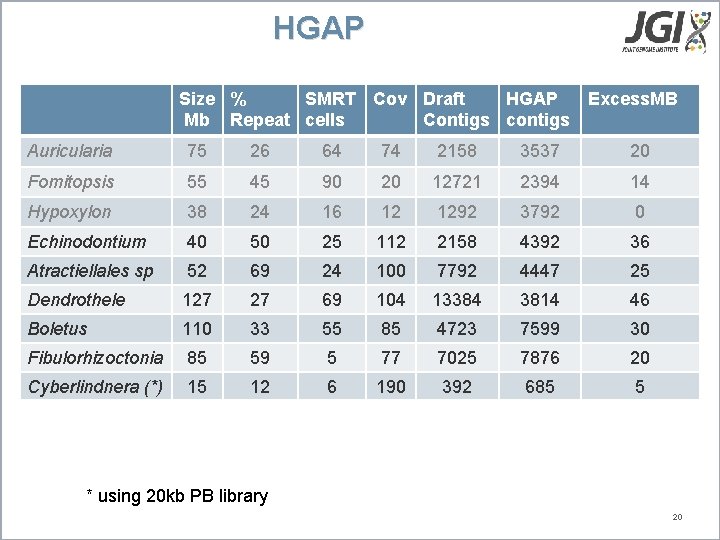
HGAP Size % SMRT Cov Draft HGAP Excess. MB Mb Repeat cells Contigs contigs Auricularia 75 26 64 74 2158 3537 20 Fomitopsis 55 45 90 20 12721 2394 14 Hypoxylon 38 24 16 12 1292 3792 0 Echinodontium 40 50 25 112 2158 4392 36 Atractiellales sp 52 69 24 100 7792 4447 25 Dendrothele 127 27 69 104 13384 3814 46 Boletus 110 33 55 85 4723 7599 30 Fibulorhizoctonia 85 59 5 77 7025 7876 20 Cyberlindnera (*) 15 12 6 190 392 685 5 * using 20 kb PB library 20
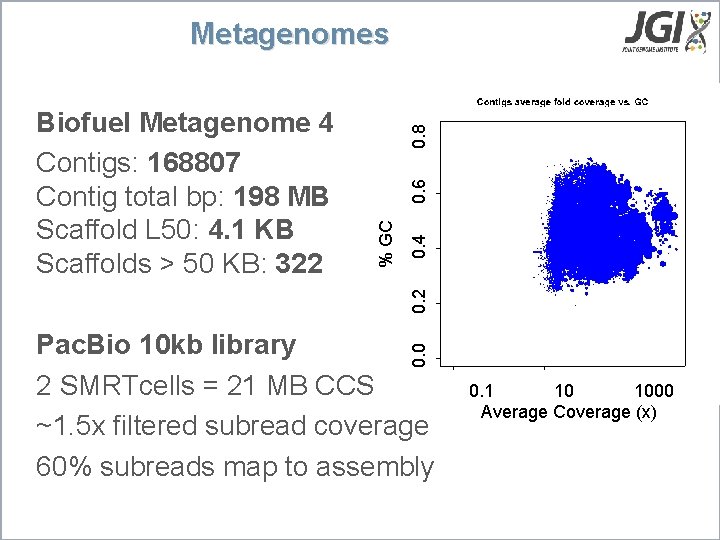
0. 6 0. 4 0. 2 % GC Biofuel Metagenome 4 Contigs: 168807 Contig total bp: 198 MB Scaffold L 50: 4. 1 KB Scaffolds > 50 KB: 322 0. 8 Metagenomes 0. 0 Pac. Bio 10 kb library 2 SMRTcells = 21 MB CCS ~1. 5 x filtered subread coverage 60% subreads map to assembly 0. 1 10 1000 Average Coverage (x)
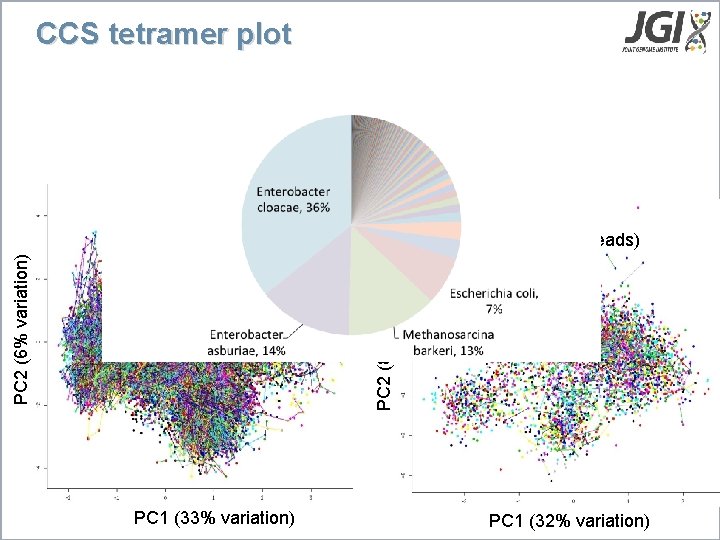
CCS tetramer plot PCA (Illumina contigs) PC 2 (8% variation) PC 2 (6% variation) PCA (CCS reads) PC 1 (33% variation) PC 1 (32% variation)
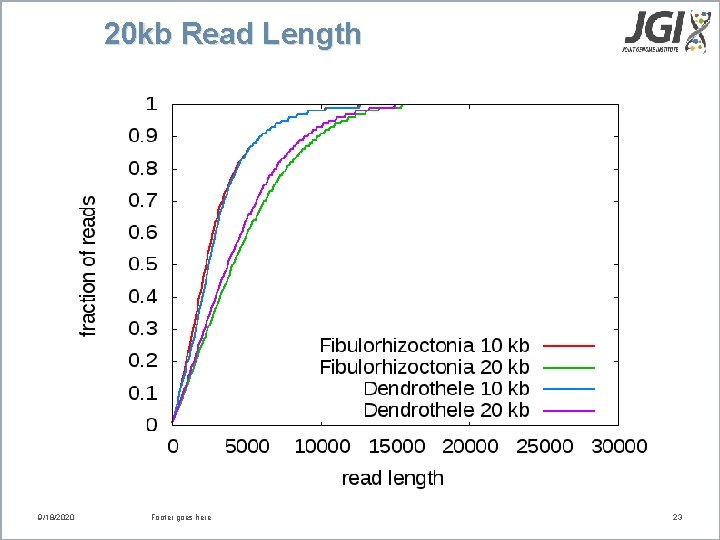
20 kb Read Length 9/18/2020 Footer goes here 23

Acknowledgements JGI Len Pennacchio, Tanja Woyke Genome Assembly Group: Alicia Clum, Kurt Labutti, Hui Sun Platforms: Chris Daum, Katy Munson QAQC: James Han, Alex Spunde, Stephan Trong, Kecia Duffy Pacific Biosciences Luke Hickey, Jason Chin 24
- Slides: 23