Oncogene and Tumor Suppressor gene networks in cell






![q. NET graphs from M • mij= [ xi/ xj]o mij 0 Cycle 1 q. NET graphs from M • mij= [ xi/ xj]o mij 0 Cycle 1](https://slidetodoc.com/presentation_image/c7157af527378301a85d47f1fd3c12a3/image-20.jpg)


![Transcritical Bifurcation in Positively Coupled Cycles [Y 2]ss 1 r 0 Y 1 X Transcritical Bifurcation in Positively Coupled Cycles [Y 2]ss 1 r 0 Y 1 X](https://slidetodoc.com/presentation_image/c7157af527378301a85d47f1fd3c12a3/image-29.jpg)





- Slides: 56

Oncogene and Tumor Suppressor gene networks in cell cycle checkpoints, apoptosis & cell survival Baltazar D. Aguda Mathematical Biosciences Institute Ohio State University bdaguda@mbi. osu. edu Seminar given at Indiana University-Purdue University at Indianapolis, 6 Oct 2006

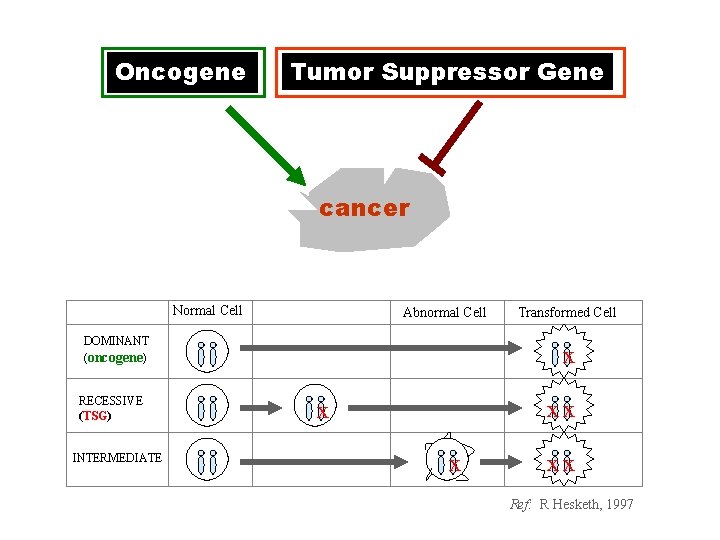
Oncogene Tumor Suppressor Gene cancer Normal Cell Abnormal Cell DOMINANT (oncogene) RECESSIVE (TSG) INTERMEDIATE Transformed Cell x xx x x xx Ref: R Hesketh, 1997

Oncogenes & TSGs associated with some human cancers breast carcinoma : MYB, MYC, cyclin D 1, cyclin D 3, EGFR, HER 2, HRAS, HSTF 1, INT 2, YES 1, P 53, RB 1, BRCA 1 cervical carcinoma : P 53, RB 1, MYCN, HRAS colorectal carcinoma : APC, MCC, DCC, P 53, TGFBR 2, KRAS 2 CML : BCR-ABL, MYC, NRAS, RB 1, P 53, ERG, TLS, HOXA 9, NUP 98, AML 1, EAP, EVI 1, MN 1, TEL, MDS 1

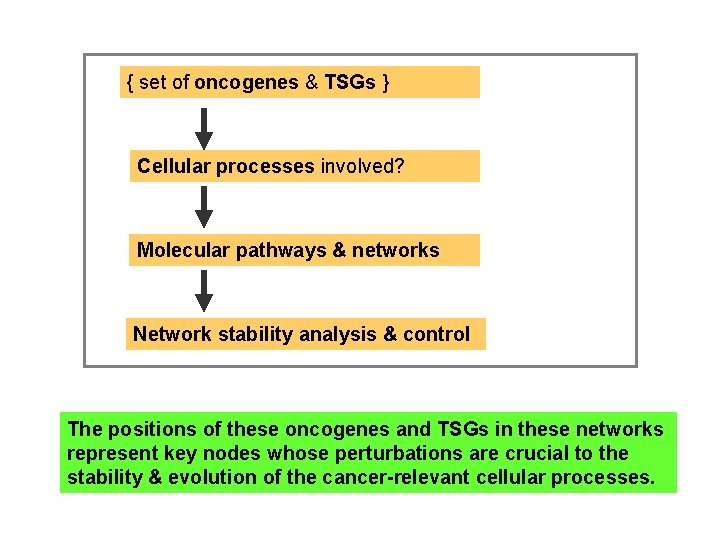
{ set of oncogenes & TSGs } Cellular processes involved? Molecular pathways & networks Network stability analysis & control The positions of these oncogenes and TSGs in these networks represent key nodes whose perturbations are crucial to the stability & evolution of the cancer-relevant cellular processes.

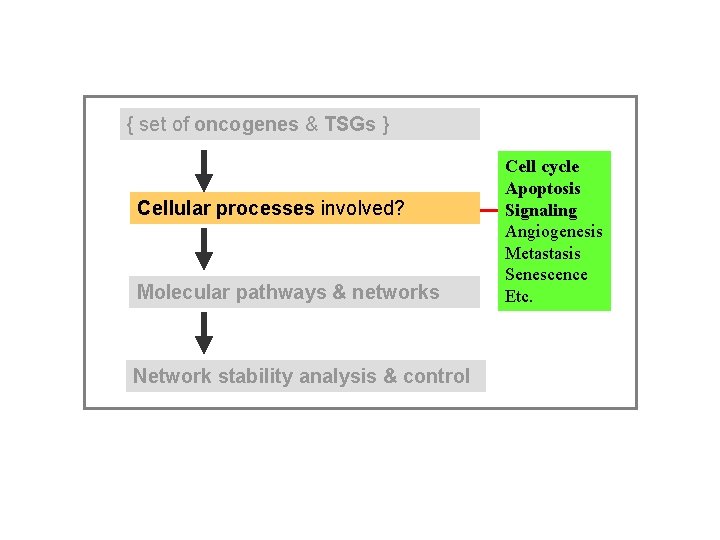
{ set of oncogenes & TSGs } Cellular processes involved? Molecular pathways & networks Network stability analysis & control Cell cycle Apoptosis Signaling Angiogenesis Metastasis Senescence Etc.

{ set of oncogenes & TSGs } Cellular processes involved? Molecular pathways & networks Pathway ontology (BIOPAX) DBs/KBs (e. g. BIND, Biocarta, Gen. MAPP, Reactome) Network stability analysis & control Proposal: Library of Pathway Modules (functional modules according to specific cellular processes; e. g. Biocarta) Repository of Models: Biomodels Database: http: //www. ebi. ac. uk/biomodels/ Cell. ML model repository: http: //www. cellml. org/examples/repository/index. html

{ set of oncogenes & TSGs } Topological analysis of q. NETs (qualitative networks) Cellular processes involved? Network modularization schemes [e. g. Aguda & Algar (2003) Cell Cycle 2: 538 ] Model extraction & algorithms Molecular pathways networks Network perturbation and control protocols Network stability analysis & control

Oncogenes & TSGs in G 1 -S Network

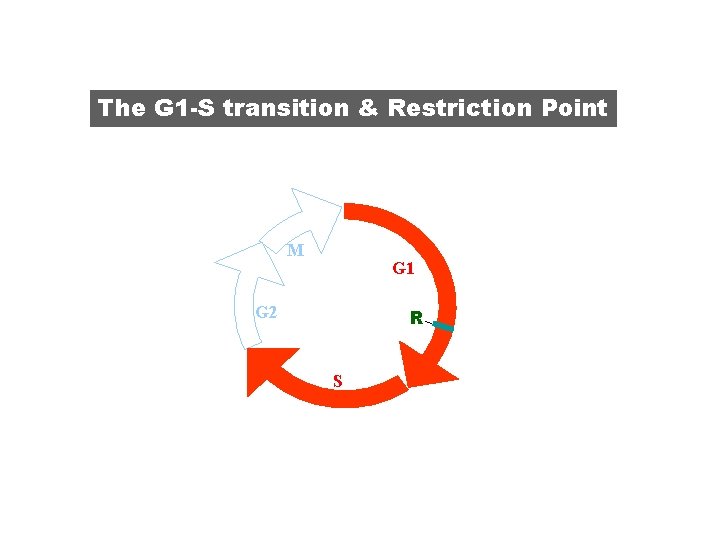
The G 1 -S transition & Restriction Point M G 1 G 2 R S

