Overexpression and purification of recombinant murine wild type
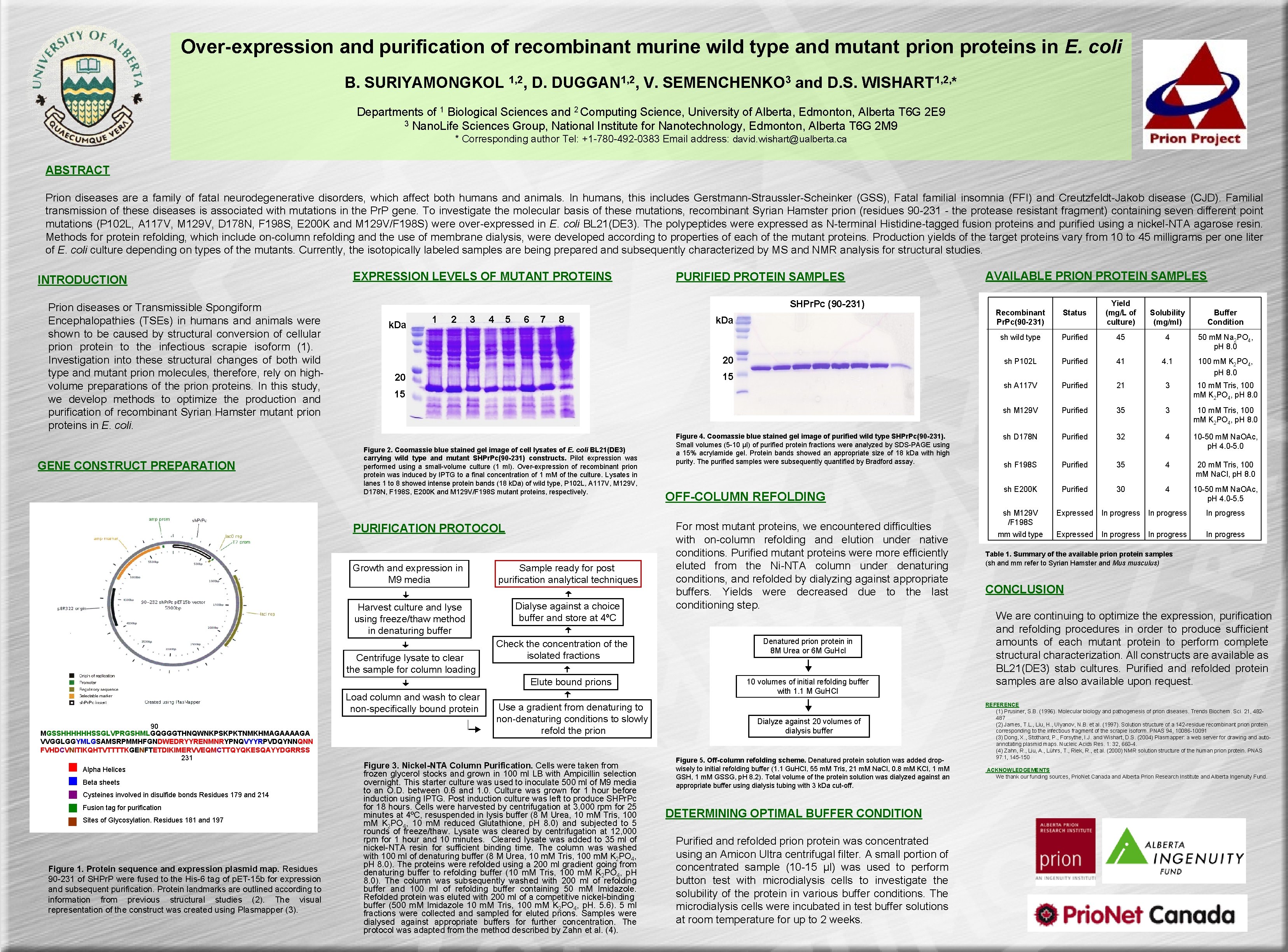
Over-expression and purification of recombinant murine wild type and mutant prion proteins in E. coli B. SURIYAMONGKOL 1, 2, D. DUGGAN 1, 2, V. SEMENCHENKO 3 and D. S. WISHART 1, 2, * Departments of 1 Biological Sciences and 2 Computing Science, University of Alberta, Edmonton, Alberta T 6 G 2 E 9 3 Nano. Life Sciences Group, National Institute for Nanotechnology, Edmonton, Alberta T 6 G 2 M 9 * Corresponding author Tel: +1 -780 -492 -0383 Email address: david. wishart@ualberta. ca ABSTRACT Prion diseases are a family of fatal neurodegenerative disorders, which affect both humans and animals. In humans, this includes Gerstmann-Straussler-Scheinker (GSS), Fatal familial insomnia (FFI) and Creutzfeldt-Jakob disease (CJD). Familial transmission of these diseases is associated with mutations in the Pr. P gene. To investigate the molecular basis of these mutations, recombinant Syrian Hamster prion (residues 90 -231 - the protease resistant fragment) containing seven different point mutations (P 102 L, A 117 V, M 129 V, D 178 N, F 198 S, E 200 K and M 129 V/F 198 S) were over-expressed in E. coli BL 21(DE 3). The polypeptides were expressed as N-terminal Histidine-tagged fusion proteins and purified using a nickel-NTA agarose resin. Methods for protein refolding, which include on-column refolding and the use of membrane dialysis, were developed according to properties of each of the mutant proteins. Production yields of the target proteins vary from 10 to 45 milligrams per one liter of E. coli culture depending on types of the mutants. Currently, the isotopically labeled samples are being prepared and subsequently characterized by MS and NMR analysis for structural studies. INTRODUCTION Prion diseases or Transmissible Spongiform Encephalopathies (TSEs) in humans and animals were shown to be caused by structural conversion of cellular prion protein to the infectious scrapie isoform (1). Investigation into these structural changes of both wild type and mutant prion molecules, therefore, rely on highvolume preparations of the prion proteins. In this study, we develop methods to optimize the production and purification of recombinant Syrian Hamster mutant prion proteins in E. coli. GENE CONSTRUCT PREPARATION EXPRESSION LEVELS OF MUTANT PROTEINS SHPr. Pc (90 -231) k. Da 1 2 3 4 5 6 7 8 15 20 15 Figure 2. Coomassie blue stained gel image of cell lysates of E. coli BL 21(DE 3) carrying wild type and mutant SHPr. Pc(90 -231) constructs. Pilot expression was performed using a small-volume culture (1 ml). Over-expression of recombinant prion protein was induced by IPTG to a final concentration of 1 m. M of the culture. Lysates in lanes 1 to 8 showed intense protein bands (18 k. Da) of wild type, P 102 L, A 117 V, M 129 V, D 178 N, F 198 S, E 200 K and M 129 V/F 198 S mutant proteins, respectively. Growth and expression in M 9 media Sample ready for post purification analytical techniques Harvest