Oncogenic Mutation Screening in Solitary Fibrous Tumors Elizabeth

Oncogenic Mutation Screening in Solitary Fibrous Tumors Elizabeth G. Demicco, Khalida Wani, Kenneth Aldape, Alexander J. Lazar, and Wei-Lien Wang
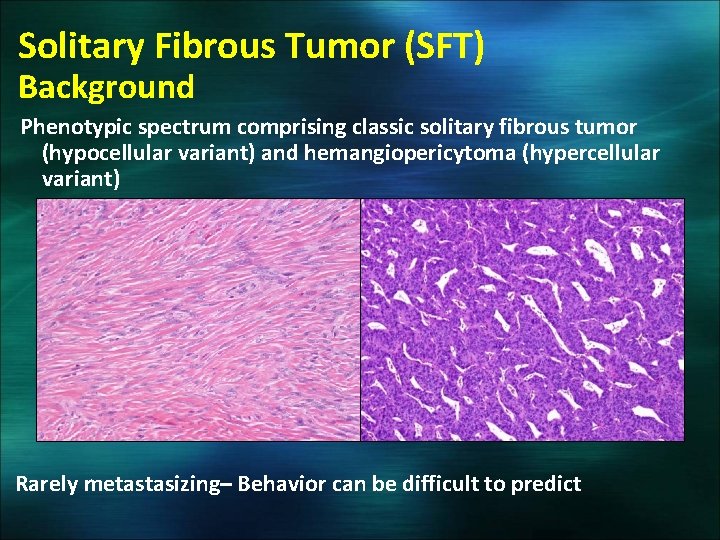
Solitary Fibrous Tumor (SFT) Background Phenotypic spectrum comprising classic solitary fibrous tumor (hypocellular variant) and hemangiopericytoma (hypercellular variant) Rarely metastasizing– Behavior can be difficult to predict

Solitary Fibrous Tumor (SFT) Background Limited data on cytogenetic abnormalities. Relatively simple genomic alterations Few recurrent features Limited data on genetic abnormalities PDGFRB mutations, 2/88 pleural SFT No mutations in DDR 1 (n=8), ERBB 2 (n=10), FGFR 1 (n=15), PDGFRB (n=39) TP 53 one case, dedifferentiated SFT of nasal cavity
Purpose Examine a large series of solitary fibrous tumors for common oncogenic mutations.

