Ontology The New Era Barry Smith http ncbo

Ontology: The New Era Barry Smith http: //ncbo. us 1
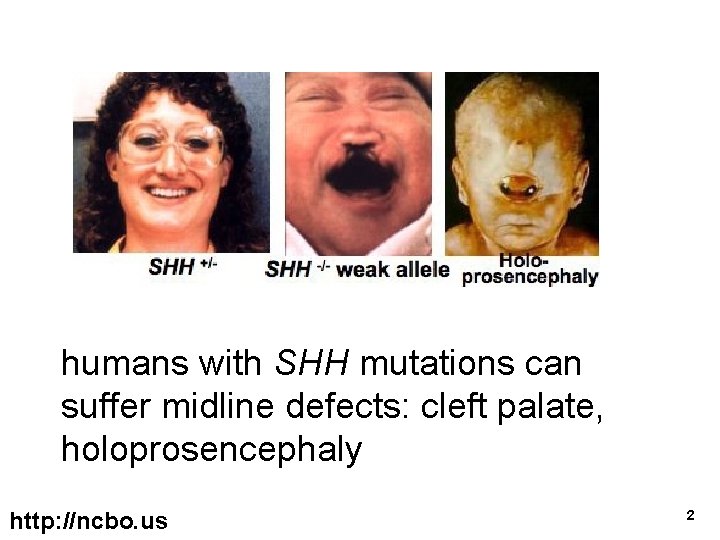
humans with SHH mutations can suffer midline defects: cleft palate, holoprosencephaly http: //ncbo. us 2

but holoprosencephaly can also appear in individuals with normal SHH Question: due to what other factors? Answer: Let’s look at orthologs of SHH in other model organisms http: //ncbo. us 3

what we find mutations in shh, the zebrafish ortholog of SHH, yield analogous defects but so do mutations in oep, another zebrafish gene molecular identification of oep allowed discovery of mutations in the human oep ortholog (TDGF 1) which could be shown to cause holoprosencephaly http: //ncbo. us 4

… but this took four years http: //ncbo. us 5

The holy grail What would it take to detect patterns of similarity between human phenotypes and those model organism phenotypes for which we have potentially useful molecule-level data and to isolate those automatically ? First: good (realistic, scientific) data sources http: //ncbo. us 6
genes associated with cleft palate 445 genes http: //ncbo. us 7

abnormal proteins associated with cleft palate http: //ncbo. us 8
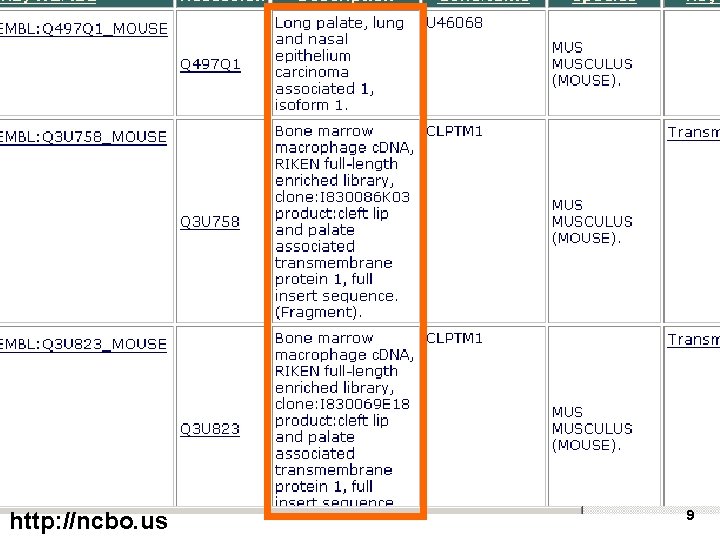
http: //ncbo. us 9

Second: good (realistic, scientific) ontologies http: //ncbo. us 10

Finding shared cross-species phenotypic features with implications for our understanding of human diseases is like finding needles in haystacks http: //ncbo. us 11

haystack with needle http: //ncbo. us 12

haystack without needle http: //ncbo. us 13

http: //ncbo. us 14

Needle with haystack as represented in a good, realist ontology http: //ncbo. us 15