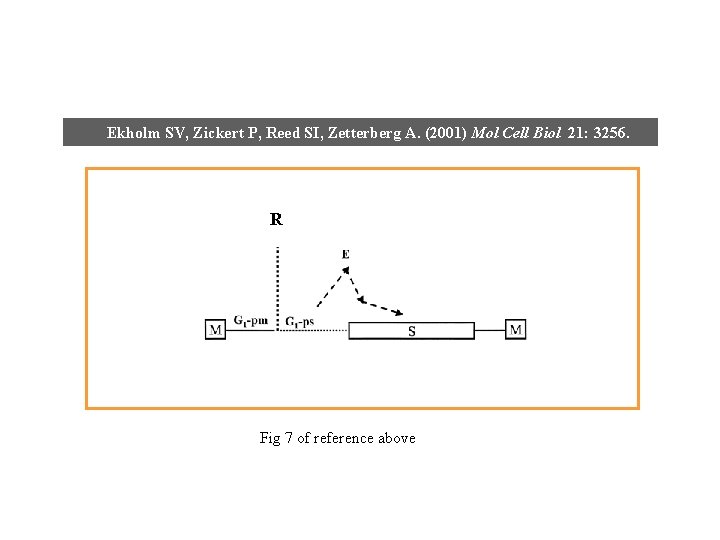
Ekholm SV, Zickert P, Reed SI, Zetterberg A. (2001) Mol Cell Biol 21: 3256. R G 1 Fig 7 of reference above

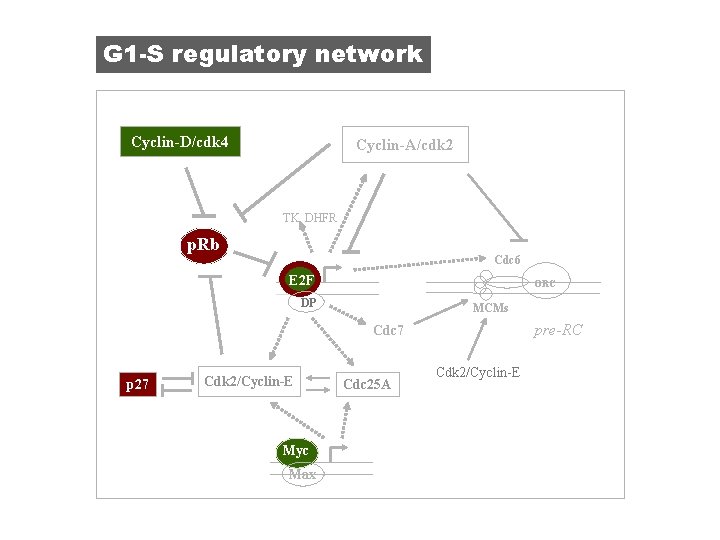
G 1 -S regulatory network Cyclin-D/cdk 4 Cyclin-A/cdk 2 TK, DHFR p. Rb p. RB Cdc 6 E 2 F ORC DP MCMs pre-RC Cdc 7 p 27 Cdk 2/Cyclin-E Myc Max Cdc 25 A Cdk 2/Cyclin-E

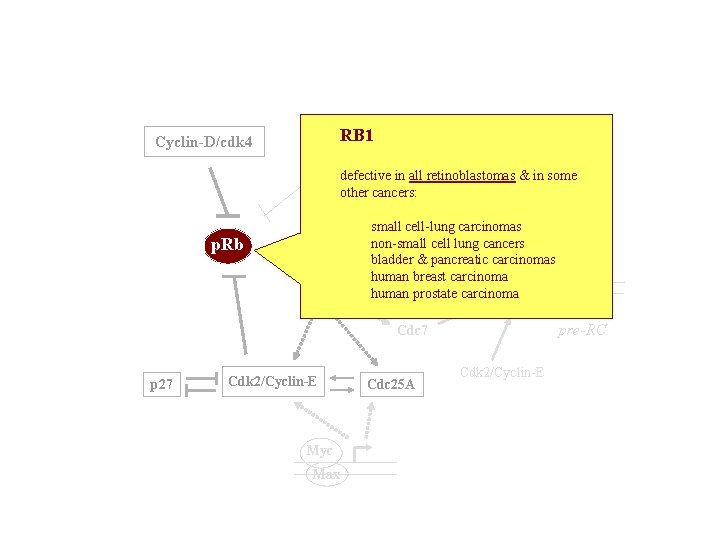
RB 1 Cyclin-A/cdk 2 Cyclin-D/cdk 4 defective in all retinoblastomas & in some TK, other cancers: DHFR p. Rb p. RB E 2 F DP small cell-lung carcinomas non-small cell lung cancers Cdc 6 bladder & pancreatic carcinomas human breast carcinoma ORC human prostate carcinoma MCMs pre-RC Cdc 7 p 27 Cdk 2/Cyclin-E Myc Max Cdc 25 A Cdk 2/Cyclin-E

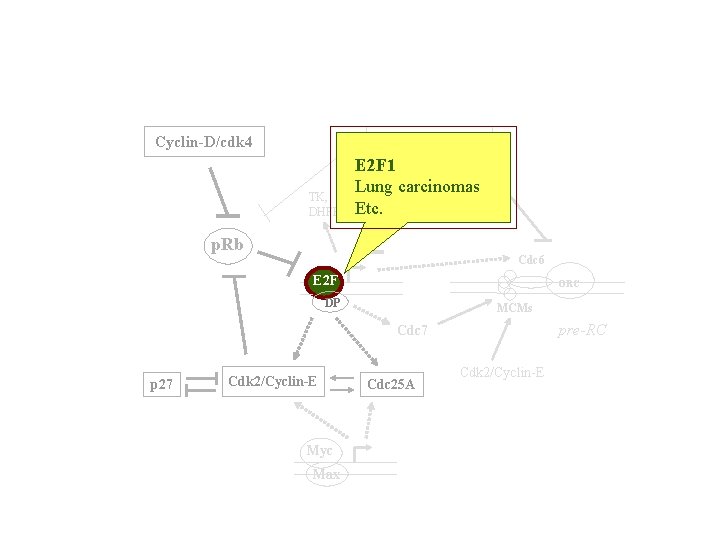
Cyclin-D/cdk 4 Cyclin-A/cdk 2 TK, DHFR E 2 F 1 Lung carcinomas Etc. p. Rb p. RB Cdc 6 E 2 F ORC DP MCMs pre-RC Cdc 7 p 27 Cdk 2/Cyclin-E Myc Max Cdc 25 A Cdk 2/Cyclin-E

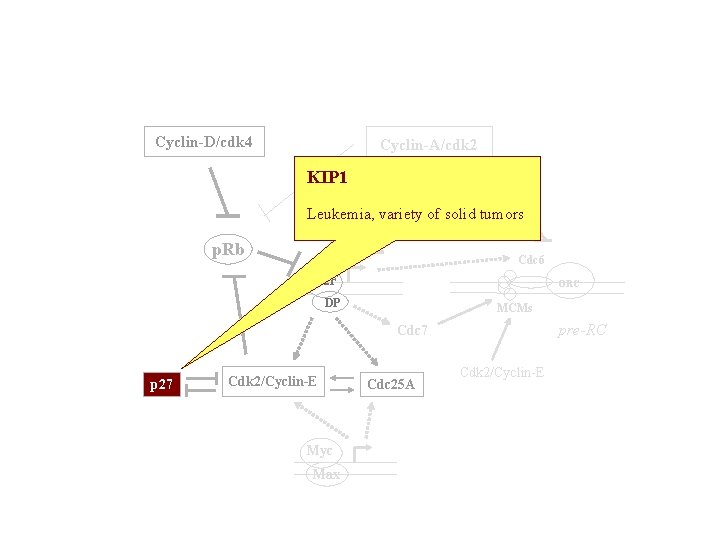
Cyclin-D/cdk 4 Cyclin-A/cdk 2 KIP 1 TK, DHFR Leukemia, variety of solid tumors p. Rb Cdc 6 E 2 F ORC DP MCMs pre-RC Cdc 7 p 27 Cdk 2/Cyclin-E Myc Max Cdc 25 A Cdk 2/Cyclin-E