Screening Approach to Mutational Analysis (Sequenom Mass. Array) Detection method for single nucleotide polymorphisms (SNP) Step 1. PCR-based Allele (WT vs. mutant) specific probes Single nucleotide extension across site of SNP Different probe sizes allow for allele differentiation and multiplexed reaction Step 2. MALDI-TOF mass spectrometry Analyze primer extension product Time of flight mass spectrometry differentiates based on probe size
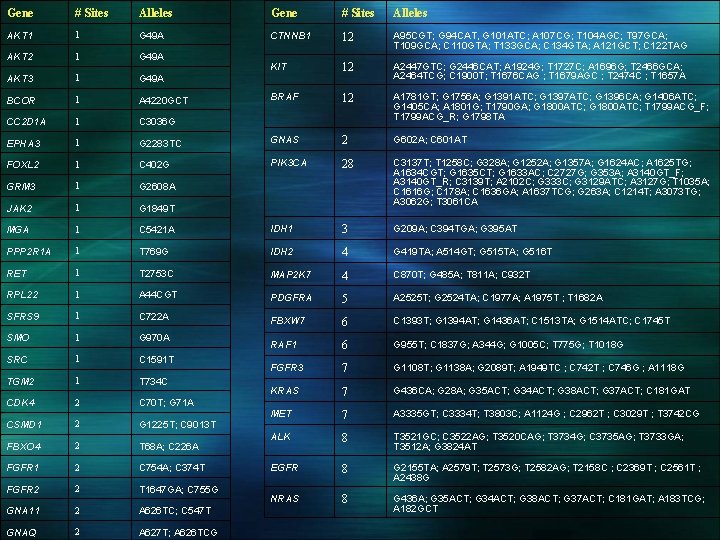
Gene # Sites Alleles AKT 1 1 G 49 A CTNNB 1 12 A 95 CGT; G 94 CAT, G 101 ATC; A 107 CG; T 104 AGC; T 97 GCA; T 109 GCA; C 110 GTA; T 133 GCA; C 134 GTA; A 121 GCT; C 122 TAG AKT 2 1 G 49 A KIT AKT 3 1 G 49 A 12 A 2447 GTC; G 2446 CAT; A 1924 G; T 1727 C; A 1696 G; T 2466 GCA; A 2464 TCG; C 1900 T; T 1676 CAG ; T 1679 AGC ; T 2474 C ; T 1657 A BCOR 1 A 4220 GCT BRAF 12 CC 2 D 1 A 1 C 3036 G A 1781 GT; G 1756 A; G 1391 ATC; G 1397 ATC; G 1396 CA; G 1406 ATC; G 1405 CA; A 1801 G; T 1790 GA; G 1800 ATC; T 1799 ACG_F; T 1799 ACG_R; G 1798 TA EPHA 3 1 G 2283 TC GNAS 2 G 602 A; C 601 AT FOXL 2 1 C 402 G PIK 3 CA 28 GRM 3 1 G 2608 A JAK 2 1 G 1849 T C 3137 T; T 1258 C; G 328 A; G 1252 A; G 1357 A; G 1624 AC; A 1625 TG; A 1634 CGT; G 1635 CT; G 1633 AC; C 2727 G; G 353 A; A 3140 GT_F; A 3140 GT_R; C 3139 T; A 2102 C; G 333 C; G 3129 ATC; A 3127 G; T 1035 A; C 1616 G; C 178 A; C 1636 GA; A 1637 TCG; G 263 A; C 1214 T; A 3073 TG; A 3062 G; T 3061 CA MGA 1 C 5421 A IDH 1 3 G 209 A; C 394 TGA; G 395 AT PPP 2 R 1 A 1 T 769 G IDH 2 4 G 419 TA; A 514 GT; G 515 TA; G 516 T RET 1 T 2753 C MAP 2 K 7 4 C 870 T; G 485 A; T 811 A; C 932 T RPL 22 1 A 44 CGT PDGFRA 5 A 2525 T; G 2524 TA; C 1977 A; A 1975 T ; T 1682 A SFRS 9 1 C 722 A FBXW 7 6 C 1393 T; G 1394 AT; G 1436 AT; C 1513 TA; G 1514 ATC; C 1745 T SMO 1 G 970 A RAF 1 6 G 955 T; C 1837 G; A 344 G; G 1005 