Organization of DNA Andy Howard Introductory Biochemistry 10

Organization of DNA Andy Howard Introductory Biochemistry 10 October 2013 Nucleic Acid Structure II 10/10/2013
DNA structure informs its function n We will revisit several aspects of DNA structure that help us understand how it operates. 10/10/2013 Nucleic Acid Structure II p. 2 of 58
What we’ll discuss n n n DNA versus RNA DNA & RNA cleavage Restriction Enzymes Review of A, B, Z DNA Intercalation 10/10/2013 Nucleic Acid Structure II n n n Denaturation and renaturation of DNA density Tertiary structure Supercoiling n Gyrases n Nucleosomes n Higher levels n p. 3 of 58
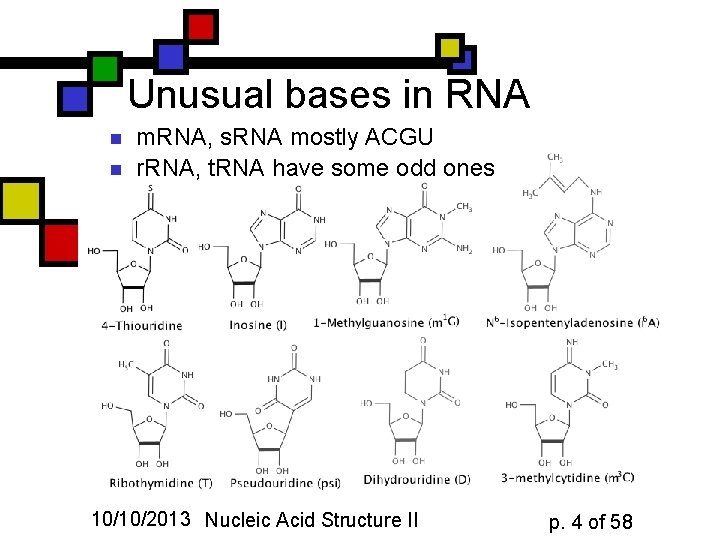
Unusual bases in RNA n n m. RNA, s. RNA mostly ACGU r. RNA, t. RNA have some odd ones 10/10/2013 Nucleic Acid Structure II p. 4 of 58
Other small RNAs n n n 21 -28 nucleotides Target RNA or DNA through complementary base-pairing Several types, based on function: n n n Small interfering RNAs (q. v. ) micro. RNA: control developmental timing Small nucleolar RNA: catalysts that (among other things) create the oddball bases sno. RNA 77 courtesy Wikipedia 10/10/2013 Nucleic Acid Structure II p. 5 of 58
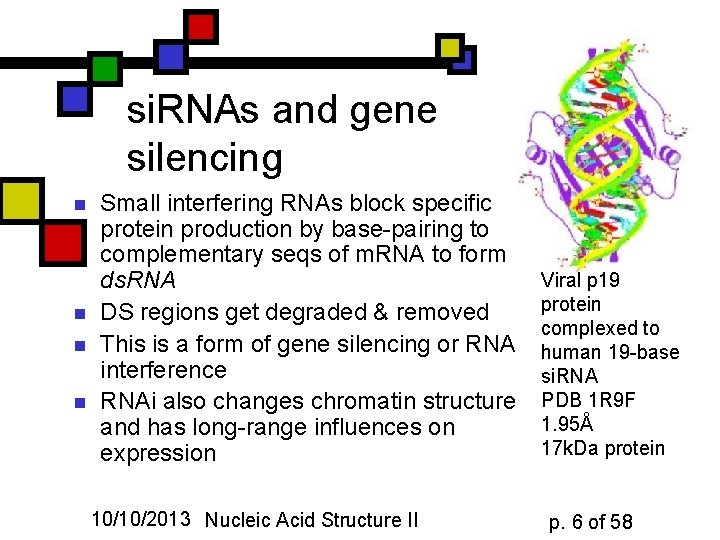
si. RNAs and gene silencing n n Small interfering RNAs block specific protein production by base-pairing to complementary seqs of m. RNA to form ds. RNA DS regions get degraded & removed This is a form of gene silencing or RNA interference RNAi also changes chromatin structure and has long-range influences on expression 10/10/2013 Nucleic Acid Structure II Viral p 19 protein complexed to human 19 -base si. RNA PDB 1 R 9 F 1. 95Å 17 k. Da protein p. 6 of 58

Do the differences between RNA and DNA matter? Yes! n DNA has deoxythymidine, RNA has uridine: n n n cytidine spontaneously degrades to uridine d. C spontaneously degrades to d. U The only d. U found in DNA is there because of degradation: d. T goes with d. A So when a cell finds d. U in its DNA, it knows it should replace it with d. C or else synthesize d. G opposite the d. U instead of d. A Deoxy sugars don’t degrade as readily! … see next slide 10/10/2013 Nucleic Acid Structure II p. 7 of 58
