Organization of Biological Data and Databases Pramod Wangikar

Organization of Biological Data and Databases Pramod Wangikar Dept. of Chemical Engineering IIT Bombay
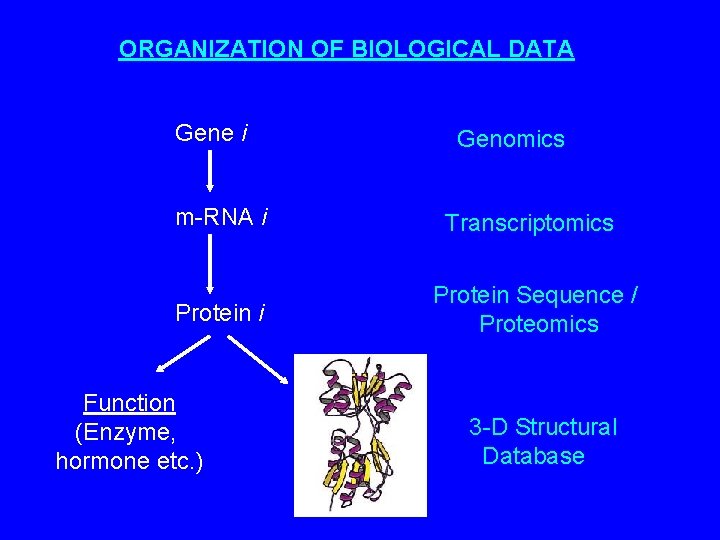
ORGANIZATION OF BIOLOGICAL DATA Gene i Genomics m-RNA i Transcriptomics Protein i Protein Sequence / Proteomics Function (Enzyme, hormone etc. ) 3 -D Structural Database
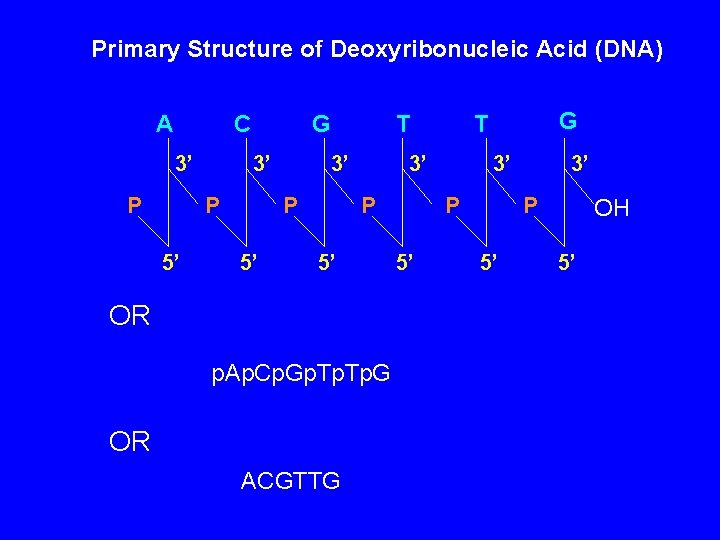
Primary Structure of Deoxyribonucleic Acid (DNA) A C 3’ P G 3’ P 5’ T 3’ P 5’ OR p. Ap. Cp. Gp. Tp. G OR ACGTTG G T 3’ P 5’ OH 5’
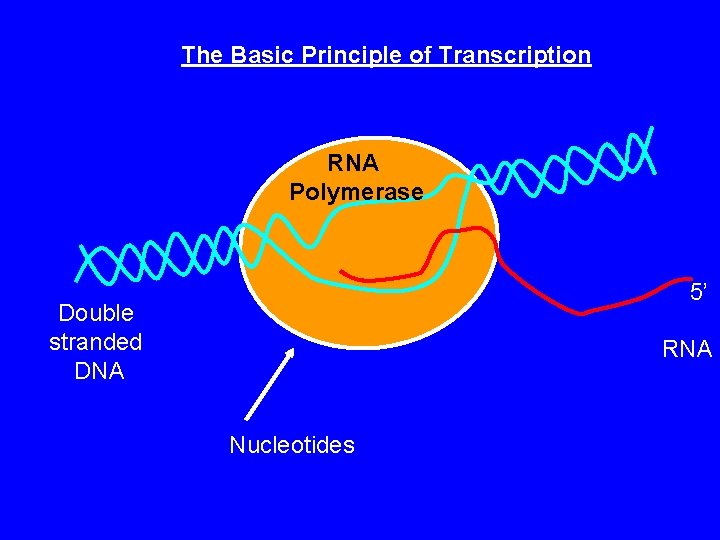
The Basic Principle of Transcription RNA Polymerase 5’ Double stranded DNA RNA Nucleotides
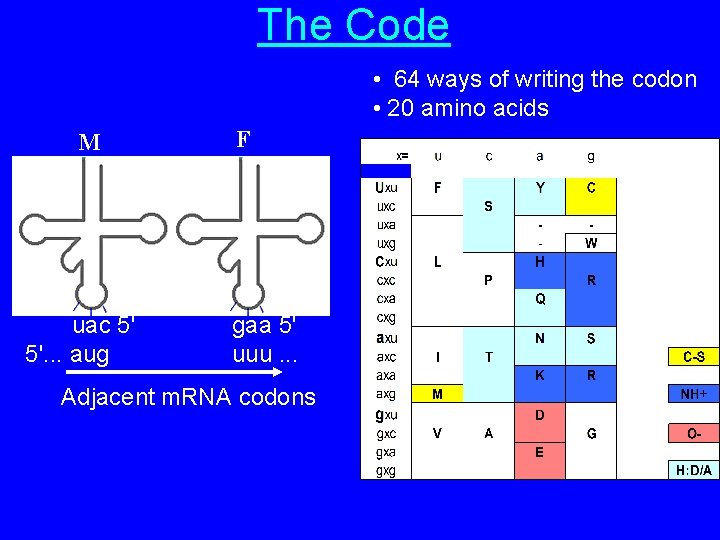
The Code • 64 ways of writing the codon • 20 amino acids M uac 5' 5'. . . aug F gaa 5' uuu. . . Adjacent m. RNA codons
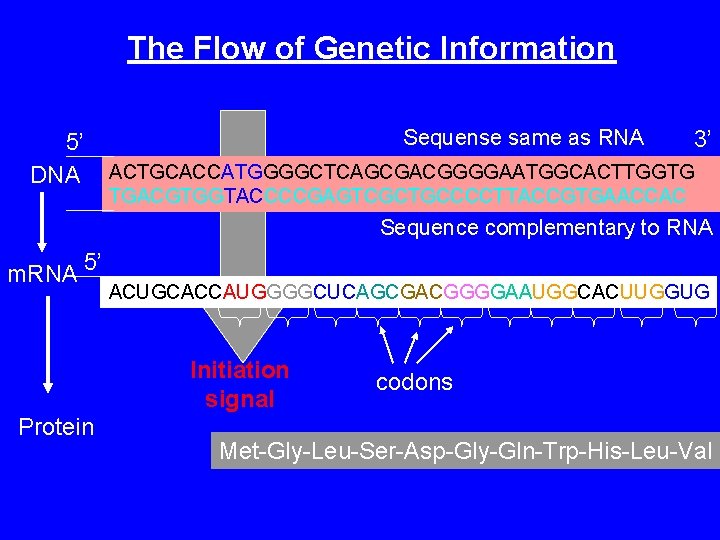
The Flow of Genetic Information 5’ DNA Sequense same as RNA 3’ ACTGCACCATGGGGCTCAGCGACGGGGAATGGCACTTGGTG TGACGTGGTACCCCGAGTCGCTGCCCCTTACCGTGAACCAC Sequence complementary to RNA m. RNA 5’ ACUGCACCAUGGGGCUCAGCGACGGGGAAUGGCACUUGGUG Initiation signal Protein codons Met-Gly-Leu-Ser-Asp-Gly-Gln-Trp-His-Leu-Val

