On Pulse The progressive Upper Level Set Scan



![ULS Nodal Tree [3] 3 18 [8] 8 [8, 7] 7 [3, 18, 0, ULS Nodal Tree [3] 3 18 [8] 8 [8, 7] 7 [3, 18, 0,](https://slidetodoc.com/presentation_image_h2/3926b52c11d208d3d11c3f0492c24aec/image-12.jpg)



















- Slides: 52

On Pulse: The progressive Upper Level Set Scan Statistic System for Geospatial and Spatio. Temporal Hotspot Detection Dr. G P Patil Dr. Luiz Duczmal Dr. Mural Haran Pushkar Patankar

Progressive Upper Level Set Scan (PULSE) l l l Pulse is a variant of Upper Level Set Scan (ULS) hotspot detection algorithm to detect arbitrary shaped hotspots in geospatial and spatial temporal data PULSE aims to increase the nodal quantity and quality of the upper level set tree Cells are ranked not just by a single response variable like disease rate but response variable such as likelihood values are also used to rank the cells

Demonstration Example

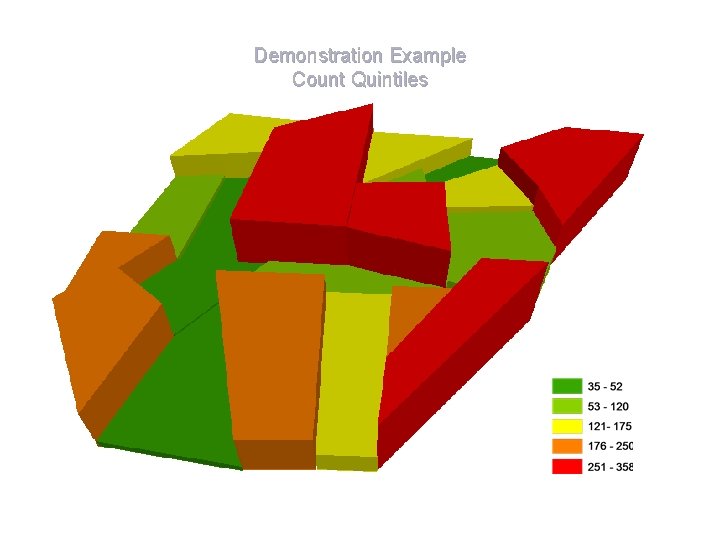
Demonstration Example Count Quintiles

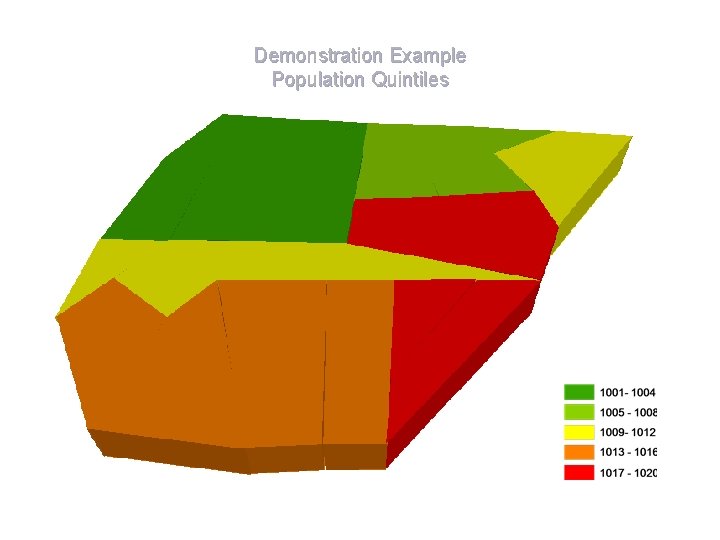
Demonstration Example Population Quintiles