Ribose vs. deoxyribose n n n Presence of -OH on 2’ position makes the 3’ position in RNA more susceptible to nonenzymatic cleavage than the 3’ in DNA The ribose vs. deoxyribose distinction also influences enzymatic degradation of nucleic acids I can carry DNA in my shirt pocket, but not RNA 10/10/2013 Nucleic Acid Structure II p. 8 of 58
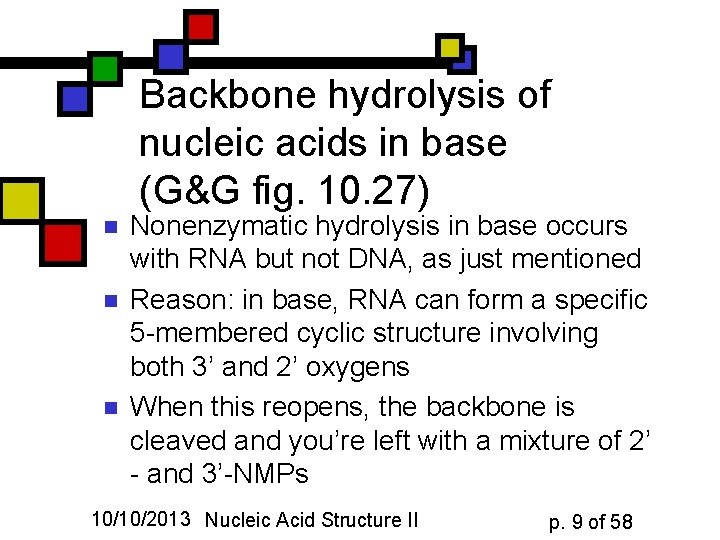
Backbone hydrolysis of nucleic acids in base (G&G fig. 10. 27) n n n Nonenzymatic hydrolysis in base occurs with RNA but not DNA, as just mentioned Reason: in base, RNA can form a specific 5 -membered cyclic structure involving both 3’ and 2’ oxygens When this reopens, the backbone is cleaved and you’re left with a mixture of 2’ - and 3’-NMPs 10/10/2013 Nucleic Acid Structure II p. 9 of 58
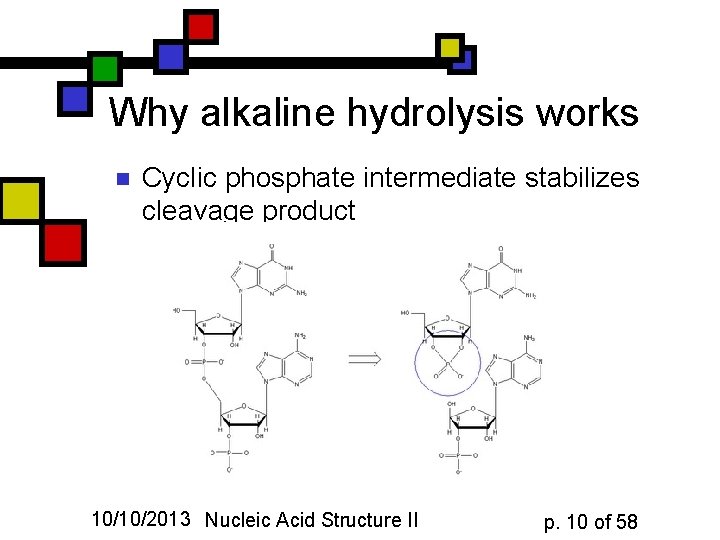
Why alkaline hydrolysis works n Cyclic phosphate intermediate stabilizes cleavage product 10/10/2013 Nucleic Acid Structure II p. 10 of 58
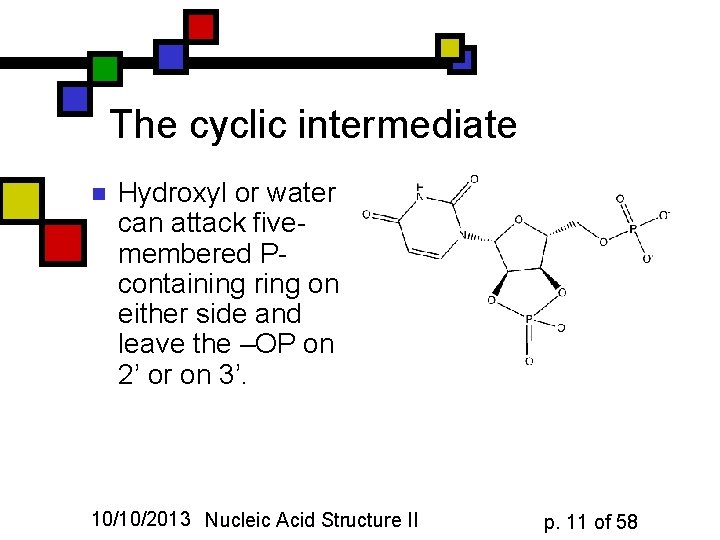
The cyclic intermediate n Hydroxyl or water can attack fivemembered Pcontaining ring on either side and leave the –OP on 2’ or on 3’. 10/10/2013 Nucleic Acid Structure II p. 11 of 58

Consequences n n n So RNA is considerably less stable compared to DNA, owing to the formation of this cyclic phosphate intermediate DNA can’t form this because it doesn’t have a 2’ hydroxyl In fact, deoxyribose has no free hydroxyls! 10/10/2013 Nucleic Acid Structure II p. 12 of 58
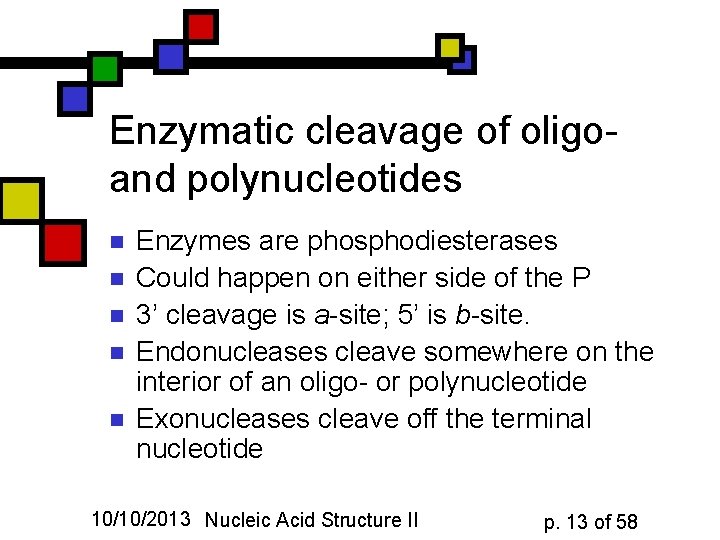
Enzymatic cleavage of oligoand polynucleotides n n n Enzymes are phosphodiesterases Could happen on either side of the P 3’ cleavage is a-site; 5’ is b-site. Endonucleases cleave somewhere on the interior of an oligo- or polynucleotide Exonucleases cleave off the terminal nucleotide 10/10/2013 Nucleic Acid Structure II p. 13 of 58
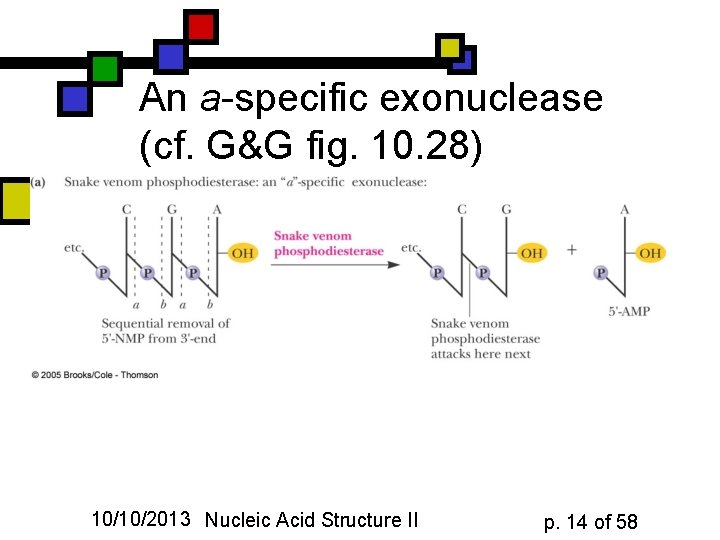
An a-specific exonuclease (cf. G&G fig. 10. 28) 10/10/2013 Nucleic Acid Structure II p. 14 of 58
