Ontologies in Biomedicine AMIA Tutorial T 21 November

Ontologies in Biomedicine AMIA Tutorial T 21 November 12, 2006 Mark A. Musen Stanford University

What Is An Ontology? • The study of being • A discipline co-opted by computer science to enable the explicit specification of – Entities – Properties and attributes of entities – Relations between entities • A theory that provides a common vocabulary for an application domain

Porphyry’s depiction of Aristotle’s Categories Supreme genus: Differentiae: SUBSTANCE material Subordinate genera: Differentiae: BODY animate Subordinate genera: Differentiae: inanimate sensitive MINERAL insensitive ANIMAL rational Species: Individuals: SPIRIT LIVING Proximate genera: Differentiae: immaterial irrational HUMAN Socrates PLANT Plato BEAST Aristotle …


Why develop an ontology? • To share a common understanding of the entities in a given domain – among people – among software agents – between people and software • To enable reuse of data and information – to avoid re-inventing the wheel – to introduce standards to allow interoperability and automatic reasoning • To create communities of researchers
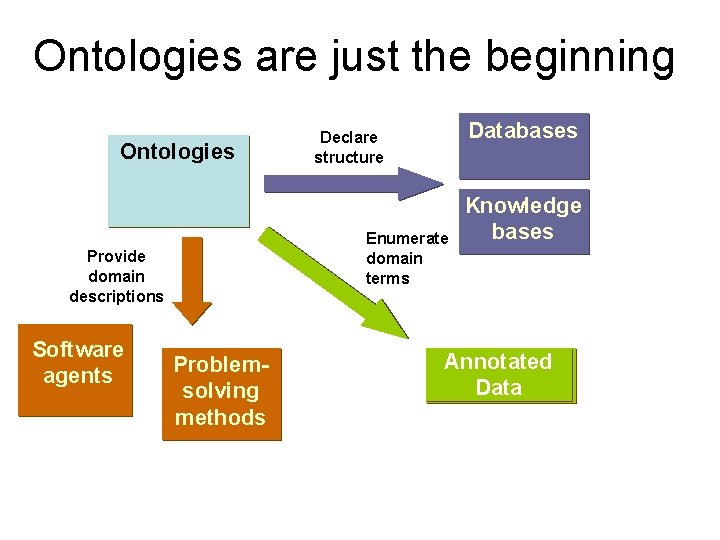
Ontologies are just the beginning Ontologies Enumerate domain terms Provide domain descriptions Software agents Databases Declare structure Problemsolving methods Knowledge bases Annotated Data

Open Directory Project • Started in 1998 as a volunteer effort to develop an open-content directory of Web pages • In its first year, 4500 editors had indexed 100 K Web sites • By July 2005, 69 K editors had indexed 4. 6 M sites using 580 K categories • On average, between 9 K and 10 K volunteer editors are working on ODP at any given time

Foundational Model of Anatomy • Long-term project at University of Washington to create a comprehensive ontology of human anatomy • 72 K concepts, 1. 9 M relationships • One of the largest and best developed ontologies in biomedicine
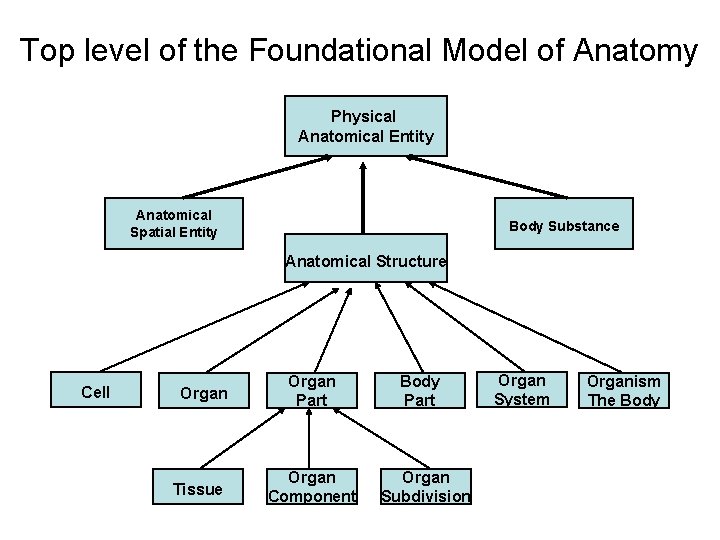
Top level of the Foundational Model of Anatomy Physical Anatomical Entity Anatomical Spatial Entity Body Substance Anatomical Structure Cell Organ Tissue Organ Part Organ Component Body Part Organ Subdivision Organ System Organism The Body
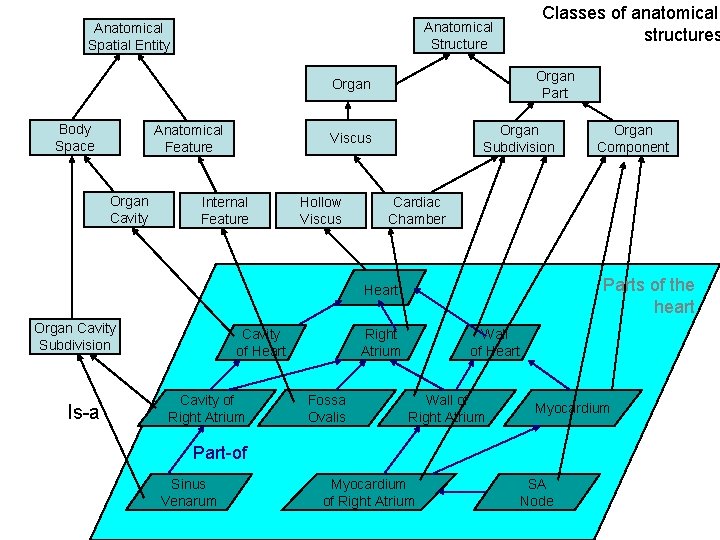
Anatomical Structure Anatomical Spatial Entity Organ Part Organ Body Space Anatomical Feature Organ Cavity Organ Subdivision Viscus Internal Feature Hollow Viscus Classes of anatomical structures Cardiac Chamber Parts of the heart Heart Organ Cavity Subdivision Is-a Cavity of Heart Cavity of Right Atrium Fossa Ovalis Wall of Heart Wall of Right Atrium Myocardium Part-of Sinus Venarum Organ Component Myocardium of Right Atrium SA Node

We really want ontologies in electronic form • Ontology contents can be processed and interpreted by computers • Interactive tools can assist developers in ontology authoring

The FMA demonstrates that distinctions are not universal • Blood is not a tissue, but rather a body substance (like saliva or sweat) • The pericardium is not part of the heart, but rather an organ in and of itself • Each joint, each tendon, each piece of fascia is a separate organ These views are not shared by many anatomists!

Creating Ontologies in Machine-Processable Form • Provides a mechanism for developers to codify salient distinctions about the world or some application area • Provides a structure for knowledge bases that can enable – – Information retrieval Information integration Automated translation Decision support

Ontologies are cropping up everywhere! • Indexing of online information for access by humans or search engines • Reference terminologies for machine translation and data interchange • Standard terms for describing experimental data • Frameworks for structuring knowledge for decision support

The New Philosophers • Categorizing what exists in machineunderstandable form • Providing a structure that enables – developers to locate and update relevant descriptions – computers to infer relationships and properties – quantitative data to be annotated in such a way as to become available for semantically meaningful search

Lots of ontology builders are not very good philosophers • Nearly always, ontologies are created to address pressing practical needs • The people who have the most insight into professional knowledge of a given biomedical domain may have little appreciation for metaphysics, principles of knowledge representation, or computational logic • There simply aren’t enough good philosophers to go around

A case in point: The International Classification of Diseases • An enumeration of diseases that forms the basis for all medical claims and reimbursements in the United States • A “legacy” terminology that has its roots in 19 th century epidemiology • Created initially by biostatisticians with a pressing need to compare death statistics in different European countries • A system that won’t go away—and yet we would never create anything like it again

The notion of disease emerged in the 17 th century • Thomas Sydenham: “The selfsame phenomena that you observe in the sickness of a Socrates you would observe in the sickness of a simpleton” – Description of a plant applies to its species; – Description of disease should be similar • Over the next 100 years, diverse classification systems for diseases emerged • Emphasis became to understand disease in terms of associate pathological findings at autopsy

The Origins of the International Classification of Diseases • First International Statistical Congress, 1853, initially commissioned development of a nomenclature for coding causes of death • The resulting nomenclature was revised in 1855, 1864, 1874, 1880, 1886, with continued confusion as to how diseases should be classified • In 1891, Jacques Bertillon generated a synthesis organized largely anatomically; adopted by the International Statistical Institute in 1893 as the International List of Causes of Death

The ICD becomes an International Standard • In 1948, the World Health Organization became responsible for the International List of Causes of Death • Nonfatal diseases were then added to the list, in the course of multiple revisions to what then became the ICD • The Ninth revision, ICD-9 was released in 1977 • In 1978, after “clinical modification, ” ICD-9 -CM became a source taxonomy “for statistics concerning the planning, monitoring, and evaluation of health services”