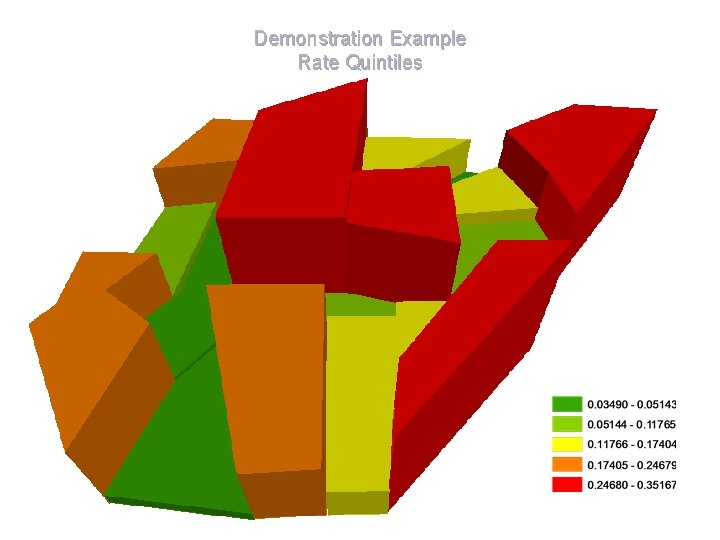
Demonstration Example Rate Quintiles

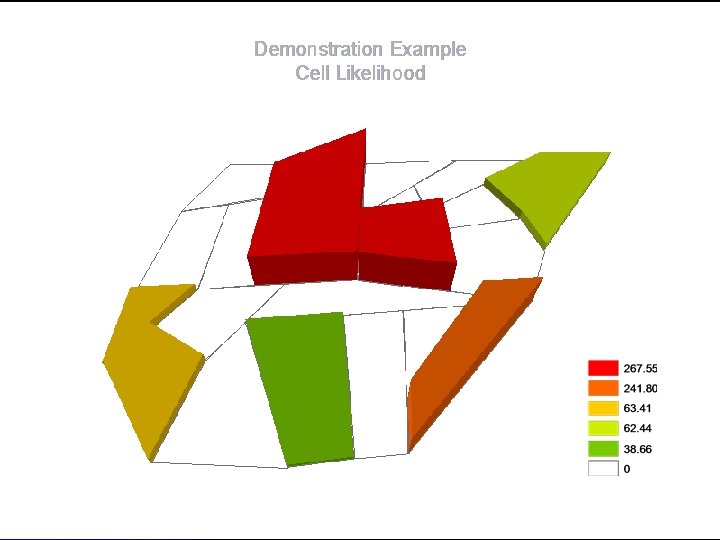
Demonstration Example Cell Likelihood

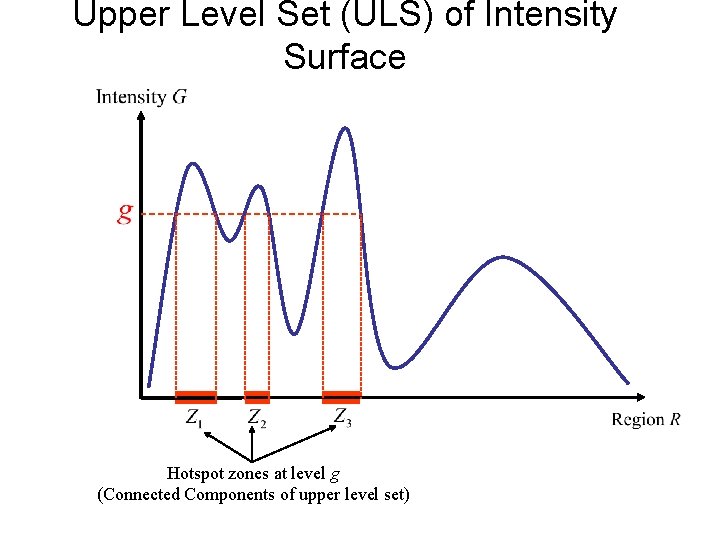
Upper Level Set (ULS) of Intensity Surface Hotspot zones at level g (Connected Components of upper level set)

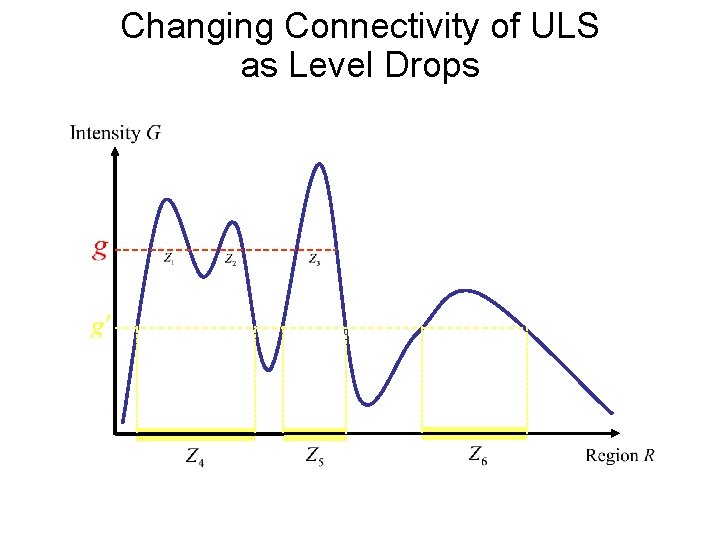
Changing Connectivity of ULS as Level Drops

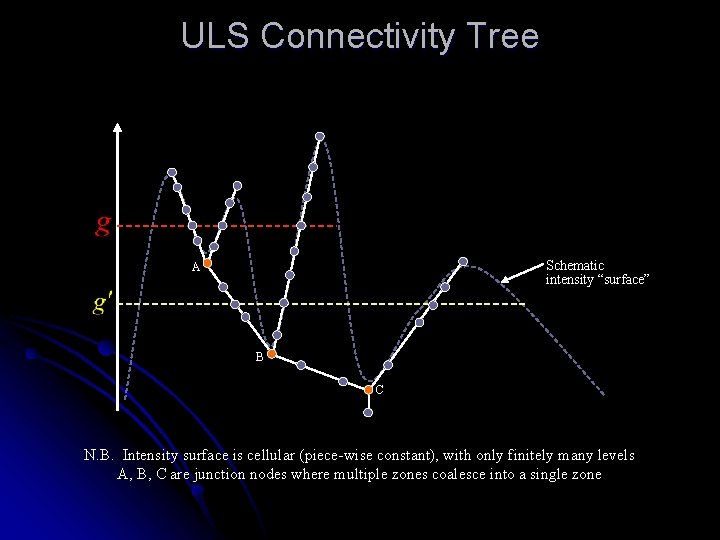
ULS Connectivity Tree Schematic intensity “surface” A B C N. B. Intensity surface is cellular (piece-wise constant), with only finitely many levels A, B, C are junction nodes where multiple zones coalesce into a single zone

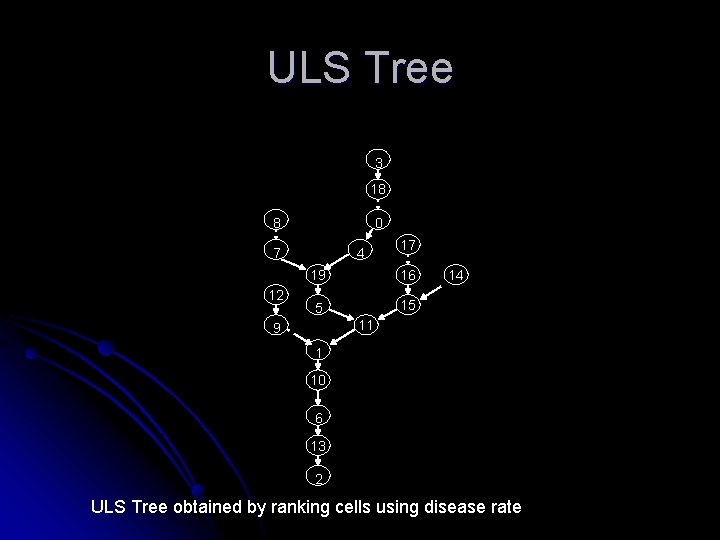
ULS Tree 3 18 0 8 4 7 12 17 19 16 5 15 14 11 9 1 10 6 13 2 ULS Tree obtained by ranking cells using disease rate
![ULS Nodal Tree 3 3 18 8 8 8 7 7 3 18 0 ULS Nodal Tree [3] 3 18 [8] 8 [8, 7] 7 [3, 18, 0,](https://slidetodoc.com/presentation_image_h2/3926b52c11d208d3d11c3f0492c24aec/image-12.jpg)
ULS Nodal Tree [3] 3 18 [8] 8 [8, 7] 7 [3, 18, 0, 4; 8, 7; 19] 12 0 [3, 18, 0, 4] 4 [3, 18] [3, 18, 0] [17] 17 19 [17, 16] 16 5 15 11 9 1 10 6 13 2 14 [14] [17, 16; 14; 15] [3, 18, 0, 4, 8, 7, 19, 5; 17, 16, 14, 15; 11]