culture and lyse using freeze/thaw method in denaturing buffer Dialyse against a choice buffer and store at 4ºC Centrifuge lysate to clear the sample for column loading Check the concentration of the isolated fractions Elute bound prions Load column and wash to clear non-specifically bound protein Alpha Helices Beta sheets Cysteines involved in disulfide bonds Residues 179 and 214 Fusion tag for purification Sites of Glycosylation. Residues 181 and 197 Figure 1. Protein sequence and expression plasmid map. Residues 90 -231 of SHPr. P were fused to the His-6 tag of p. ET-15 b for expression and subsequent purification. Protein landmarks are outlined according to information from previous structural studies (2). The visual representation of the construct was created using Plasmapper (3). k. Da 20 PURIFICATION PROTOCOL 90 MGSSHHHHHHSSGLVPRGSHMLGQGGGTHNQWNKPSKPKTNMKHMAGAAAAGA VVGGLGGYMLGSAMSRPMMHFGNDWEDRYYRENMNRYPNQVYYRPVDQYNNQNN FVHDCVNITIKQHTVTTTTKGENFTETDIKIMERVVEQMCTTQYQKESQAYYDGRRSS 231 PURIFIED PROTEIN SAMPLES Use a gradient from denaturing to non-denaturing conditions to slowly refold the prion Figure 3. Nickel-NTA Column Purification. Cells were taken from frozen glycerol stocks and grown in 100 ml LB with Ampicillin selection overnight. This starter culture was used to inoculate 500 ml of M 9 media to an O. D. between 0. 6 and 1. 0. Culture was grown for 1 hour before induction using IPTG. Post induction culture was left to produce SHPr. Pc for 18 hours. Cells were harvested by centrifugation at 3, 000 rpm for 25 minutes at 4ºC, resuspended in lysis buffer (8 M Urea, 10 m. M Tris, 100 m. M K 2 PO 4, 10 m. M reduced Glutathione, p. H 8. 0) and subjected to 5 rounds of freeze/thaw. Lysate was cleared by centrifugation at 12, 000 rpm for 1 hour and 10 minutes. Cleared lysate was added to 35 ml of nickel-NTA resin for sufficient binding time. The column washed with 100 ml of denaturing buffer (8 M Urea, 10 m. M Tris, 100 m. M K 2 PO 4, p. H 8. 0). The proteins were refolded using a 200 ml gradient going from denaturing buffer to refolding buffer (10 m. M Tris, 100 m. M K 2 PO 4, p. H 8. 0). The column was subsequently washed with 200 ml of refolding buffer and 100 ml of refolding buffer containing 50 m. M Imidazole. Refolded protein was eluted with 200 ml of a competitive nickel-binding buffer (500 m. M Imidazole 10 m. M Tris, 100 m. M K 2 PO 4, p. H. 5. 6). 5 ml fractions were collected and sampled for eluted prions. Samples were dialysed against appropriate buffers for further concentration. The protocol was adapted from the method described by Zahn et al. (4). Figure 4. Coomassie blue stained gel image of purified wild type SHPr. Pc(90 -231). Small volumes (5 -10 µl) of purified protein fractions were analyzed by SDS-PAGE using a 15% acrylamide gel. Protein bands showed an appropriate size of 18 k. Da with high purity. The purified samples were subsequently quantified by Bradford assay. OFF-COLUMN REFOLDING For most mutant proteins, we encountered difficulties with on-column refolding and elution under native conditions. Purified mutant proteins were more efficiently eluted from the Ni-NTA column under denaturing conditions, and refolded by dialyzing against appropriate buffers. Yields were decreased due to the last conditioning step. Denatured prion protein in 8 M Urea or 6 M Gu. Hcl 10 volumes of initial refolding buffer with 1. 