Of Mice and Motifs and Best Laid Plans

Of Mice and Motifs and Best Laid Plans Michael Smith Genome Sciences Centre Terry Fox Laboratory British Columbia Cancer Agency, Vancouver CMMT, Vancouver Genome Sciences Centre BC Cancer Agency

Of mice… • To measure gene expression levels in tissues of developing mice to gain insight into the normal development process. • To develop supporting technologies and techniques to improve the process for generating and analysing this data Genome Sciences Centre BC Cancer Agency
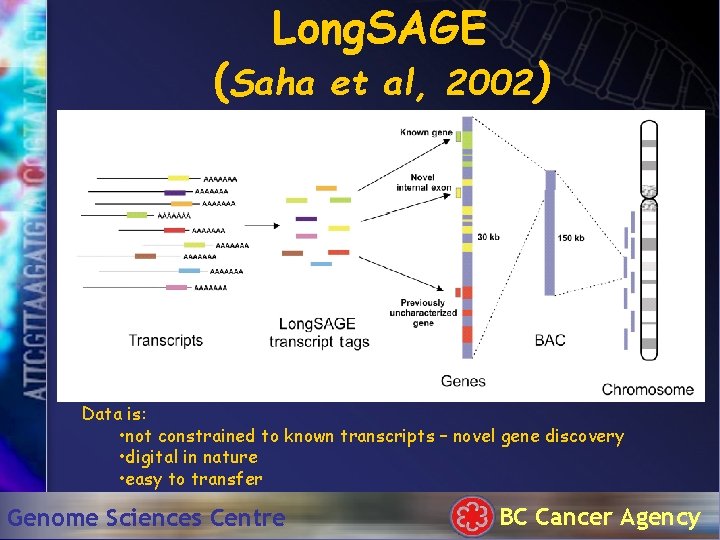
Long. SAGE (Saha et al, 2002) Data is: • not constrained to known transcripts – novel gene discovery • digital in nature • easy to transfer Genome Sciences Centre BC Cancer Agency