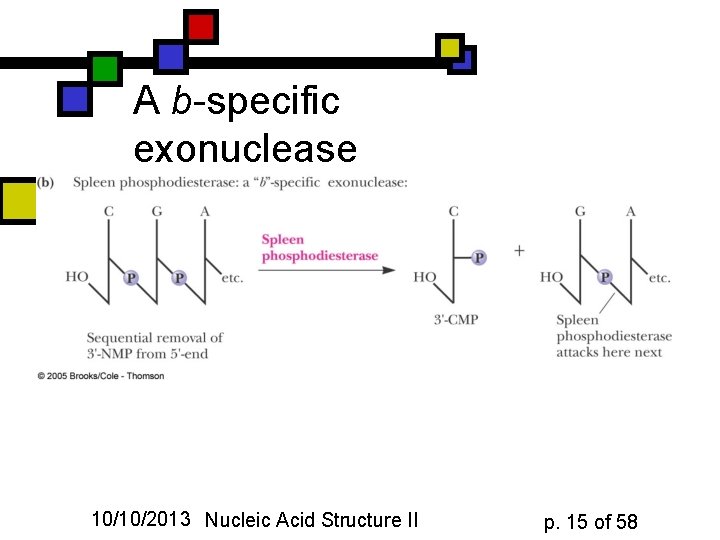
A b-specific exonuclease 10/10/2013 Nucleic Acid Structure II p. 15 of 58

Specificity in nucleases n n n Some cleave only RNA, others only DNA, some both Often a preference for a specific base or even a particular 4 -8 nucleotide sequence (restriction endonucleases) These can be used as lab tools, but they evolved for internal reasons 10/10/2013 Nucleic Acid Structure II p. 16 of 58
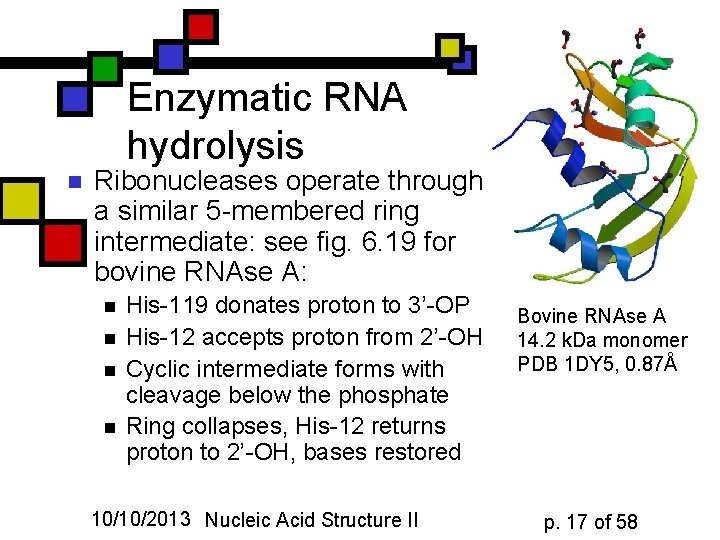
Enzymatic RNA hydrolysis n Ribonucleases operate through a similar 5 -membered ring intermediate: see fig. 6. 19 for bovine RNAse A: n n His-119 donates proton to 3’-OP His-12 accepts proton from 2’-OH Cyclic intermediate forms with cleavage below the phosphate Ring collapses, His-12 returns proton to 2’-OH, bases restored 10/10/2013 Nucleic Acid Structure II Bovine RNAse A 14. 2 k. Da monomer PDB 1 DY 5, 0. 87Å p. 17 of 58
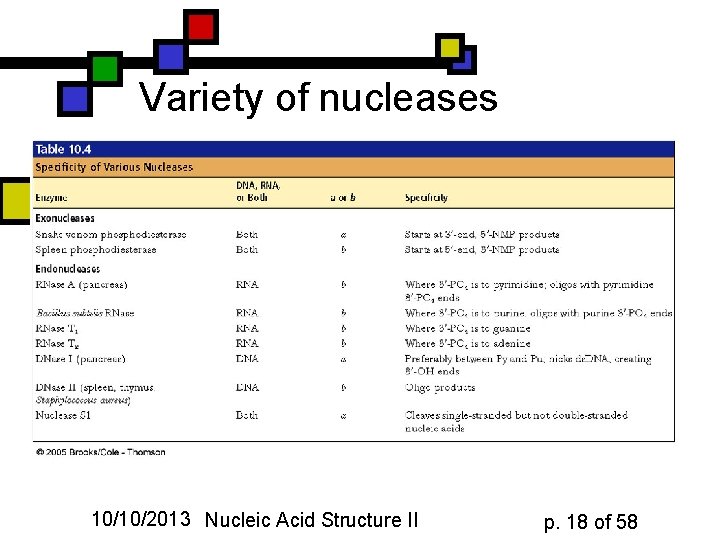
Variety of nucleases 10/10/2013 Nucleic Acid Structure II p. 18 of 58

Restriction Endonucleases (G&G § 10. 6) n n n Evolve in bacteria as antiviral tools “Restriction” because they restrict the incorporation of foreign DNA into the bacterial chromosome Recognize and bind to specific palindromic DNA sequences and cleave them Self-cleavage avoided by methylation Types I, III: II is most important I and III have inherent methylase activity; II has methylase activity in an attendant enzyme 10/10/2013 Nucleic Acid Structure II p. 19 of 58

What do we mean by palindromic? n In ordinary language, it means a phrase that reads the same forward and back: n n n Madam, I’m Adam. (Genesis 3: 20) Eve, man, am Eve. Sex at noon taxes. Able was I ere I saw Elba. (Napoleon) A man, a plan, a canal: Panama! (T. Roosevelt) With DNA it means the double-stranded sequence is identical on both strands 10/10/2013 Nucleic Acid Structure II p. 20 of 58

Palindromic DNA n n Example: G-A-A-T-T-C Single strand isn’t symmetric: but the combination with the complementary strand is: G-A-A-T-T-C C-T-T-A-A-G These kinds of sequences are the recognition sites for restriction endonucleases. This particular hexanucleotide is the recognition sequence for Eco. RI. 10/10/2013 Nucleic Acid Structure II p. 21 of 58