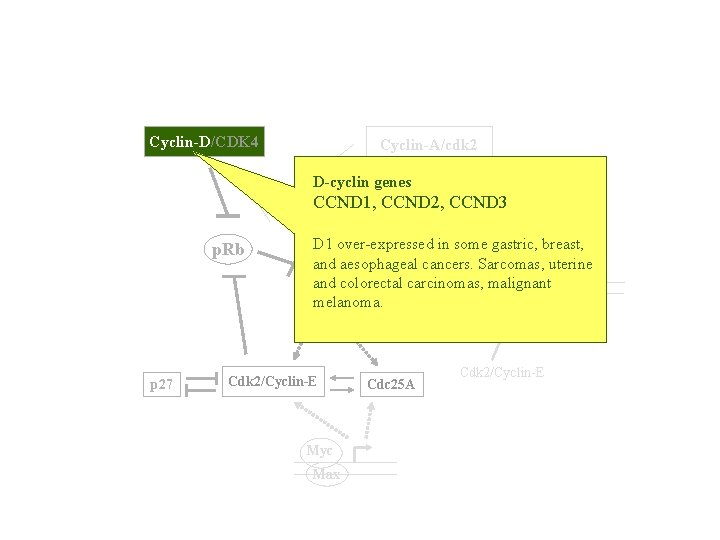
Cyclin-D/CDK 4 Cyclin-A/cdk 2 D-cyclin genes TK, CCND 1, DHFR p. Rb CCND 2, CCND 3 D 1 over-expressed in some gastric, breast, Cdc 6 uterine and aesophageal cancers. Sarcomas, E 2 F and colorectal carcinomas, malignant ORC DP melanoma. MCMs pre-RC Cdc 7 p 27 Cdk 2/Cyclin-E Myc Max Cdc 25 A Cdk 2/Cyclin-E

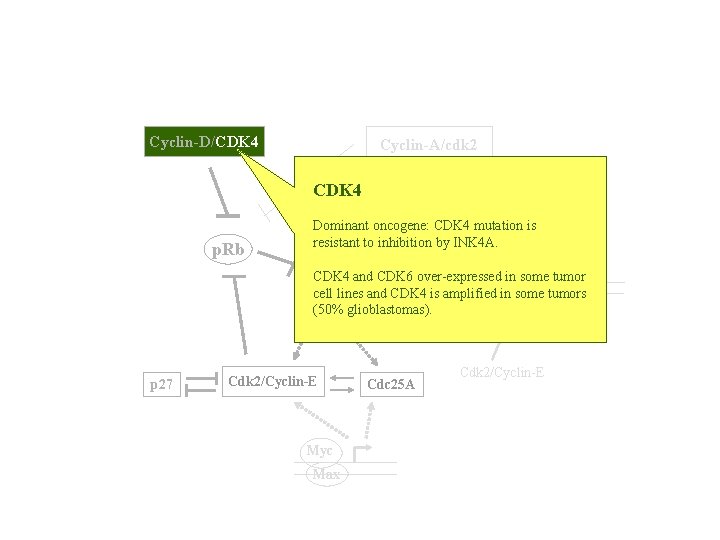
Cyclin-D/CDK 4 Cyclin-A/cdk 2 CDK 4 TK, DHFR p. Rb Dominant oncogene: CDK 4 mutation is resistant to inhibition by INK 4 A. Cdc 6 CDK 4 E 2 F and CDK 6 over-expressed in some tumor ORC cell lines and CDK 4 is amplified in some tumors DP MCMs (50% glioblastomas). pre-RC Cdc 7 p 27 Cdk 2/Cyclin-E Myc Max Cdc 25 A Cdk 2/Cyclin-E

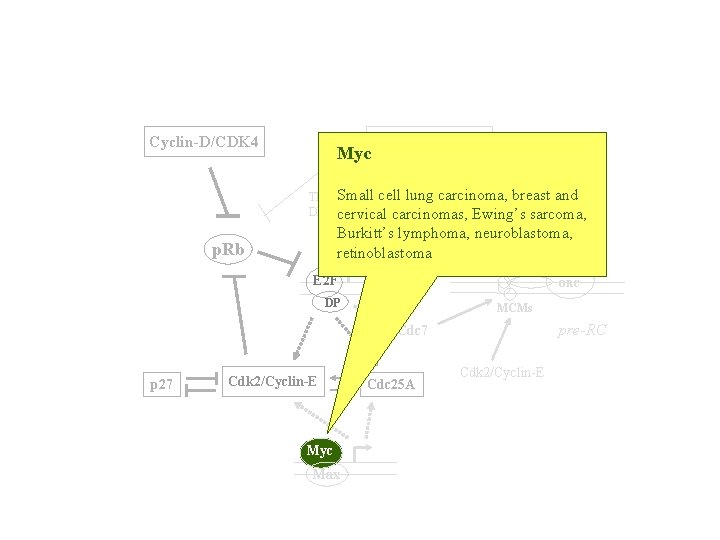
Cyclin-D/CDK 4 Myc Cyclin-A/cdk 2 TK, Small cell lung carcinoma, breast and DHFRcervical carcinomas, Ewing’s sarcoma, Burkitt’s lymphoma, neuroblastoma, retinoblastoma Cdc 6 p. Rb E 2 F ORC DP MCMs pre-RC Cdc 7 p 27 Cdk 2/Cyclin-E Myc Max Cdc 25 A Cdk 2/Cyclin-E

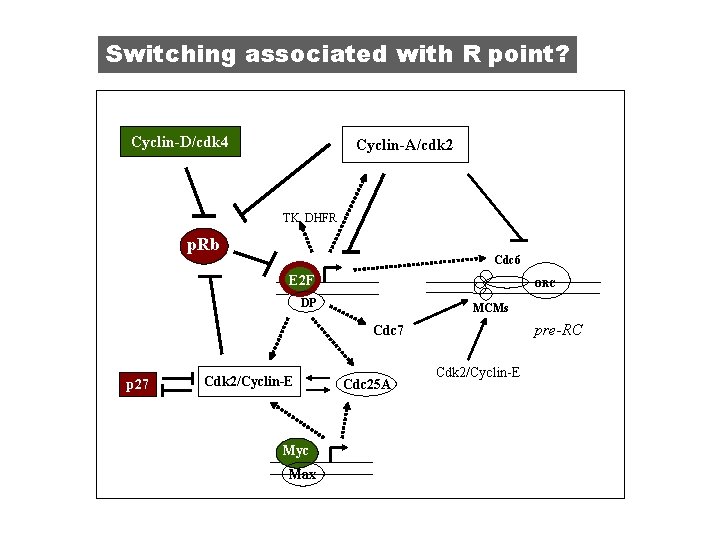
Switching associated with R point? Cyclin-D/cdk 4 Cyclin-A/cdk 2 TK, DHFR p. Rb p. RB Cdc 6 E 2 F ORC DP MCMs pre-RC Cdc 7 p 27 Cdk 2/Cyclin-E Myc Max Cdc 25 A Cdk 2/Cyclin-E

Network Structure & Instability q. NET analysis
![q NET graphs from M mij xi xjo mij 0 Cycle 1 q. NET graphs from M • mij= [ xi/ xj]o mij 0 Cycle 1](https://slidetodoc.com/presentation_image/c7157af527378301a85d47f1fd3c12a3/image-20.jpg)
q. NET graphs from M • mij= [ xi/ xj]o mij 0 Cycle 1 -cycle Xj Xj activates inhibits Xi Xi ( Xj Xi ) strength graph mii Xi • 2 -cycle mijmji 3 -cycle mijmjkmki Xi • Xi • • Xk • Xj Xj •

eigenvalues are functions of cycles only dx/dt = f(x) d(Dx)/dt = M(Dx) det(λI-M) = ln + a 1 ln-1 + a 2 ln-2 + … + an-1 l + an = 0 where a 1 = i [-C 1(i)] a 2 = i, j [-C 1(i)][-C 1(j)] + jk [-C 2(jk)] a 3 = i, j, k [-C 1(i)][-C 1(j)][-C 1(k)] + i, jk [-C 1(i)][-C 2(jk)] + ijk [-C 3(ijk)]. . . where C 1(i) = mii (1 -cycles) C 2(jk) = mjkmkj (2 -cycles) C 3(ijk) = mijmjkmki (3 -cycles) …. . . Hurwitz Determinants, D = functions of Ck ‘s