Increasing number of zones l l For a ULS tree with m cells, at most m candidate zones will be produced To increase the number of candidate zones, multiple ULS trees need to be created Multiple ULS trees can be created by ranking cells by more than just disease rate Cells can be ordered randomly and ULS tree can be obtained from that ordering

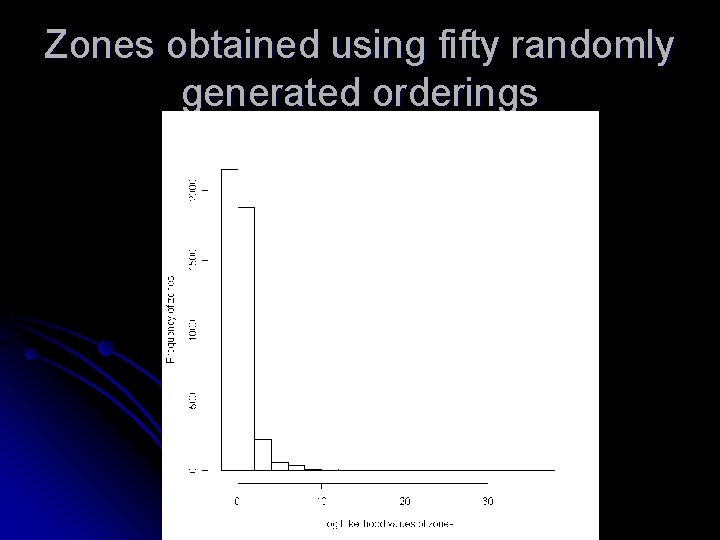
Zones obtained using fifty randomly generated orderings

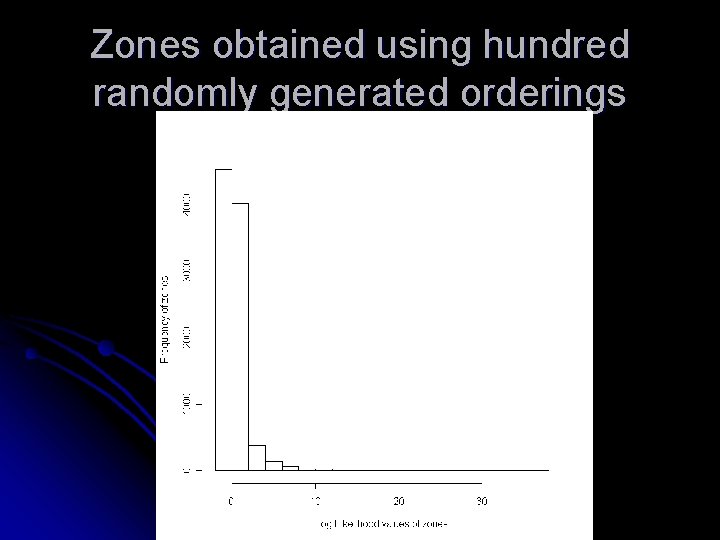
Zones obtained using hundred randomly generated orderings

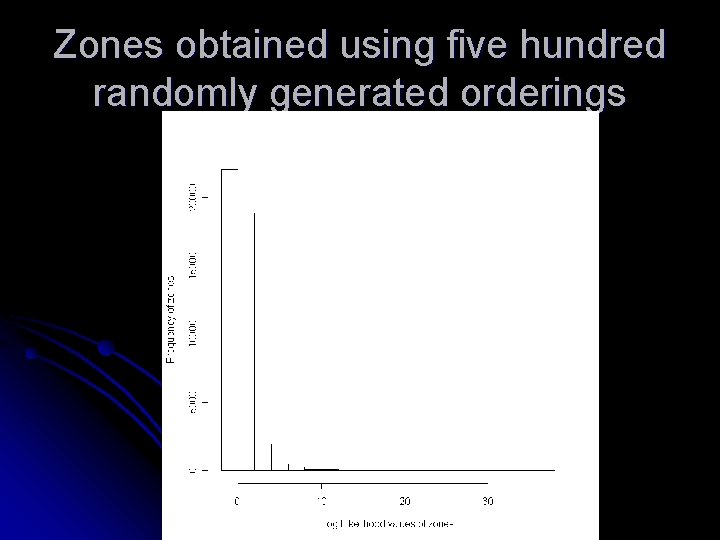
Zones obtained using five hundred randomly generated orderings

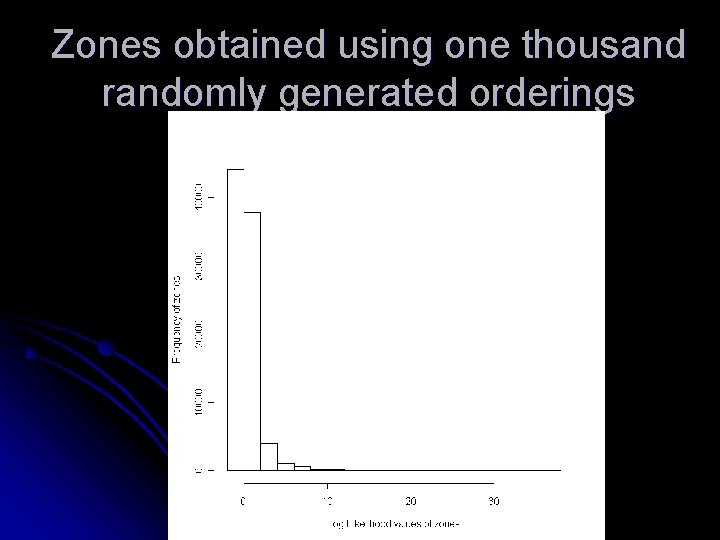
Zones obtained using one thousand randomly generated orderings

Observations Increasing the number of randomly generated orderings does not increase the quality of zones l The two orderings that generates zones with largest loglikelihood values are by disease rate and cell likelihood l Other orderings that can help in increasing the nodal quantity is still a matter of investigation l

Simulation Study l l Aims to compare three methods ULS, PULSE and SATSCAN in terms of their detection and delineation capability The parameters of interest are: l l Power of detection Overlap Missout Extraneous

