Optimization Problems for Polymorphisms of Single Nucleotides Polymorphisms

Optimization Problems for Polymorphisms of Single Nucleotides

Polymorphisms A polymorphism is a feature

Polymorphisms A polymorphism is a feature - common to everybody

Polymorphisms A polymorphism is a feature - common to everybody - not identical in everybody

Polymorphisms A polymorphism is a feature - common to everybody - not identical in everybody - the possible variants (alleles) are just a few

Polymorphisms A polymorphism is a feature - common to everybody - not identical in everybody - the possible variants (alleles) are just a few E. g. think of eye-color

Polymorphisms A polymorphism is a feature - common to everybody - not identical in everybody - the possible variants (alleles) are just a few E. g. think of eye-color Or blood-type for a feature not visible from outside blood-type

At DNA level, a polymorphism is a sequence of nucleotides varying in a population.

At DNA level, a polymorphism is a sequence of nucleotides varying in a population. The shortest possible sequence has only 1 nucleotide, hence Single Nucleotide Polymorphism (SNP)

At DNA level, a polymorphism is a sequence of nucleotides varying in a population. The shortest possible sequence has only 1 nucleotide, hence Single Nucleotide Polymorphism (SNP) atcggattagttagggcacaggacggac atcggattagttagggcacaggacggac atcggattagttagggcacaggacggac atcggattagttagggcacaggacggac

At DNA level, a polymorphism is a sequence of nucleotides varying in a population. The shortest possible sequence has only 1 nucleotide, hence Single Nucleotide Polymorphism (SNP) atcggcttagggcacaggacgtac atcggattagggcacaggacgtac atcggcttagggcacaggacgtac atcggattagttagggcacaggacggac atcggattagggcacaggac atcggcttagggcacaggac atcggattagttagggcacaggacgtac atcggattagttagggcacaggacggac atcggcttagttagggcacaggacggac

- SNPs are predominant form of human variations - On average one every 1, 000 bases - Used for drug design, study disease, forensic, evolutionary. . . atcggcttagggcacaggacgtac atcggattagggcacaggacgtac atcggcttagggcacaggacgtac atcggattagttagggcacaggacggac atcggattagggcacaggac atcggcttagggcacaggac atcggattagttagggcacaggacgtac atcggattagttagggcacaggacggac atcggcttagttagggcacaggacggac
- Multimillion dollar SNP consortium project - 1 st step: build maps of several thousand SNPs - Goal: associate SNPs (or group of SNPs) to genetic diseases atcggcttagggcacaggacgtac atcggattagggcacaggacgtac atcggcttagggcacaggacgtac atcggattagttagggcacaggacggac atcggattagggcacaggac atcggcttagggcacaggac atcggattagttagggcacaggacgtac atcggattagttagggcacaggacggac atcggcttagttagggcacaggacggac

HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS atcggcttagggcacaggacgtac atcggattagggcacaggacgtac atcggcttagggcacaggacgtac atcggattagttagggcacaggacggac atcggattagggcacaggac atcggcttagggcacaggac atcggattagttagggcacaggacgtac atcggattagttagggcacaggacggac atcggcttagttagggcacaggacggac
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS atcggcttagggcacaggacgtac atcggattagggcacaggacgtac atcggcttagggcacaggacgtac atcggattagttagggcacaggacggac atcggattagggcacaggac atcggcttagggcacaggac atcggattagttagggcacaggacgtac atcggattagttagggcacaggacggac atcggcttagttagggcacaggacggac
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS HETEROZYGOUS: different alleles HETEROZYGOUS atcggcttagggcacaggacgtac atcggattagggcacaggacgtac atcggcttagggcacaggacgtac atcggattagttagggcacaggacggac atcggattagggcacaggac atcggcttagggcacaggac atcggattagttagggcacaggacgtac atcggattagttagggcacaggacggac atcggcttagttagggcacaggacggac
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS HETEROZYGOUS: different alleles HETEROZYGOUS atcggcttagggcacaggacgtac atcggattagggcacaggacgtac atcggcttagggcacaggacgtac atcggattagttagggcacaggacggac atcggattagggcacaggac atcggcttagggcacaggac atcggattagttagggcacaggacgtac atcggattagttagggcacaggacggac atcggcttagttagggcacaggacggac

HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS HETEROZYGOUS: different alleles HETEROZYGOUS HAPLOTYPE: chromosome content at SNP sites atcggcttagggcacaggacgtac atcggattagggcacaggacgtac atcggcttagggcacaggacgtac atcggattagttagggcacaggacggac atcggattagggcacaggac atcggcttagggcacaggac atcggattagttagggcacaggacgtac atcggattagttagggcacaggacggac atcggcttagttagggcacaggacggac
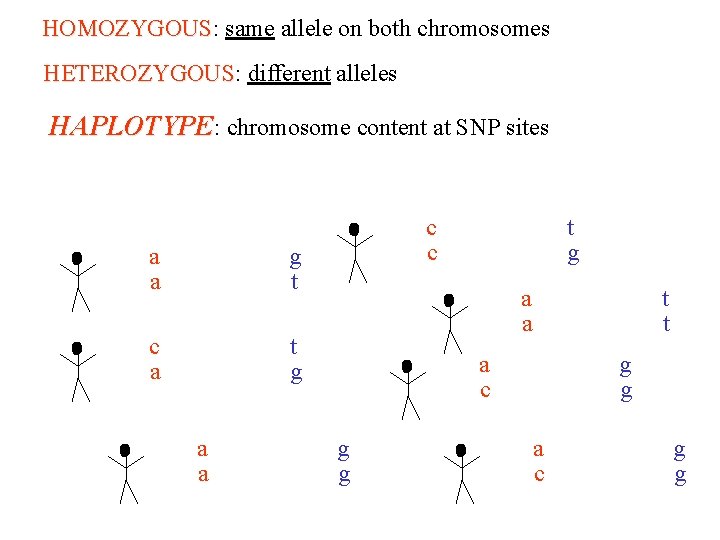
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS HETEROZYGOUS: different alleles HETEROZYGOUS HAPLOTYPE: chromosome content at SNP sites atcggcttagggcacaggacgtac atcggattagggcacaggacgtac atcggcttagggcacaggacgtac atcggattagttagggcacaggacggac atcggattagggcacaggac atcggcttagggcacaggac atcggattagggcacaggacgtac atcggattagggcacaggacgt atcggattagttagggcacaggacggac atcggcttagttagggcacaggacggac
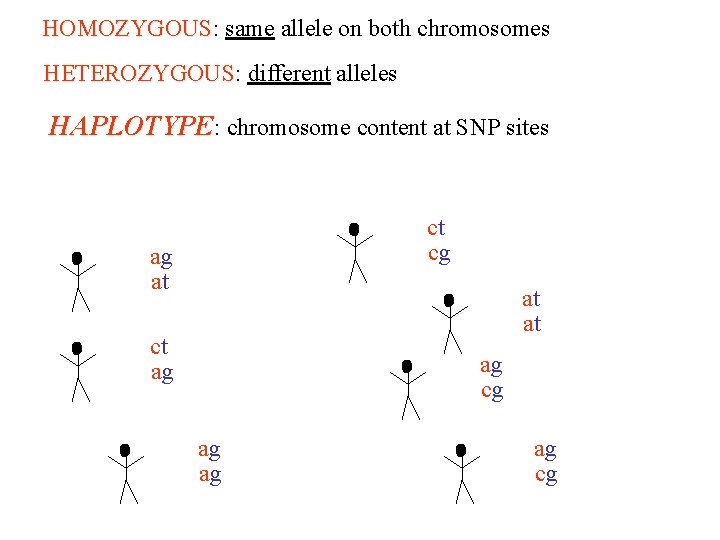
HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS HETEROZYGOUS: different alleles HETEROZYGOUS HAPLOTYPE: chromosome content at SNP sites ct ag at ct ag ag cg at ag cg