Good (scientific, realist) ontologies require hard work and staying power http: //ncbo. us 16

the haystack ontology John built last Tuesday http: //ncbo. us 17

Sally’s haystack ontology she did this morning http: //ncbo. us 18

So: to detect cross-species similarities we can’t just google our way across John’s, and Sally’s, and Bill’s, and Tom’s ontologies • they are all still fragments • they are syntactically unregimented • none are interoperable with the FMA (or with anything else) • none allows automatic error-checking or automatic reasoning http: //ncbo. us 19

all are full of weird artefacts Bill confuses portions of tissue with limbs Sally thinks gland is identical with observation of a gland Tom thinks blood pressure is an act of measurement Singupta thinks the heart is an ordered pair consisting of a preferred term and a concept unique identifier Jim thinks there are exactly 26 kinds of chemicals Olivier wastes half his life constructing mappings between these various bits of nonsense http: //ncbo. us 20

Finding shared cross-species phenotypic features is like finding needles in haystacks where our search is constrained by the need to reason back and forth across heterogeneous data sources relating to entities at different levels of granularity http: //ncbo. us 21
genes associated with cleft palate 445 genes http: //ncbo. us 22

abnormal proteins associated with cleft palate http: //ncbo. us 23
medical records Referent tracking data SNOMED codes http: //ncbo. us 24

We know that high-quality ontologies can help in creating high-quality mappings between human and model organism phenotypes http: //ncbo. us 25

OWL is not enough The use of a common syntax and logical machinery and the careful separating out of ontologies into namespaces does not solve the problem of ontology integration And it certainly does not solve the problem of ontology quality. http: //ncbo. us 26

“Alignment of Multiple Ontologies of Anatomy: Deriving Indirect Mappings from Direct Mappings to a Reference Ontology” Songmao Zhang Olivier Bodenreider AMIA 2005 http: //ncbo. us 27
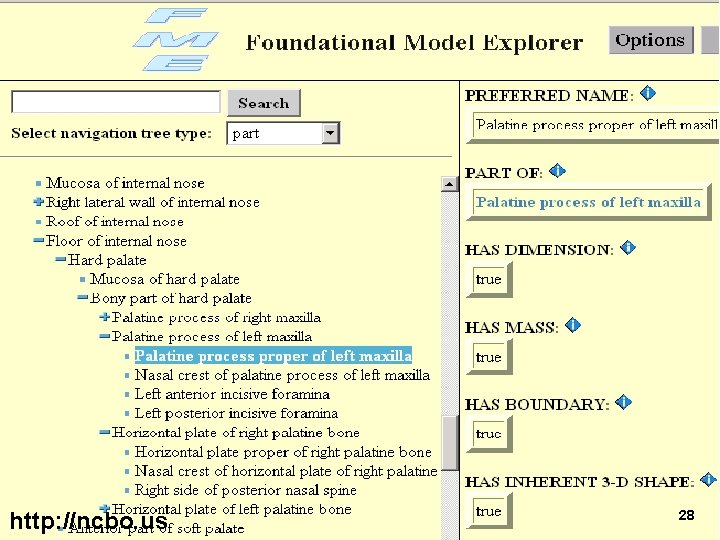
http: //ncbo. us 28

Robin Mc. Entire, GSK What we need is a strong push toward "industrial -strength" ontologies. … ontologies with a consistent and rich representation formalism that are amenable for use as an integration framework, and support reasoning capabilities. We anticipate that pharma's need to bring together mountains of data and information and to properly analyse that information all depend on having a stable, well-developed semantic framework that links information/data and that allows reasoning systems to perform some of our more "mundane" analysis work. http: //ncbo. us 29

Scientific, rigorously tested reference ontologies in anatomy in physiology in pathology in chemistry a reformed GO for genes and gene products … http: //ncbo. us 30

One Central Goal of the National Center for Biomedical Ontology apply the scientific method to the development of biomedical ontologies treat them not as word lists for hobbyists but as scientific theories which are subject to empirical testing against realworld benchmarks able to support tools for automatic reasoning and error-checking http: //ncbo. us 31

To realize this goal we will 1. organize hands-on workshops* with different community groups, coaxing and cajoling them to incorporate ontology best practices in their work and to foster mutual learning and coordination 2. *Workshop in Schloss Dagstuhl, Germany http: //ncbo. us 32

2. use top-level reference ontologies as constraints on Open Biomedical Ontologies (OBO) library* http: //obo. sourceforge. net to create the conditions for a step-by -step evolution towards high-quality interoperable reference ontologies in the biomedical domain *elite GOLD membership category http: //ncbo. us 33
- Slides: 33