Cleavages by restriction endonucleases n Breaks can be n n n cohesive (if off-center within the sequence) or non-cohesive (blunt) (if they’re at the center) Eco. RI leaves staggered 5’-termini: cleaves between initial G and A Pst. I cleaves CTGCAG between A and G, so it leaves staggered 3’-termini Bal. I cleaves TGGCCA in the middle: blunt! 10/10/2013 Nucleic Acid Structure II p. 22 of 58
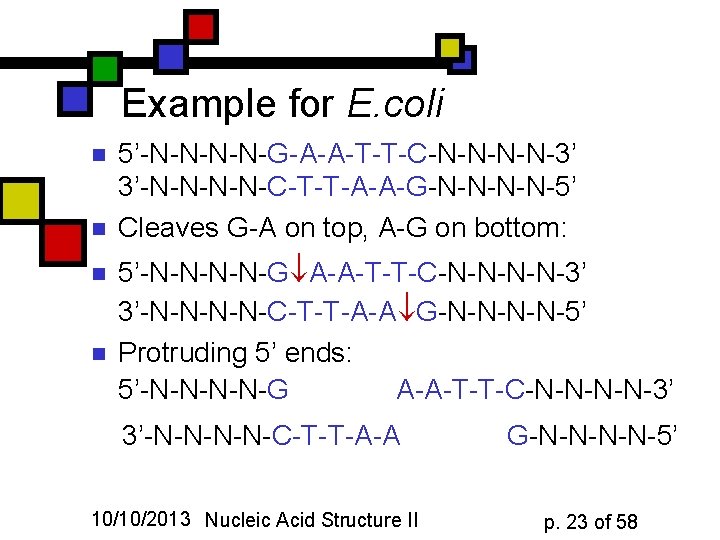
Example for E. coli n n 5’-N-N-G-A-A-T-T-C-N-N-3’ 3’-N-N-C-T-T-A-A-G-N-N-5’ Cleaves G-A on top, A-G on bottom: 5’-N-N-G A-A-T-T-C-N-N-3’ 3’-N-N-C-T-T-A-A G-N-N-5’ Protruding 5’ ends: 5’-N-N-G A-A-T-T-C-N-N-3’ 3’-N-N-C-T-T-A-A 10/10/2013 Nucleic Acid Structure II G-N-N-5’ p. 23 of 58
How often? n n n 4 types of bases So a recognition site that is 4 bases long will occur once every 44 = 256 bases on either strand, on average 6 -base site: every 46= 4096 bases, which is roughly one gene’s worth 10/10/2013 Nucleic Acid Structure II p. 24 of 58
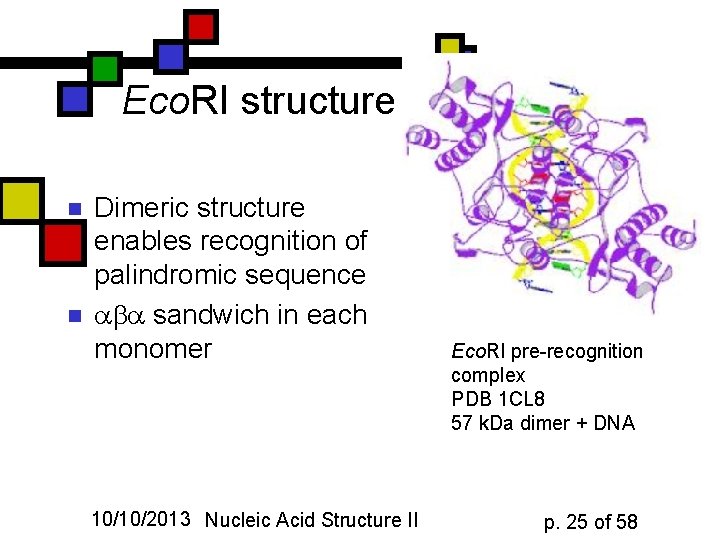
Eco. RI structure n n Dimeric structure enables recognition of palindromic sequence sandwich in each monomer 10/10/2013 Nucleic Acid Structure II Eco. RI pre-recognition complex PDB 1 CL 8 57 k. Da dimer + DNA p. 25 of 58
The biology problem n n n How does the bacterium mark its own DNA so that it does replicate its own DNA but not the foreign DNA? Answer: by methylating specific bases in its DNA prior to replication Unmethylated DNA from foreign source gets cleaved by restriction endonuclease Only the methylated DNA survives to be replicated Most methylations are of A & G, but sometimes C gets it too 10/10/2013 Nucleic Acid Structure II p. 26 of 58
How this works n n n When an unmethylated specific sequence appears in the DNA, the enzyme cleaves it When the corresponding methylated sequence appears, it doesn’t get cleaved and remains available for replication The restriction endonucleases only bind to palindromic sequences 10/10/2013 Nucleic Acid Structure II p. 27 of 58