HOMOZYGOUS: same allele on both chromosomes HOMOZYGOUS HETEROZYGOUS: different alleles HETEROZYGOUS HAPLOTYPE: chromosome content at SNP sites GENOTYPE: “union” of 2 haplotypes ct ag at ct ag cg Oa. E Oc. E at EE ag Oa. Og ag cg Oa. Ot EOg ag cg Og. E
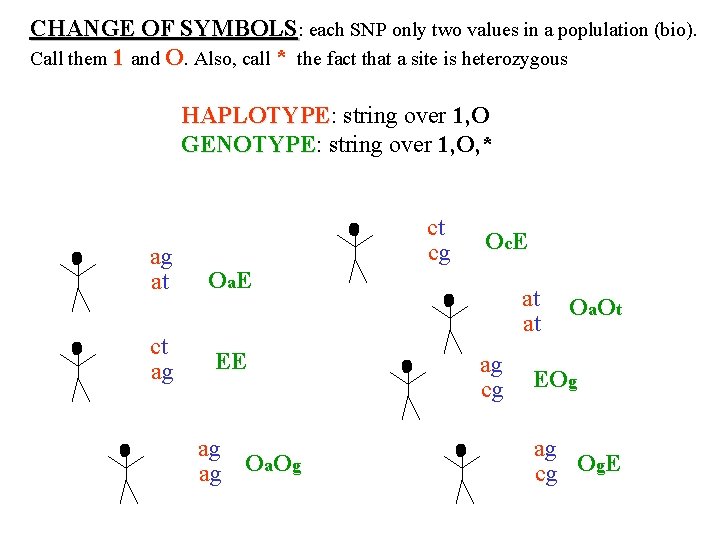
CHANGE OF SYMBOLS: each SNP only two values in a poplulation (bio). Call them 1 and O. Also, call * the fact that a site is heterozygous HAPLOTYPE: string over 1, O HAPLOTYPE GENOTYPE: string over 1, O, * GENOTYPE ct ag at ct ag cg Oa. E Oc. E at EE ag Oa. Og ag cg Oa. Ot EOg ag cg Og. E
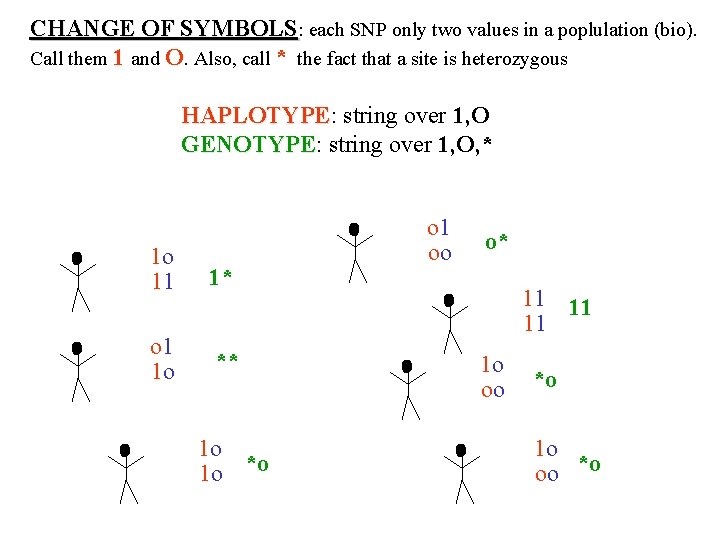
CHANGE OF SYMBOLS: each SNP only two values in a poplulation (bio). Call them 1 and O. Also, call * the fact that a site is heterozygous HAPLOTYPE: string over 1, O HAPLOTYPE GENOTYPE: string over 1, O, * GENOTYPE o 1 1 o 11 o 1 1 o 1* oo o* 11 ** 1 o *o 1 o oo 11 *o 1 o oo *o

THE HAPLOTYPING PROBLEM Single Individual: Given genomic data of one individual, determine Individual 2 haplotypes (one per chromosome) Population : Given genomic data of k individuals, determine (at most) 2 k haplotypes (one per chromosome/indiv. ) For the individual problem, input is erroneous haplotype data, from sequencing For the population problem, data is ambiguous genotype data, from screening OBJ is lead by Occam’s razor: find minimum explanation of observed data under given hypothesis (a. k. a. parsimony principle)

Theory and Results Single individual - Polynomial Algorithms for gapless haplotyping (L, Bafna, Istrail, Lippert, Schwartz 01 & Bafna, L, Istrail, Rizzi 02) - Polynomial Algorithms for bounded-length gapped haplotyping (BLIR 02) - NP-hardness for general gapped haplotyping (LBILS 01) Population - APX-hardness (Gusfield 00) - Reduction to Graph-Theoretic model and I. P. approach (Gusfield 01) -New formulations and Disease Detection (L, Ravi, Rizzi, 02) - Exact algorithms for min-size solution (L, Serafini 2011) - Heuristics (Tininini, L, Bertolazzi 2010)

The Single-Individual Haplotyping problem
![Shotgun Assembly of a Chromosome [ Webber and Myers, 1997] fragmentation sequencing TGAGCCTAG GATTT Shotgun Assembly of a Chromosome [ Webber and Myers, 1997] fragmentation sequencing TGAGCCTAG GATTT](http://slidetodoc.com/presentation_image_h/16620208203029879d966c141039d73f/image-27.jpg)
Shotgun Assembly of a Chromosome [ Webber and Myers, 1997] fragmentation sequencing TGAGCCTAG GATTT GCCTAG CTATCTT ATAGATA GAGATTTCTAGAAATC ACTGA TAGAGATTTC TCCTAAAGAT CGCATAGATA assembly ACTGCAGCCTAGAGATTCTCAGATATTTCTAGGCGTATCTATCTT

MAIN ERROR SOURCES -Sequencing errors: ACTGCCTGGCCAATGGAACGGACAAG CTGGCCAAT CATTGGAAC AATGGAACGGA -Contaminants

Given errors, errors the data may be inconsistent with exactly 2 haplotypes Hence, assembler is unable to build 2 chromosomes PROBLEM: Find and remove the errors so that the data becomes consistent with exactly 2 haplotypes
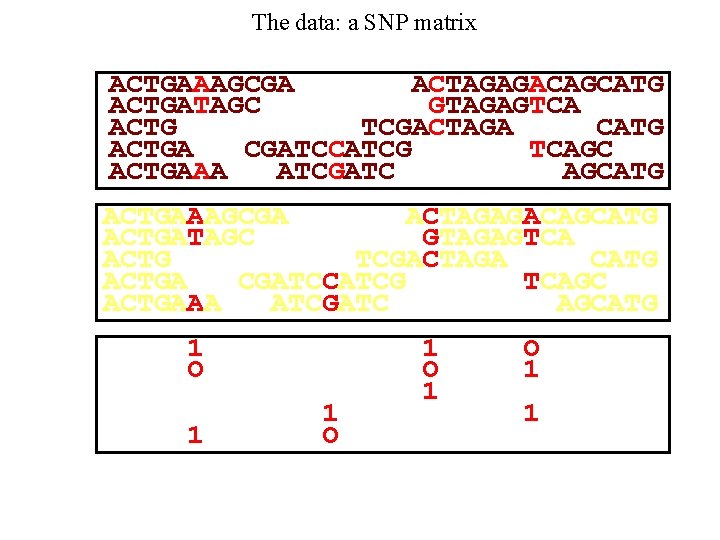
The data: a SNP matrix ACTGAAAGCGA ACTAGAGACAGCATG ACTGATAGC GTAGAGTCA ACTG TCGACTAGA CATG ACTGA CGATCCATCG TCAGC ACTGAAA ATCGATC AGCATG 1 O 1 O 1 1