Overview SAGE data • 2 dataset – October Freeze • • 72 21 -mer libraries 8. 55 million tags 924, 392 unique tag types 49 tissues, 25 developmental stages – January Freeze • 105 21 -mer libraries (92 fully sequenced) • 11. 65 million • 1, 235, 833 unique tag types Genome Sciences Centre BC Cancer Agency
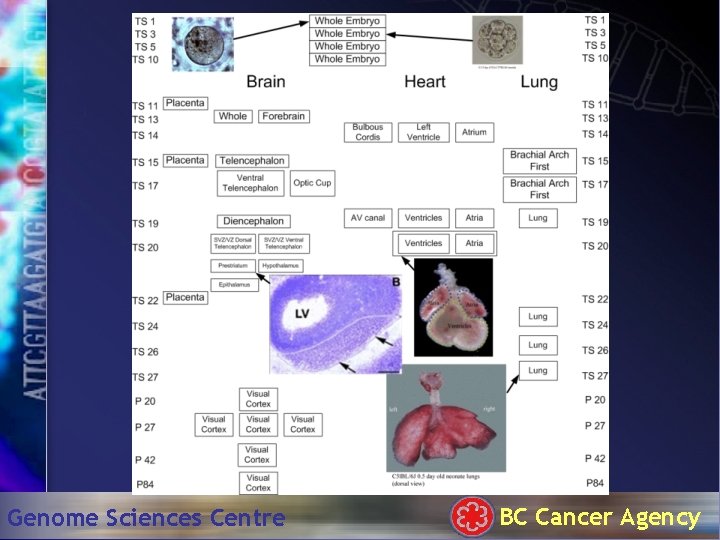
Genome Sciences Centre BC Cancer Agency
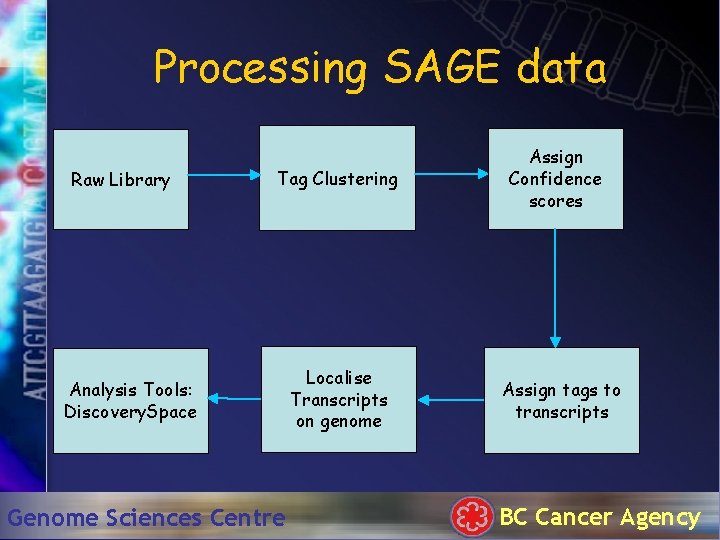
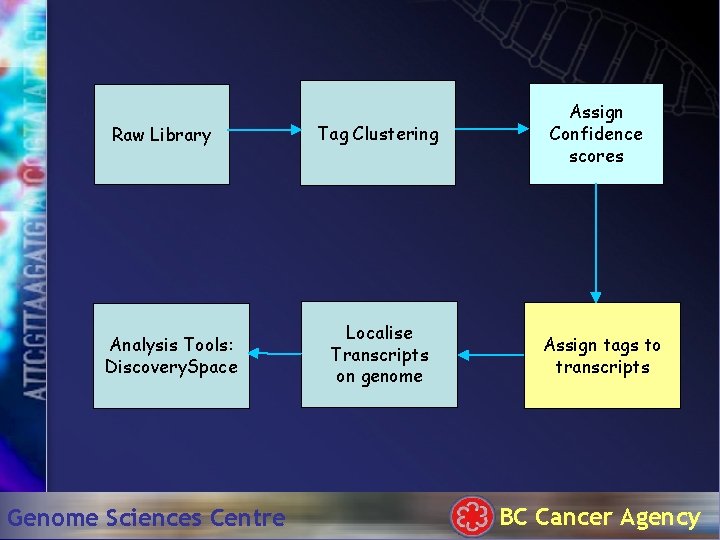
Processing SAGE data Raw Library Tag Clustering Analysis Tools: Discovery. Space Genome Sciences Centre Localise Transcripts on genome Assign Confidence scores Assign tags to transcripts BC Cancer Agency
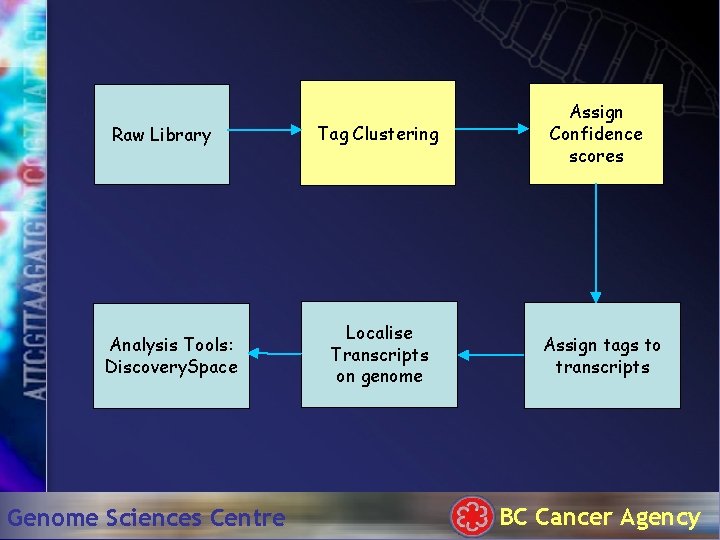
Raw Library Analysis Tools: Discovery. Space Genome Sciences Centre Tag Clustering Localise Transcripts on genome Assign Confidence scores Assign tags to transcripts BC Cancer Agency

