Omics informatics Biotechnology Genomics Proteomics Metabolomics Genome informatics



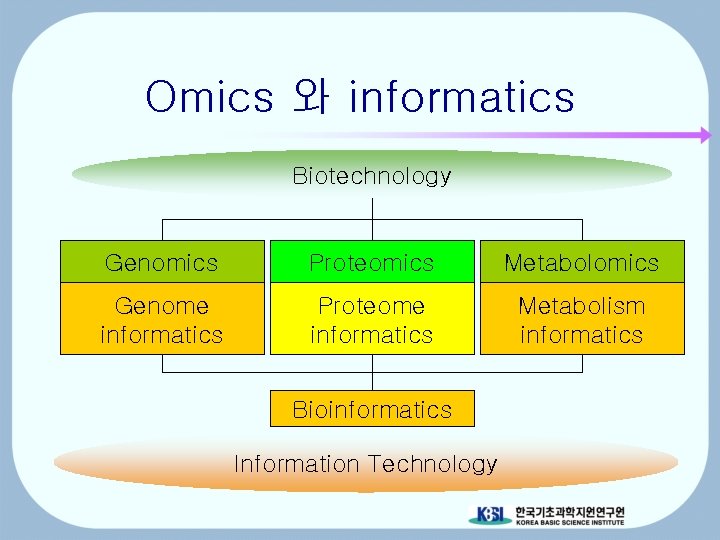
Omics 와 informatics Biotechnology Genomics Proteomics Metabolomics Genome informatics Proteome informatics Metabolism informatics Bioinformatics Information Technology

Omics 의 관계 metabolomics Molecular machinery of cellular processes proteomics Characterization of proteome genomics Genome sequencing 1989 Peptide mass fingerprinting 1986 peptide sequencing by MS/MS