Snips 1, . . , n 1 2 3 4 5 6 1 1 O - 2 O 1 O - 3 O 1 - 4 O O 1 - 5 1 1 1 O 6 1 O 7 O O 8 O O 1 1 9 1 O - Fragments 1, . . , m
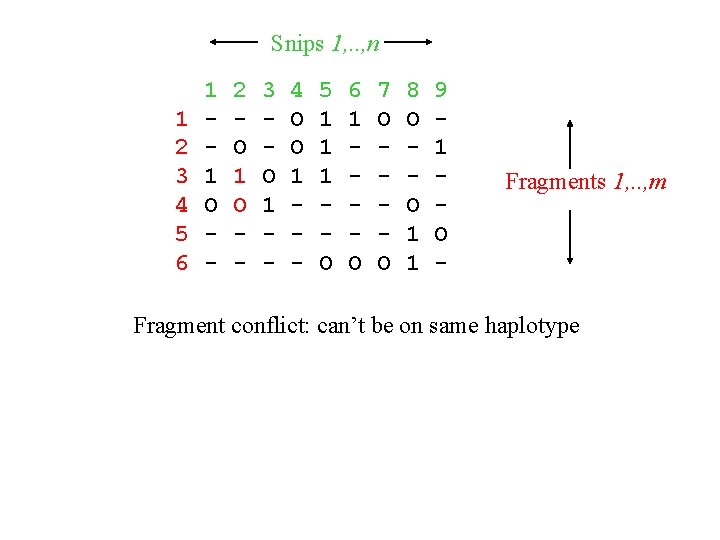
Snips 1, . . , n 1 2 3 4 5 6 1 1 O - 2 O 1 O - 3 O 1 - 4 O O 1 - 5 1 1 1 O 6 1 O 7 O O 8 O O 1 1 9 1 O - Fragments 1, . . , m Fragment conflict: can’t be on same haplotype
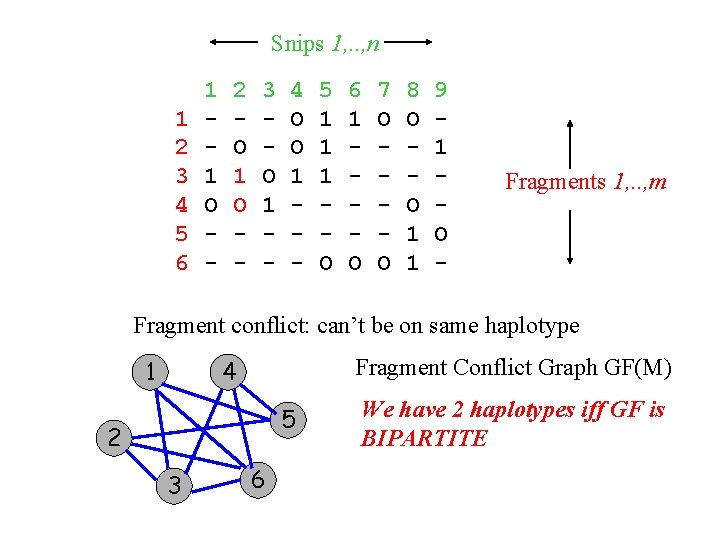
Snips 1, . . , n 1 2 3 4 5 6 1 1 O - 2 O 1 O - 3 O 1 - 4 O O 1 - 5 1 1 1 O 6 1 O 7 O O 8 O O 1 1 9 1 O - Fragments 1, . . , m Fragment conflict: can’t be on same haplotype 1 Fragment Conflict Graph GF(M) 4 5 2 3 6 We have 2 haplotypes iff GF is BIPARTITE

Snips 1, . . , n 1 2 3 4 5 6 1 1 O - 2 O 1 O - 3 O 1 - 4 O O 1 - 5 1 1 1 O 6 1 O 7 O O 8 O O 1 1 9 1 O - Fragments 1, . . , m PROBLEM (Fragment Removal): make GF Bipartite 1 4 5 2 3 6
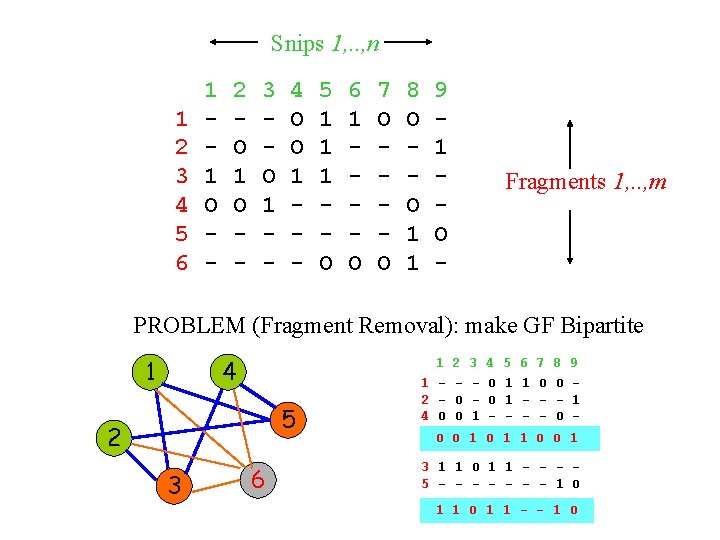
Snips 1, . . , n 1 2 3 4 5 6 1 1 O - 2 O 1 O - 3 O 1 - 4 O O 1 - 5 1 1 1 O 6 1 O 7 O O 8 O O 1 1 9 1 O - Fragments 1, . . , m PROBLEM (Fragment Removal): make GF Bipartite 1 4 5 2 3 6 1 1 2 4 O 2 O O 3 1 4 O O - 5 1 1 - 6 1 - 7 O - 8 O O 9 1 - O O 1 1 O O 1 3 1 1 O 1 1 - - - 5 - - - - 1 O 1 1 - - 1 O

Removing fewest fragments is equivalent to maximum induced bipartite subgraph NP-complete [Yannakakis, 1978 a, 1978 b; Lewis, 1978] O(|V|(log |V|/log |V|)2)-approximable [Halldórsson, 1999] not O(|V| )-approximable for some [Lund and Yannakakis, 1993] Are there cases of M for which GF(M) is easier? YES: the gapless M ---O 11 OO 1 O 1 O 1 OO 1 --- gapless ---O 11 OO---O 1 OO 1 --- gap ---O 11 --1 O----O 1 --- 2 gaps

Why gaps? Sequencing errors (don’t call with low confidence) ---OO 11? 11 --- ===> ---OO 11 -11 ---