A Small Portion of ICD 9 -CM 724. 01 724. 02 724. 09 724. 1 724. 2 724. 3 724. 4 724. 5 724. 6 724. 70 724. 71 724. 8 724. 9 Unspecified disorders of the back Spinal stenosis, other than cervical Spinal stenosis, unspecified region Spinal stenosis, thoracic region Spinal stenosis, lumbar region Spinal stenosis, other Pain in thoracic spine Lumbago Sciatica Thoracic or lumbosacral neuritis Backache, unspecified Disorders of sacrum Disorders of coccyx Unspecified disorder of coccyx Hypermobility of coccyx Coccygodynia Other symptoms referable to back Other unspecified back disorders
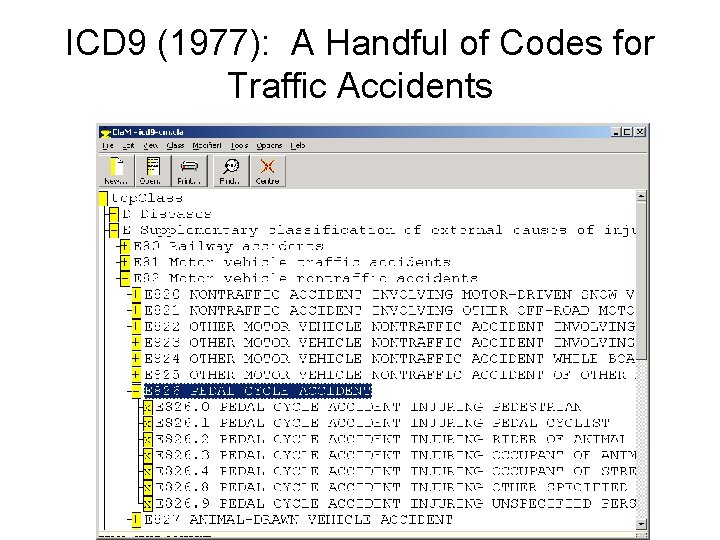
ICD 9 (1977): A Handful of Codes for Traffic Accidents

ICD 10 (1999): 587 codes for such accidents • V 31. 22 Occupant of three-wheeled motor vehicle injured in collision with pedal cycle, person on outside of vehicle, nontraffic accident, while working for income • W 65. 40 Drowning and submersion while in bath-tub, street and highway, while engaged in sports activity • X 35. 44 Victim of volcanic eruption, street and highway, while resting, sleeping, eating or engaging in other vital activities

ICD is used for lots of (too many? ) things! • ICD is used to code all patient encounters with the health-care system for purposes of – – – Billing and reimbursement Institutional planning Disease surveillance and public health Quality assurance Economic modeling by third-party payors • ICD was never intended to make the distinctions relevant to all these tasks!

If real ontologists could build the ICD from scratch … • Diseases would be organized with well-defined relationships • Diseases would be associated with computerunderstandable definitions • There would be well-defined rules for ensuring that descriptions are sensible • There would be well-defined mechanisms for creating use-specific views of the ICD • There would be a well-defined path to integration with bioinformatics resources that describe the molecular underpinnings of disease

The components of ontologies • Classes: The primary entities in the world being models (e. g. , “organ”) • Attributes: The properties of classes (e. g. , “shape”, “location”) • Relations: Statements regarding how one class may relate to others (e. g. , “the heart” is-a “organ”) • Axioms: More complex logical statements (e. g. , “only paired organs can be left-sided or right-sided”)
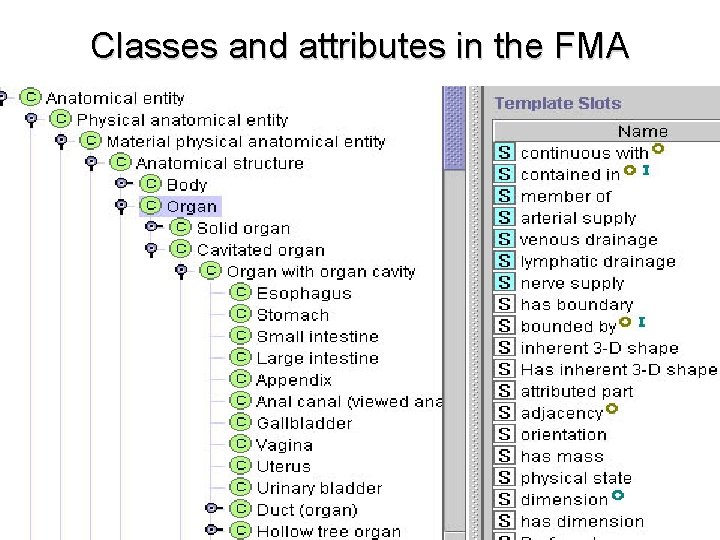
Classes and attributes in the FMA
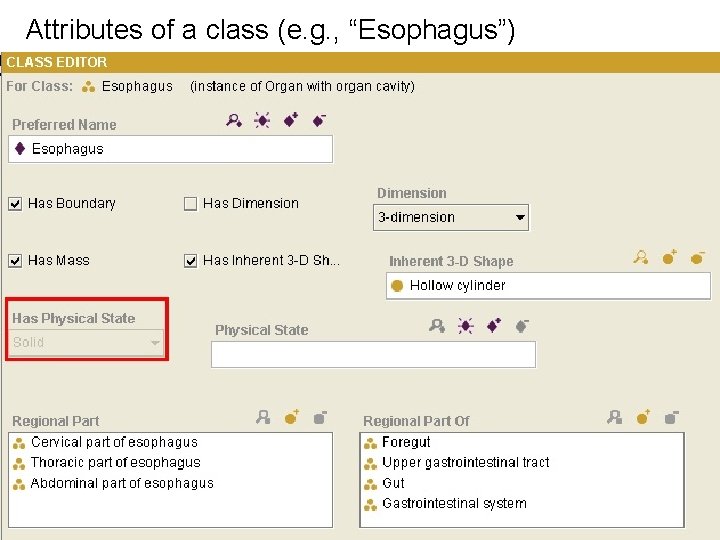
Attributes of a class (e. g. , “Esophagus”)

is-a is a special relation If a sub-class is-a member of a super-class, then – every instance of the subclass is also an instance of the super-class (e. g. , every member of the set aorta is necessarily a member of the set artery) – values of attributes of the super-class are inherited by every instance of the subclass (e. g. , if arteries have cylindrical shape, then aorta has cylindrical shape)

Modeling part-of relationships is tricky • Inheritance is not necessarily transitive – In an is-a relation, if a stomach is an organ and an organ has a volume, then a stomach has volume – In a part-of relation, if an eyebrow is part of the head and the head has a volume, then does an eyebrow have a volume? • There are many kinds of part-of relationships, each with slightly different semantics

Kinds of part-of relationships (after Winston and Odell) • • Component (e. g. , handle of a car door) Stuff (e. g. , flour in bread) Portion (e. g. , a slice from a loaf of bread) Area (e. g. , city in a country) Member (e. g. , ship in a fleet of ships) Partner (e. g. , Laurel in Laurel & Hardy) Piece (e. g. , handle when removed from the door)

“Frame-based” knowledgerepresentation systems • Allow developers to encode – Taxonomic hierarchies of classes – Other relations among classes (e. g. , “part-of”) in addition to the is-a hierarchy – Attributes of classes that take on particular values to define instances of the classes • Support inheritance of attributes and values along taxonomic relations

Distinctions about ontologies • “Light” versus “heavy”: Is the ontology a simple taxonomy or does the ontology additional detail regarding the nature of classes? • “Upper-level” versus “domain-oriented”: Does the ontology try to describe general, abstract concepts or concepts tied to a particular application area?

Suggested Upper Merged Ontology (SUMO)
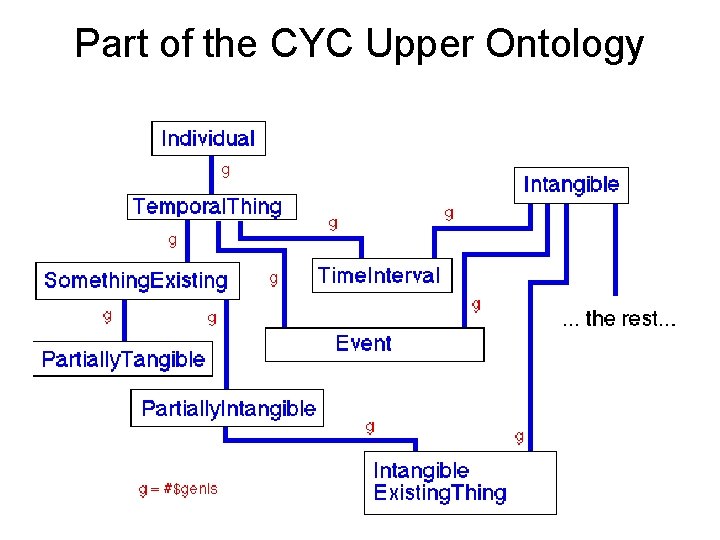
Part of the CYC Upper Ontology

The CYC Project • Started in 1984 by Doug Lenat • Original goal was to encode all the knowledge in the Encyclopedia Britannica • Now the goal is to encode all the commonsense knowledge needed to read the Encyclopedia Britannica • Blurs the distinction between ontology and knowledge base • Has formed the foundation for several commercial biomedical applications

Some CYC Definitions #$Cancer isa: #$Physiological. Condition. Type genls: #$Ailment. Condition #$Terminal. Physiological Condition #$Tumor isa: #$Existing. Object. Type genls: #$Biological. Living. Object #$Infection isa: #$Physiological. Condition. Type genls: #$Ailment. Condition