프로테옴 데이터 electrophoresis Mass spectrometry Database search 단백질 분리 펩타이드 분자량 측정 단백질 동정
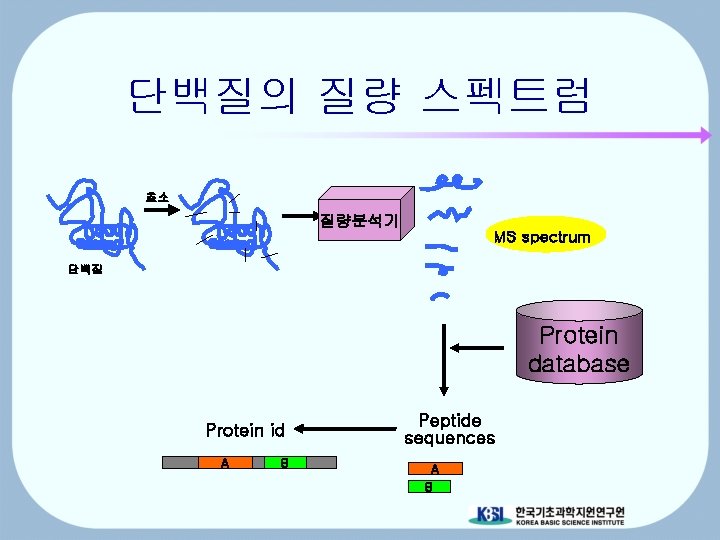
단백질의 질량 스펙트럼 효소 질량분석기 MS spectrum 단백질 Protein database Protein id A B Peptide sequences A B
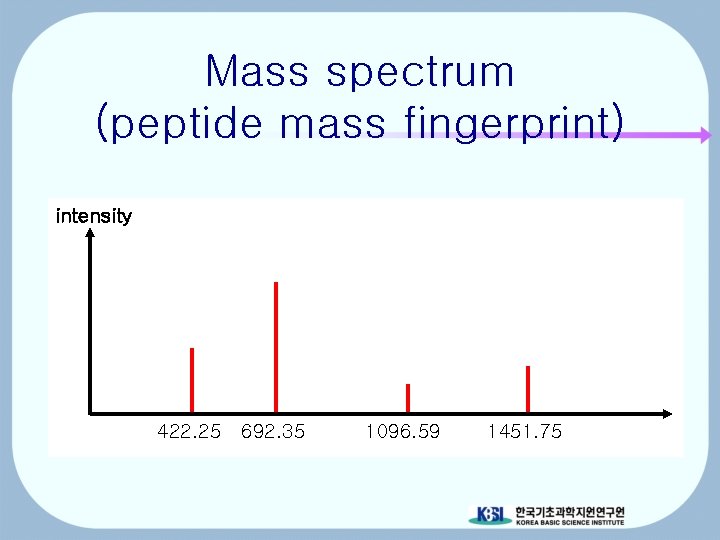
Mass spectrum (peptide mass fingerprint) intensity 422. 25 692. 35 1096. 59 1451. 75
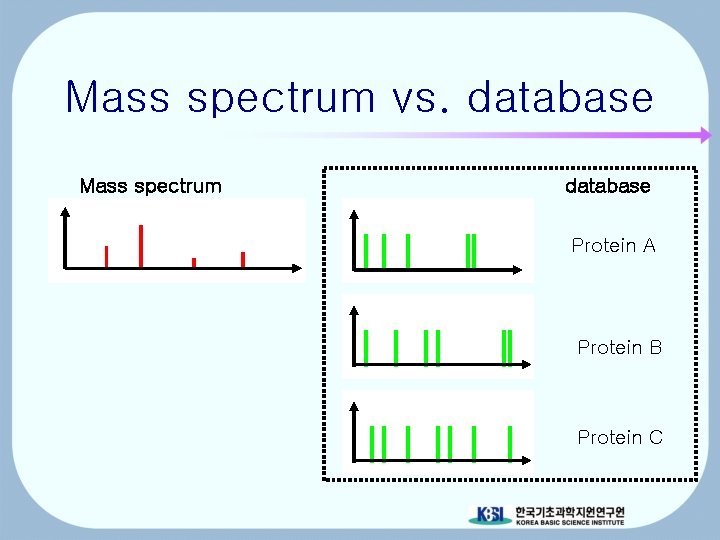
Mass spectrum vs. database Mass spectrum database Protein A Protein B Protein C
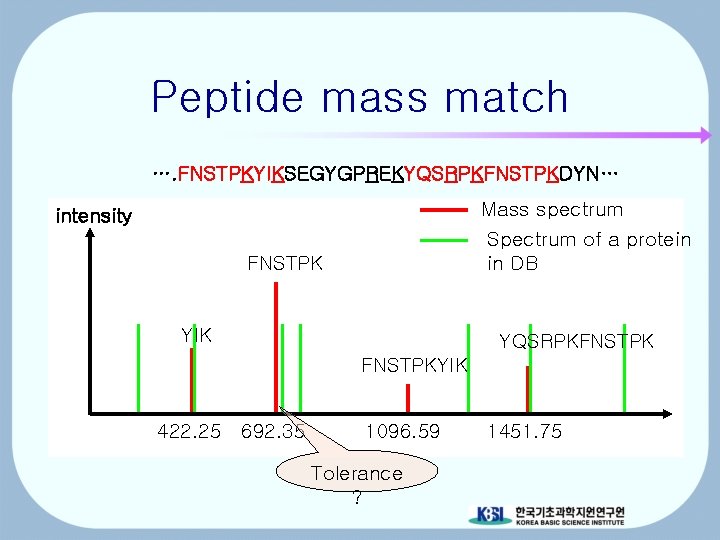
Peptide mass match …. FNSTPKYIKSEGYGPREKYQSRPKFNSTPKDYN… Mass spectrum Spectrum of a protein in DB intensity FNSTPK YIK YQSRPKFNSTPKYIK 422. 25 692. 35 1096. 59 Tolerance ? 1451. 75

단백질 데이터베이스 non-redundant protein DB 효소: trypsin 0 Protein mass (0 -150 k. Da) Peptide mass (0 -12. 5 k. Da)