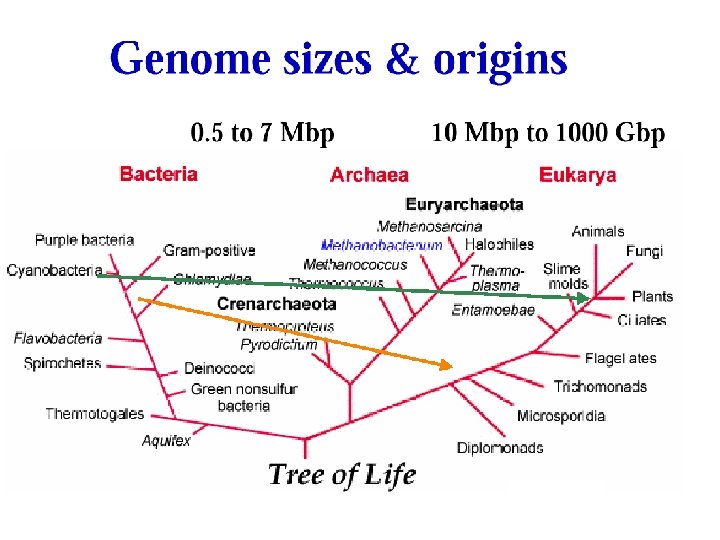
Memory Requirements for Storing Genomes 00 01 10 11 Prokaryotic Eukaryotic = = a c g t 0. 5 -7. 0 Mbp 10 Mbp - 1000 Gbp