Basic Formal Ontology An example of a top-level ontology being used in biomedicine to structure ontologies for specific domains that are marked by internal coherence and external interoperability
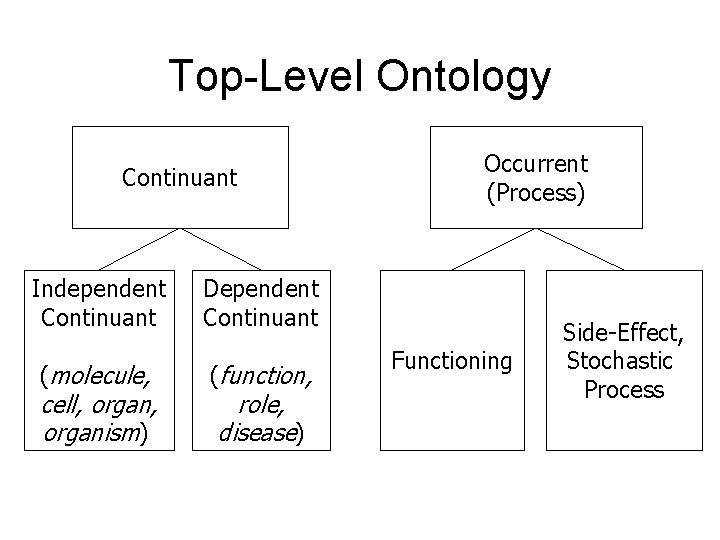
Top-Level Ontology Continuant Independent Continuant Dependent Continuant (molecule, (function, cell, organism) role, disease) Occurrent (Process) Functioning Side-Effect, Stochastic Process
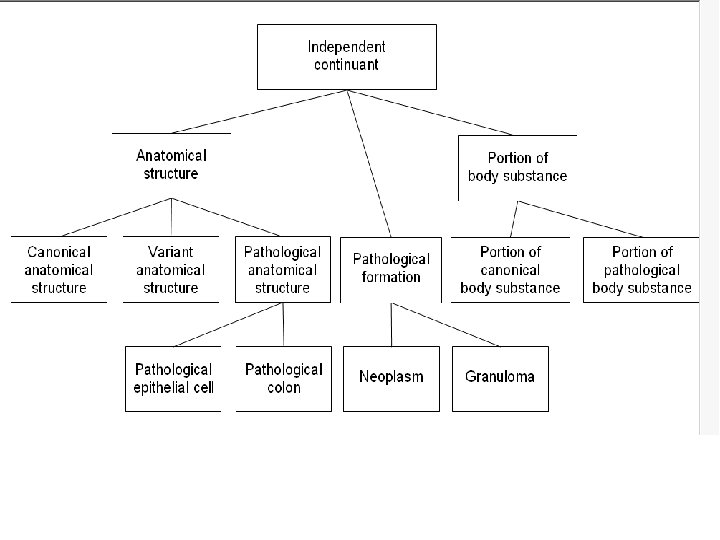

Distinctions among ontologies • “Light” versus “heavy”: Is the ontology a simple taxonomy or does the ontology provide additional detail regarding the nature of entities? • “Upper-level” versus “domain-oriented”: Does the ontology try to describe general, abstract entities or entities tied to a particular application area?

Taxonomies are “Light-Weight” Ontologies 724. 01 724. 02 724. 09 724. 1 724. 2 724. 3 724. 4 724. 5 724. 6 724. 70 724. 71 724. 8 724. 9 Unspecified disorders of the back Spinal stenosis, other than cervical Spinal stenosis, unspecified region Spinal stenosis, thoracic region Spinal stenosis, lumbar region Spinal stenosis, other Pain in thoracic spine Lumbago Sciatica Thoracic or lumbosacral neuritis Backache, unspecified Disorders of sacrum Disorders of coccyx Unspecified disorder of coccyx Hypermobility of coccyx Coccygodynia Other symptoms referable to back Other unspecified back disorders

“Heavy weight” ontologies make explicit: • Relationships among entities (e. g. , is-akind-of; is-a-part-of) • Properties of entities (e. g. , all organs have the property size) • Constraints on relationships and properties (e. g. , only organs are paired may have laterality)
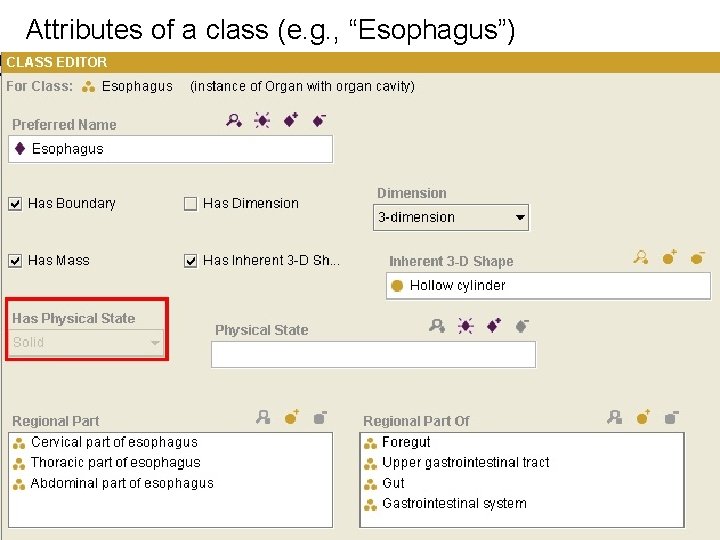
Attributes of a class (e. g. , “Esophagus”)

The story so far … • Ontologies define the entities—and relationships among entities—in some application area • The authors’ point of view determines which distinctions are appropriate in a particular ontology • Ontologies often use frame-based representations (including classes, attributes, relationships, and axioms) to encode knowledge • People are building ontologies for nearly every niche of biomedicine
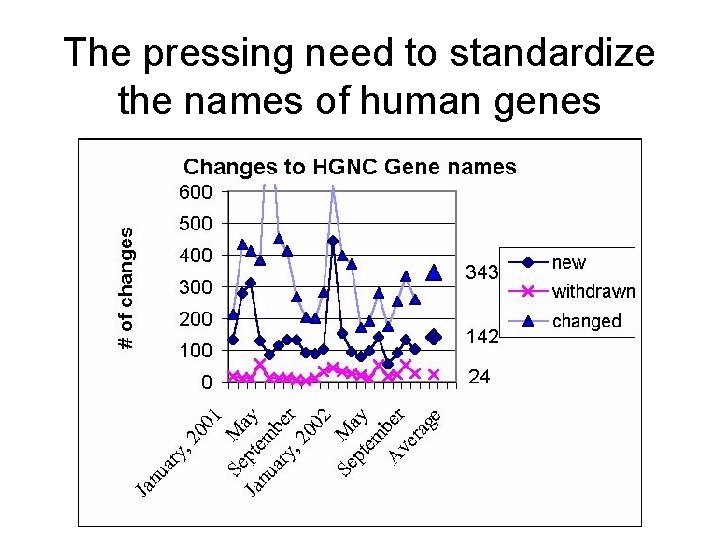
The pressing need to standardize the names of human genes

But the human genome is only part of the problem … • Biologists maintain huge databases of gene sequences and gene expression for a wide range of “model organisms” (e. g. , mouse, rat, yeast, fruit fly, round worm, slime mold) • Database entries are annotated with entries such as the name of a gene, the function of the gene, and so on • How do you ensure uniformity of these annotations?

Gene Ontology Consortium • Founded in 1998 as a collaboration among scientists responsible for developing different databases of genomic data for model organisms (fruit fly, yeast, mouse) • Now, essentially all developers of all model-organism databases participate • Goal: To produce a dynamic, controlled vocabulary that can be applied to all organism databases even as knowledge of gene and protein roles in cells is accumulating and changing

Gene Ontology (GO) • Comprises three independent “ontologies” – molecular function of gene products – cellular component of gene products – biological process representing the gene product’s higher order role. • Uses these terms as attributes of gene products in the collaborating databases (gene product associations) • Allows queries across databases using GO terms, providing linkage of biological information across species

GO = Three Ontologies • Molecular Function – elemental activity or task – example: DNA binding • Cellular Component – location or complex – example: cell nucleus • Biological Process – goal or objective within cell – example: secretion
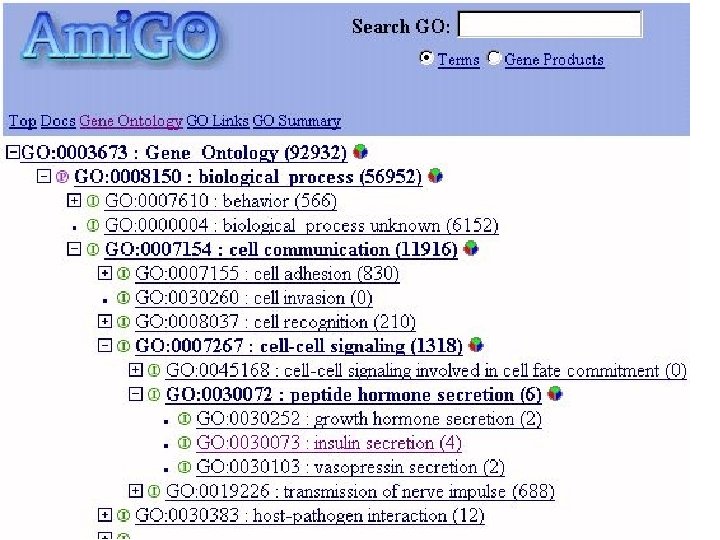

GO has been wildly successful!! • Dozens of biologists around the world contribute to GO on a regular basis • The ontology is updated every 30 minutes! • It’s now impossible to work in most areas of computational biology without making use of GO terms