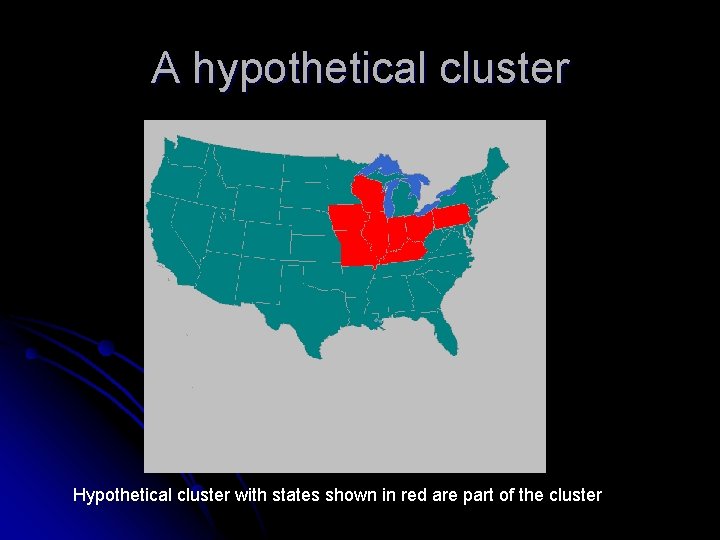
A hypothetical cluster Hypothetical cluster with states shown in red are part of the cluster

Cluster detected by a certain method States shown in yellow are part of a cluster obtained using a certain hotspot detection methodology

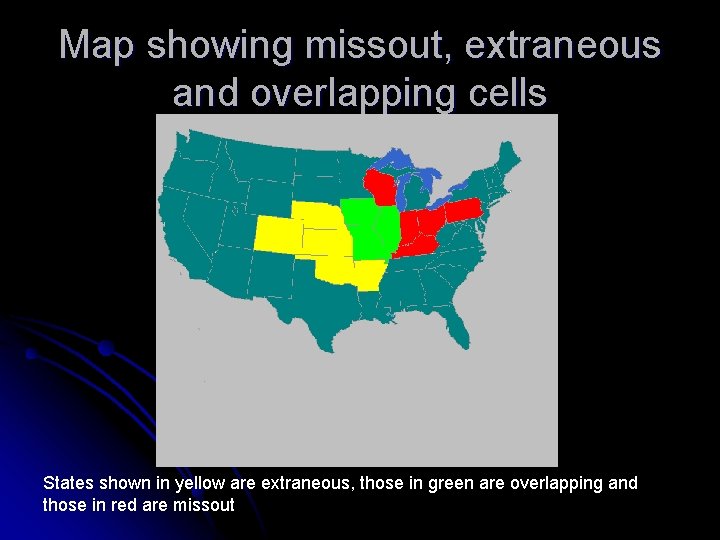
Map showing missout, extraneous and overlapping cells States shown in yellow are extraneous, those in green are overlapping and those in red are missout

Simulation Setup Region R divided into m cells l For each cell a є M, l l a cell intensity l ya cell count/cases l Na cell population l Average intensity = (1/N)∑ a Na l Total number of cases, ∑ya = y l Total population, ∑Na = N

Simulation Setup l ya ~ Poisson( a. Na) l f(ya) = e-( a. Na)ya/ya! y ~ Poisson( N) l ya|y ~Multinomial(y; {pa = raπa}) l Where, = a/ , relative risk of cell a l πa = Na /N, population fraction of cell a l ra This constitutes multinomial characterization of cell a

ULS Scan and Pulse Setup H 0: No cluster/hotspot: a= , i. e. ra = 1 for all a є M l HA: There is a cluster/hotspot z, i. e. l l a= z, a є z l a= z’, a є z’ l z > z’, which implies rz > rz’ Where, z = ∑ a Na, a є z, z’ = R - z

Circular Scan Setup H 0: No cluster/hotspot: a= , i. e. ra = 1 for all a є M l HA: rz > 1 and rz’ < 1 l Alternate hypothesis compares zone intensity z with average intensity l Alternative hypothesis of circular scan implies rz > rz’, but not on the components of inside and outside homogeneities l

Conditional Simulation with Hotspot z assumed known l l For z assumed known we should be able to reject H 0 for z at α =. 05 We should be able to control false positives for z at 1 – β =. 999 under HA Use the binomial of yz under H 0 to obtain the cutoff for α =. 05 and under HA to obtain the detectable RR rz for 1 – β =. 999 Use the binomial of yz’ under H 0 to obtain the cutoff for α =. 05 and under HA to obtain the detectable RR rz’ for 1 – β =. 999

Obtaining the cutoff using simulation H 0: ya|y ~Multinomial(y; {pa = raπa}) l Perform 100, 000 simulations to obtain 100, 000 values of yz l k. 05 = 95 th percentile of simulated yz l Solve for rz using, k. 05 = μA + 3. 09σA, where μA = rzπz and σ2 A = rzπz(1 - rzπz)/y l HA: ya~Multinomial(y; {rzπa, aєz}, {rz’πa, aєz’}) l

Simulation Data Simulation study has been performed on 1988 -1992 breast cancer mortality data on Northeastern united states. l Northeastern US map contains 245 counties spanning 10 states and district of Columbia l It has a total population of 29, 535, 201 women and has 58, 943 cases l