Why gaps? Sequencing errors (don’t call with low confidence) ---OO 11? 11 --- ===> ---OO 11 -11 --Celera’s mate pairs attcgttgtagtggtagcctaaatgtcggtagaccttga

THEOREM For a gapless M, the Min Fragment Removal Problem is Polynomial NOTE: Does not need to be gapless. Enough if it can be NOTE sorted to become such (Consecutive Ones Property, Booth and Lueker, 1976)
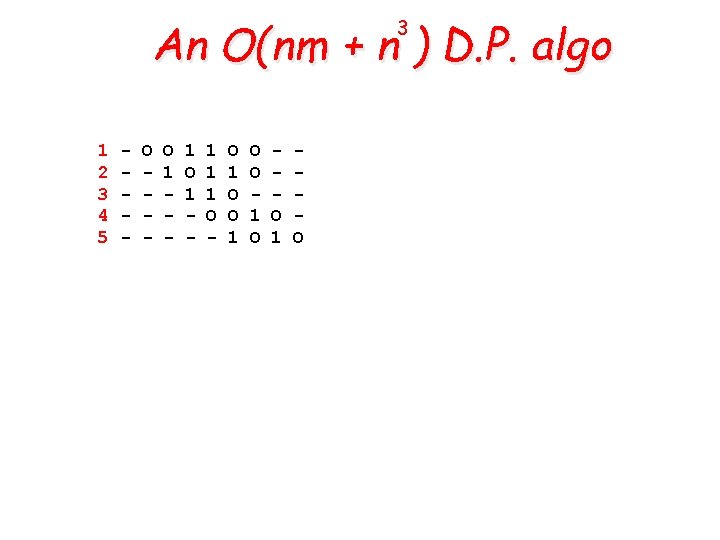
An O(nm + n ) D. P. algo 3 1 2 3 4 5 - O 1 - 1 1 1 O - O 1 O
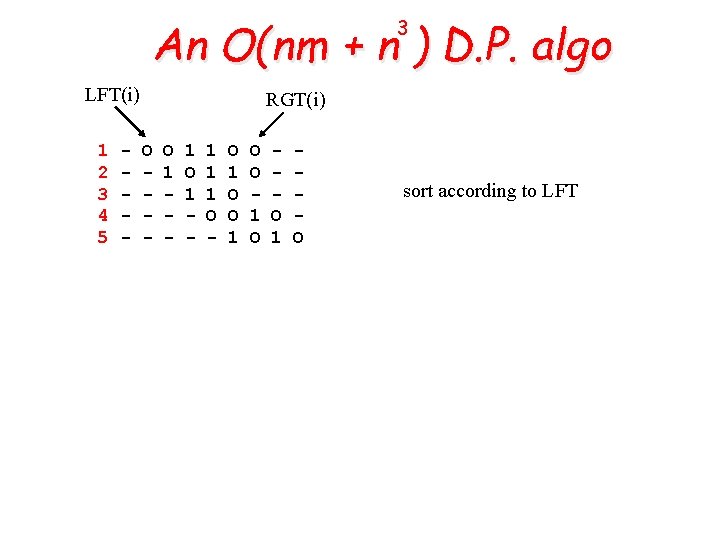
An O(nm + n ) D. P. algo 3 LFT(i) 1 2 3 4 5 - RGT(i) O - O 1 - 1 1 1 O - O 1 O sort according to LFT
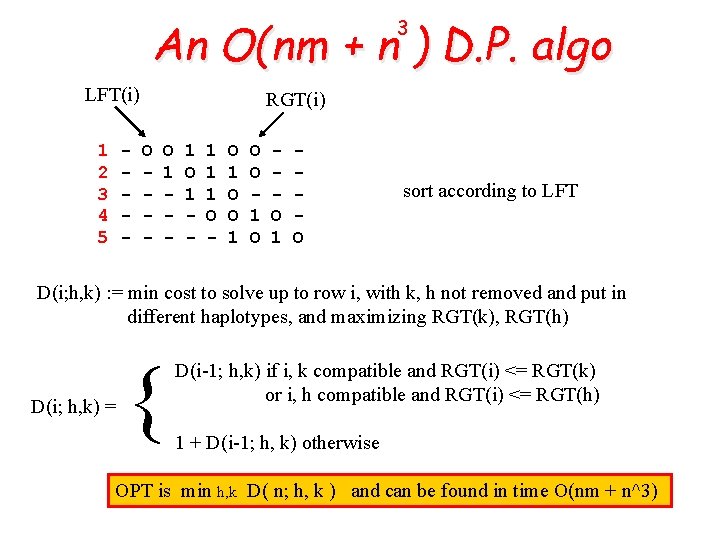
An O(nm + n ) D. P. algo 3 LFT(i) 1 2 3 4 5 - RGT(i) O - O 1 - 1 1 1 O - O 1 O sort according to LFT D(i; h, k) : = min cost to solve up to row i, with k, h not removed and put in different haplotypes, and maximizing RGT(k), RGT(h) D(i; h, k) = { D(i-1; h, k) if i, k compatible and RGT(i) <= RGT(k) or i, h compatible and RGT(i) <= RGT(h) 1 + D(i-1; h, k) otherwise OPT is min h, k D( n; h, k ) and can be found in time O(nm + n^3)

WITH GAPS…. . Th: NP-Hard if 2 gaps per fragment proof: (simple) use fact that for every G there is M s. t. G = GF(M) and reduce from Max Bip. Induced Subgraph on 3 -regular graphs (in each row, max 3 non-bit, hence max 2 gaps)

WITH GAPS…. . Th: NP-Hard if 2 gaps per fragment proof: (simple) use fact that for every G there is M s. t. G = GF(M) and reduce from Max Bip. Induced Subgraph on 3 -regular graphs (in each row, max 3 non-bit, hence max 2 gaps) Th : NP-Hard if even 1 gap per fragment proof: technical. reduction from MAX 2 SAT

WITH GAPS…. . Th: NP-Hard if 2 gaps per fragment proof: (simple) use fact that for every G there is M s. t. G = GF(M) and reduce from Max Bip. Induced Subgraph on 3 -regular graphs (in each row, max 3 non-bit, hence max 2 gaps) Th : NP-Hard if even 1 gap per fragment proof: technical. reduction from MAX 2 SAT But, gaps must be long for problem to be difficult. 2 L 3 L 3 We have O( 2 mn + 2 n ) D. P. for MFR on matrix with total gaps length L