How Much Data Does a Bacteria (E. coli) contain?
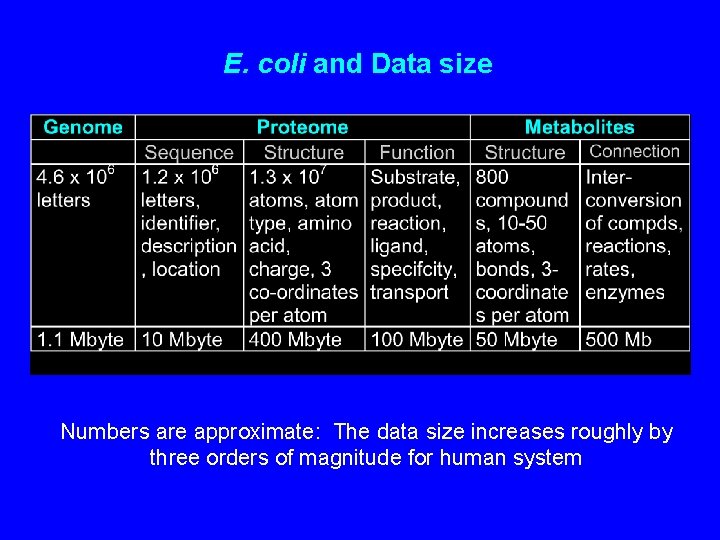
E. coli and Data size Numbers are approximate: The data size increases roughly by three orders of magnitude for human system
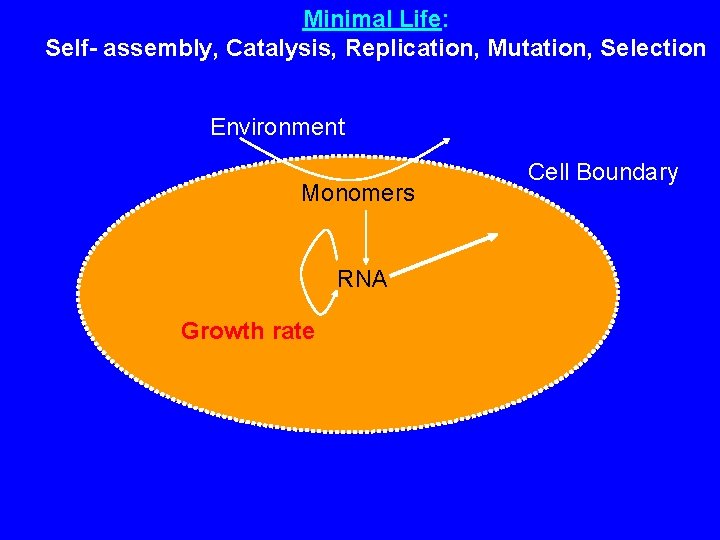
Minimal Life: Self- assembly, Catalysis, Replication, Mutation, Selection Environment Monomers RNA Growth rate Cell Boundary
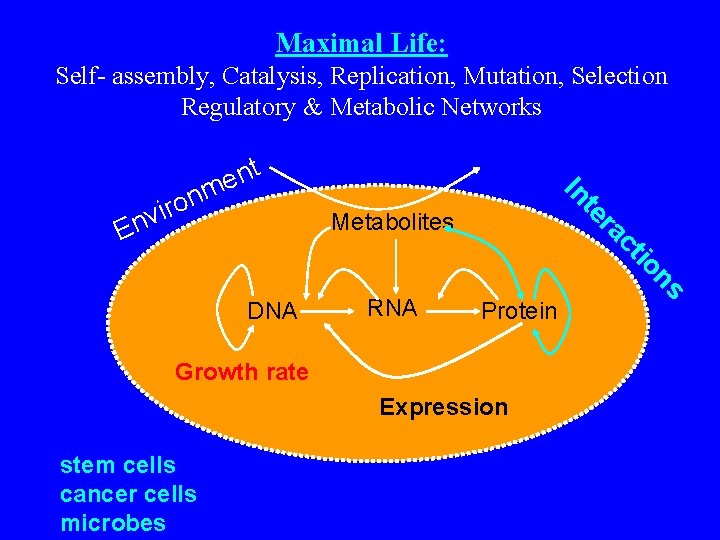
Maximal Life: Self- assembly, Catalysis, Replication, Mutation, Selection Regulatory & Metabolic Networks te Metabolites Protein Growth rate Expression stem cells cancer cells microbes ns RNA io ct ra DNA In v n E nt e m n o ir