PMF search programs MS-Fit http: //prospector. ucsf. edu/ MASCOT http: //www. matrixscience. com/ Pept. Ident http: //www. expasy. org/ Peptide. Search http: //www. mann. embl-heidelberg. de/
MS-Fit
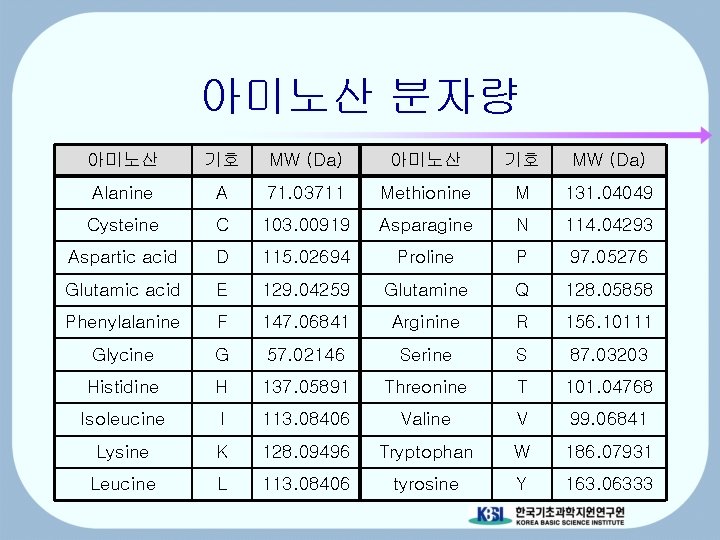
아미노산 분자량 아미노산 기호 MW (Da) Alanine A 71. 03711 Methionine M 131. 04049 Cysteine C 103. 00919 Asparagine N 114. 04293 Aspartic acid D 115. 02694 Proline P 97. 05276 Glutamic acid E 129. 04259 Glutamine Q 128. 05858 Phenylalanine F 147. 06841 Arginine R 156. 10111 Glycine G 57. 02146 Serine S 87. 03203 Histidine H 137. 05891 Threonine T 101. 04768 Isoleucine I 113. 08406 Valine V 99. 06841 Lysine K 128. 09496 Tryptophan W 186. 07931 Leucine L 113. 08406 tyrosine Y 163. 06333