Tag Clustering • Tag Types cluster in tag space – Colinge and Ferge, 2001 • PCR error + Sequencing error – Akmaev and Wong, 2004 • We have used real PHRED values to quantify p-values per tag type Genome Sciences Centre BC Cancer Agency
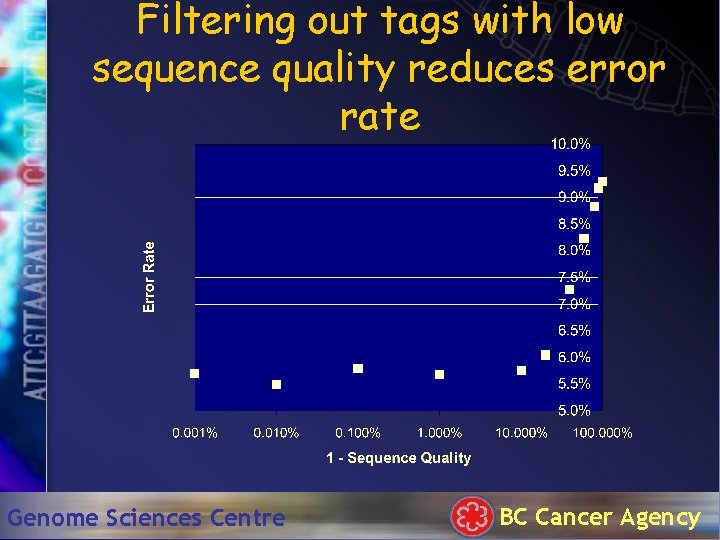
Filtering out tags with low sequence quality reduces error rate Genome Sciences Centre BC Cancer Agency

Tag/ Tag Type Confidence • Individual Tag Error = (Base Library Error) combined with (Tag Sequence Error) • Combine Individual Tag Errors to generate Tag Type errors for each library • Combine Tag Type errors from each library to generate Tag Type error for the metalibrary Genome Sciences Centre BC Cancer Agency
Raw Library Analysis Tools: Discovery. Space Genome Sciences Centre Tag Clustering Localise Transcripts on genome Assign Confidence scores Assign tags to transcripts BC Cancer Agency
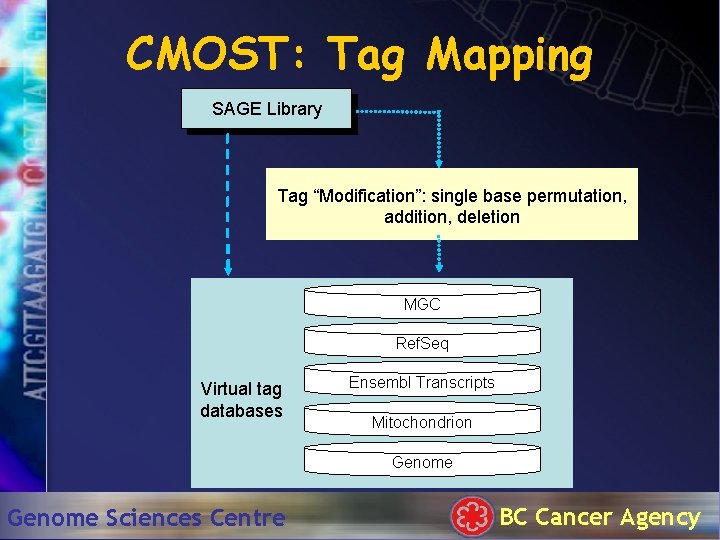