What for MFR with gaps? Why not ILP. . .
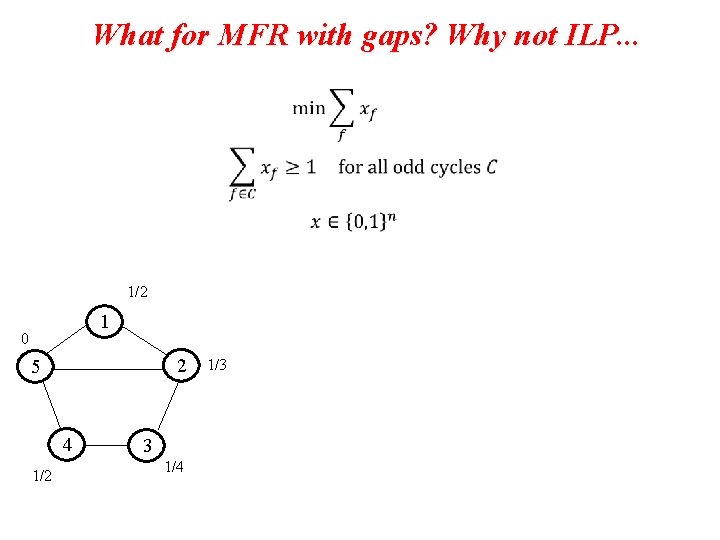
What for MFR with gaps? Why not ILP. . . 1/2 1 0 2 5 4 1/2 3 1/4 1/3
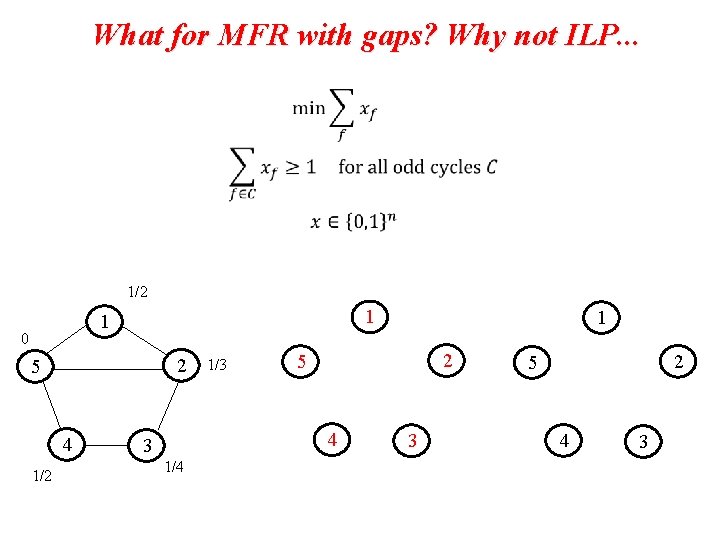
What for MFR with gaps? Why not ILP. . . 1/2 1 1 0 2 5 4 1/2 1/3 2 5 4 3 1/4 1 3 2 5 4 3
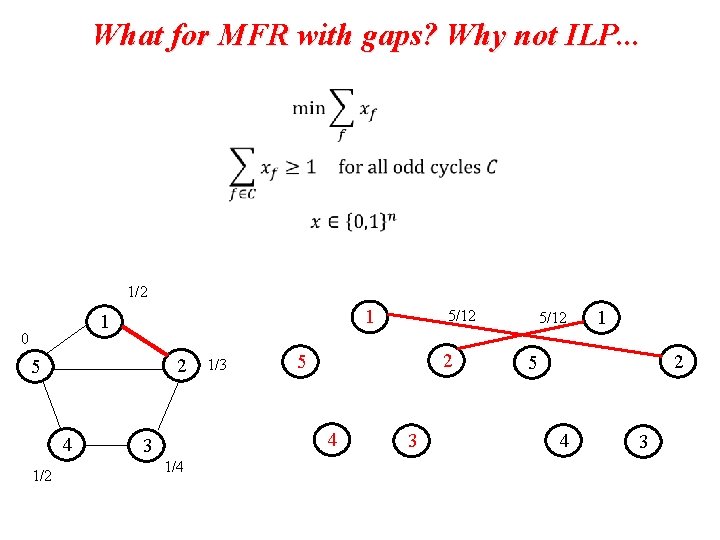
What for MFR with gaps? Why not ILP. . . 1/2 1 1 0 2 5 4 1/2 1/3 2 5 4 3 1/4 5/12 3 5/12 1 2 5 4 3
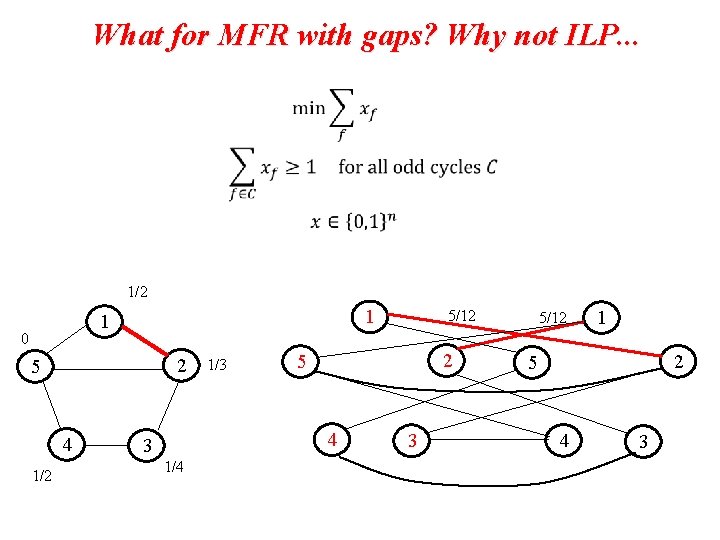
What for MFR with gaps? Why not ILP. . . 1/2 1 1 0 2 5 4 1/2 1/3 2 5 4 3 1/4 5/12 3 5/12 1 2 5 4 3

What for MFR with gaps? Why not ILP. . . 1/2 1 1 0 2 5 4 1/2 1/3 2 5 4 3 1/4 5/12 3 5/12 1 2 5 4 3
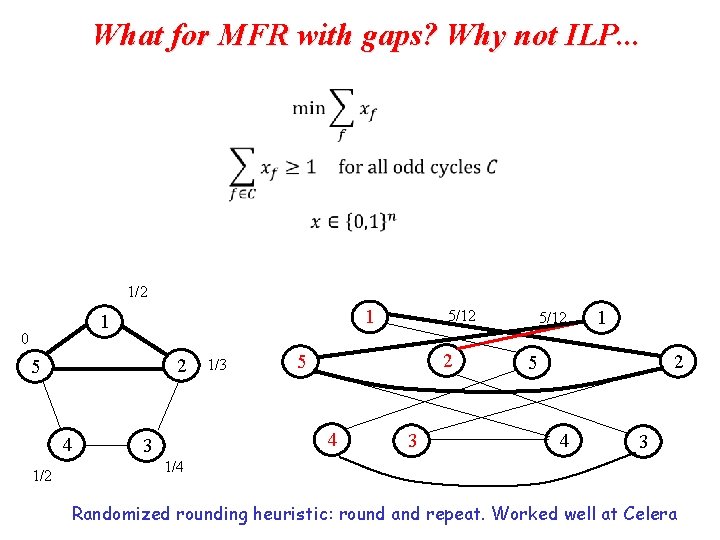
What for MFR with gaps? Why not ILP. . . 1/2 1 1 0 2 5 4 1/2 1/3 2 5 4 3 5/12 1 2 5 4 3 1/4 Randomized rounding heuristic: round and repeat. Worked well at Celera