1 M Gu. HCl Dialyze against 20 volumes of dialysis buffer Figure 5. Off-column refolding scheme. Denatured protein solution was added dropwisely to initial refolding buffer (1. 1 Gu. HCl, 55 m. M Tris, 21 m. M Na. Cl, 0. 8 m. M KCl, 1 m. M GSH, 1 m. M GSSG, p. H 8. 2). Total volume of the protein solution was dialyzed against an appropriate buffer using dialysis tubing with 3 k. Da cut-off. DETERMINING OPTIMAL BUFFER CONDITION Purified and refolded prion protein was concentrated using an Amicon Ultra centrifugal filter. A small portion of concentrated sample (10 -15 µl) was used to perform button test with microdialysis cells to investigate the solubility of the protein in various buffer conditions. The microdialysis cells were incubated in test buffer solutions at room temperature for up to 2 weeks. AVAILABLE PRION PROTEIN SAMPLES Recombinant Pr. Pc(90 -231) Status Yield (mg/L of culture) sh wild type Purified 45 4 50 m. M Na 2 PO 4, p. H 8. 0 sh P 102 L Purified 41 4. 1 100 m. M K 2 PO 4, p. H 8. 0 sh A 117 V Purified 21 3 10 m. M Tris, 100 m. M K 2 PO 4, p. H 8. 0 sh M 129 V Purified 35 3 10 m. M Tris, 100 m. M K 2 PO 4, p. H 8. 0 sh D 178 N Purified 32 4 10 -50 m. M Na. OAc, p. H 4. 0 -5. 0 sh F 198 S Purified 35 4 20 m. M Tris, 100 m. M Na. Cl, p. H 8. 0 sh E 200 K Purified 30 4 10 -50 m. M Na. OAc, p. H 4. 0 -5. 5 sh M 129 V /F 198 S Expressed In progress mm wild type Expressed In progress Solubility (mg/ml) Buffer Condition Table 1. Summary of the available prion protein samples (sh and mm refer to Syrian Hamster and Mus musculus) CONCLUSION We are continuing to optimize the expression, purification and refolding procedures in order to produce sufficient amounts of each mutant protein to perform complete structural characterization. All constructs are available as BL 21(DE 3) stab cultures. Purified and refolded protein samples are also available upon request. REFERENCE (1) Prusiner, S. B. (1996). Molecular biology and pathogenesis of prion diseases. Trends Biochem. Sci. 21, 482487 (2) James, T. L. , Liu, H. , Ulyanov, N. B. et al. (1997). Solution structure of a 142 -residue recombinant prion protein corresponding to the infectious fragment of the scrapie isoform. PNAS 94, 10086 -10091 (3) Dong, X. , Stothard, P. , Forsythe, I. J. and Wishart, D. S. (2004) Plasmapper: a web server for drawing and autoannotating plasmid maps. Nucleic Acids Res. 1: 32, 660 -4. (4) Zahn, R. , Liu, A. , Lührs, T. , Riek, R. , et al. (2000) NMR solution structure of the human prion protein. PNAS 97: 1, 145 -150 ACKNOWLEDGEMENTS We thank our funding sources, Prio. Net Canada and Alberta Prion Research Institute and Alberta Ingenuity Fund.
- Slides: 1