CMOST: Tag Mapping SAGE Library Tag “Modification”: single base permutation, addition, deletion MGC Ref. Seq Virtual tag databases Ensembl Transcripts Mitochondrion Genome Sciences Centre BC Cancer Agency
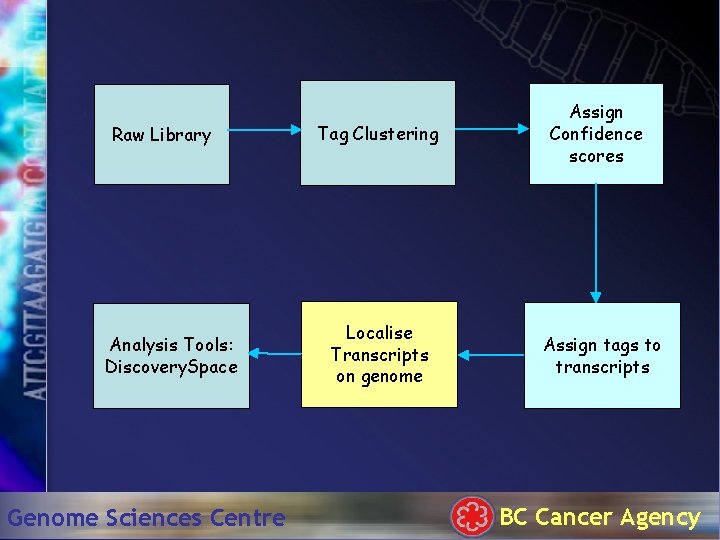
Raw Library Analysis Tools: Discovery. Space Genome Sciences Centre Tag Clustering Localise Transcripts on genome Assign Confidence scores Assign tags to transcripts BC Cancer Agency
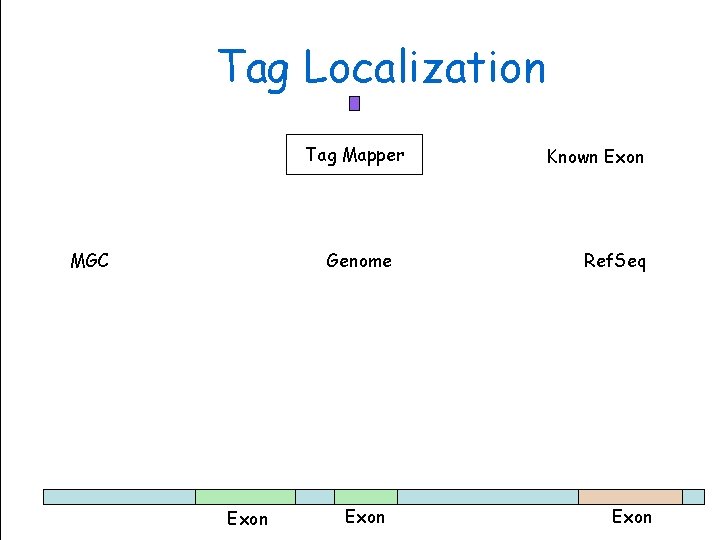
Tag Localization Tag Mapper MGC Exon Genome Sciences Centre Genome Exon Known Exon Ref. Seq BC Cancer. Exon Agency
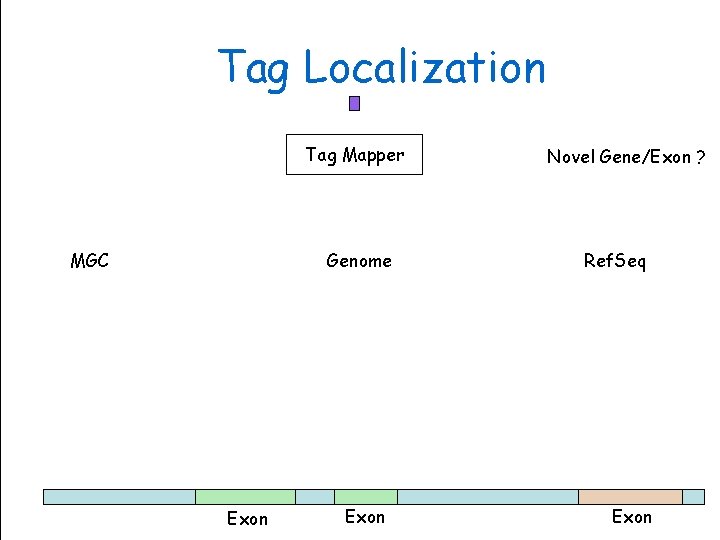
Tag Localization Tag Mapper MGC Exon Genome Sciences Centre Genome Exon Novel Gene/Exon ? Ref. Seq BC Cancer. Exon Agency
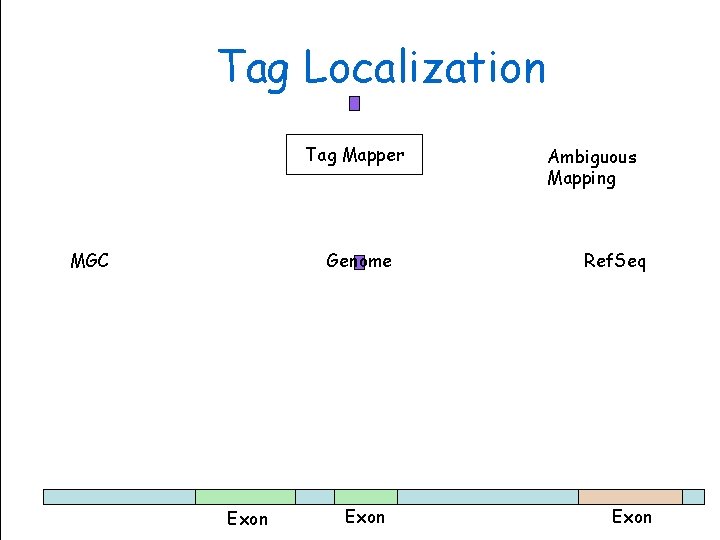
Tag Localization Tag Mapper MGC Exon Genome Sciences Centre Genome Exon Ambiguous Mapping Ref. Seq BC Cancer. Exon Agency