Assumed Hotspots of Different Types

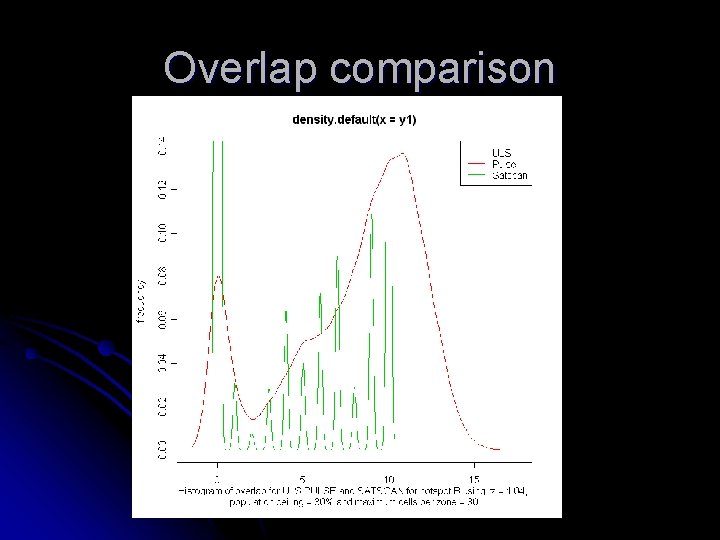
Overlap comparison

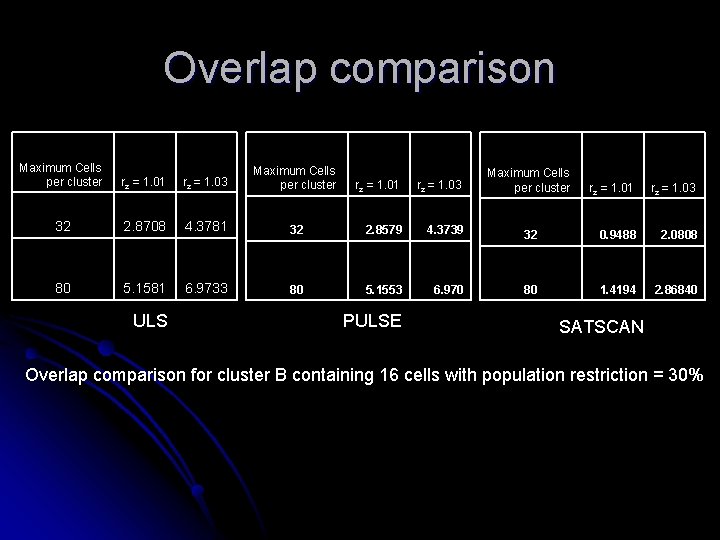
Overlap comparison Maximum Cells per cluster rz = 1. 01 rz = 1. 03 Maximum Cells per cluster 32 2. 8708 4. 3781 80 5. 1581 6. 9733 ULS rz = 1. 01 rz = 1. 03 32 2. 8579 4. 3739 80 5. 1553 6. 970 PULSE Maximum Cells per cluster rz = 1. 01 rz = 1. 03 32 0. 9488 2. 0808 80 1. 4194 2. 86840 SATSCAN Overlap comparison for cluster B containing 16 cells with population restriction = 30%

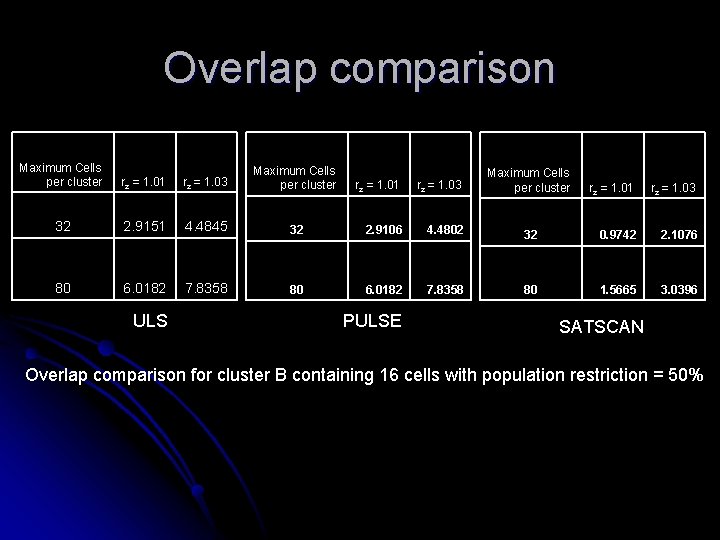
Overlap comparison Maximum Cells per cluster rz = 1. 01 rz = 1. 03 Maximum Cells per cluster 32 2. 9151 4. 4845 80 6. 0182 7. 8358 ULS rz = 1. 01 rz = 1. 03 32 2. 9106 4. 4802 80 6. 0182 7. 8358 PULSE Maximum Cells per cluster rz = 1. 01 rz = 1. 03 32 0. 9742 2. 1076 80 1. 5665 3. 0396 SATSCAN Overlap comparison for cluster B containing 16 cells with population restriction = 50%

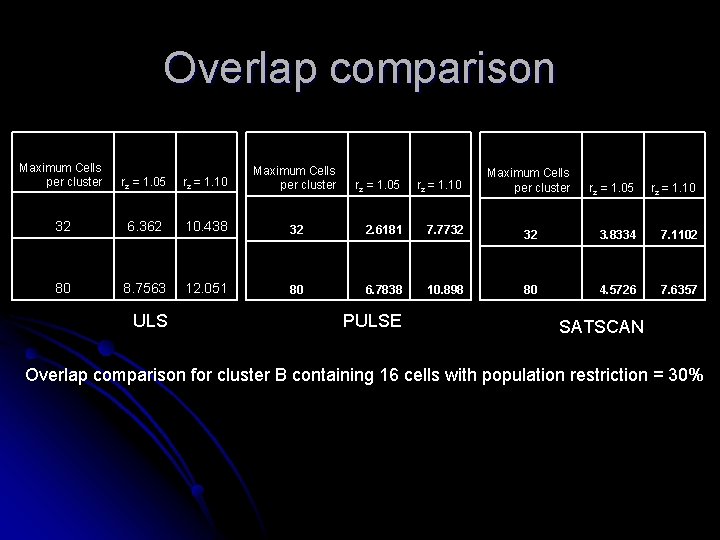
Overlap comparison Maximum Cells per cluster rz = 1. 05 rz = 1. 10 Maximum Cells per cluster 32 6. 362 10. 438 80 8. 7563 12. 051 ULS rz = 1. 05 rz = 1. 10 32 2. 6181 7. 7732 80 6. 7838 10. 898 PULSE Maximum Cells per cluster rz = 1. 05 rz = 1. 10 32 3. 8334 7. 1102 80 4. 5726 7. 6357 SATSCAN Overlap comparison for cluster B containing 16 cells with population restriction = 30%

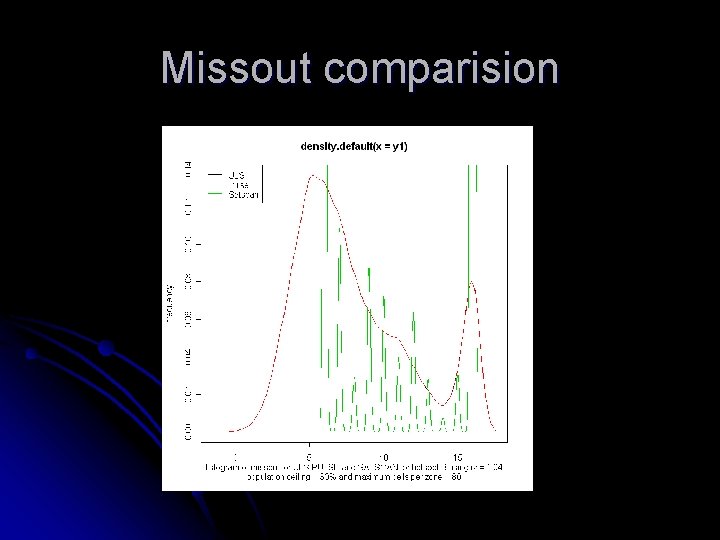
Missout comparision

Missout comparision Maximum Cells per cluster rz = 1. 01 rz = 1. 03 11. 6219 32 13. 1421 11. 6261 32 15. 0512 13. 9192 9. 0267 80 10. 8447 9. 0300 80 14. 5806 13. 1316 rz = 1. 01 rz = 1. 03 32 13. 1292 80 10. 8419 ULS PULSE SATSCAN Missout comparison for cluster B containing 16 cells with population restriction = 30%