The fragment removal is good to get rid of contaminants. However, we may want to keep all fragments and correct errors otherwise A dual point of view is to disregard some SNPs and keep the largest subset sufficient to reconstruct the haplotypes All fragments get assigned to one of the two haplotypes. We describe the min SNP removal problem: remove the fewest number of columns from M so that the fragment graph becomes bipartite.
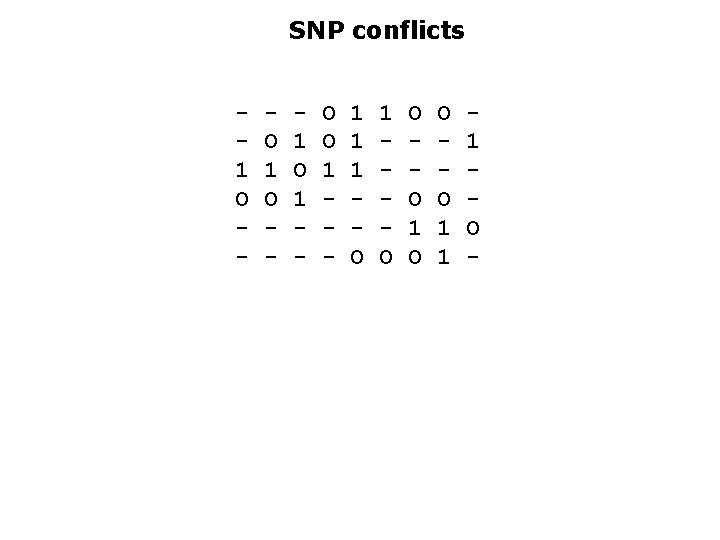
SNP conflicts 1 O - O 1 O - 1 O 1 - O O 1 - 1 1 1 O O O 1 1 1 O -
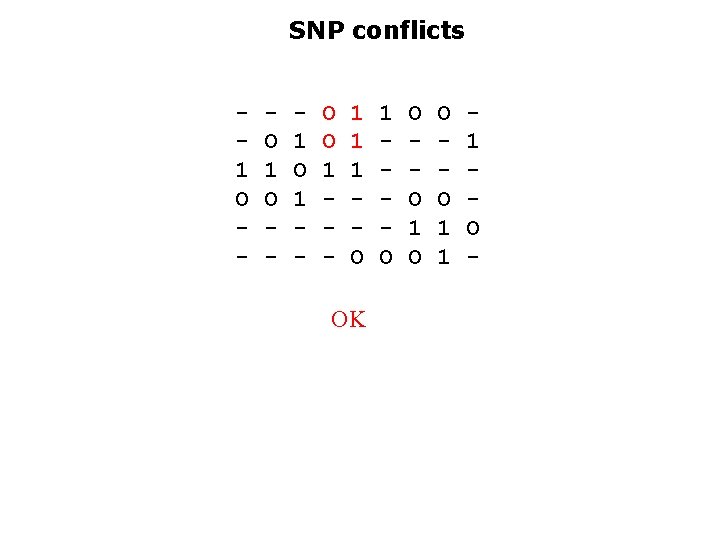
SNP conflicts 1 O - O 1 O - 1 O 1 - O O 1 - 1 1 1 O OK 1 O O O 1 1 1 O -
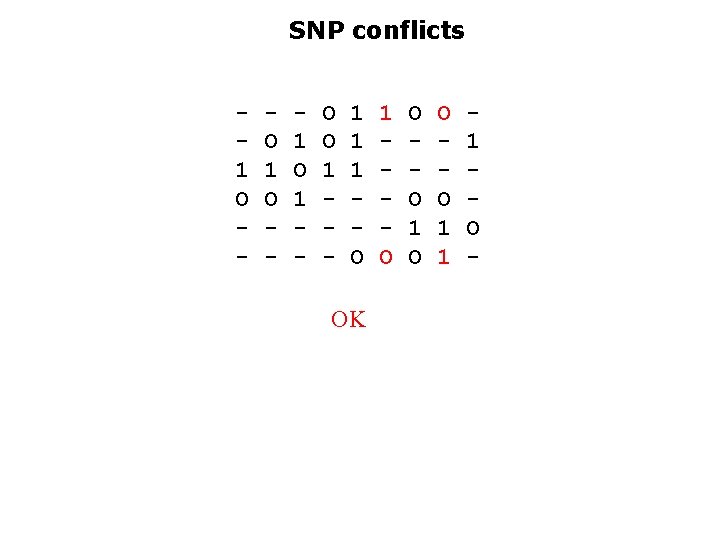
SNP conflicts 1 O - O 1 O - 1 O 1 - O O 1 - 1 1 1 O OK 1 O O O 1 1 1 O -
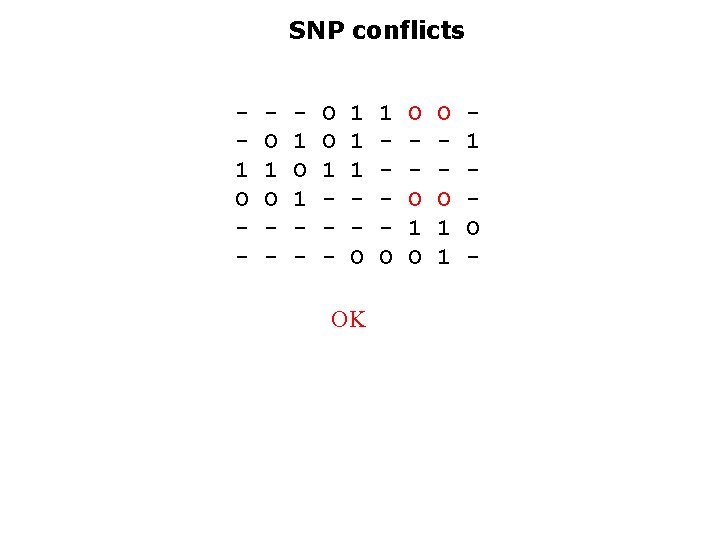
SNP conflicts 1 O - O 1 O - 1 O 1 - O O 1 - 1 1 1 O OK 1 O O O 1 1 1 O -