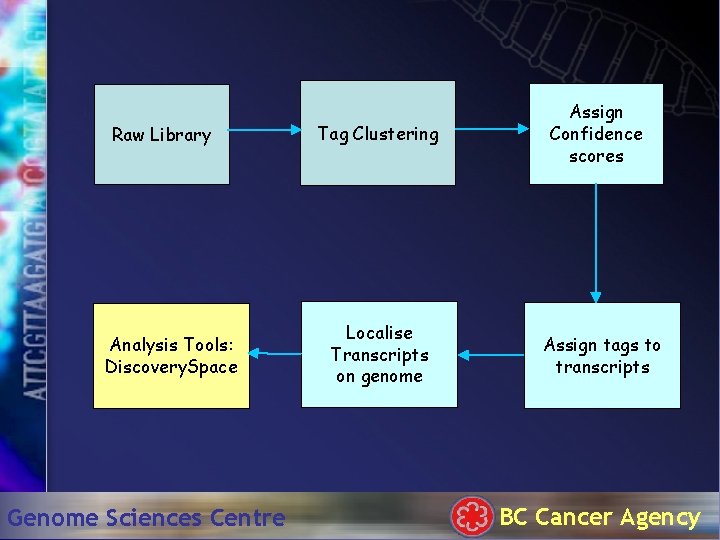
Raw Library Analysis Tools: Discovery. Space Genome Sciences Centre Tag Clustering Localise Transcripts on genome Assign Confidence scores Assign tags to transcripts BC Cancer Agency
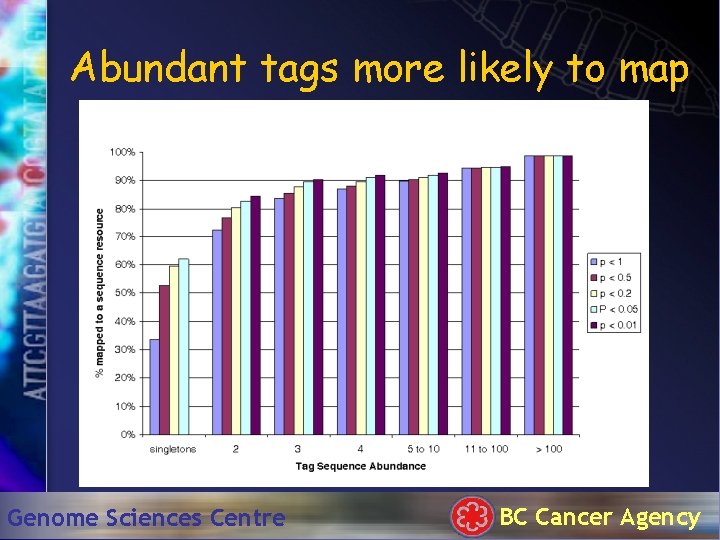
Abundant tags more likely to map Genome Sciences Centre BC Cancer Agency
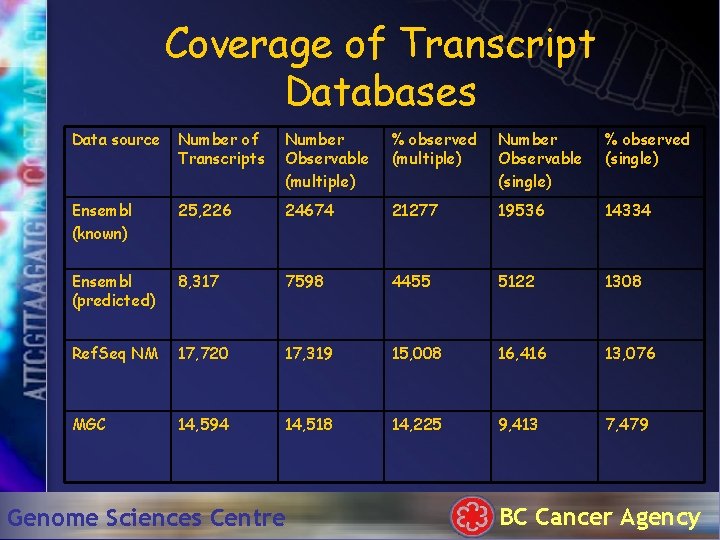
Coverage of Transcript Databases Data source Number of Transcripts Number Observable (multiple) % observed (multiple) Number Observable (single) % observed (single) Ensembl (known) 25, 226 24674 21277 19536 14334 Ensembl (predicted) 8, 317 7598 4455 5122 1308 Ref. Seq NM 17, 720 17, 319 15, 008 16, 416 13, 076 MGC 14, 594 14, 518 14, 225 9, 413 7, 479 Genome Sciences Centre BC Cancer Agency
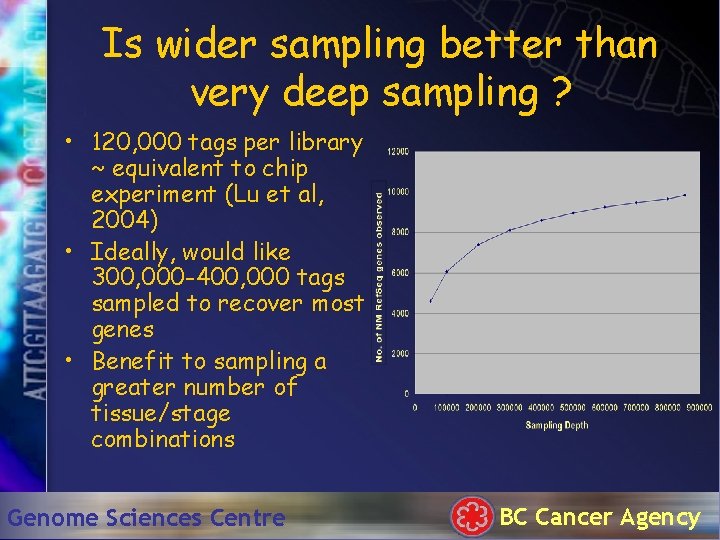