But GO has had real problems … • Ontologies initially were represented in an idiosyncratic format that was not compatible with standard knowledge-representation systems (DAG-Edit) • The format was based on directed acyclic graphs of concepts, without the general ability to specify machine interpretable properties of concepts or definitions of concepts • Because of the informal knowledge-representation system, lots of errors crept into GO – Terms that were duplicated in different places – Terms with no superclasses – Uncertain relationships between terms • The GO consortium is working hard to rectify these problems by means of a new representation (OBO-Edit) and enhanced quality control
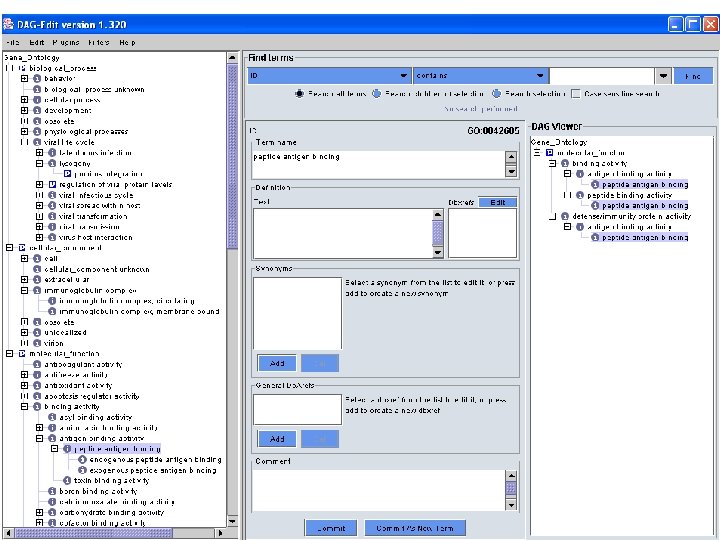

Tension in the GO Community • Biologists around the world who have pressing needs to integrate research databases work together to add terms to GO nearly continuously • Computer scientists bemoan a tendency toward ad-hoc-ery and worry that GO eventually will become unusable and unmaintainable

A wonderful keynote talk from the meeting on Standards and Ontologies for Functional Genomics in 2004 The Capulets and Montagues A plague on both your houses? Professor Carole Goble University of Manchester, UK Warning: This talk contains sweeping generalisations

Carole Goble Prologue Two households, both alike in dignity, In fair genomics, where we lay our scene, (One, comforted by its logic’s rigour, Claims ontology for the realm of pure, The other, with blessed scientist’s vigour, Acts hastily on models that endure), From ancient grudge break to new mutiny, When “being” drives a fly-man to blaspheme. From forth the fatal loins of these two foes Researchers to unlock the book of life; Whole misadventured piteous overthrows Can with their work bury their clans’ strife. The fruitful passage of their GO-mark'd love, And the continuance of their studies sage, Which, united, yield ontologies undreamed-of, Is now the hours' traffic of our stage; The which if you with patient ears attend, What here shall miss, our toil shall strive to mend. Based on an idea by Shakespeare

Creating ontologies has become a widespread cottage industry • Professional Societies – HL 7: Reference Information Model – MGED: Microarray Gene Expression Data Society Ontology – HUPO: Human Protein Organization Ontology • Government – NCI Thesaurus – NIST: Process Specification Language • Open Biological Ontologies – GO – Three dozen (and growing) other ontologies – Mostly in DAG-Edit, some in Protégé format

A Portion of the OBO Library
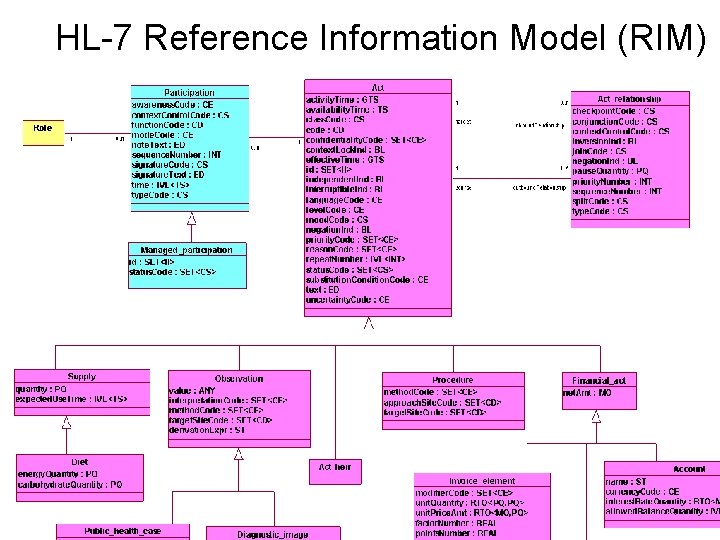
HL-7 Reference Information Model (RIM)

HL 7 RIM • Provides a uniform framework for specification of information required by health-care information systems • Based on six top-level, very general classes: Act, Entity, Role, Participation, Act_relationship, and Role_link • Designed to facilitate information exchange among distributed elements of clinical information systems • Has the same limitations that all “upper level” ontologies share: – Abstract concepts are hard to define – It’s hard to know what should be “in” and what should be “out”

Barry Smith cites the RIM as an example of “current chaos” in ontology development • Animal Definition: A subtype of Living Subject representing any animal-of-interest to the Personnel Management domain. • Living. Subject Definition: A subtype of Entity representing an organism or complex animal, alive or not.

How to impose order on the chaos • Use of upper-level ontologies (e. g. , SUMO) • Use of modularization (e. g. , GO) • Use rich knowledge-representation systems to increase interconnections (e. g. , CYC) • Use standardized relations among classes (e. g. , new entries in the OBO library) • Use description logic to clarify inferred relationships (e. g. , NCI Thesaurus)
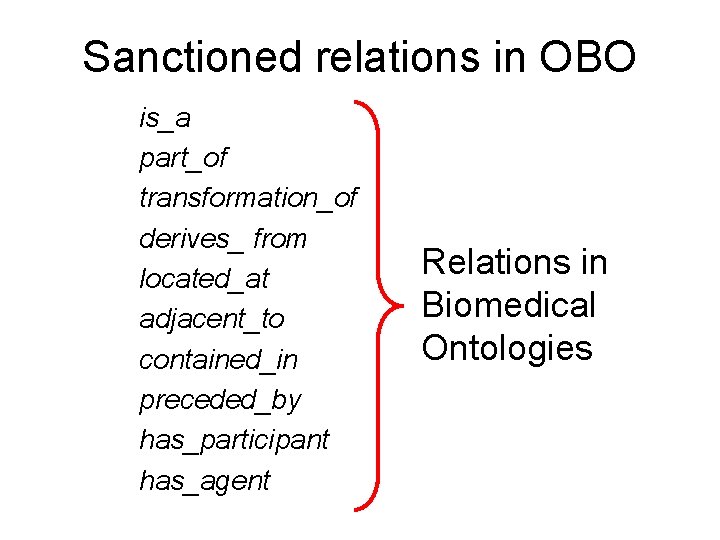
Sanctioned relations in OBO is_a part_of transformation_of derives_ from located_at adjacent_to contained_in preceded_by has_participant has_agent Relations in Biomedical Ontologies

NCI Enterprise Vocabulary Services 1997: R. Klausner, Director NCI, wanted a “science management system” • Know about everything funded by NCI • Goals and results – “bench to bedside” - Thereby improve and speed translation of research Approach: 1. Create integrative terminology 2. Evolve terminology scope from supporting grants management to supporting science 3. Build Web-accessible infrastructure – ca. CORE


More than 37, 000 concepts are represented with extremely detailed granularity in many areas
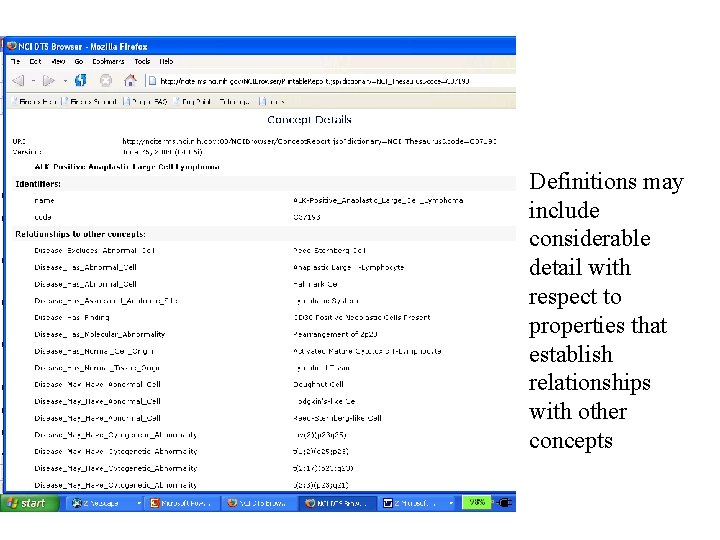
Definitions may include considerable detail with respect to properties that establish relationships with other concepts
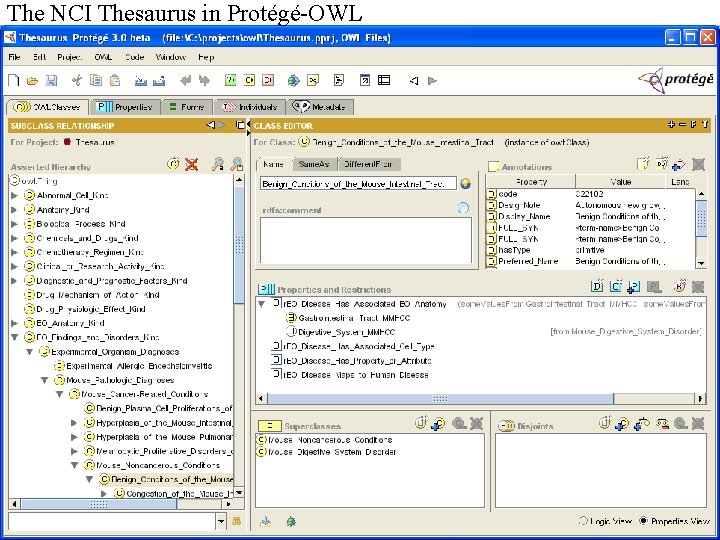
The NCI Thesaurus in Protégé-OWL

NCI uses an elaborate process for editing and maintenance

Description Logic (DL) • A subset of logic designed to focus on categories and their definitions in terms of existing relations • More expressive than frame-based representations systems (as in FMA) but less expressive than first-order logic (as in CYC) • Major inference tasks: – Subsumption Is category C 1 a subset of C 2? – Classification Does Object O belong to C?