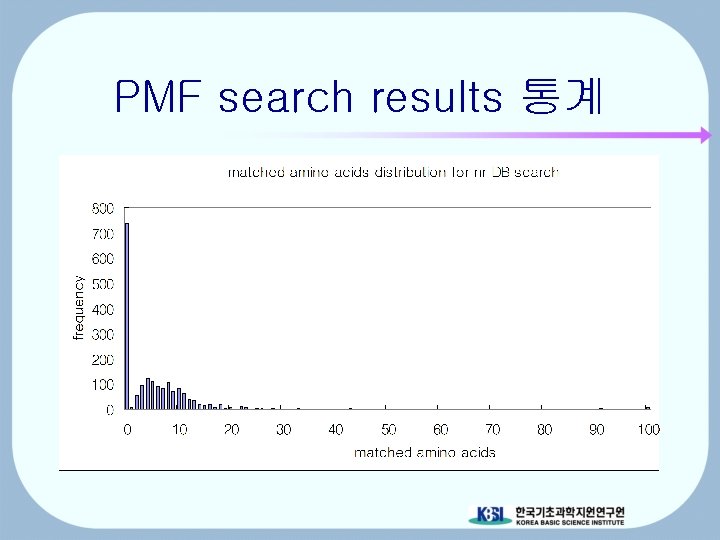
PMF search results 통계

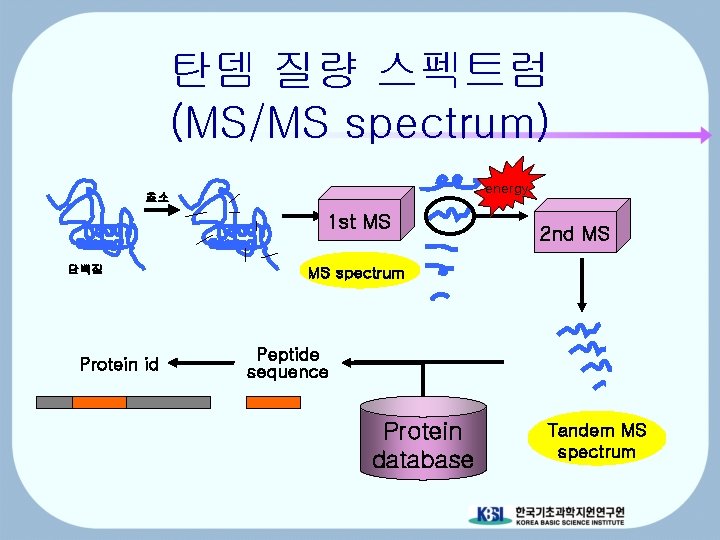
탄뎀 질량 스펙트럼 (MS/MS spectrum) energy 효소 1 st MS 단백질 Protein id 2 nd MS MS spectrum Peptide sequence Protein database Tandem MS spectrum
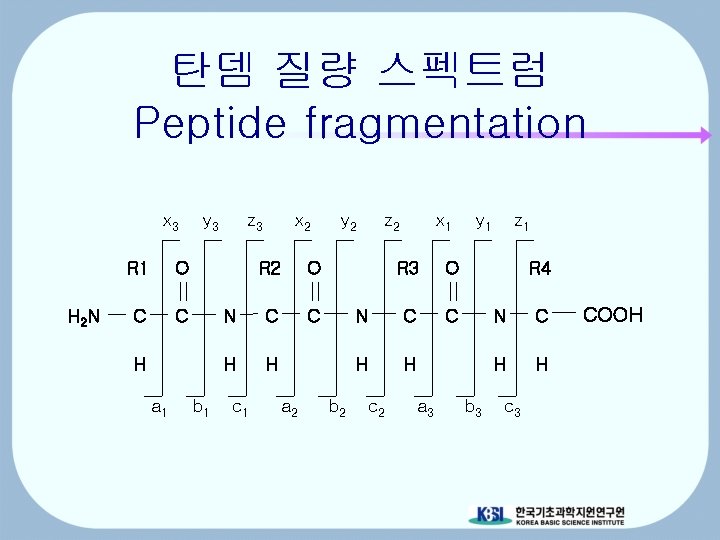
탄뎀 질량 스펙트럼 Peptide fragmentation x 3 H 2 N R 1 O C C y 3 H a 1 b 1 z 3 x 2 R 2 O N C C H H c 1 a 2 y 2 b 2 z 2 x 1 R 3 O N C C H H c 2 a 3 y 1 z 1 R 4 b 3 N C H H c 3 COOH

단백질 동정 (protein identification) intensity Peptide 의 탄뎀 질량 스펙트럼 compare intensity m/z a spectrum of a peptide in database m/z
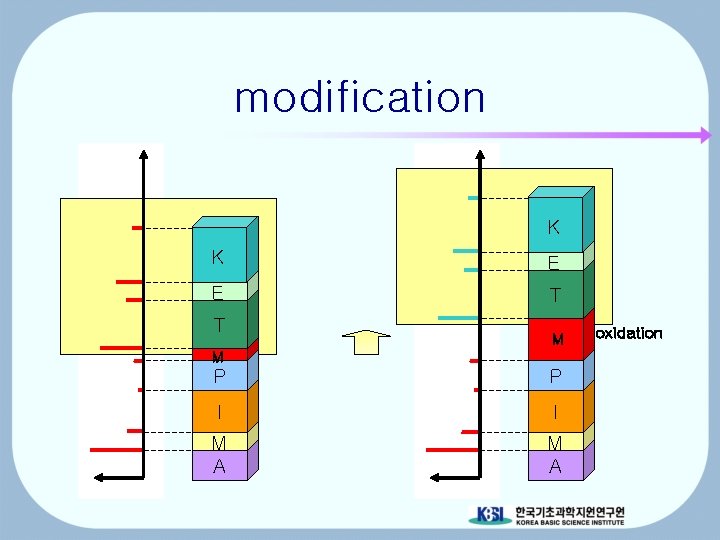
modification K K E E T T M M P P I I M A oxidation