Missout comparision Maximum Cells per cluster rz = 1. 01 rz = 1. 03 32 13. 0849 11. 515 32 13. 0849 11. 519 32 15. 0258 13. 892 80 9. 9818 8. 1642 80 9. 9817 8. 1642 80 14. 433 12. 9604 ULS PULSE SATSCAN Missout comparison for cluster B containing 16 cells with population restriction = 50%

Missout comparision Maximum Cells per cluster rz = 1. 05 rz = 1. 10 32 9. 638 5. 5616 32 13. 3819 80 7. 2437 3. 9495 80 9. 2162 ULS PULSE Maximum Cells per cluster rz = 1. 05 rz = 1. 10 8. 2268 32 12. 1666 8. 8898 5. 1019 80 11. 4274 8. 3643 SATSCAN Missout comparison for cluster B containing 16 cells with population restriction = 30%

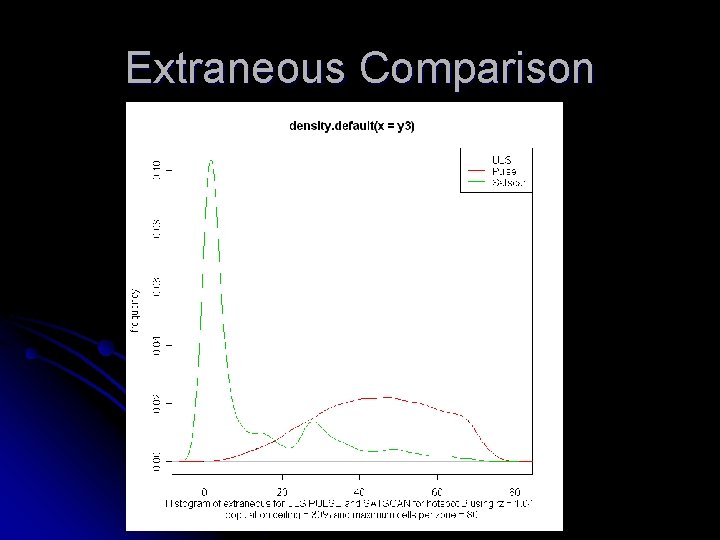
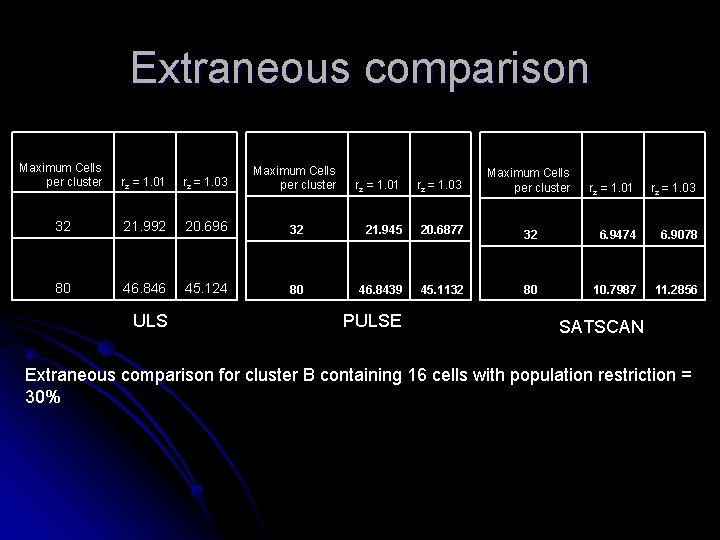
Extraneous Comparison

Extraneous comparison Maximum Cells per cluster rz = 1. 01 rz = 1. 03 Maximum Cells per cluster 32 21. 992 20. 696 80 46. 846 45. 124 ULS rz = 1. 01 rz = 1. 03 Maximum Cells per cluster 32 21. 945 20. 6877 80 46. 8439 45. 1132 PULSE rz = 1. 01 rz = 1. 03 32 6. 9474 6. 9078 80 10. 7987 11. 2856 SATSCAN Extraneous comparison for cluster B containing 16 cells with population restriction = 30%

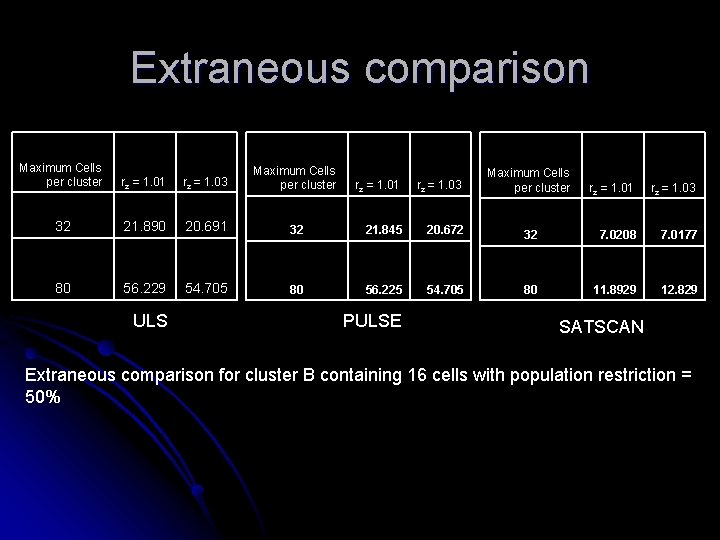
Extraneous comparison Maximum Cells per cluster rz = 1. 01 rz = 1. 03 Maximum Cells per cluster 32 21. 890 20. 691 80 56. 229 54. 705 ULS rz = 1. 01 rz = 1. 03 32 21. 845 20. 672 80 56. 225 54. 705 PULSE Maximum Cells per cluster rz = 1. 01 rz = 1. 03 32 7. 0208 7. 0177 80 11. 8929 12. 829 SATSCAN Extraneous comparison for cluster B containing 16 cells with population restriction = 50%