Kinds of concepts • Defined – Have explicit necessary and sufficient properties – Often are specializations of primitive concepts • Primitive – Have no sufficient properties – May have other, necessary properties – Correspond to natural kinds
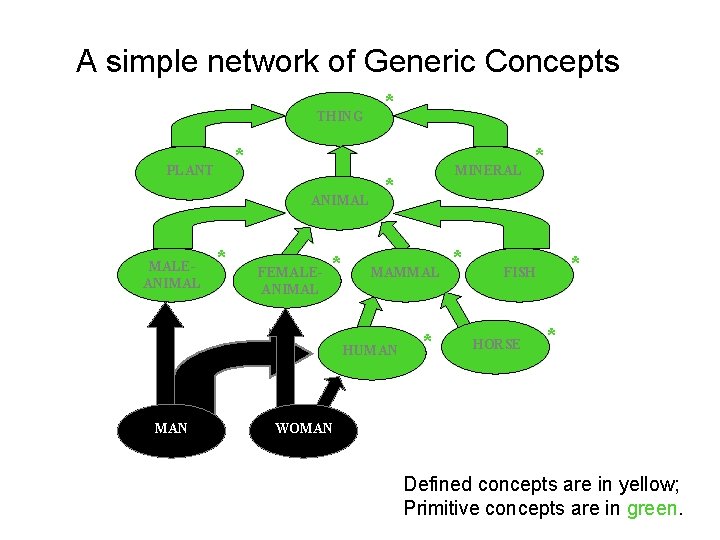
A simple network of Generic Concepts THING * PLANT ANIMAL MALEANIMAL * * FEMALEANIMAL * * MAMMAL HUMAN MINERAL * * FISH HORSE * WOMAN Defined concepts are in yellow; Primitive concepts are in green.

A classifier is a program that can use DL to conclude: • • All WOMEN are FEMALE ANIMALS A HORSE may not also be a PLANT HUMAN subsumes MAN and WOMAN A MAN may not also be a WOMAN
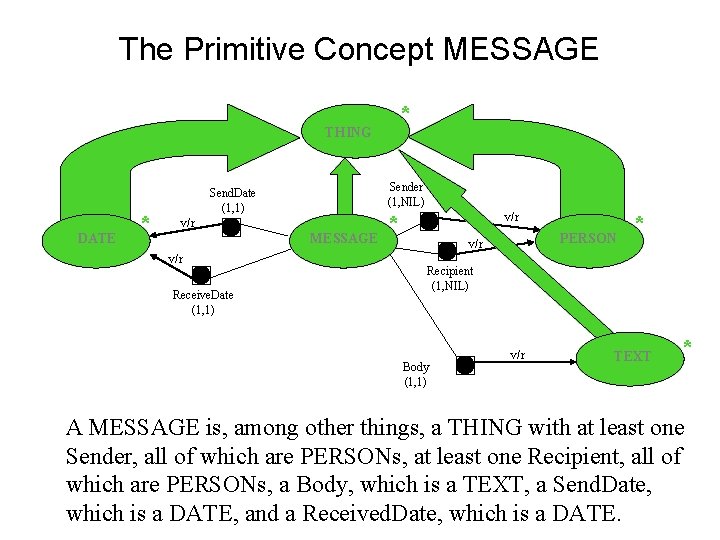
The Primitive Concept MESSAGE * THING DATE * Sender (1, NIL) Send. Date (1, 1) v/r MESSAGE v/r Receive. Date (1, 1) * v/r PERSON v/r * Recipient (1, NIL) Body (1, 1) v/r TEXT * A MESSAGE is, among other things, a THING with at least one Sender, all of which are PERSONs, at least one Recipient, all of which are PERSONs, a Body, which is a TEXT, a Send. Date, which is a DATE, and a Received. Date, which is a DATE.
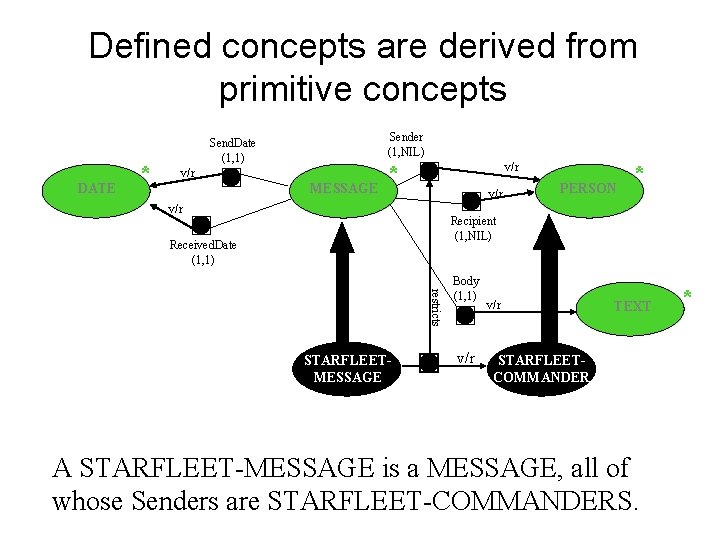
Defined concepts are derived from primitive concepts DATE * Sender (1, NIL) Send. Date (1, 1) v/r MESSAGE * v/r v/r PERSON * Recipient (1, NIL) Received. Date (1, 1) restricts STARFLEETMESSAGE Body (1, 1) v/r TEXT STARFLEETCOMMANDER A STARFLEET-MESSAGE is a MESSAGE, all of whose Senders are STARFLEET-COMMANDERS. *

A DL Classifier • Takes a new Concept and automatically determines all subsumption relations between it and all other Concepts in the network • Adds new links when new subsumption relations are discovered • Automates the placement of new Concepts in the taxonomy
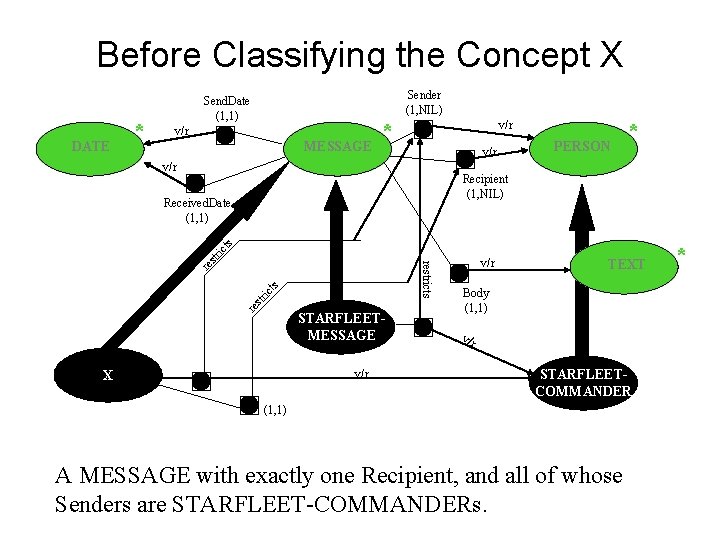
Before Classifying the Concept X DATE * Sender (1, NIL) Send. Date (1, 1) v/r MESSAGE * v/r v/r PERSON * Recipient (1, NIL) re str ict s re restricts str i cts Received. Date (1, 1) STARFLEETMESSAGE v/r X v/r TEXT Body (1, 1) v/r STARFLEETCOMMANDER (1, 1) A MESSAGE with exactly one Recipient, and all of whose Senders are STARFLEET-COMMANDERs. *
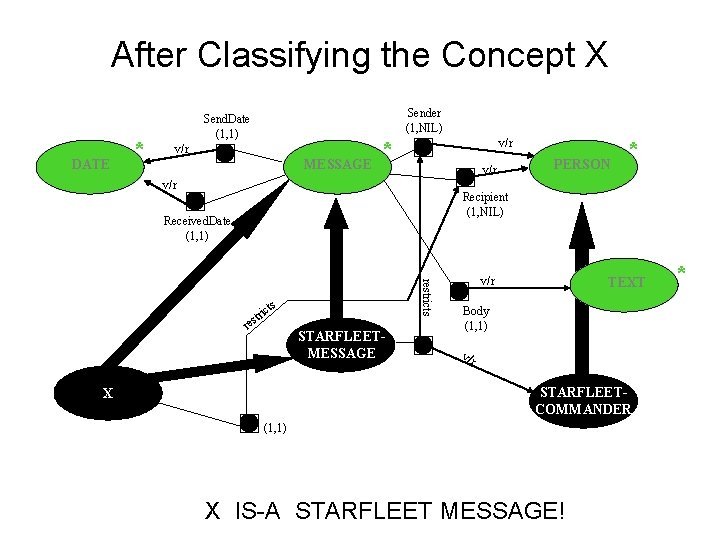
After Classifying the Concept X DATE * Sender (1, NIL) Send. Date (1, 1) v/r MESSAGE * v/r v/r PERSON * Recipient (1, NIL) Received. Date (1, 1) restricts s ict tr res STARFLEETMESSAGE v/r TEXT Body (1, 1) v/r STARFLEETCOMMANDER X (1, 1) X IS-A STARFLEET MESSAGE! *