MS/MS Search programs MASCOT http: //www. matrixscience. com/ MS-TAG http: //prospector. ucsf. edu/ SEQUEST http: //fields. scripps. edu/sequest/
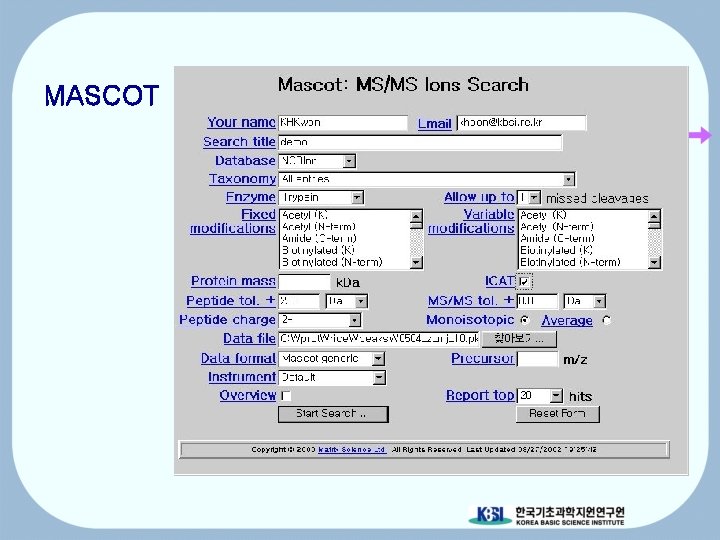
MASCOT
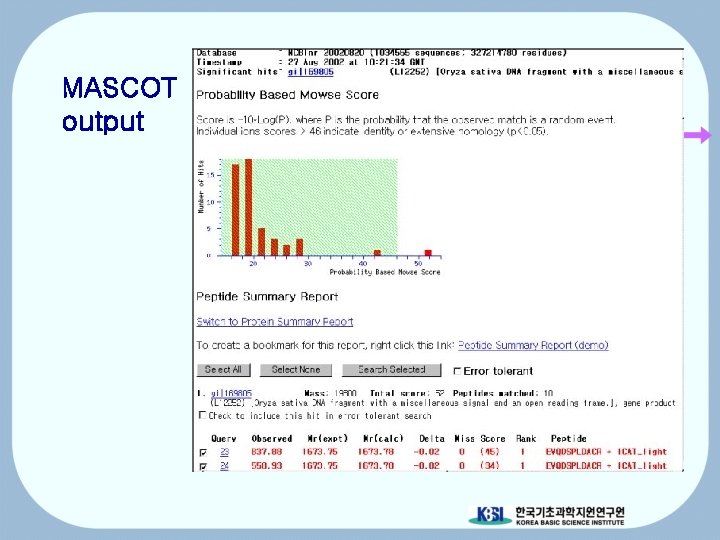
MASCOT output
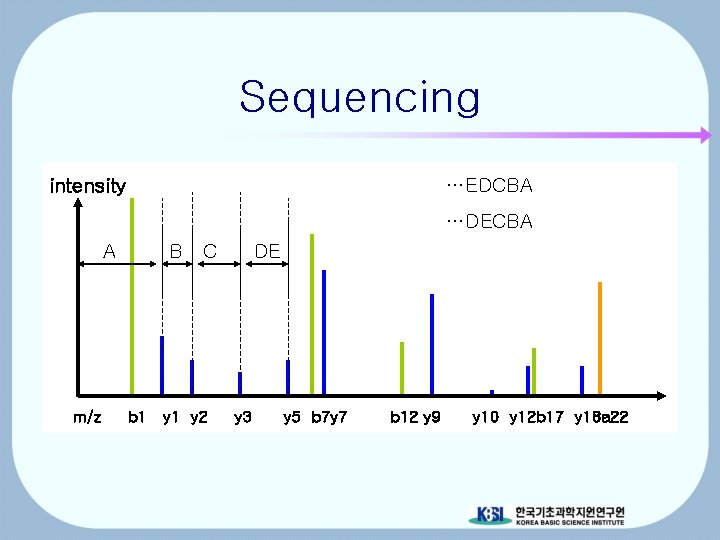
Sequencing intensity …EDCBA …DECBA A m/z B C b 1 y 2 DE y 3 y 5 b 7 y 7 b 12 y 9 y 10 y 12 b 17 y 18 a 22
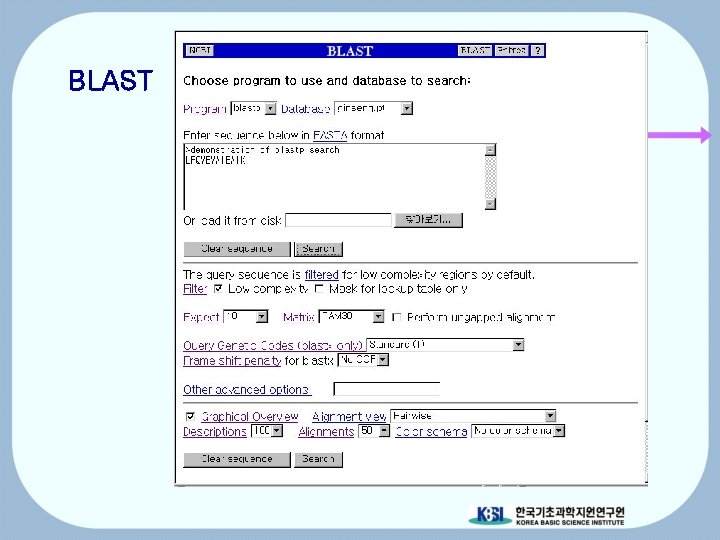
BLAST