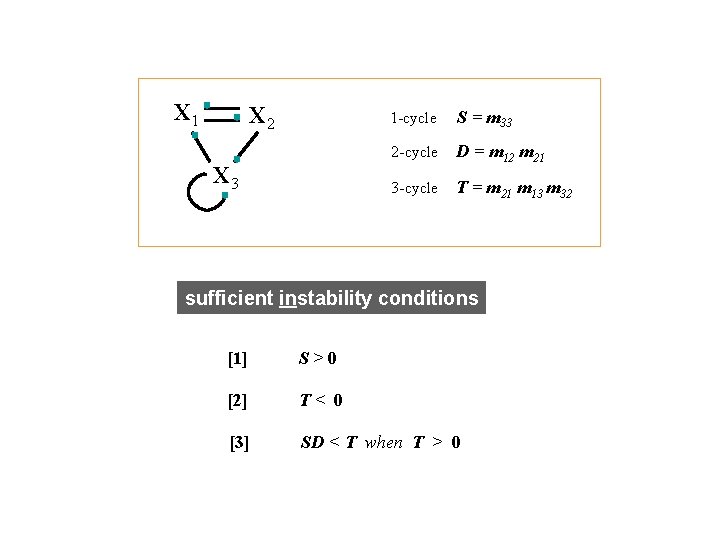
X 1 X 2 X 3 1 -cycle S = m 33 2 -cycle D = m 12 m 21 3 -cycle T = m 21 m 13 m 32 sufficient instability conditions [1] S>0 [2] T< 0 [3] SD < T when T > 0

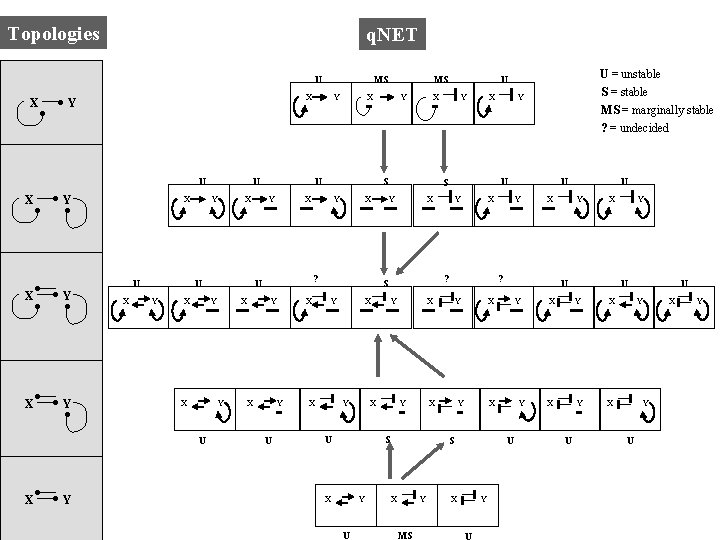
Topologies q. NET U X X Y U X Y Y X X Y X U X Y X U Y X Y Y X ? U Y X X Y Y Y U Y X Y MS Y X X Y U Y X U X S X U ? X Y Y U X S X X ? X U Y S Y U = unstable S = stable MS = marginally stable ? = undecided U X S X Y MS X U U Y X Y U X MS U X Y X U Y X Y U Y

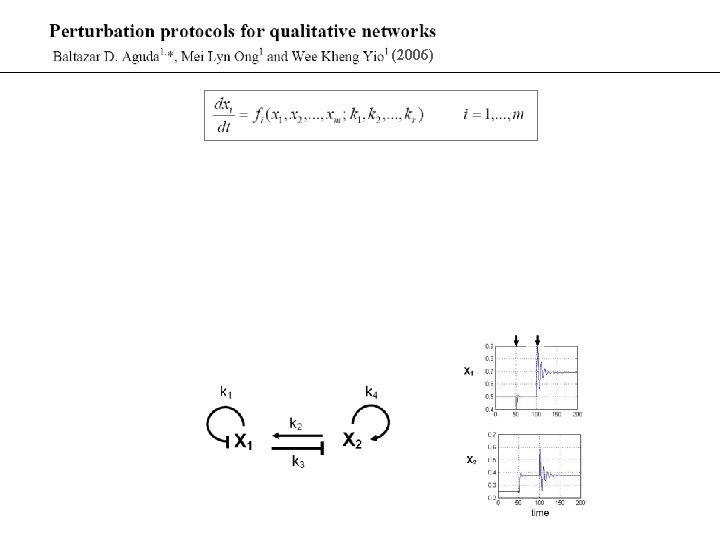
(2006)

Modeling the G 1 Checkpoint

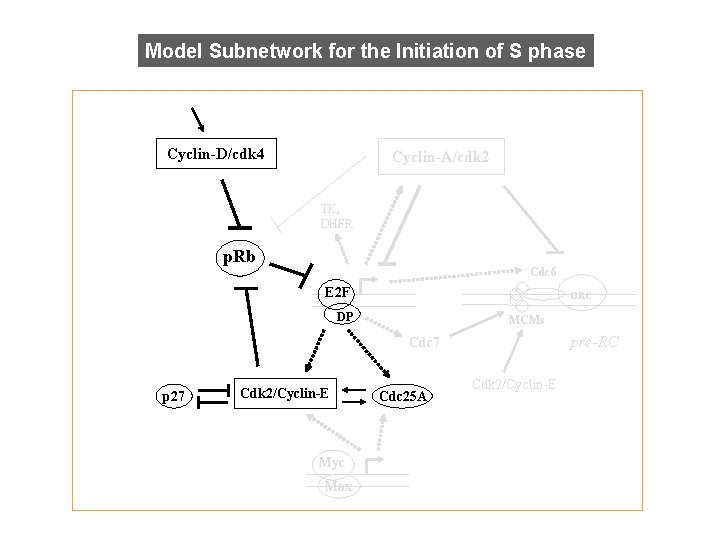
Model Subnetwork for the Initiation of S phase Cyclin-D/cdk 4 Cyclin-A/cdk 2 TK, DHFR p. Rb p. RB Cdc 6 E 2 F ORC DP MCMs pre-RC Cdc 7 p 27 Cdk 2/Cyclin-E Myc Max Cdc 25 A Cdk 2/Cyclin-E

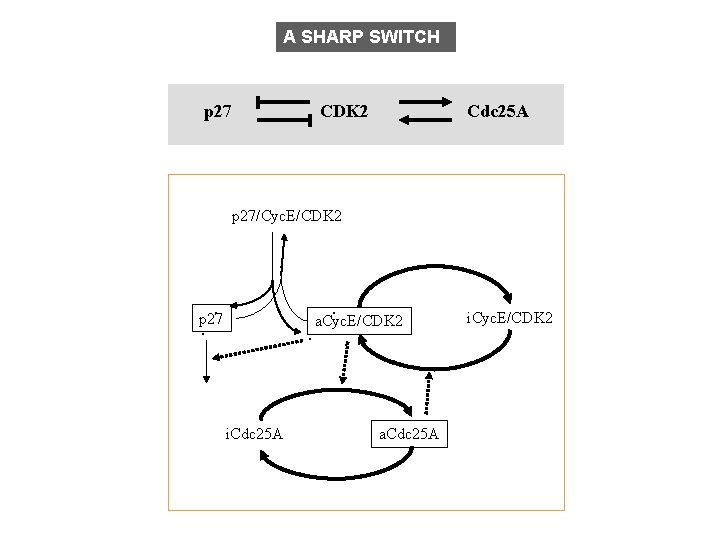
A SHARP SWITCH p 27 CDK 2 Cdc 25 A p 27/Cyc. E/CDK 2 . . . p 27 . . i. Cdc 25 A a. Cyc. E/CDK 2 . . a. Cdc 25 A i. Cyc. E/CDK 2