The Beauty of Classification for Ontologies • The classifier takes care of where to place a new concept in the hierarchy • All inheritance relationships are automatically propagated to the new concept • Relationships among a new concept and other entities are automatically simplified by classifying the new concept as a specialization of existing concepts
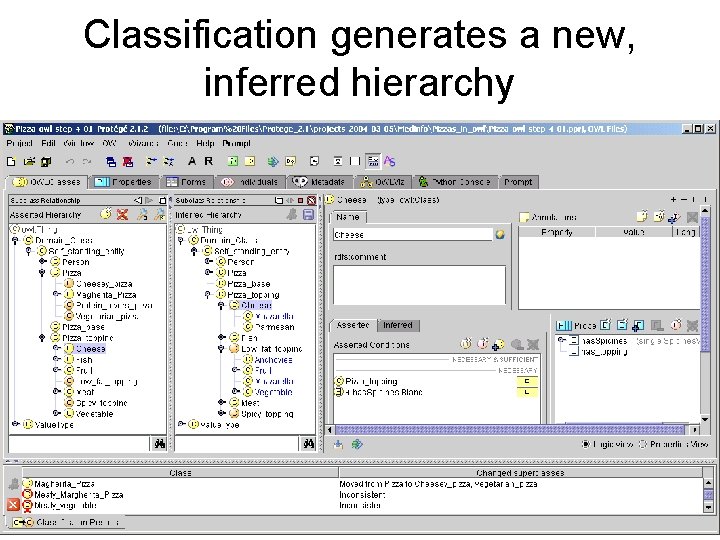
Classification generates a new, inferred hierarchy

The Ontology Web Language (OWL) • Comes in three flavors: – OWL Lite (frame-based) – OWL DL (decription logic) – OWL Full (first-order logic and then some) • Rapidly being adopted for use in biomedical ontologies, including: – NCI Thesaurus (cancer biology and oncology) – MGED Ontology (DNA micro-array experiments) – Bio. PAX (metabolic pathways) • The new editor and representation system for OBO ontologies (OBO-Edit) uses a subset of OWL

DL and Ontologies • There is not just one “description logic”; DLs come in different varieties with different expressivity • DLs are of value primarily to ontology developers, to see the implications of modeling decisions • DLs also can be used by end users, when reasoning about systems that ontologies model
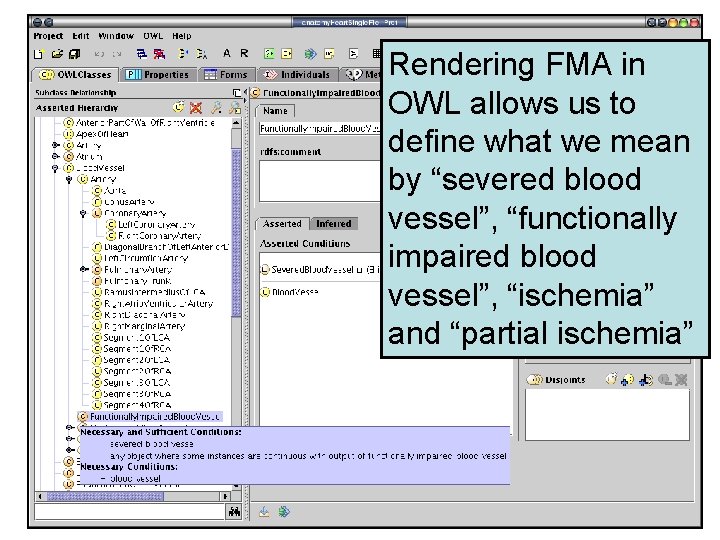
Rendering FMA in OWL allows us to define what we mean by “severed blood vessel”, “functionally impaired blood vessel”, “ischemia” and “partial ischemia”
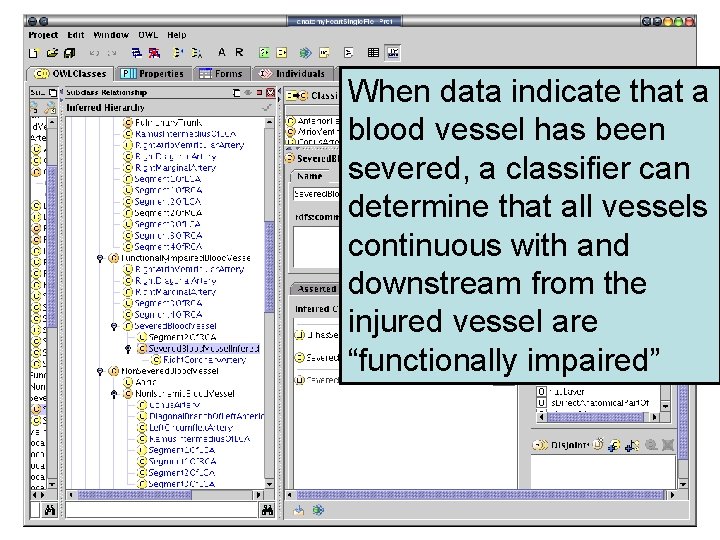
When data indicate that a blood vessel has been severed, a classifier can determine that all vessels continuous with and downstream from the injured vessel are “functionally impaired”
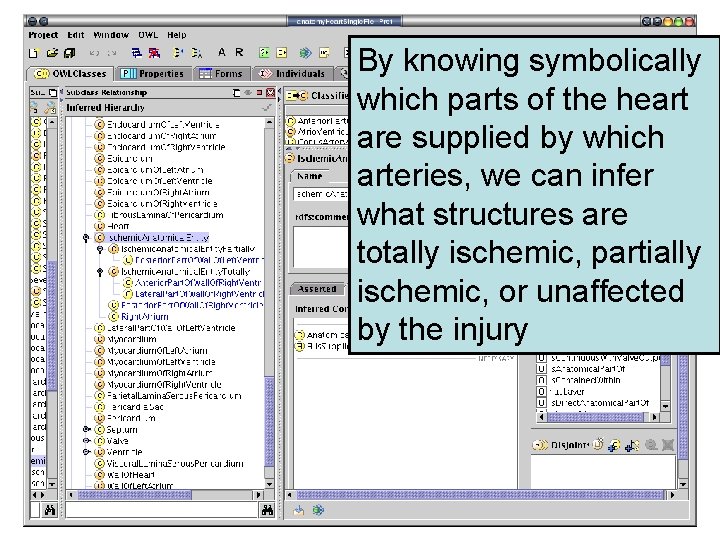
By knowing symbolically which parts of the heart are supplied by which arteries, we can infer what structures are totally ischemic, partially ischemic, or unaffected by the injury

The story so far … • Everyone and his brother is working on new ontologies for biomedicine • There is increasing awareness of the need to overcome flaws in modeling and representation • Use of Description Logics has been advanced as a means to identify modeling errors and to improve the quality of ontologies • OWL is being promoted as a knowledgerepresentation standard and is getting considerable traction
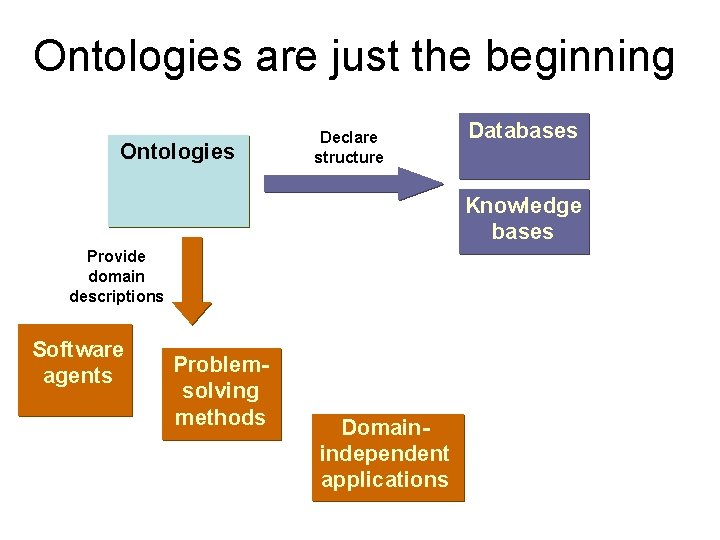
Ontologies are just the beginning Ontologies Declare structure Databases Knowledge bases Provide domain descriptions Software agents Problemsolving methods Domainindependent applications

Goals of Biomedical Ontologies • • • To provide a classification of biomedical entities To summarize and annotate data To mediate among different social groups To mediate among different software components To simplify the engineering of complex software systems To provide a formal specification of biomedical knowledge

Classification of biomedical entities • To classify is human … • But biomedicine did not really get into the act until Linnaeus and the advent of the ICD • The classifications that drive much of health care do not describe natural kinds: LOINC, CPT, ICD, DSM • Many classifications have huge societal implications – “Premenstrual syndrome” and “Homosexuality” as disorders in DSM – “Menopause” as a disease in ICD

Racial classifications under apartheid reinforced perceived biological “truths” • • Europeans Asiatics Persons of mixed race (coloureds) Bantus – Xhosa – Zulu – and six other groups …

Many classifications • Enforce or preserve existing social conventions • Are motivated because making particular distinctions is to the advantage of some social group • Reinforce or even create a perceived “reality” by legitimizing certain distinctions

The Proliferation of Nursing Vocabularies • International Classification of Nursing Practice (ICNP) • Nursing Intervention Lexicon and Taxonomy • The Omaha System: Nursing diagnoses, interventions, and clinical outcomes • Nursing Interventions Classifications

Some classes from the Nursing Intervention Classification • Cultural Brokerage – Bridging, negotiating, or linking the orthodox health care system with a patient and family of a different culture • Spiritual support – Assisting the patient to feel balance and connection with a greater power • Humor – Facilitating the patient to perceive, appreciate, and express what is funny, amusing, or ludicrous in order to establish relationships, relieve tension, release anger, facilitate learning, or cope with personal feelings