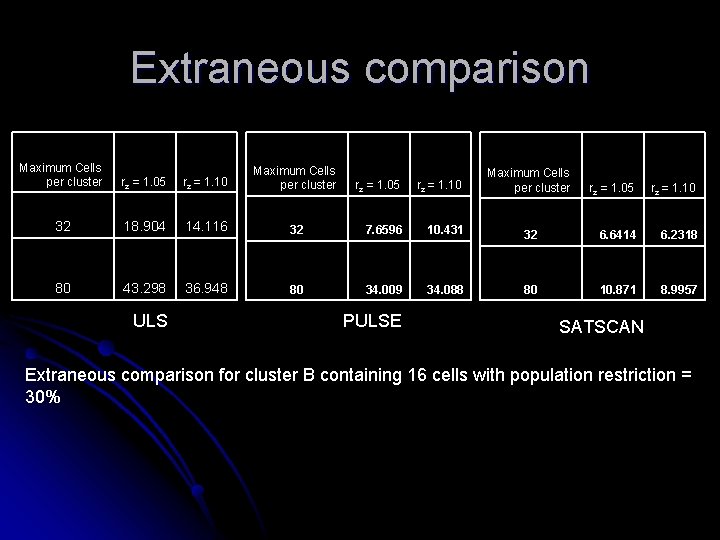
Extraneous comparison Maximum Cells per cluster rz = 1. 05 rz = 1. 10 Maximum Cells per cluster 32 18. 904 14. 116 80 43. 298 36. 948 ULS rz = 1. 05 rz = 1. 10 32 7. 6596 10. 431 80 34. 009 34. 088 PULSE Maximum Cells per cluster rz = 1. 05 rz = 1. 10 32 6. 6414 6. 2318 80 10. 871 8. 9957 SATSCAN Extraneous comparison for cluster B containing 16 cells with population restriction = 30%

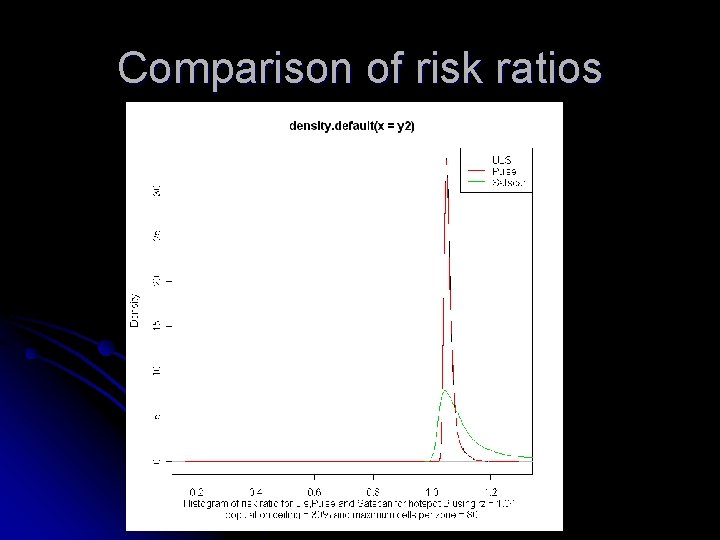
Comparison of risk ratios

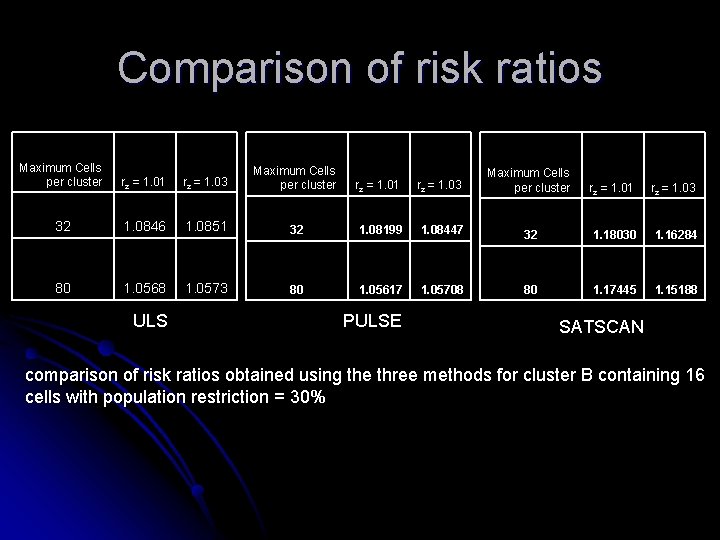
Comparison of risk ratios Maximum Cells per cluster rz = 1. 01 rz = 1. 03 32 1. 0846 1. 0851 32 1. 08199 1. 08447 32 1. 18030 1. 16284 80 1. 0568 1. 0573 80 1. 05617 1. 05708 80 1. 17445 1. 15188 ULS PULSE SATSCAN comparison of risk ratios obtained using the three methods for cluster B containing 16 cells with population restriction = 30%

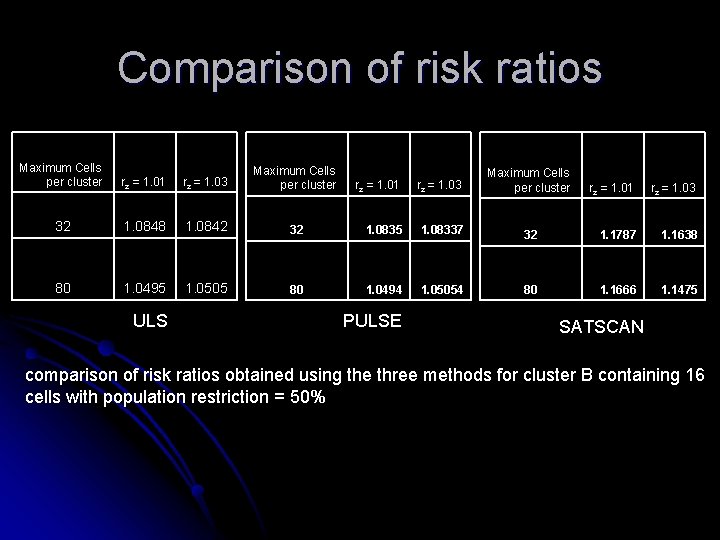
Comparison of risk ratios Maximum Cells per cluster rz = 1. 01 rz = 1. 03 Maximum Cells per cluster 32 1. 0848 1. 0842 80 1. 0495 1. 0505 ULS rz = 1. 01 rz = 1. 03 Maximum Cells per cluster 32 1. 0835 1. 08337 80 1. 0494 1. 05054 PULSE rz = 1. 01 rz = 1. 03 32 1. 1787 1. 1638 80 1. 1666 1. 1475 SATSCAN comparison of risk ratios obtained using the three methods for cluster B containing 16 cells with population restriction = 50%