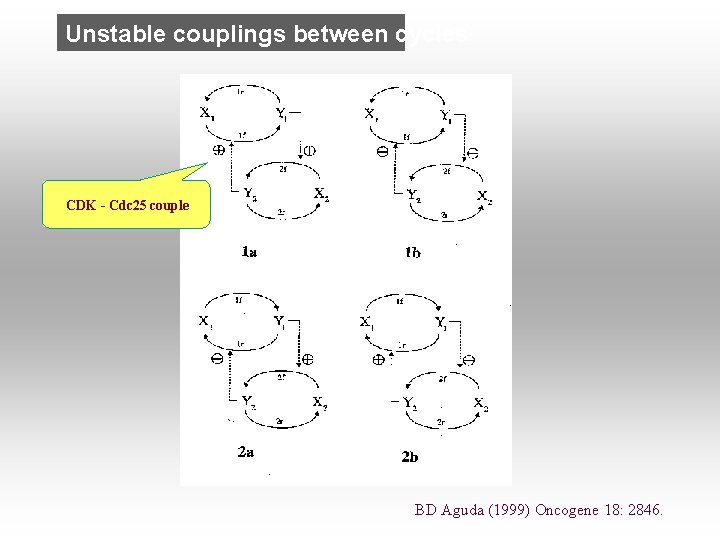
Unstable couplings between cycles CDK - Cdc 25 couple BD Aguda (1999) Oncogene 18: 2846.
![Transcritical Bifurcation in Positively Coupled Cycles Y 2ss 1 r 0 Y 1 X Transcritical Bifurcation in Positively Coupled Cycles [Y 2]ss 1 r 0 Y 1 X](https://slidetodoc.com/presentation_image/c7157af527378301a85d47f1fd3c12a3/image-29.jpg)
Transcritical Bifurcation in Positively Coupled Cycles [Y 2]ss 1 r 0 Y 1 X 1 s u s E 2 1 f [Y 2]ss 2 f X 2 Y 2 0 E 1 2 r Y 1 & Y 2 turned ‘on’ only if E 1*E 2 > (k 1 r/k 1 f)*(k 2 r/k 2 f) mass-action kinetics in graphs shown; similar for Michaelis-Menten kinetics Y 1 ss [ ]ss ss Y 2 0 E 1

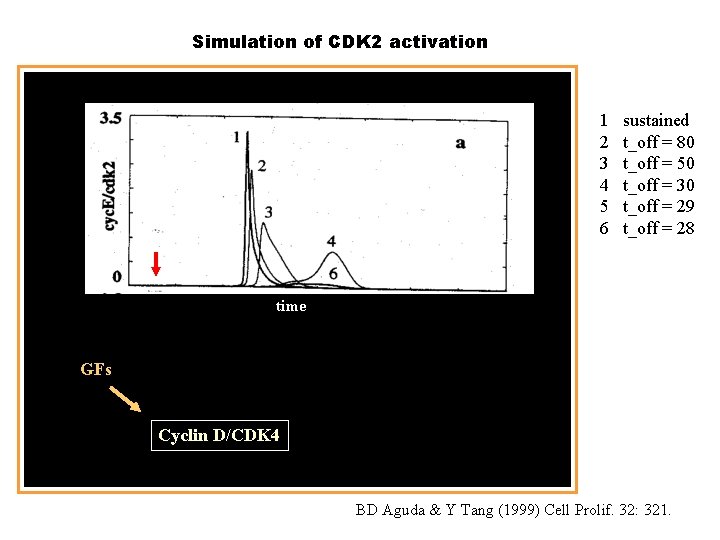
Simulation of CDK 2 activation 1 2 3 4 5 6 sustained t_off = 80 t_off = 50 t_off = 30 t_off = 29 t_off = 28 time GFs Cyclin D/CDK 4 BD Aguda & Y Tang (1999) Cell Prolif. 32: 321.

Ekholm SV, Zickert P, Reed SI, Zetterberg A. (2001) Mol Cell Biol 21: 3256. R Fig 7 of reference above

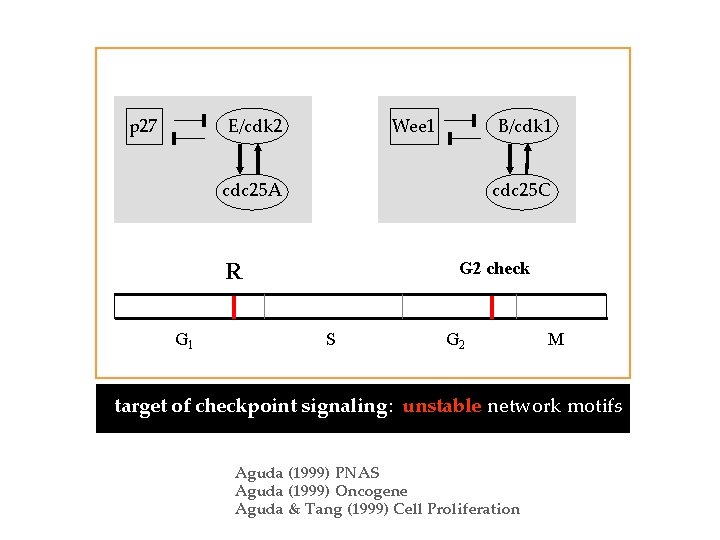
E/cdk 2 p 27 B/cdk 1 Wee 1 cdc 25 A cdc 25 C R G 1 G 2 check S G 2 M target of checkpoint signaling: unstable network motifs Aguda (1999) PNAS Aguda (1999) Oncogene Aguda & Tang (1999) Cell Proliferation

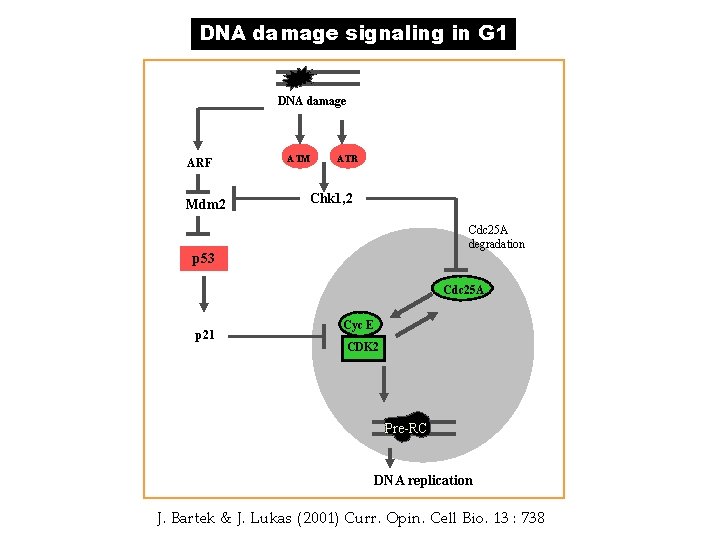
DNA damage signaling in G 1 DNA damage ARF Mdm 2 ATM ATR Chk 1, 2 Cdc 25 A degradation p 53 Cdc 25 A p 21 Cyc E CDK 2 Pre-RC DNA replication J. Bartek & J. Lukas (2001) Curr. Opin. Cell Bio. 13 : 738

Signaling, Cell cycle & Apoptosis

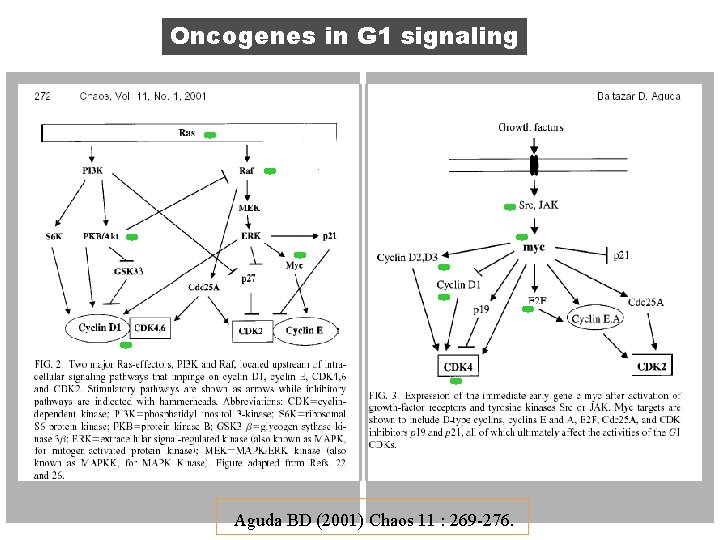
Oncogenes in G 1 signaling Aguda BD (2001) Chaos 11 : 269 -276.