Goals of Biomedical Ontologies • • • To provide a classification of biomedical entities To summarize and annotate data To mediate among different social groups To mediate among different software components To simplify the engineering of complex software systems To provide a formal specification of biomedical knowledge

Summarization and annotation of data • Biologists generally don’t care about modeling reality beyond their data • Biologists care about – Making sense of terabytes of data – Accessing and indexing data – Comparing data sets with one another • The goal is to create annotations that make distinctions about the data, not about the world
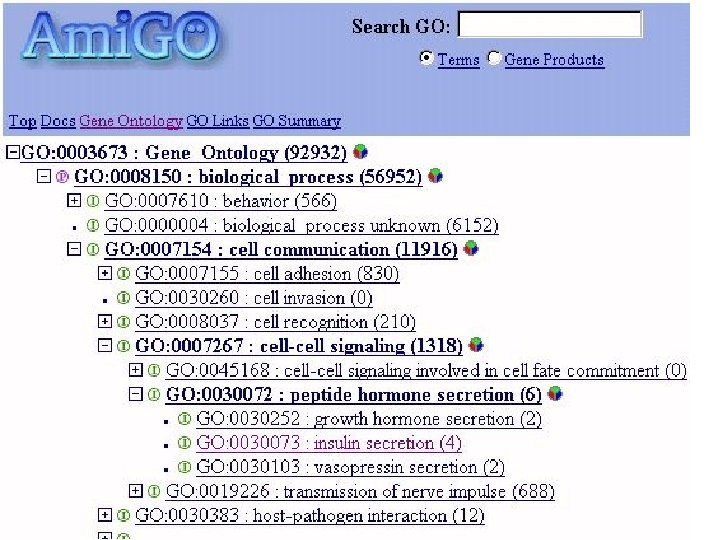

Goals of Biomedical Ontologies • • • To provide a classification of biomedical entities To summarize and annotate data To mediate among different social groups To mediate among different software components To simplify the engineering of complex software systems To provide a formal specification of biomedical knowledge

Mediation among different social groups • ICD and CPT codes propagate from clinicians to healthcare organizations to payors to epidemiologists to policy makers • Within institutions, ontologies provide the basis for getting our work done. – No coded lab test results, no treatment – No ICD code, no reimbursement

Goals of Biomedical Ontologies • • • To provide a classification of biomedical entities To summarize and annotate data To mediate among different social groups To mediate among different software components To simplify the engineering of complex software systems To provide a formal specification of biomedical knowledge

Mediation among different software components • HL 7 was founded to get individual departmental information systems to talk to one another • The implied ontology is one of messages, not of entities in the real world • The HL 7 message standard builds on longstanding work on machine interoperability

An ontology for CAD/CAM: STEP • Provides an international standard for interacting computer-aided design and manufacturing applications • Defines over 1300 classes of objects, addressing areas such as – – Geometry and topology Product configuration Form features Tolerances

Sample STEP Class Definition ENTITY part_model SUBTYPE OF (design_model); nominal_shape: model_units: part_features: part_tolerances: equivalents: shape_model; units; OPTIONAL LIST (0: #) OF form_features; OPTIONAL LIST (0: #) OF shape_tolerances; OPTIONAL LIST (0: #) OF part_model_structure; WHERE NOT (part_model IN equivalents. model_element); END ENTITY;

Goals of Biomedical Ontologies • • • To provide a classification of biomedical entities To summarize and annotate data To mediate among different social groups To mediate among different software components To simplify the engineering of complex software systems To provide a formal specification of biomedical knowledge

Engineering of complex software systems • Object-oriented design and programming is well entrenched in current software-engineering practices • OOP owes considerable legacy to frame -based knowledge-representation systems developed in AI in the 1970 s • Ontologies are now at the core of advanced software engineering
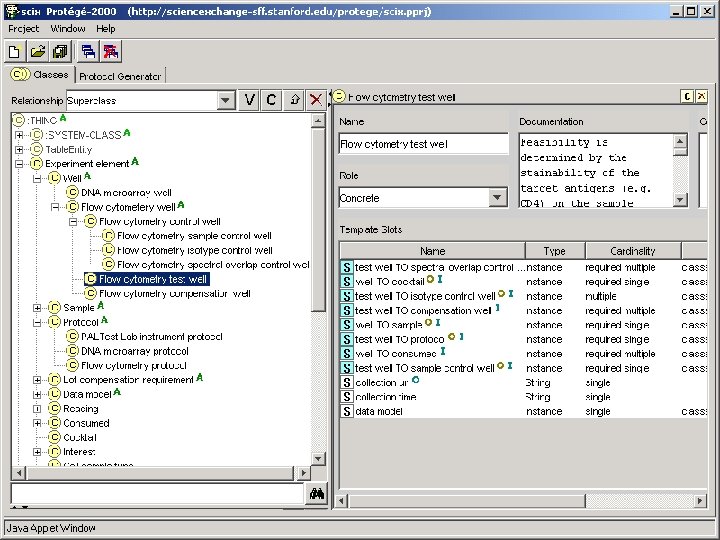
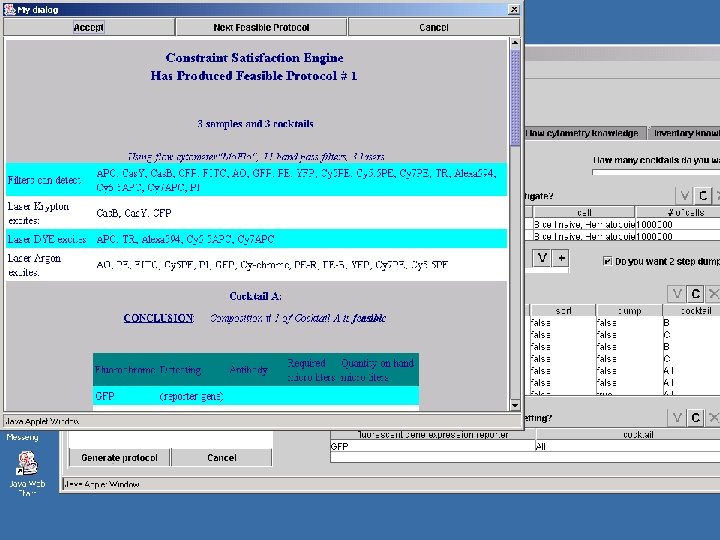


Model-Directed Architecture • Platform-independent UML model contains elements of – Domain ontology (in terms of UML classes) – Problem-solving methods (in terms of pre- and postconditions and action semantics) • CASE tools (under development) perform transformation of UML model into program code • MDA does not cleanly separate domain ontology from problem-solving components • Platform-independent models have potential to outlive particular software implementations

Goals of Biomedical Ontologies • • • To provide a classification of biomedical entities To summarize and annotate data To mediate among different social groups To mediate among different software components To simplify the engineering of complex software systems To provide a formal specification of biomedical knowledge

Formal specification of biomedical knowledge • In the “information society, ” there will be increasing motivation for representing human knowledge in machineprocessable form • Ontologies of professional knowledge are being seen as having value even for their own sake
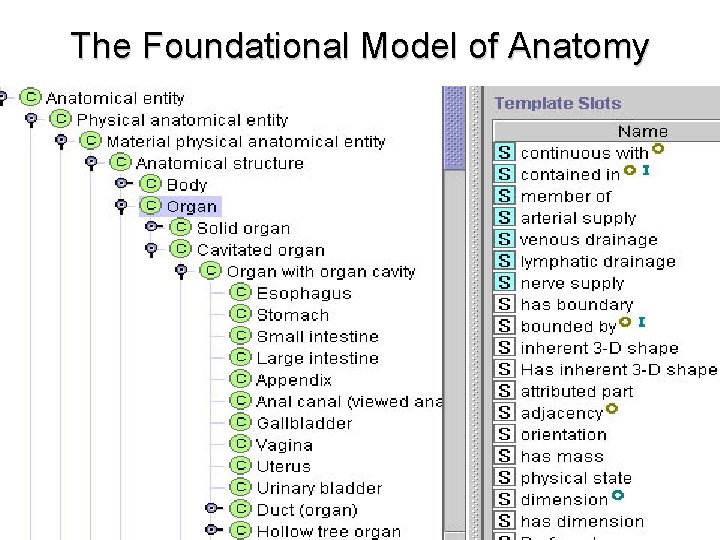
The Foundational Model of Anatomy

Goals of Biomedical Ontologies • • • To provide a classification of biomedical entities To summarize and annotate data To mediate among different social groups To mediate among different software components To simplify the engineering of complex software systems To provide a formal specification of biomedical knowledge

Ontologies are built with purposes in mind • These purposes reflect scientific, political, social, economic, and engineering goals that have little to do with metaphysics • Making explicit these additional considerations will lead to “purer” and more useful ontologies—at the risk of exposing issues that developers might rather leave buried

A Portion of the OBO Library

Ontologies need metadata • • • To clarify intentions of developers To define anticipated context of use To aid collaborative development To enable users to provide feedback Ultimately, to support formal peer review

Ontologies are not like journal articles • It is difficult to judge methodological soundness simply by inspection • We may wish to use an ontology even though some portions – Are not well designed – Make distinctions that are different from those that we might want

Ontologies are not like journal articles II • The utility of ontologies – Depends on the task – May be highly subjective • The expertise and biases of reviewers may vary widely with respect to different portions of an ontology • Users should want the opinions of more than 2– 3 hand-selected reviewers • Peer review needs to scale to the entire user community
n o i lut hot o S s p a Sn

In an “open” rating system: • Anyone can annotate an ontology to say anything that one would like • Users can “rate the raters” to express preferences for those reviewers whom they trust • A “web of trust” may allow users to create transitive trust relationships to filter unwanted reviews

Possible Review Criteria • What is the level of user support? • What documentation is available? • What is the granularity of the ontology content in specific areas? • How well does the ontology cover a particular domain? • In what applications has the ontology been used successfully? Where has it failed?