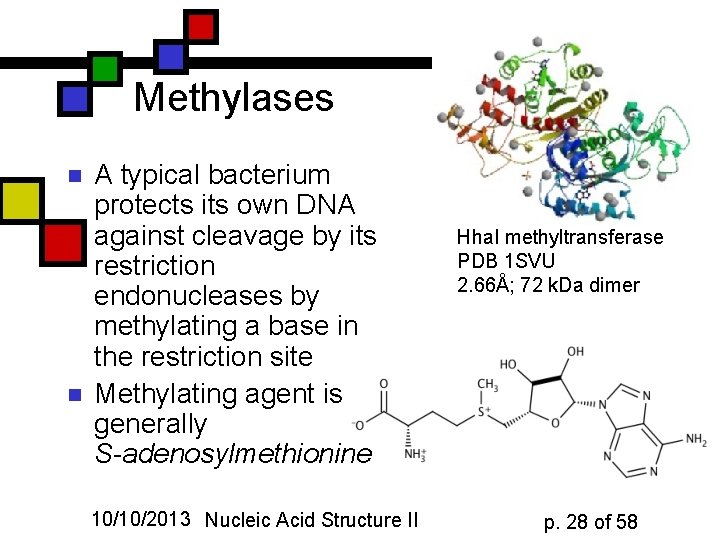
Methylases n n A typical bacterium protects its own DNA against cleavage by its restriction endonucleases by methylating a base in the restriction site Methylating agent is generally S-adenosylmethionine 10/10/2013 Nucleic Acid Structure II Hha. I methyltransferase PDB 1 SVU 2. 66Å; 72 k. Da dimer p. 28 of 58
Use of restriction enzymes n Nature made these to protect bacteria; we use them to cleave DNA in analyzable ways n n Similar to proteolytic digestion of proteins Having a variety of nucleases means we can get fragments in multiple ways We can amplify our DNA first Can also be used in synthesis of inserts that we can incorporate into plasmids that enable us to make appropriate DNA molecules in bacteria 10/10/2013 Nucleic Acid Structure II p. 29 of 58
Intercalating agents (G&G fig. 11. 14) n n Generally: aromatic compounds that can form -stack interactions with bases Bases must be forced apart to fit them in Results in an almost ladderlike structure for the sugar-phosphate backbone locally Conclusion: it must be easy to do local unwinding to get those in! 10/10/2013 Nucleic Acid Structure II p. 30 of 58
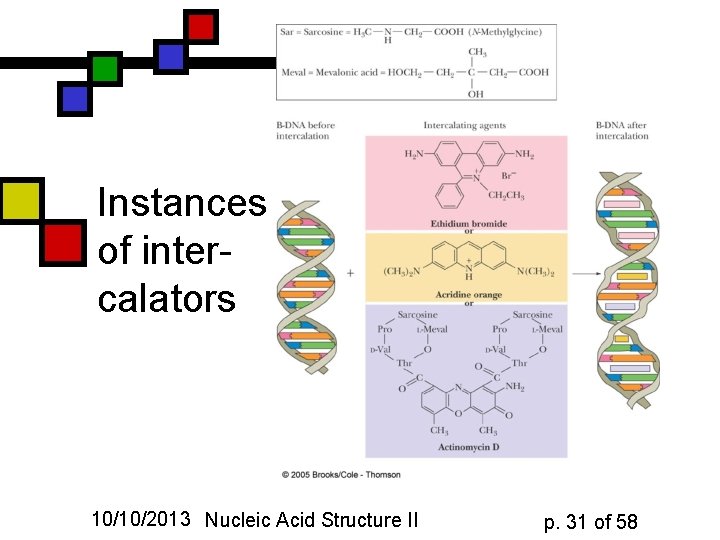
Instances of intercalators 10/10/2013 Nucleic Acid Structure II p. 31 of 58
Denaturing and Renaturing DNA (G&G § 11. 3) n n See Figure 11. 17 When DNA is heated to > 80ºC, its UV absorbance increases by 30 -40% This hyperchromic shift reflects the unwinding of the DNA double helix Stacked base pairs in native DNA absorb less light When T is lowered, the absorbance drops, reflecting the re-establishment of stacking 10/10/2013 Nucleic Acid Structure II p. 32 of 58
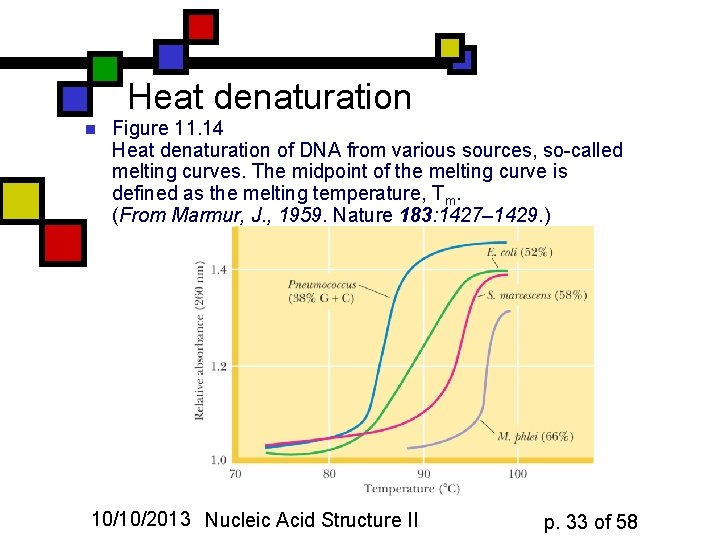
Heat denaturation n Figure 11. 14 Heat denaturation of DNA from various sources, so-called melting curves. The midpoint of the melting curve is defined as the melting temperature, Tm. (From Marmur, J. , 1959. Nature 183: 1427– 1429. ) 10/10/2013 Nucleic Acid Structure II p. 33 of 58
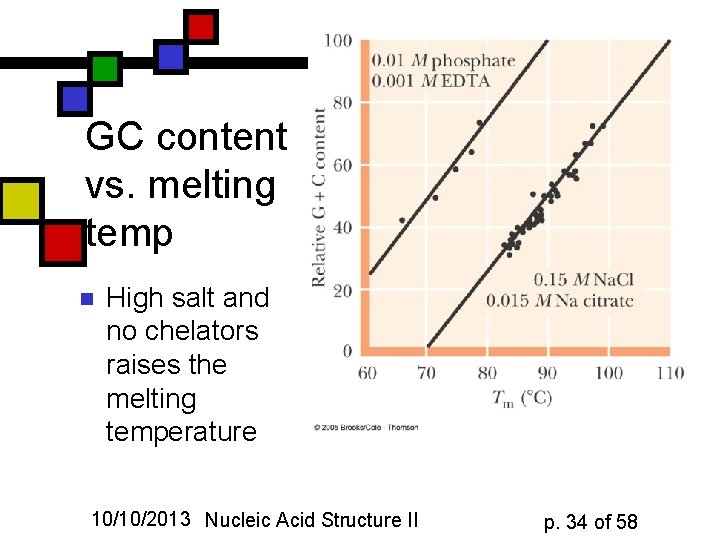