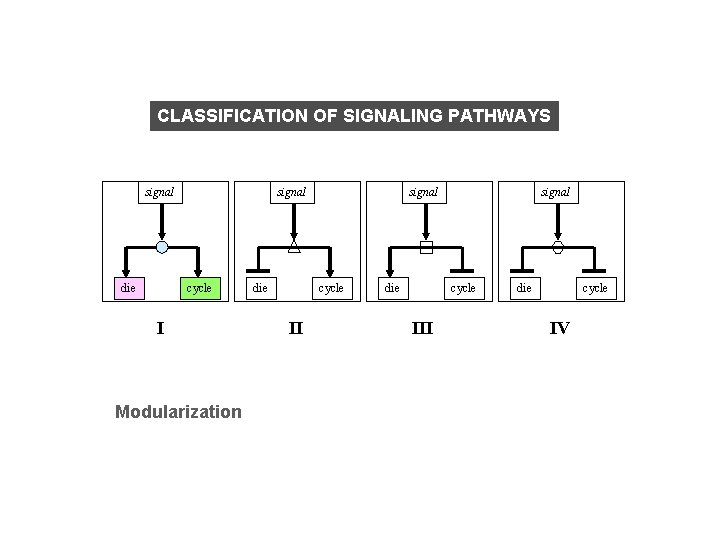
CLASSIFICATION OF SIGNALING PATHWAYS signal die signal cycle I Modularization die signal cycle III die cycle IV

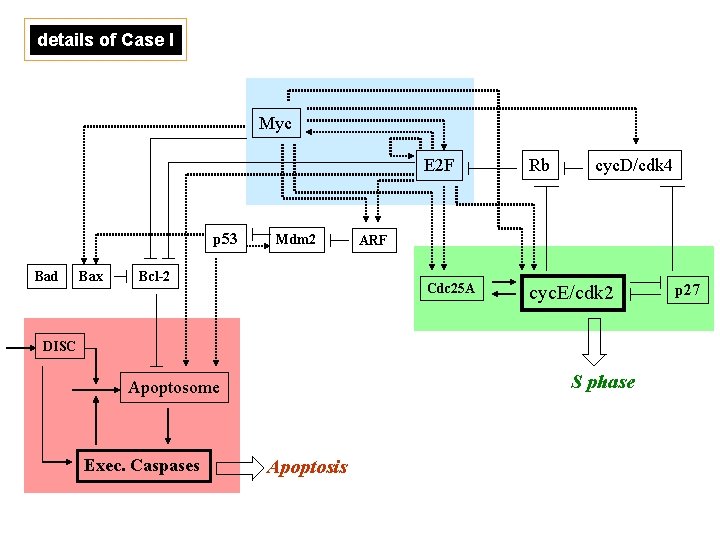
details of Case I Myc p 53 Bad Bax Mdm 2 Bcl-2 E 2 F Rb cyc. D/cdk 4 Cdc 25 A cyc. E/cdk 2 ARF DISC S phase Apoptosome Exec. Caspases Apoptosis p 27

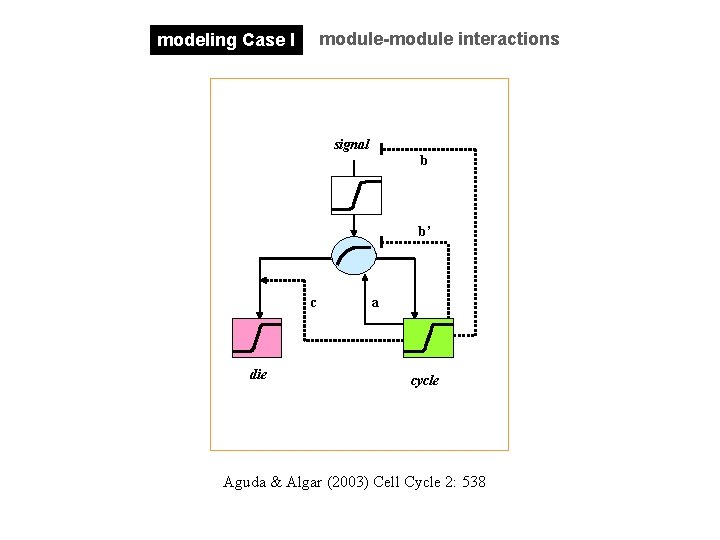
module-module interactions modeling Case I signal b b’ c die a cycle Aguda & Algar (2003) Cell Cycle 2: 538

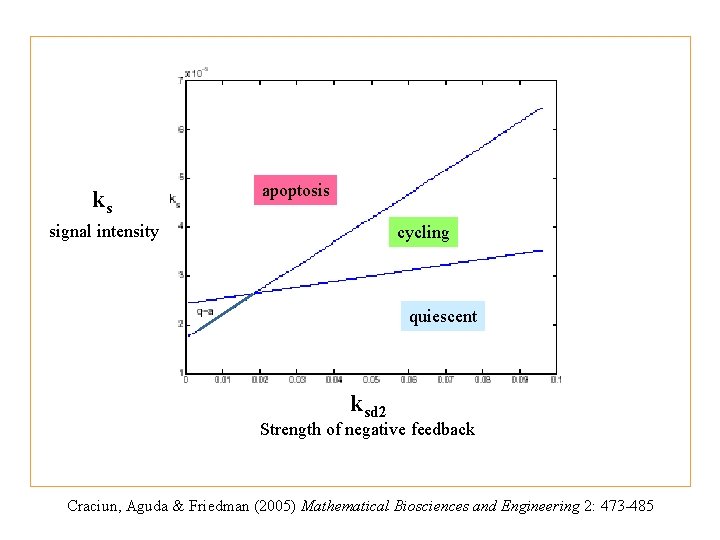
ks apoptosis signal intensity cycling quiescent ksd 2 Strength of negative feedback Craciun, Aguda & Friedman (2005) Mathematical Biosciences and Engineering 2: 473 -485

Cell Survival and Death p 53 vs Akt with K. B. (Dave) Wee, Ph. D student

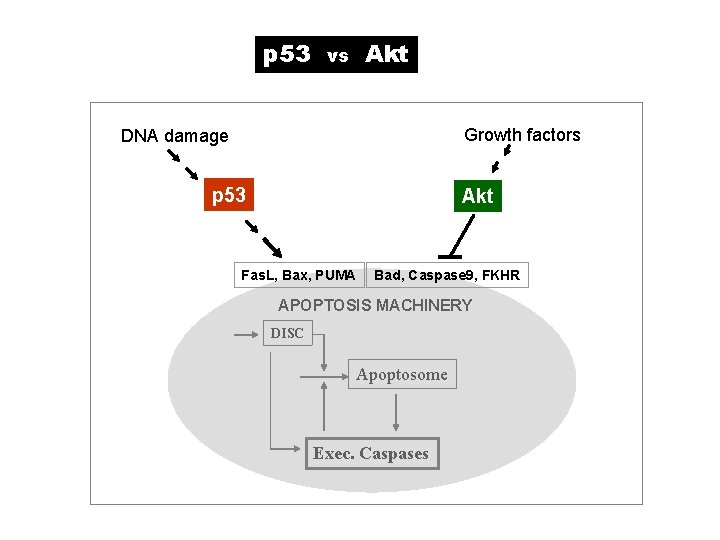
p 53 Akt vs Growth factors DNA damage p 53 Akt Fas. L, Bax, PUMA Bad, Caspase 9, FKHR APOPTOSIS MACHINERY DISC Apoptosome Exec. Caspases

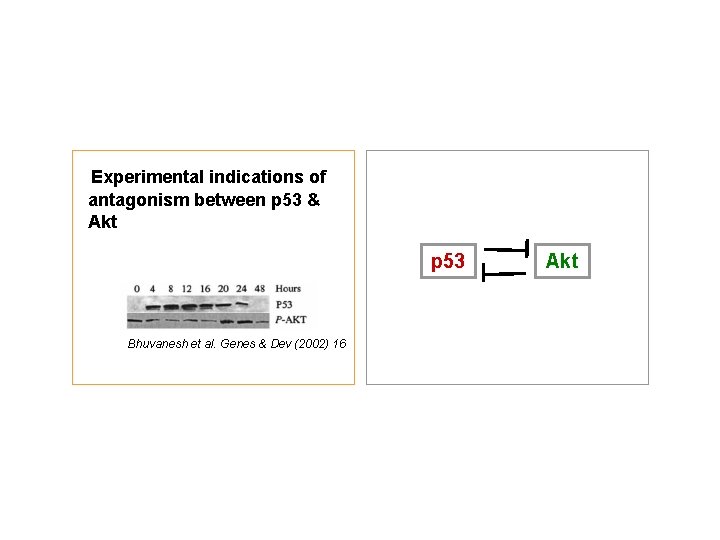
Experimental indications of antagonism between p 53 & Akt p 53 Bhuvanesh et al. Genes & Dev (2002) 16 Akt