Scores Homology search (BLAST) alignment score e-value Peptide Mass Fingerprint Number of matched peptides Coverage MOWSE score Probability based on MOWSE score Tandem mass search Cross-correlation coverage


Proteomics Trends 1. High-throughput Proteomics 2. High-throughput automated identification of proteins 3. 2. Quantitation 4. Quantitation of relative protein abundance
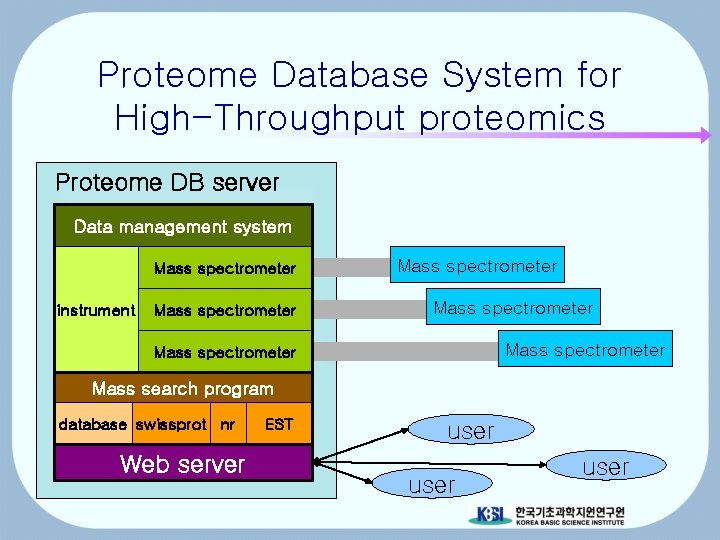
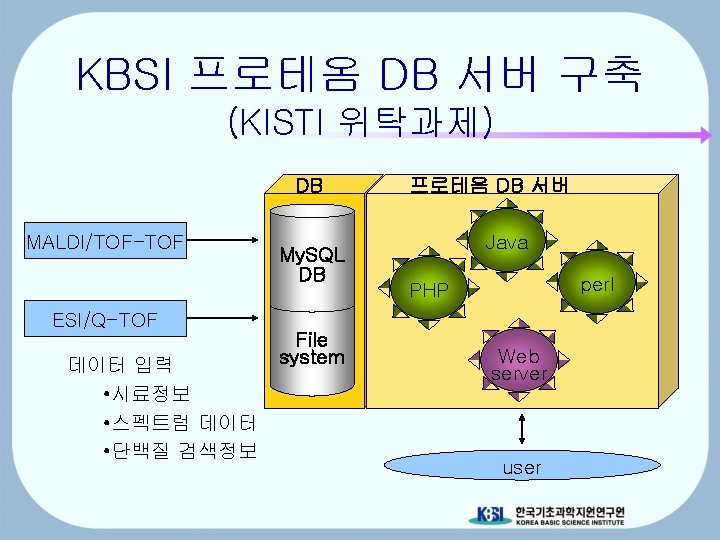
Proteome Database System for High-Throughput proteomics Proteome DB server Data management system Mass spectrometer instrument Mass spectrometer Mass spectrometer Mass search program database swissprot nr Web server EST user