SNP conflicts 1 O - O 1 O - 1 O 1 - O O 1 - 1 1 1 O O O 1 O CONFLICT ! O O 1 1 1 O -
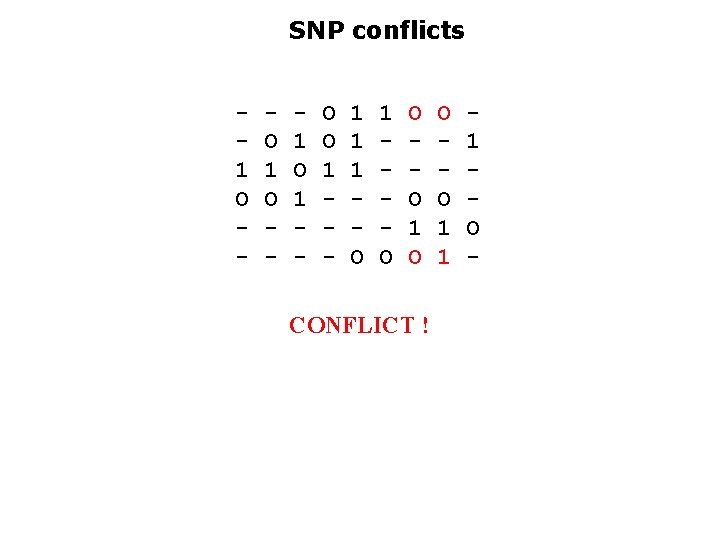
SNP conflicts 1 O - O 1 O - 1 O 1 - O O 1 - 1 1 1 O O O 1 O CONFLICT ! O O 1 1 1 O -
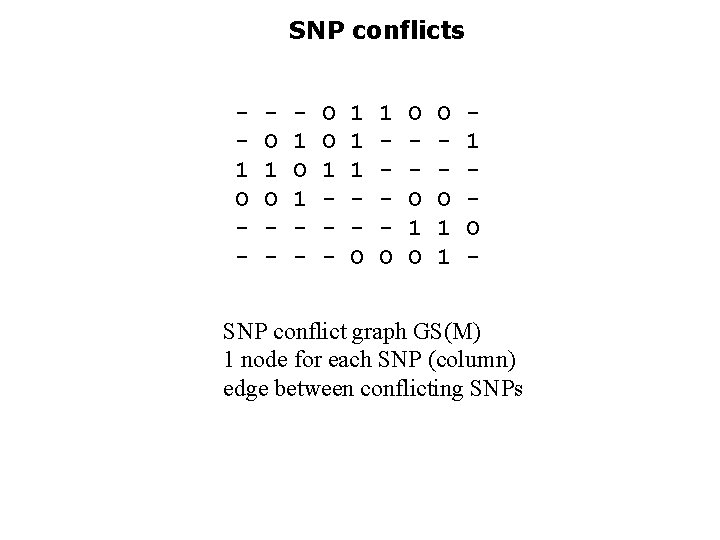
SNP conflicts 1 O - O 1 O - 1 O 1 - O O 1 - 1 1 1 O O O 1 1 1 O - SNP conflict graph GS(M) 1 node for each SNP (column) edge between conflicting SNPs
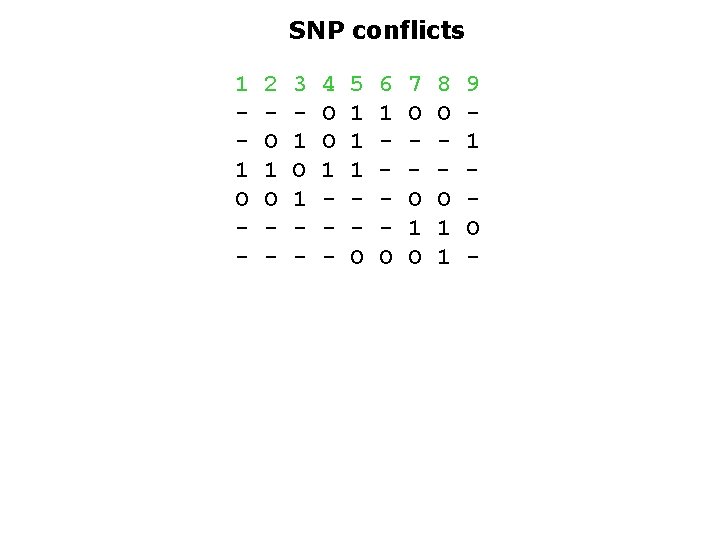
SNP conflicts 1 1 O - 2 O 1 O - 3 1 O 1 - 4 O O 1 - 5 1 1 1 O 6 1 O 7 O O 1 O 8 O O 1 1 9 1 O -
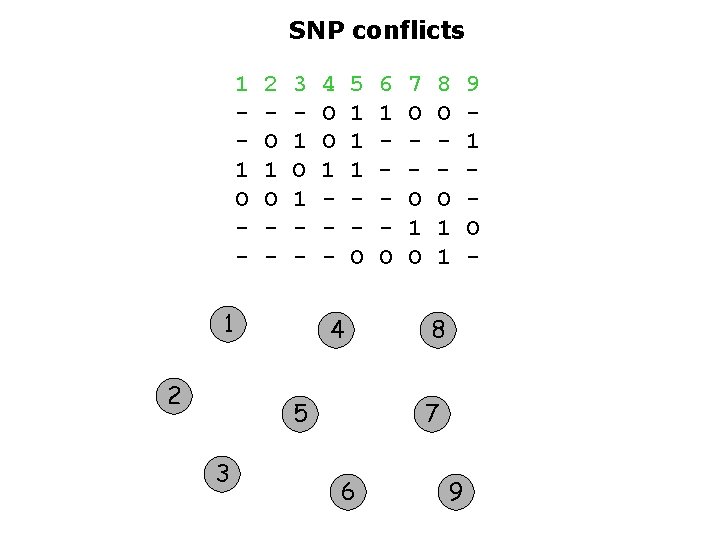
SNP conflicts 1 1 O - 2 O 1 O - 3 1 O 1 - 1 2 4 O O 1 - 5 1 1 1 O 4 5 3 6 1 O 7 O O 1 O 8 O O 1 1 8 7 6 9 9 1 O -

SNP conflicts 1 1 O - 2 O 1 O - 3 1 O 1 - 1 2 4 O O 1 - 5 1 1 1 O 4 5 3 6 1 O 7 O O 1 O 8 O O 1 1 8 7 6 9 9 1 O -

THEOREM 1 For a gapless M, GF(M) is bipartite if and only if GS(M) is an independent set THEOREM 2 For a gapless M, GS(M) is a perfect graph COROLLARY For a gapless M, the min SNP removal problem is polynomial

THEOREM 1 For a gapless M, GF(M) is bipartite if and only if GS(M) is an independent set PROOF (sketch): by minimal counterexample --OO 11 OO------OO 1 O 11 O-----11 O 1 O 111 ----11 OO 1 O 11 O-----1 OOO 1 -----11111 O------11 O 1 OO-----Assume M gapless, GS(M) an independent set, but GF(M) not bipartite. Take an odd cycle in GF
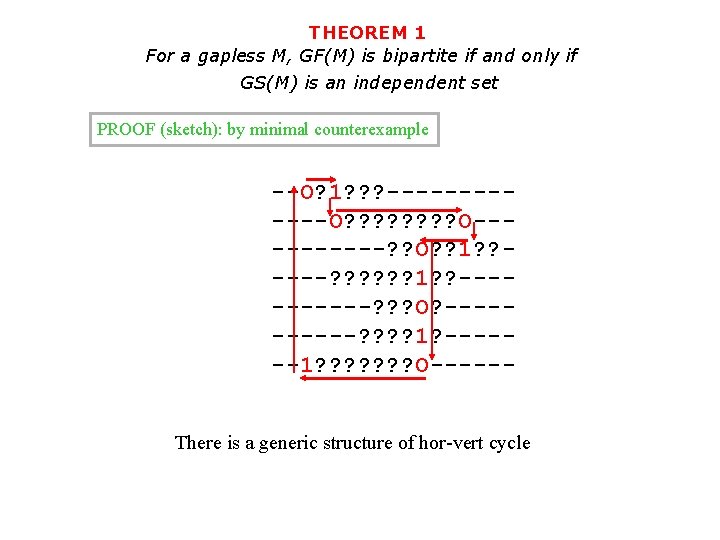
THEOREM 1 For a gapless M, GF(M) is bipartite if and only if GS(M) is an independent set PROOF (sketch): by minimal counterexample --O? 1? ? ? ------O? ? ? ? O-----? ? O? ? 1? ? ----? ? ? 1? ? -----? ? ? O? -----? ? 1? ------1? ? ? ? O-----There is a generic structure of hor-vert cycle

THEOREM 1 For a gapless M, GF(M) is bipartite if and only if GS(M) is an independent set PROOF (sketch): by minimal counterexample --O? 1? ? ? ------O? ? ? ? O-----? ? O? ? 1? ? ----? ? ? 1? ? -----? ? ? O? -----? ? 1? ------1? ? ? ? O-----“vertical lines” There cannot be only one vertical line in odd cycle We merge rightmost and next to reduce them by 1 Hence, there cannot be a minimal (in n. of vertical lines) counterexample