GC content vs. melting temp n High salt and no chelators raises the melting temperature 10/10/2013 Nucleic Acid Structure II p. 34 of 58
How else can we melt DNA? n n High p. H deprotonates the bases so the Hbonds disappear Low p. H hyper-protonates the bases so the H-bonds disappear Alkalai is better: it doesn’t break the glycosidic linkages Urea, formamide make better H-bonds than the DNA itself so they denature DNA 10/10/2013 Nucleic Acid Structure II p. 35 of 58
What happens if we separate the strands? n n We can renature the DNA into a double helix Requires re-association of 2 strands: reannealing The realignment can go wrong Association is 2 nd-order, zippering is first order and therefore faster 10/10/2013 Nucleic Acid Structure II p. 36 of 58
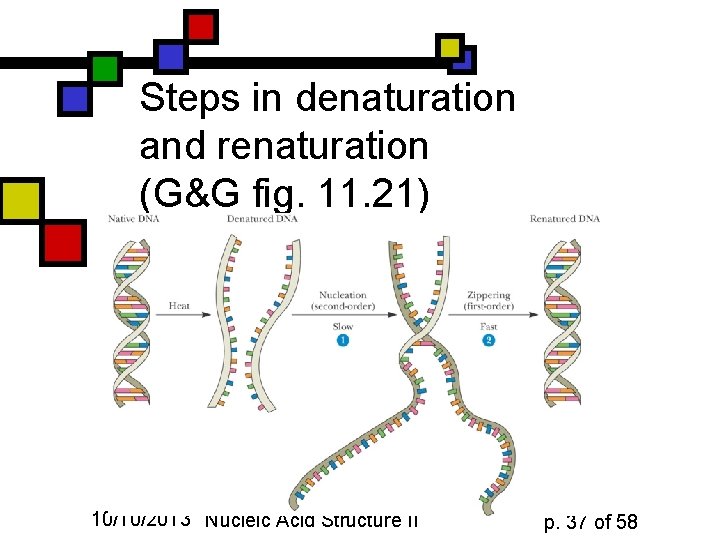
Steps in denaturation and renaturation (G&G fig. 11. 21) 10/10/2013 Nucleic Acid Structure II p. 37 of 58

Rate depends on complexity n n The more complex DNA is, the longer it takes for nucleation of renaturation to occur “Complex” can mean “large”, but complexity is influenced by sequence randomness: poly(AT) is faster than a random sequence 10/10/2013 Nucleic Acid Structure II p. 38 of 58
Second-order kinetics n n n Rate of association: -dc/dt = k 2 c 2 Boundary condition is fully denatured concentration c 0 at time t=0: c / c 0 = (1+k 2 c 0 t)-1 Half time is t 1/2 = (k 2 c 0)-1 Routine depiction: plot c 0 t vs. fraction reassociated (c /c 0) and find the halfway point. 10/10/2013 Nucleic Acid Structure II p. 39 of 58
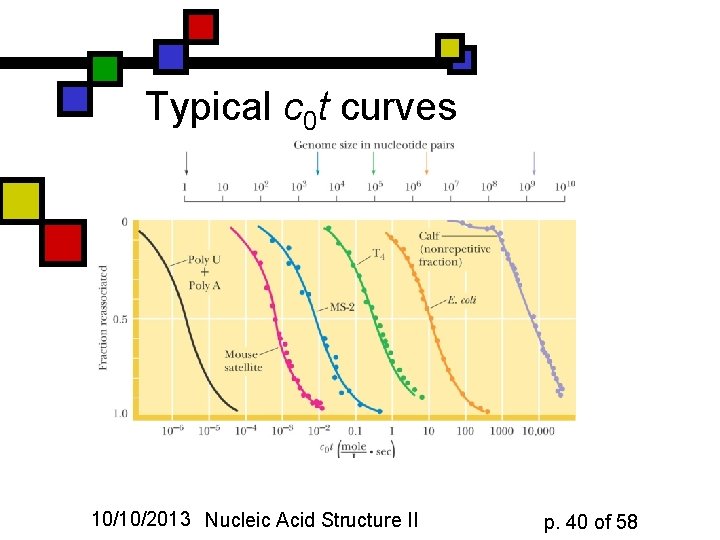
Typical c 0 t curves 10/10/2013 Nucleic Acid Structure II p. 40 of 58
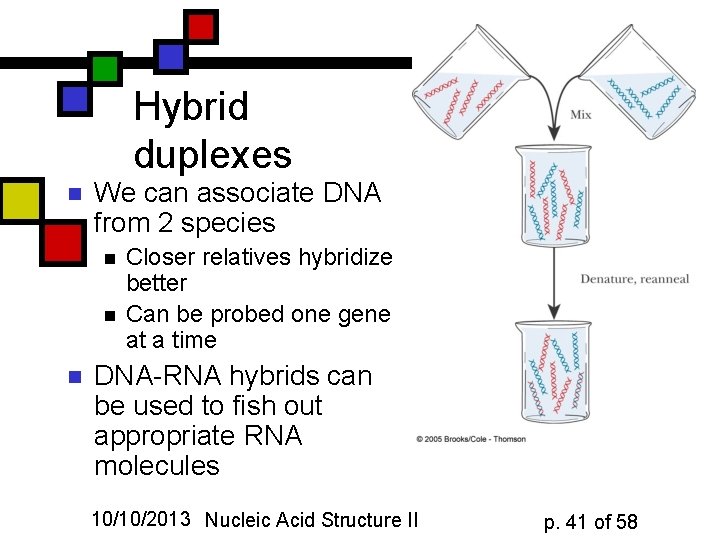
Hybrid duplexes n We can associate DNA from 2 species n n n Closer relatives hybridize better Can be probed one gene at a time DNA-RNA hybrids can be used to fish out appropriate RNA molecules 10/10/2013 Nucleic Acid Structure II p. 41 of 58