The OBO Foundry • A proposal to create a family of interoperable gold standard biomedical reference ontologies • Formulated by Barry Smith and members of the GO Consortium • A Good Housekeeping Seal of Approval for biomedical contologies The OBO Foundry http: //obofoundry. org/

OBO Foundry Criteria I • • • The ontology is open and available to be used by all The ontology is in, or can be instantiated in, a common formal language The developers of the ontology agree in advance to collaborate with developers of other OBO Foundry ontology where domains overlap The OBO Foundry http: //obofoundry. org/

OBO Foundry Criteria II 1. The developers of each ontology commit to its maintenance in light of scientific advance, and to soliciting community feedback for its improvement 2. The developers commit to working with other Foundry members to ensure that, for any particular domain, there is community convergence on a single controlled vocabulary The OBO Foundry http: //obofoundry. org/

OBO Foundry Criteria III 1. The ontology has a clearly specified and clearly delineated content 2. The ontology possesses a unique identifier space within OBO 3. The ontology provider has procedures for identifying distinct successive versions 4. The ontology includes textual definitions and, where possible, equivalent formal definitions of its terms 5. The ontology has a plurality of independent users The OBO Foundry http: //obofoundry. org/

OBO Foundry Criteria IV 1. The ontology uses relations which are unambiguously defined following the pattern of definitions laid down in the OBO Relation Ontology The OBO Foundry http: //obofoundry. org/

The story so far … • Ontologies may be developed with many different goals in mind • Different goals may necessitate different modeling choices • Modern software engineering often requires ontology engineering • Evaluation and peer-review of ontology content pose significant challenges

Ontologies are meeting an urgent need • Ontologies are being developed by interested groups from every sector of academia, industry, and government • Many of these ontologies have been proven to be extraordinarily useful to wide communities • We finally have tools and representation languages that can enable us to create durable and maintainable ontologies with rich semantic content

The National Center for Biomedical Ontology • One of three National Centers for Biomedical Computing launched by NIH in 2005 • Collaboration of Stanford, Berkeley, Mayo, Buffalo, Victoria, UCSF, Oregon, and Cambridge • Primary goal is to make ontologies accessible and usable • Research will develop technologies for ontology indexing, alignment, and peer review
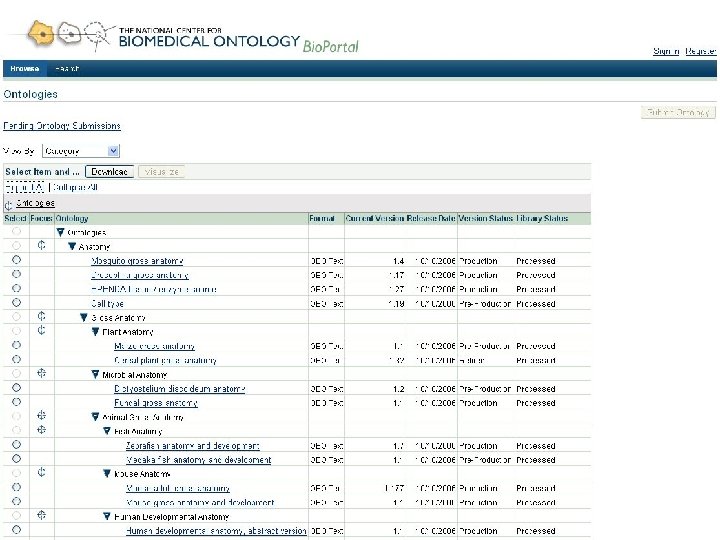
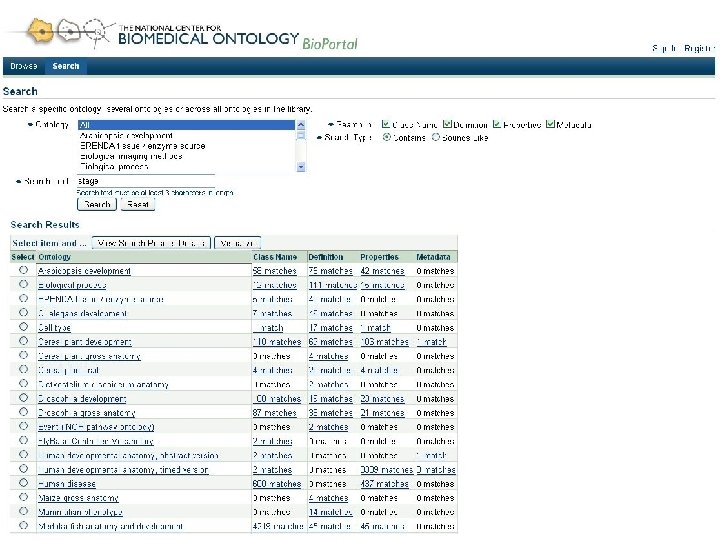
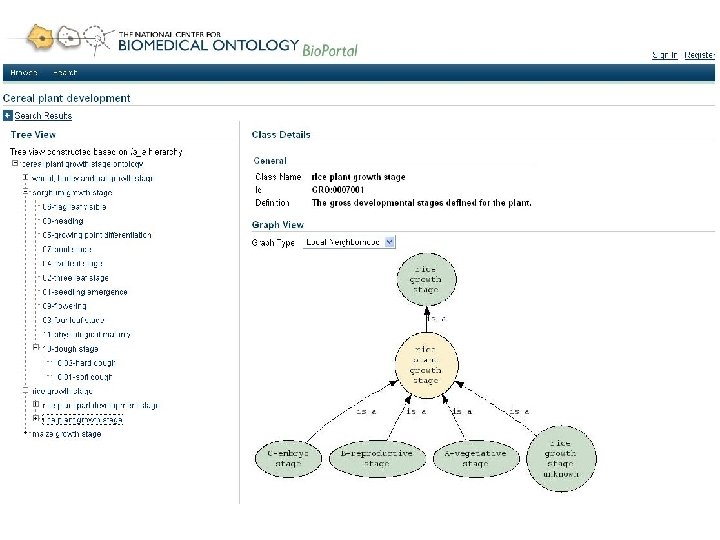
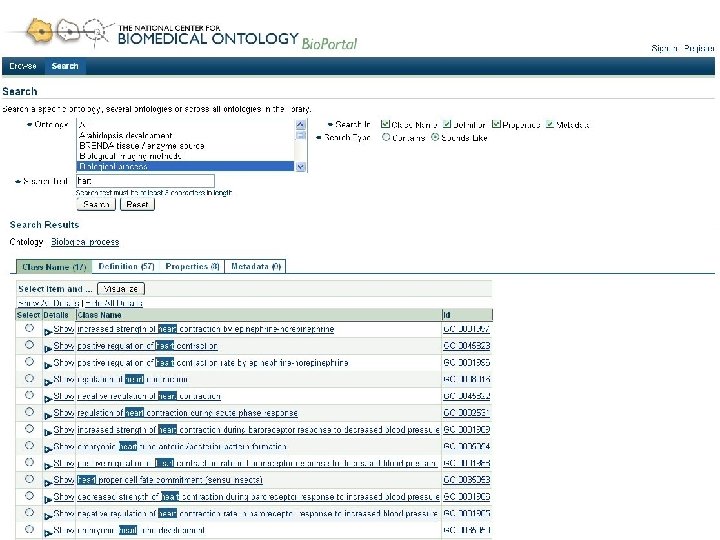

Goals for Bio. Portal • Web accessible repository of ontologies for the biomedical community • Support for ontology – Peer review – Annotation (marginalia) – Versioning – Alignment – Search

Other Center Activities • Biological Driving Projects that will use Bio. Portal ontologies to annotate biomedical data • Collaborating projects that will use Bio. Portal ontologies for – natural-language processing – information integration – data and knowledge visualization • Outreach activities to help different communities to build better ontologies and to utilize the Center’s technology
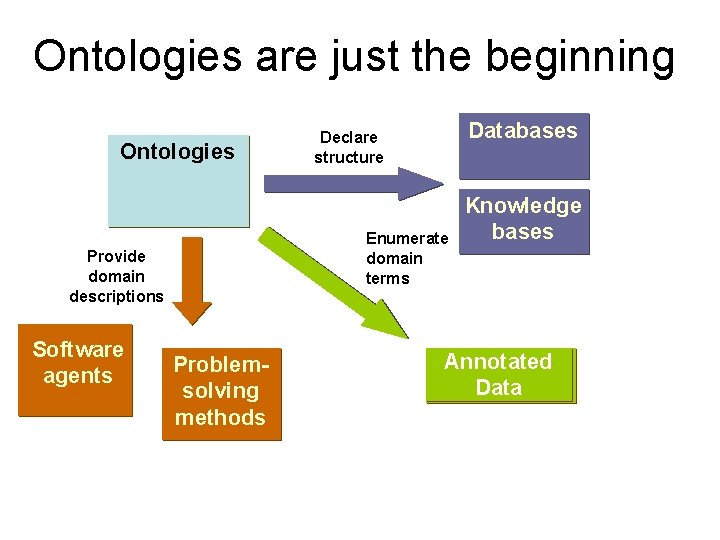
Ontologies are just the beginning Ontologies Enumerate domain terms Provide domain descriptions Software agents Databases Declare structure Problemsolving methods Knowledge bases Annotated Data

Our Center will offer • Technology for uploading, browsing, and using biomedical ontologies • Methods to make the online “publication” of ontologies more like that of journal articles • Tools to enable the biomedical community to put ontologies to work on a daily basis

http: //bioontology. org
- Slides: 139