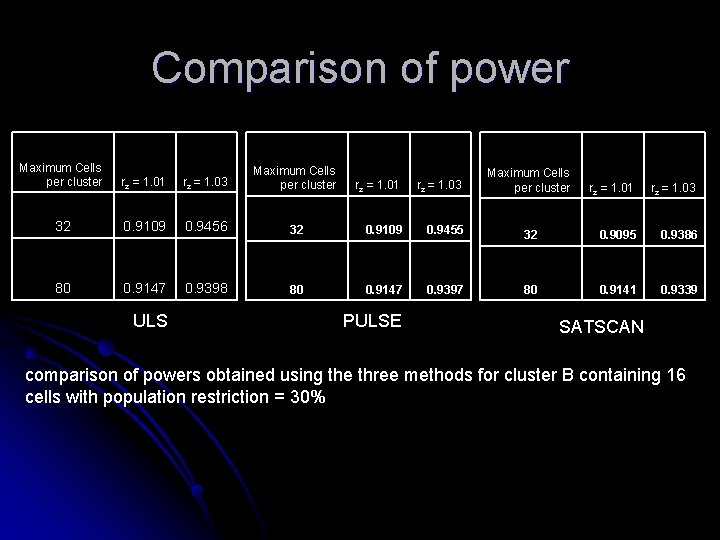
Comparison of power Maximum Cells per cluster rz = 1. 01 rz = 1. 03 Maximum Cells per cluster 32 0. 9109 0. 9456 80 0. 9147 0. 9398 ULS rz = 1. 01 rz = 1. 03 32 0. 9109 0. 9455 80 0. 9147 0. 9397 PULSE Maximum Cells per cluster rz = 1. 01 rz = 1. 03 32 0. 9095 0. 9386 80 0. 9141 0. 9339 SATSCAN comparison of powers obtained using the three methods for cluster B containing 16 cells with population restriction = 30%

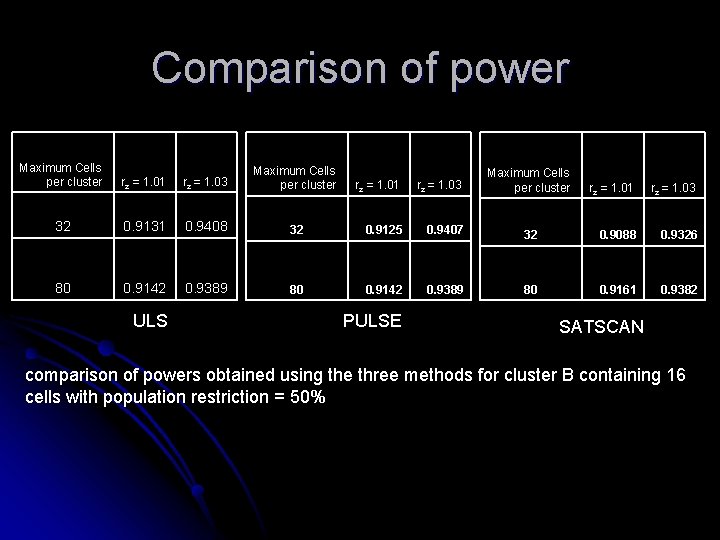
Comparison of power Maximum Cells per cluster rz = 1. 01 rz = 1. 03 Maximum Cells per cluster 32 0. 9131 0. 9408 80 0. 9142 0. 9389 ULS rz = 1. 01 rz = 1. 03 32 0. 9125 0. 9407 80 0. 9142 0. 9389 PULSE Maximum Cells per cluster rz = 1. 01 rz = 1. 03 32 0. 9088 0. 9326 80 0. 9161 0. 9382 SATSCAN comparison of powers obtained using the three methods for cluster B containing 16 cells with population restriction = 50%

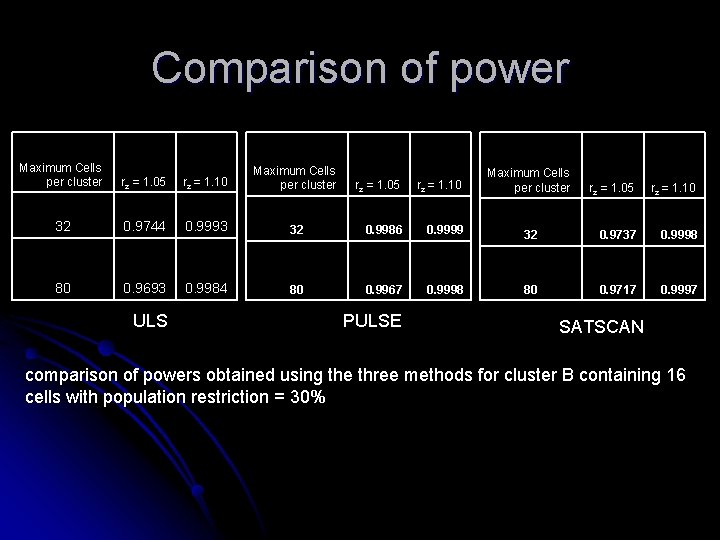
Comparison of power Maximum Cells per cluster rz = 1. 05 rz = 1. 10 Maximum Cells per cluster 32 0. 9744 0. 9993 80 0. 9693 0. 9984 ULS rz = 1. 05 rz = 1. 10 32 0. 9986 0. 9999 80 0. 9967 0. 9998 PULSE Maximum Cells per cluster rz = 1. 05 rz = 1. 10 32 0. 9737 0. 9998 80 0. 9717 0. 9997 SATSCAN comparison of powers obtained using the three methods for cluster B containing 16 cells with population restriction = 30%

Conclusions ULS and PULSE has a greater power of detection as compared to SATSCAN l Power of detection increases as the risk ratio increases l ULS and PULSE has lower missouts as compared to SATSCAN l As the risk ratio increases, missout decreases l

Conclusions l l ULS and PULSE has greater extra cells in the hotspot as compared to SATSCAN The number of extra cells identified by the three methods decreases as risk ratio increases ULS and PULSE has greater number of overlapping cells as compared to SATSCAN Number of overlapping cells increases as the risk ratio increases

Conclusions l l l Zones identified by SATSCAN tend to have larger risk ratios than those identified by ULS and PULSE Missouts are not controlled by allowable maximum population in the zone or by controlling the maximum number of cells that can be a part of a zone The three methods generally have a pretty high detection power, but as the value of risk ratio decreases, their delineation capability is greatly reduced

Thank You