Regulation: More biological data What is regulation: A catalogue of possible scenarios and respective course of action. The information for regulation can be stored in the form of: • Protein-protein interaction • Protein-DNA interaction • Protein-metabolite interaction • Molecular switches, controls, set-points, etc. Genome + Environment: Input file Biological Machinery: Executable program Observations: Output file Can we crack the executable program?
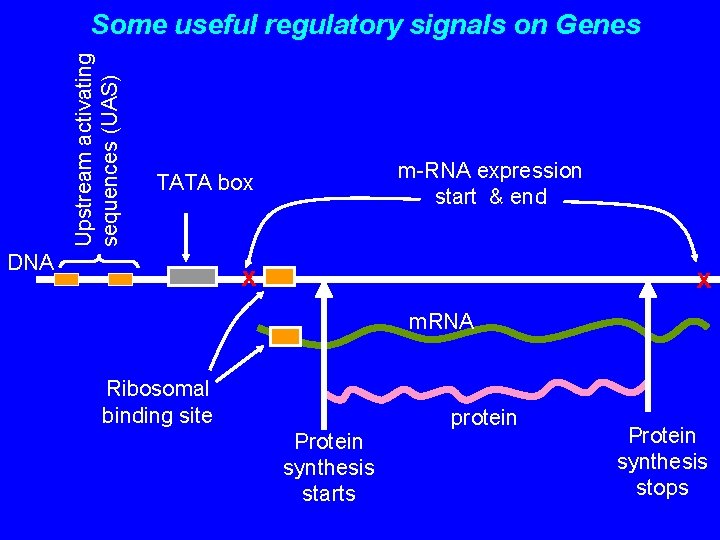
Upstream activating sequences (UAS) Some useful regulatory signals on Genes m-RNA expression start & end TATA box DNA x x m. RNA Ribosomal binding site Protein synthesis starts protein Protein synthesis stops