GC-rich DNA is denser n n DNA is denser than RNA or protein, period, because it can coil up so compactly Therefore density-gradient centrifugation separates DNA from other cellular macromolecules GC-rich DNA is 3% denser than AT-rich Can be used as a quick measure of GC content 10/10/2013 Nucleic Acid Structure II p. 42 of 58
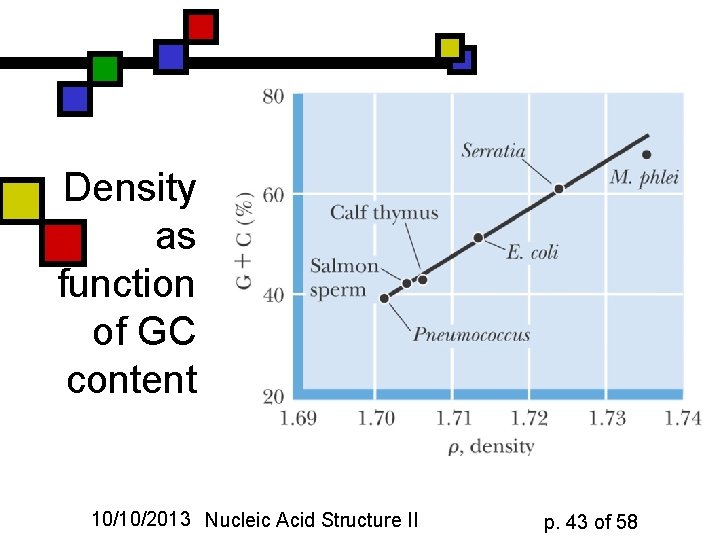
Density as function of GC content 10/10/2013 Nucleic Acid Structure II p. 43 of 58
Tertiary Structure of DNA n n n In duplex DNA, ten bp per turn of helix Circular DNA sometimes has more or less than 10 bp per turn - a supercoiled state Enzymes called topoisomerases or gyrases can introduce or remove supercoils Cruciforms occur in palindromic regions of DNA Negative supercoiling may promote cruciforms 10/10/2013 Nucleic Acid Structure II p. 44 of 58
DNA is wound (G&G § 11. 4) n n Standard is one winding per helical turn, i. e. 1 winding per 10 bp Fewer coils or more coils can happen: This introduces stresses that favors unwinding Both underwound and overwound DNA compact the DNA so it sediments faster than relaxed DNA 10/10/2013 Nucleic Acid Structure II p. 45 of 58
Linking, twists, and writhe n n n T=Twist=number of helical turns W=Writhe=number of supercoils L=T+W = Linking number is constant unless you break covalent bonds 10/10/2013 Nucleic Acid Structure II p. 46 of 58
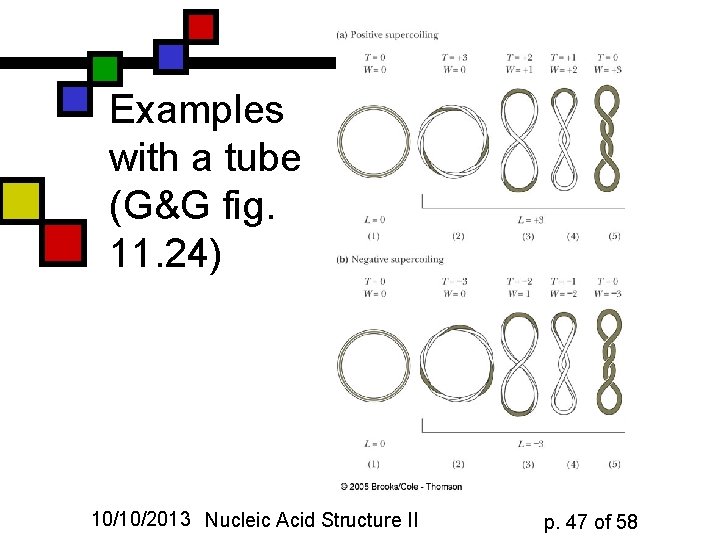
Examples with a tube (G&G fig. 11. 24) 10/10/2013 Nucleic Acid Structure II p. 47 of 58
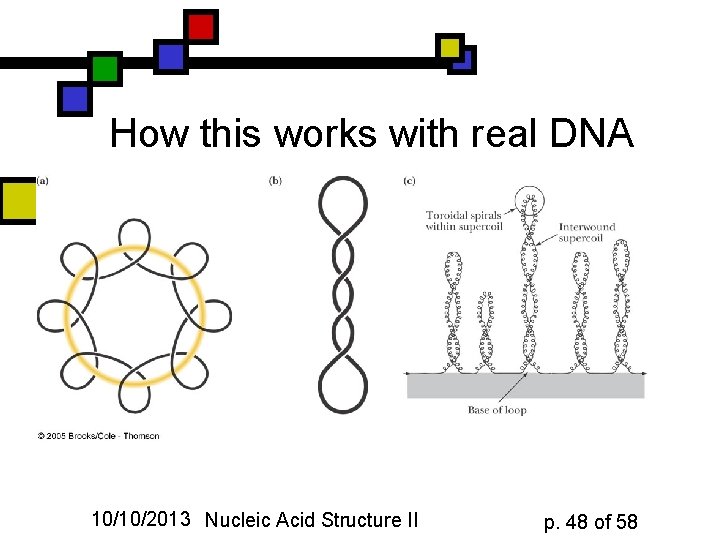
How this works with real DNA 10/10/2013 Nucleic Acid Structure II p. 48 of 58
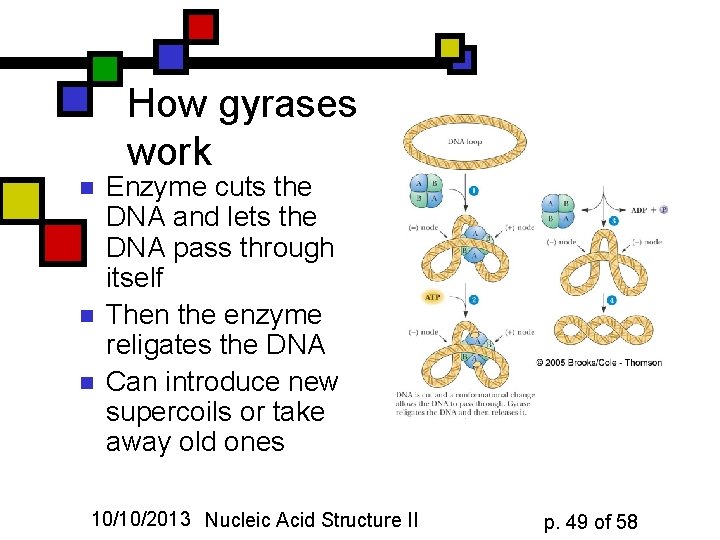
How gyrases work n n n Enzyme cuts the DNA and lets the DNA pass through itself Then the enzyme religates the DNA Can introduce new supercoils or take away old ones 10/10/2013 Nucleic Acid Structure II p. 49 of 58
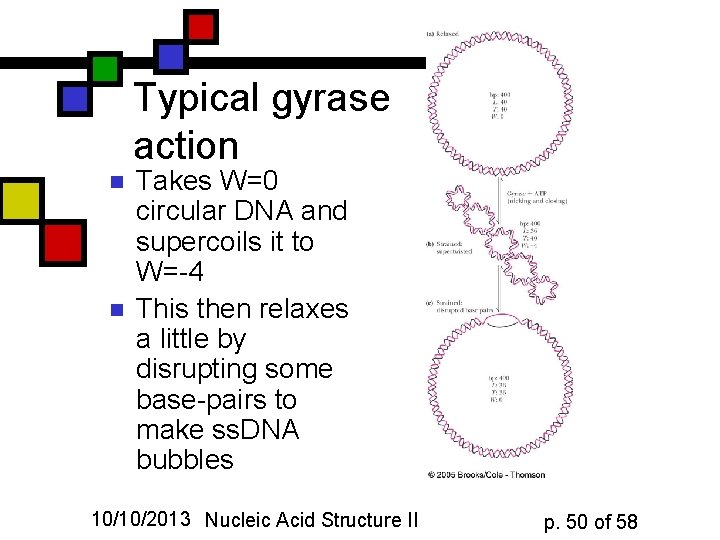
Typical gyrase action n n Takes W=0 circular DNA and supercoils it to W=-4 This then relaxes a little by disrupting some base-pairs to make ss. DNA bubbles 10/10/2013 Nucleic Acid Structure II p. 50 of 58