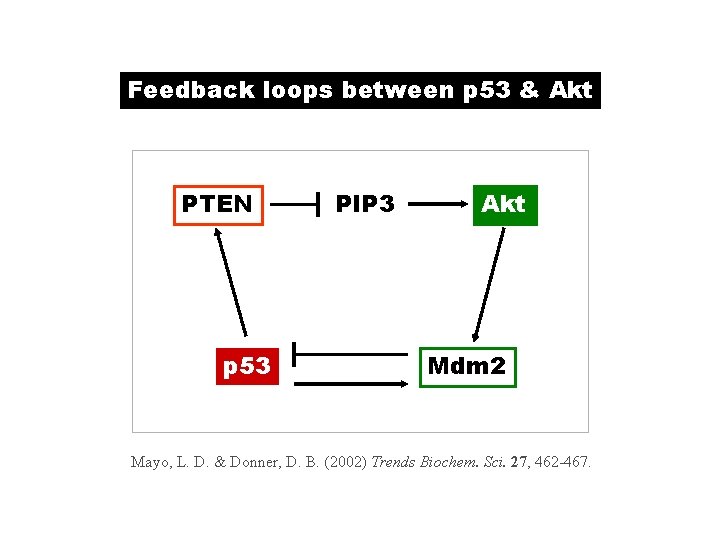
Feedback loops between p 53 & Akt PTEN p 53 PIP 3 Akt Mdm 2 Mayo, L. D. & Donner, D. B. (2002) Trends Biochem. Sci. 27, 462 -467.

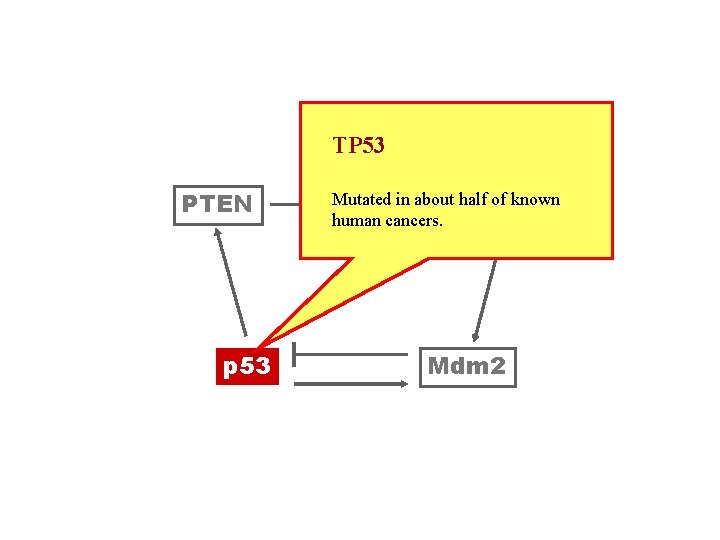
TP 53 PTEN p 53 Mutated of known PIP 3 in about half. Akt human cancers. Mdm 2

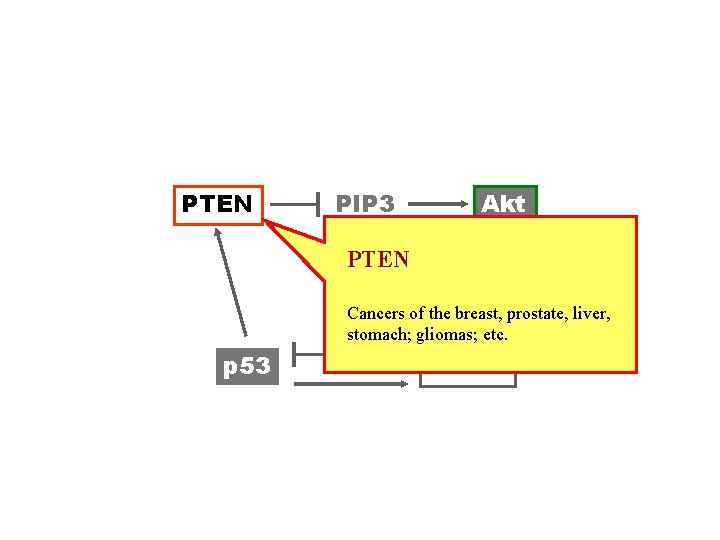
PTEN PIP 3 Akt PTEN Cancers of the breast, prostate, liver, stomach; gliomas; etc. p 53 Mdm 2

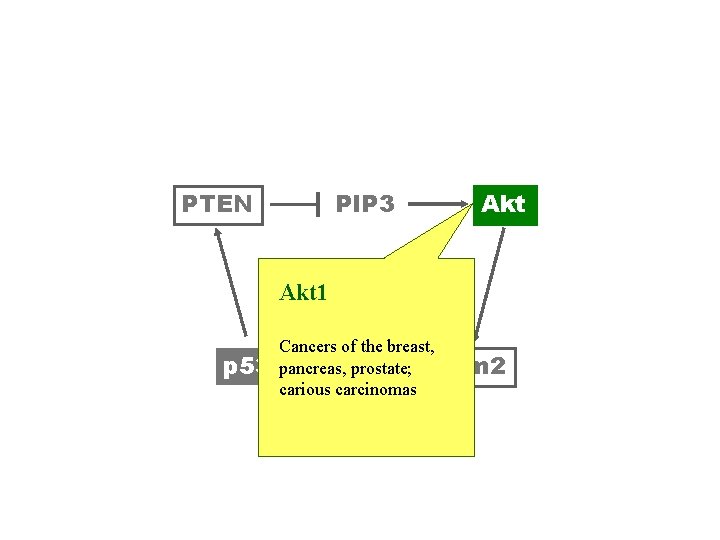
PTEN PIP 3 Akt 1 p 53 Cancers of the breast, pancreas, prostate; Mdm 2 carious carcinomas

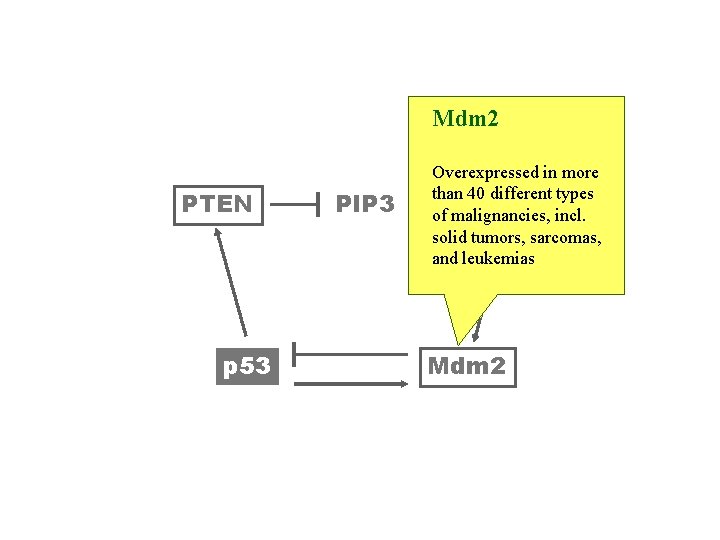
Mdm 2 PTEN p 53 PIP 3 Overexpressed in more than 40 different types Akt incl. of malignancies, solid tumors, sarcomas, and leukemias Mdm 2