Minimal Gene Complement of Mycoplasma genitalium
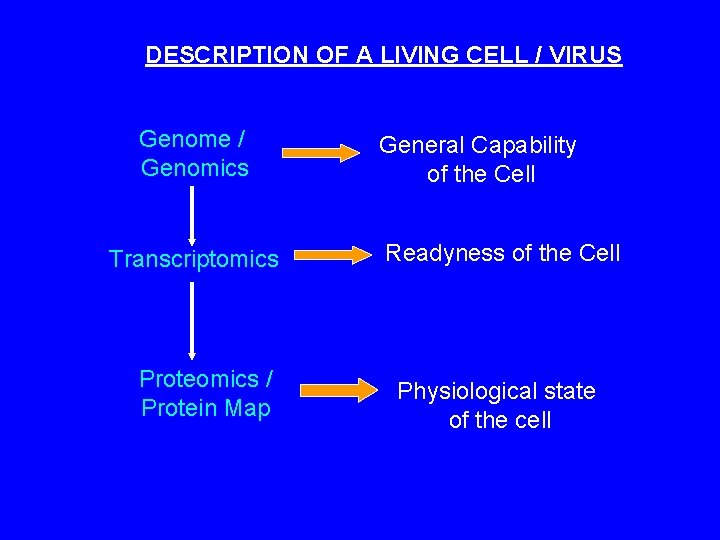
DESCRIPTION OF A LIVING CELL / VIRUS Genome / Genomics Transcriptomics Proteomics / Protein Map General Capability of the Cell Readyness of the Cell Physiological state of the cell
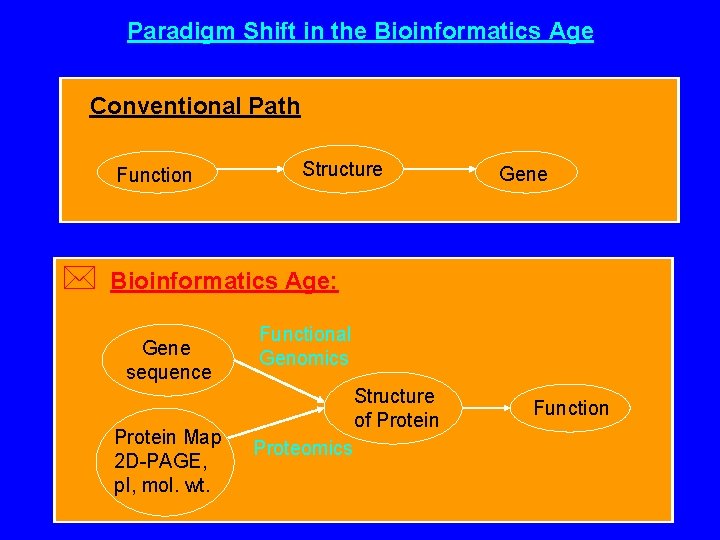
Paradigm Shift in the Bioinformatics Age Conventional Path Function * Structure Gene Bioinformatics Age: Gene sequence Protein Map 2 D-PAGE, p. I, mol. wt. Functional Genomics Structure of Protein Proteomics Function
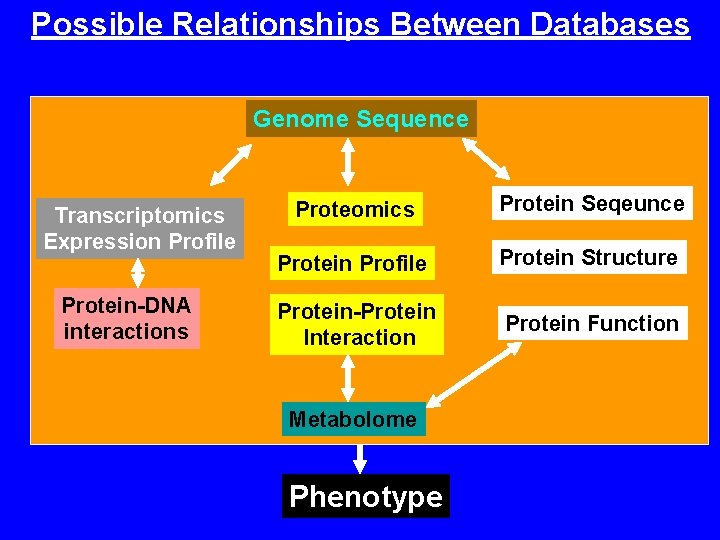
Possible Relationships Between Databases Genome Sequence Transcriptomics Expression Profile Protein-DNA interactions Proteomics Protein Seqeunce Protein Profile Protein Structure Protein-Protein Interaction Protein Function Metabolome Phenotype