THEOREM 1 For a gapless M, GF(M) is bipartite if and only if GS(M) is an independent set PROOF (sketch): by minimal counterexample Must be 1 --O? 1? ? ? ------O? ? ? ? O-----? ? O? ? 1? ? ----? ? ? 1? ? -----? ? ? O? -----? ? 1? ------1? ? ? ? O-----“vertical lines”
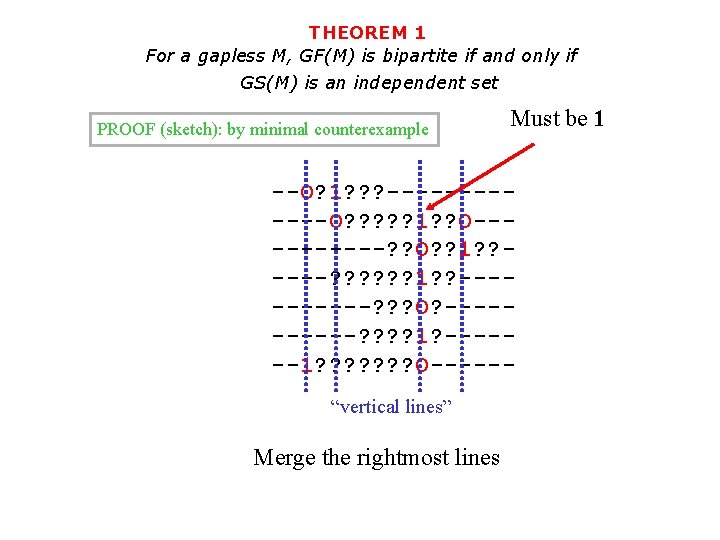
THEOREM 1 For a gapless M, GF(M) is bipartite if and only if GS(M) is an independent set PROOF (sketch): by minimal counterexample Must be 1 --O? 1? ? ? ------O? ? ? 1? ? O-----? ? O? ? 1? ? ----? ? ? 1? ? -----? ? ? O? -----? ? 1? ------1? ? ? ? O-----“vertical lines” Merge the rightmost lines

THEOREM 1 For a gapless M, GF(M) is bipartite if and only if GS(M) is an independent set PROOF (sketch): by minimal counterexample --O? 1? ? ? ------O? ? ? 1 -------? ? O-----? ? ? 1 ------? ? ? O------? ? 1 -------1? ? ? ? O-----“vertical lines” Merge the rightmost lines Still a counterexample!
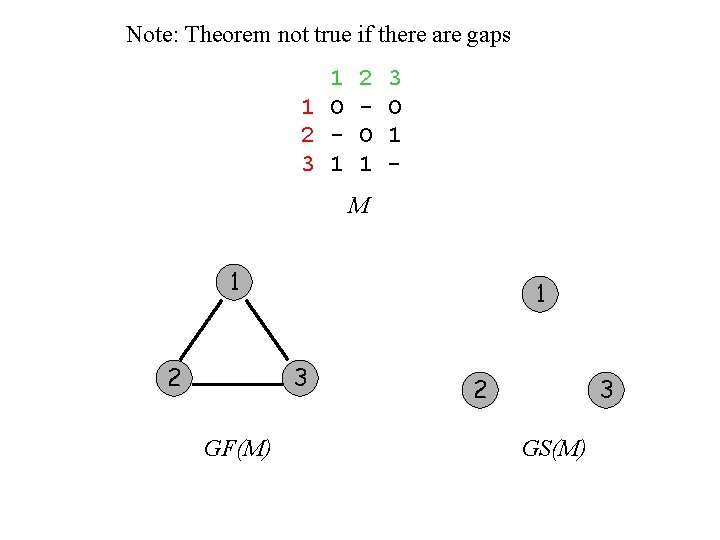
Note: Theorem not true if there are gaps 1 1 O 2 3 1 2 O 1 3 O 1 - M 1 2 1 3 GF(M) 2 3 GS(M)
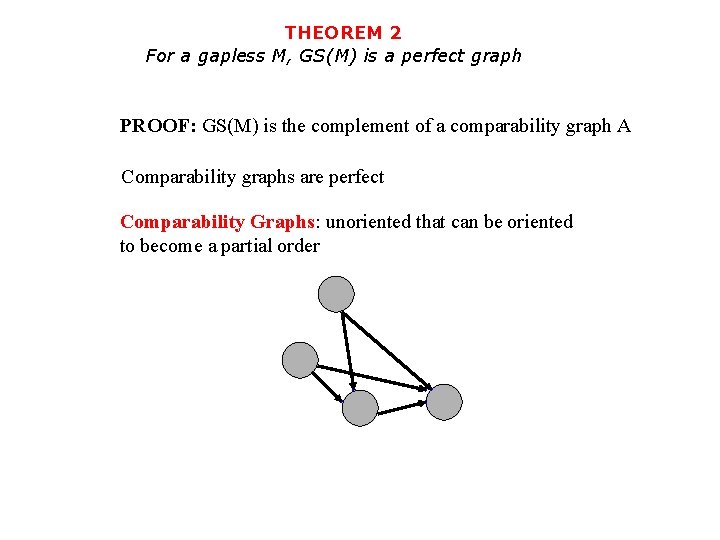
THEOREM 2 For a gapless M, GS(M) is a perfect graph PROOF: GS(M) is the complement of a comparability graph A Comparability graphs are perfect Comparability Graphs: unoriented that can be oriented to become a partial order
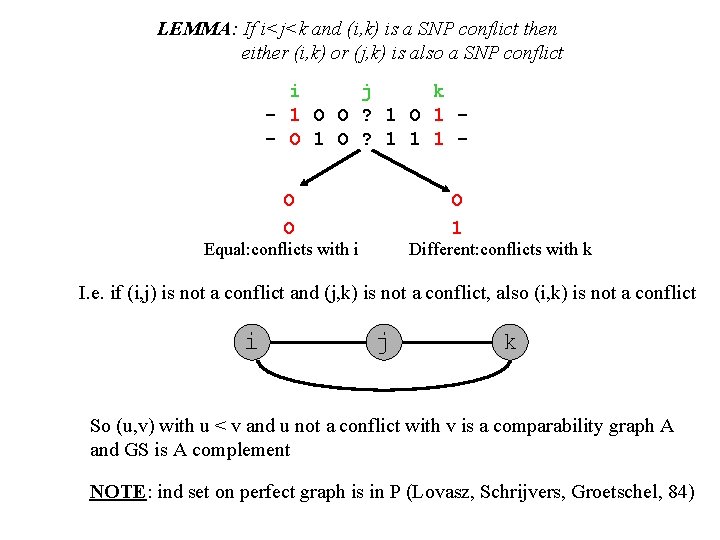
LEMMA: If i<j<k and (i, k) is a SNP conflict then either (i, k) or (j, k) is also a SNP conflict i j k - 1 O O ? 1 O 1 - O 1 O ? 1 1 1 O O O 1 Equal: conflicts with i Different: conflicts with k I. e. if (i, j) is not a conflict and (j, k) is not a conflict, also (i, k) is not a conflict i j k So (u, v) with u < v and u not a conflict with v is a comparability graph A and GS is A complement NOTE: ind set on perfect graph is in P (Lovasz, Schrijvers, Groetschel, 84)

Hence gapless MSR is polynomial (max stable set on perfect graph). There are better, D. P. , algorithms, O(mn + m^2) What if gaps ? THEOREM: The min SNP removal is NP-hard if there can be gaps (Reduction from MAXCUT) Again, gaps must be long for problem to be difficult. 2 L + 1 2 L + 2 We have O(mn + n ) D. P. for MSR on matrix with total gaps length L
- Slides: 74