Superhelix density n n Compare L for real DNA to what it would be if it were relaxed (W=0): That’s L = L - L 0 Sometimes we want = superhelix density = specific linking difference = L / L 0 Natural circular DNA always has < 0 10/10/2013 Nucleic Acid Structure II p. 51 of 58
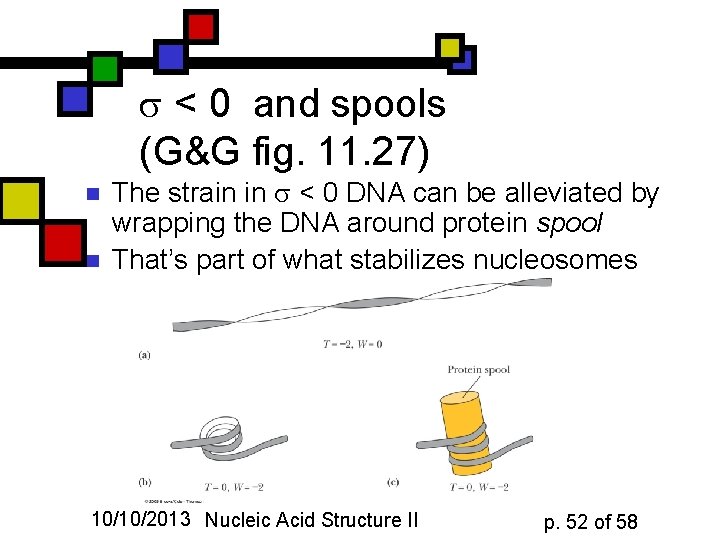
< 0 and spools (G&G fig. 11. 27) n n The strain in < 0 DNA can be alleviated by wrapping the DNA around protein spool That’s part of what stabilizes nucleosomes 10/10/2013 Nucleic Acid Structure II p. 52 of 58
Cruciform DNA n n Cross-shaped structures arise from palindromic structures, including interrupted palindromes like this example These are less stable than regular duplexes but they are common, and they do create recognition sites for DNA-binding proteins, including restriction enzymes 10/10/2013 Nucleic Acid Structure II p. 53 of 58
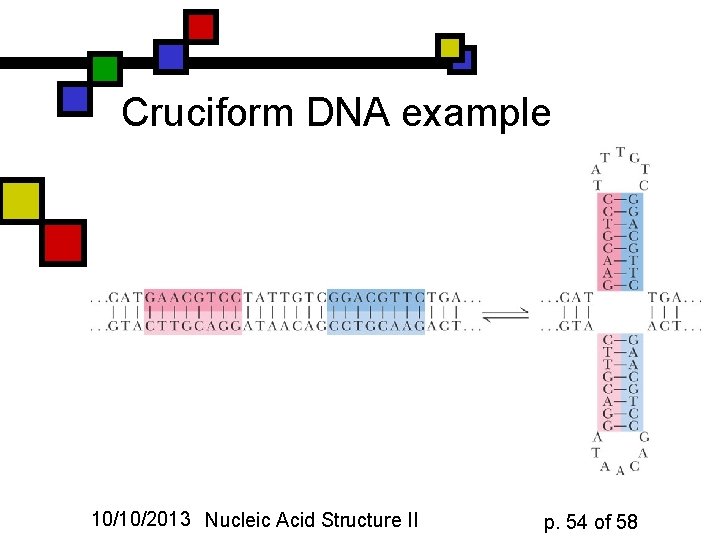
Cruciform DNA example 10/10/2013 Nucleic Acid Structure II p. 54 of 58

Eukaryotic chromosome structure (G&G § 11. 5) n n Human DNA’s total length is ~2 meters! This must be packaged into a nucleus that is about 5 micrometers in diameter This represents a compression of more than 100, 000! It is made possible by wrapping the DNA around protein spools called nucleosomes and then packing these in helical filaments 10/10/2013 Nucleic Acid Structure II p. 55 of 58
Chromatin n Discovered long before we understood molecular biology Seen to be banded objects in nuclei of stained eukaryotic cells In resting cell it exists as long slender threads, 30 nm diameter 10/10/2013 Nucleic Acid Structure II From answers. com p. 56 of 58
Squishing the DNA n n n If the double helix were fully extended, the largest human chromosome (2. 4*108 bp) would be 2. 4*108 *0. 33 nm ~ 0. 8*108 nm=80 mm; much bigger than the cell! So we have to coil it up a lot to make it fit. Longest chromosome is 10µm long So the packing ratio is 80 mm/10µm = 8000 10/10/2013 Nucleic Acid Structure II p. 57 of 58
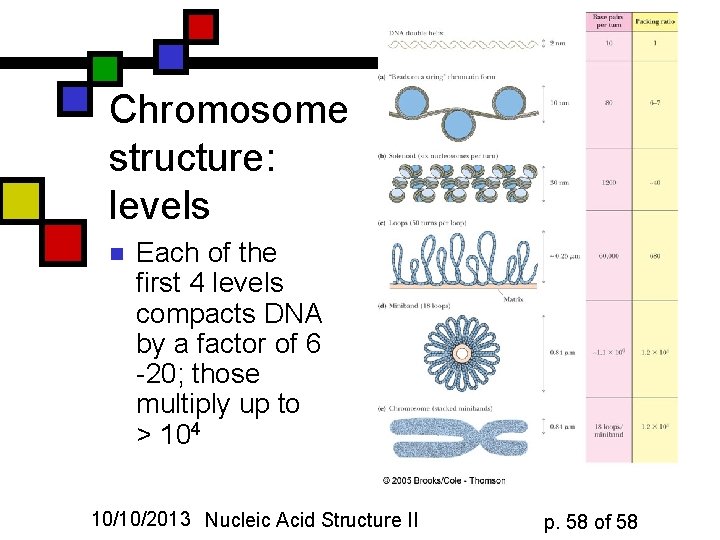
Chromosome structure: levels n Each of the first 4 levels compacts DNA by a factor of 6 -20; those multiply up to > 104 10/10/2013 Nucleic Acid Structure II p. 58 of 58
- Slides: 58