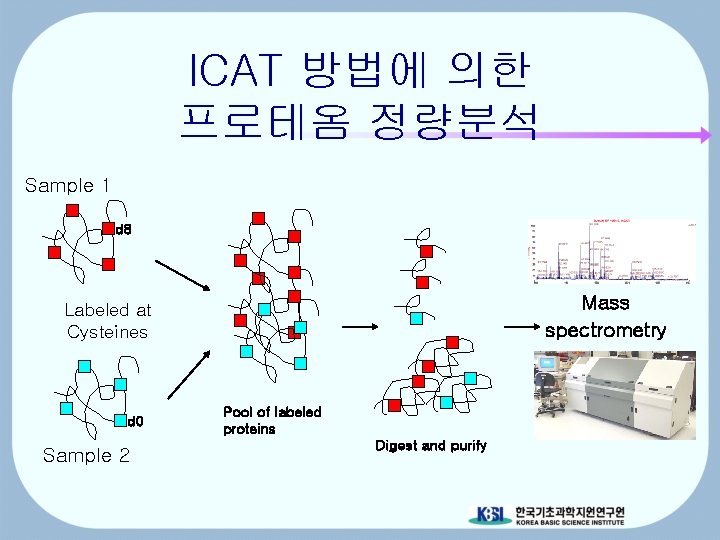
ICAT 방법에 의한 프로테옴 정량분석 Sample 1 d 8 Mass spectrometry Labeled at Cysteines d 0 Sample 2 Pool of labeled proteins Digest and purify
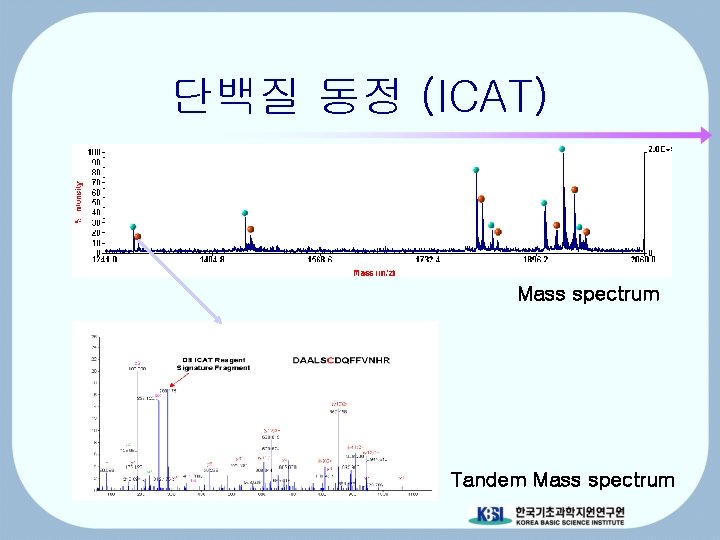
단백질 동정 (ICAT) Mass spectrum Tandem Mass spectrum


- Slides: 39