Combinatorial Problems in Biology • Prediction of ORF; gene finding • Prediction of DNA regulatory sites • DNA regulatory Proteins • Protein-Protein interactions • Protein Function • Prediction of Metabolic capability • Prediction of Genetic Regulatory Circuits

Biological Databases • Raw databases • Processed databases • Querying in databases.

Raw Databases Conventional Ones DNA / Gene / Genome Sequence Databases. EMBL, Gen. Bank, GSDB etc. > 106 genes, Doubles every 18 months. Genome Projects: E. coli, plants, Human, Mouse, etc. Protein Sequence Databases. PIR, Swiss. Prot, Gen. Bank, etc. > 105 protein sequences, Doubles every 21 months Three Dimensional structure Database. Brookhaven Protein Databank (PDB) > 20, 000 structures, doubles every 24 months.

Proteomics Database (Swiss. Prot) • Each Protein Identified by: p. I, mol wt. , mass spectra, microsequencing, peptide mass fingerprint, etc. • Entries for E. coli, yeast, human etc. Hoogland et al, Nucl. Acids Res. (2000) 28, 286

Cluster of Orthologous Groups (COG) of Proteins: A Processes Database • Compares genes from different genomes. • Forms clusters with similar sequences. • Each COG contains genes connected through vertical evolutionary descent. • 30 genomes (68, 571 genes), 2, 791 COGs with 45, 350 genes • Assignment of function for genes based on known functions for some members of the cluster. • Highly useful for functional assignments for newly sequenced genomes.
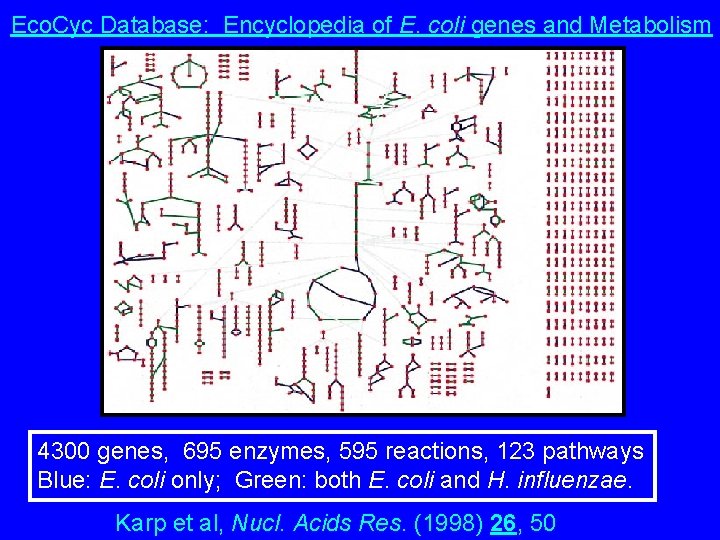
Eco. Cyc Database: Encyclopedia of E. coli genes and Metabolism 4300 genes, 695 enzymes, 595 reactions, 123 pathways Blue: E. coli only; Green: both E. coli and H. influenzae. Karp et al, Nucl. Acids Res. (1998) 26, 50

Querying in Databases • Based on sequence similarity; gives similar sequences and the similarity score or expectation value. • Normally a BLAST, FASTA search (local alignment). Can look for a sequence motif. • Gene names, biological source, functional category, cellular location / role. • Structural features (for known 3 -D structures).

Bioinformatics: A multidisciplinary effort is required • Generation of biological data • Storage and Retrieval of Data • Conversion of known biological hypotheses into mathematical/statistical models • Building models from data • Fitting new data to existing models. • Searching for patterns in data • Derive new biological knowledge from Data
- Slides: 26