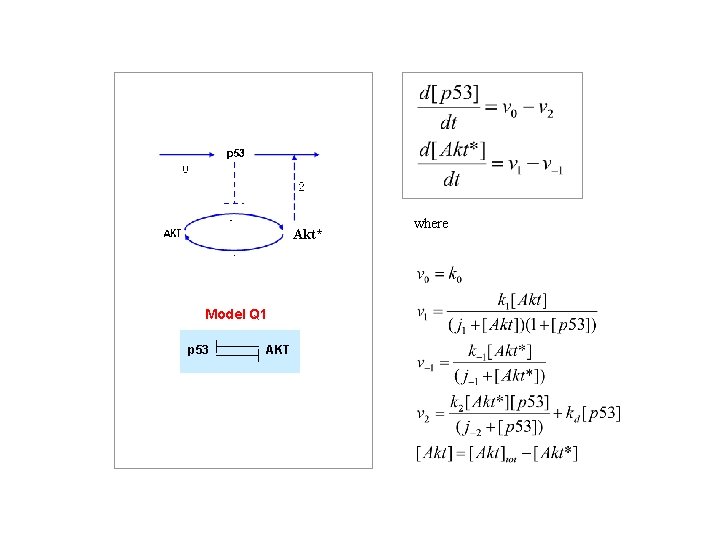
Akt* Model Q 1 p 53 AKT where

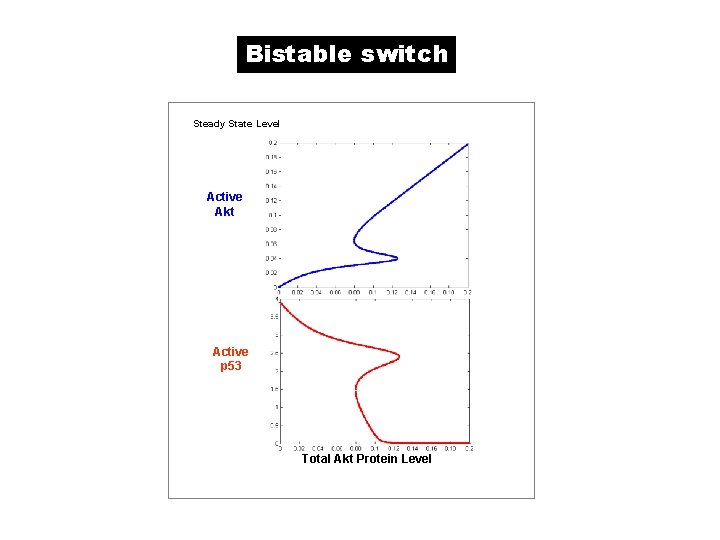
Bistable switch Steady State Level Active Akt Active p 53 Total Akt Protein Level

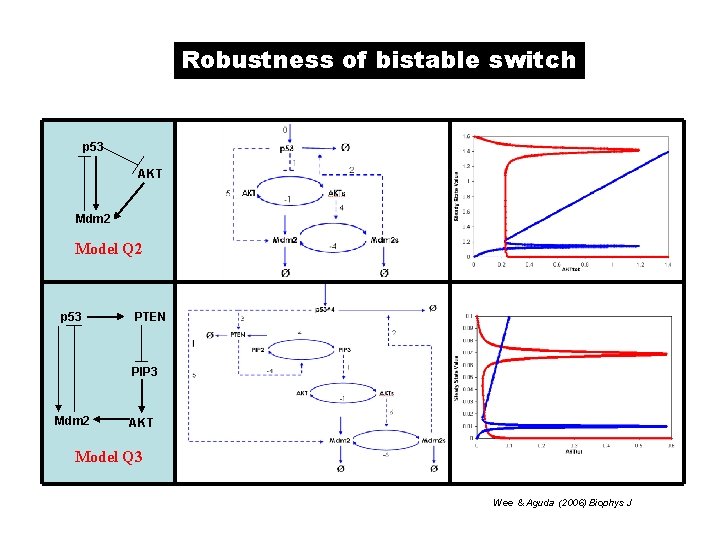
Robustness of bistable switch p 53 AKT Mdm 2 Model Q 2 p 53 PTEN PIP 3 Mdm 2 AKT Model Q 3 Wee & Aguda (2006) Biophys J

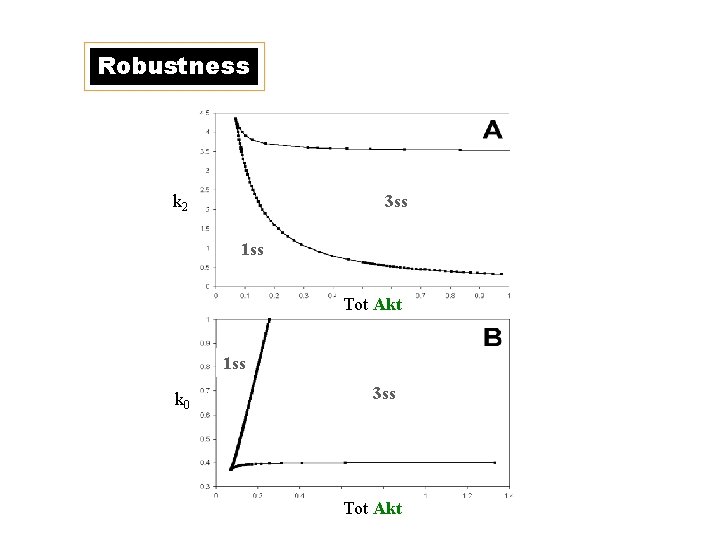
Robustness k 2 3 ss 1 ss Tot Akt 1 ss k 0 3 ss Tot Akt

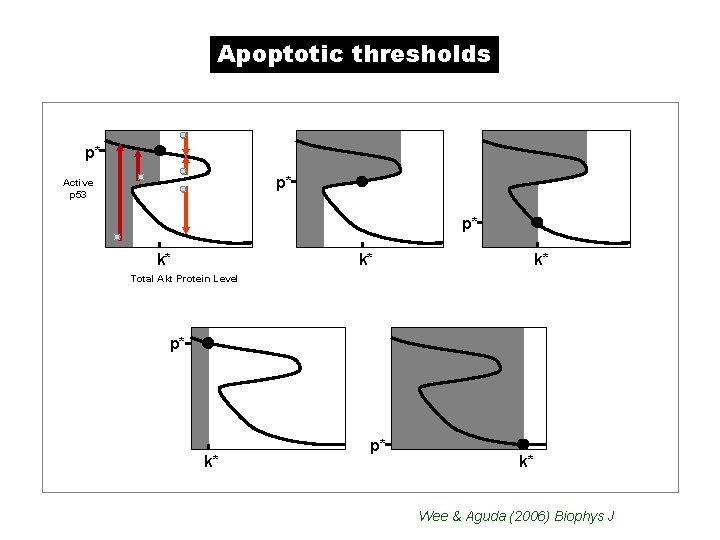
Apoptotic thresholds p* p* Active p 53 p* k* k* k* Total Akt Protein Level p* k* Wee & Aguda (2006) Biophys J

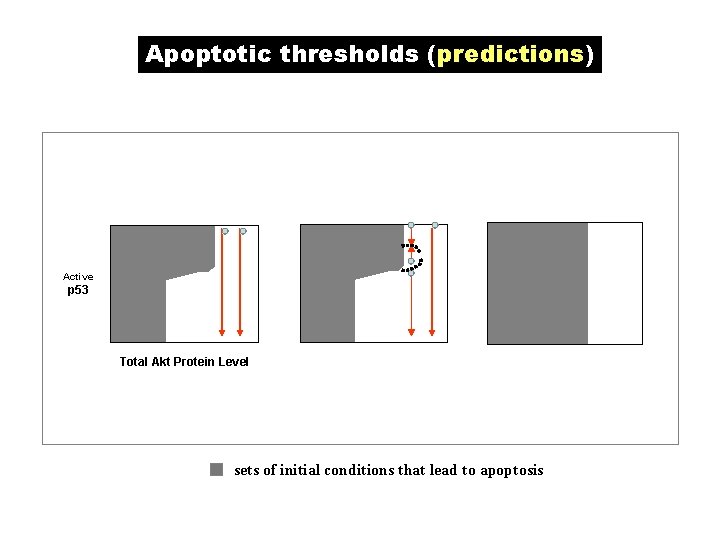
Apoptotic thresholds (predictions) Active p 53 Total Akt Protein Level sets of initial conditions that lead to apoptosis

Summary & Conclusions 1. Networks of oncogenes & TSGs in the G 1 -S cell cycle transition and p 53 -Akt interaction were analyzed. 2. There are unstable network motifs associated with G 1 and G 2 checkpoints which are targeted by signaling pathways to arrest or slow down the cell cycle. Oncogenes & TSGs are involved in these motifs and signaling. 3. The positive feedback loop between Akt and p 53 gives rise to a bistable switch that may govern a cell survival-death switch. Model predictions were given on apoptotic thresholds. 4. Modularization of networks coordinating the cell cycle and apoptosis were presented analyzed. 5. q. NET analysis is useful for network stability analysis & for model building.

ACKNOWLEDGEMENTS Collaborators: Keng Boon (Dave) Wee Ph. D Student, National University of Singapore Avner Friedman Mathematical Biosciences Institute Ohio State University, USA Gheorghe Craciun University of Wisconsin-Madison, USA

END