C; T 775 G; T 1018 G SRC 1 C 1591 T FGFR 3 7 G 1108 T; G 1138 A; G 2089 T; A 1949 TC ; C 742 T ; C 746 G ; A 1118 G TGM 2 1 T 734 C KRAS 2 C 70 T; G 71 A 7 G 436 CA; G 28 A; G 35 ACT; G 34 ACT; G 38 ACT; G 37 ACT; C 181 GAT CDK 4 CSMD 1 2 G 1225 T; C 9013 T MET 7 A 3335 GT; C 3334 T; T 3803 C; A 1124 G ; C 2962 T ; C 3029 T ; T 3742 CG FBXO 4 2 T 68 A; C 226 A ALK 8 T 3521 GC; C 3522 AG; T 3520 CAG; T 3734 G; C 3735 AG; T 3733 GA; T 3512 A; G 3824 AT FGFR 1 2 C 754 A; C 374 T EGFR 8 G 2155 TA; A 2579 T; T 2573 G; T 2582 AG; T 2158 C ; C 2369 T ; C 2561 T ; A 2438 G FGFR 2 2 T 1647 GA; C 755 G NRAS GNA 11 2 A 626 TC; C 547 T 8 G 436 A; G 35 ACT; G 34 ACT; G 38 ACT; G 37 ACT; C 181 GAT; A 183 TCG; A 182 GCT GNAQ 2 A 627 T; A 626 TCG
Tumor Characteristics 122 tumors (105 patients) 72 Primary 9 Local recurrence 41 Metastasis 13 Patients with multiple tumors tested 6 primary and metastasis(es) 5 multiple metastases 2 local recurrence(s) +/- metastasis
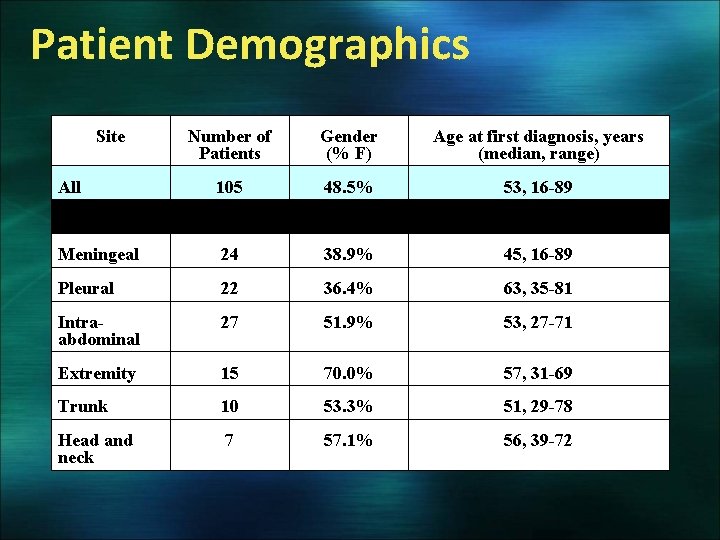
Patient Demographics Site Number of Patients Gender (% F) Age at first diagnosis, years (median, range) All 105 48. 5% 53, 16 -89 Meningeal 24 38. 9% 45, 16 -89 Pleural 22 36. 4% 63, 35 -81 Intraabdominal 27 51. 9% 53, 27 -71 Extremity 15 70. 0% 57, 31 -69 Trunk 10 53. 3% 51, 29 -78 Head and neck 7 57. 1% 56, 39 -72
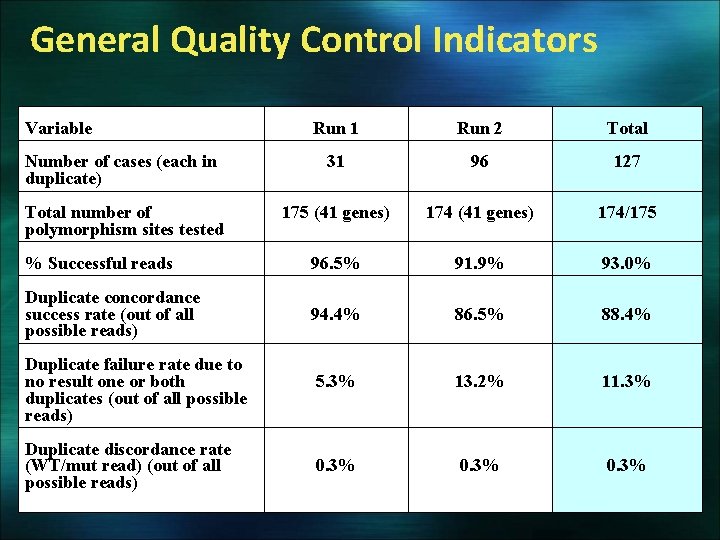
General Quality Control Indicators Variable Run 1 Run 2 Total Number of cases (each in duplicate) 31 96 127 Total number of polymorphism sites tested 175 (41 genes) 174/175 % Successful reads 96. 5% 91. 9% 93. 0% Duplicate concordance success rate (out of all possible reads) 94. 4% 86. 5% 88. 4% Duplicate failure rate due to no result one or both duplicates (out of all possible reads) 5. 3% 13. 2% 11. 3% Duplicate discordance rate (WT/mut read) (out of all possible reads) 0. 3%

Potential Mutations Identified by Sequenom Run 1 Run 2 Total 12 (0. 23%) 21 (0. 15%) 33 (0. 17%) ALK T 3733 G (p. F 1245 V) - 1 1 BRAF T 1799 A* (p. V 600 E) 2 1 3 KRAS G 35 A (p. G 12 D)** 1 10 11 KRAS G 34 A (p. G 12 S)** - 1 1 KRAS C 181 A (p. Q 61 K) - 1 1 MET C 2962 T (p. R 988 C) 1 1 2 MET C 3029 T (p. T 1010 I) - 3 3 NRAS G 35 A (p. G 12 D)** 7 3 10 NRAS G 38 A (p. G 13 D)** 1 - 1 Number SNPs detected (% of all useable reads) Site * Mutation identified on forward probe, WT on reverse ** Problems with probes later reported – Determined to be WT by CAST-PCR

Characteristics of Tumors with Identified SNPs Patient (phenotypic variant) Mutation Tumor status Primary Site this metastasis Outcome SFT 118 (hypercellular) MET C 3029 T (p. T 1010 I)* Metastasis (10 yr) Intraabdominal Peritoneum Do. D 18 yr SFT 118 (hypercellular) MET C 3029 T (p. T 1010 I)* Metastasis (14 yr) Intraabdominal Peritoneum Do. D 18 yr SFT 055 (hypocellular) MET C 3029 T (p. T 1010 I)* Primary Intraabdominal Dead other causes 30 months SFT 048 (hypercellular) MET C 2962 T (p. R 988 C)* Primary Intraabdominal Metastasized at 40 mo. D 8. 5 yr SFT 092 (hypocellular) MET C 2962 T (p. R 988 C)* Primary Pleura LTFU 1 month SFT 067 (hypercellular) ALK T 3733 G (p. F 1245 V) Primary Intraabdominal NED 69 mo SFT 112 (hypercellular) KRAS C 181 A (p. Q 61 K) Metastasis (13 yr) Intraabdominal * Presence of SNP did not correlate with c-MET expression by IHC Peritoneum Dod 16 yrs

MET C 3029 T (p. T 1010 I) and C 2962 T (p. R 988 C) Also designated as T 992 I and R 970 C Juxtamembrane domain Initially reported as rare somatic oncogenic mutations Identified in a wide range of tumors Frequency in involved cancers ~1 -10% Later proposed to represent germline polymorphisms Seen in ~1% of individuals without cancer (1/96) Germline in several patients with cancer No evidence of transformation in cell models T 1010 I as cooperative germline oncogenic mutation? Identified in ~4. 5% familial CRC Mutation proposed to indirectly activate c-MET via inhibition of inhibitor Not initiator, may promote progression
ALK T 3733 G (p. F 1245 V) Kinase-activating mutation Rare mutation Reported in neuroblastoma

KRAS C 181 A (p. Q 61 K) Activating mutation reported rarely in colonic and lung adenocarcinoma, misc. other carcinomas various sites Much more rare than the equivalent NRAS mutation

Summary We performed screening SNP analysis in a large series of SFT Confirmed that SFT have low frequency of mutations in oncogenes commonly reported in other malignancies 7/122 cases (5. 7%) Abdominal location predominant Insufficient data on relevance to status, prognosis

Conclusions Mutations in commonly identified oncogenes do not appear to function in pathogenesis of SFT. Alternative mechanisms? Mutations in uncommon genes Imprinting mi. RNA regulation Studies are ongoing

Thank You.
- Slides: 17