Is wider sampling better than very deep sampling ? • 120, 000 tags per library ~ equivalent to chip experiment (Lu et al, 2004) • Ideally, would like 300, 000 -400, 000 tags sampled to recover most genes • Benefit to sampling a greater number of tissue/stage combinations Genome Sciences Centre BC Cancer Agency
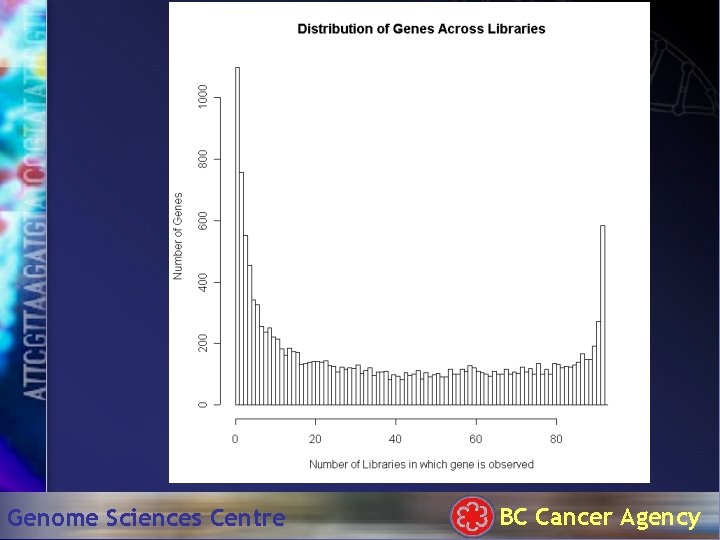
Genome Sciences Centre BC Cancer Agency
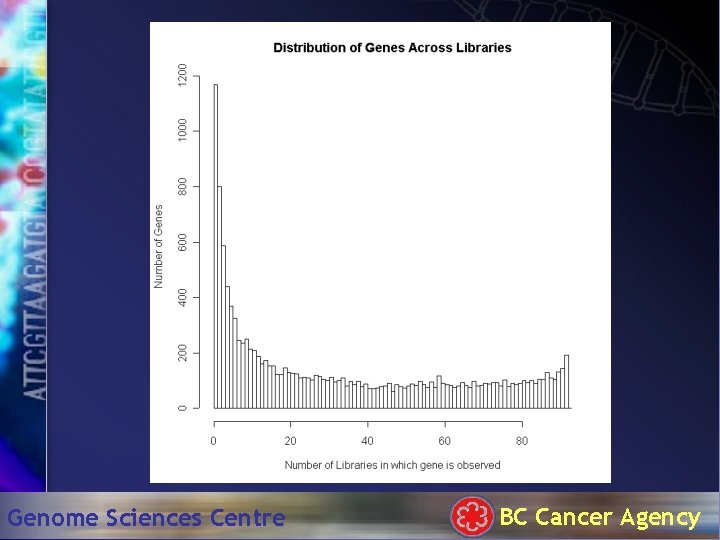
Genome Sciences Centre BC Cancer Agency

GO Analysis of 177 common genes • • 38% – metabolism 19% - cell growth and/or maintenance 13% – transport 6% - cell communication Genome Sciences Centre BC Cancer Agency
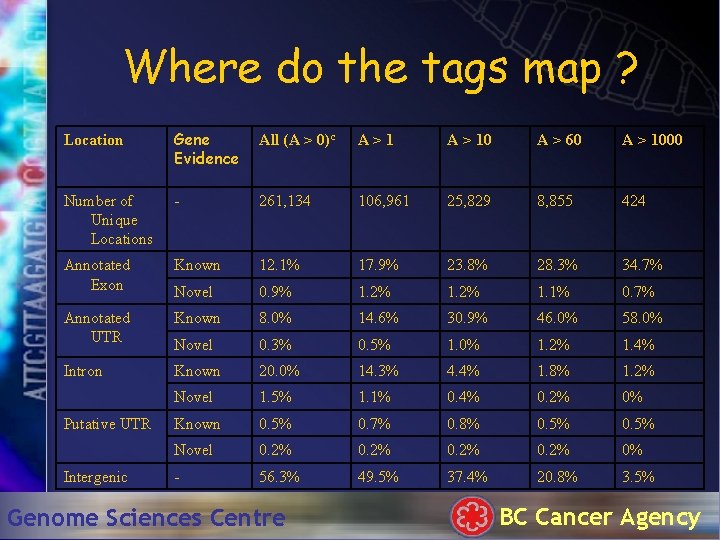
Where do the tags map ? Location Gene Evidence All (A > 0)c A>1 A > 10 A > 60 A > 1000 Number of Unique Locations - 261, 134 106, 961 25, 829 8, 855 424 Annotated Exon Known 12. 1% 17. 9% 23. 8% 28. 3% 34. 7% Novel 0. 9% 1. 2% 1. 1% 0. 7% Annotated UTR Known 8. 0% 14. 6% 30. 9% 46. 0% 58. 0% Novel 0. 3% 0. 5% 1. 0% 1. 2% 1. 4% Intron Known 20. 0% 14. 3% 4. 4% 1. 8% 1. 2% Novel 1. 5% 1. 1% 0. 4% 0. 2% 0% Known 0. 5% 0. 7% 0. 8% 0. 5% Novel 0. 2% 0% - 56. 3% 49. 5% 37. 4% 20. 8% 3. 5% Putative UTR Intergenic Genome Sciences Centre BC Cancer Agency

How many genes observed ? • 107 k transcripts covering 18. 6 k high quality annotated genes • 14 k transcripts covering 4 k predicted Ref. Seq and ENSEMBL genes • ~21 k genes observed Genome Sciences Centre BC Cancer Agency

What are the “intergenic tags” ? • 140 k tags unaccounted for… • Novel genes ? • 24 k transcripts covering 12 k UNIGENE and ENSEMBL EST genes • 36% map antisense to annotated genes • Many are singletons Genome Sciences Centre BC Cancer Agency

Singletons • Unannotated singletons – no genes, ESTs • 81% success rate for meta-singletons • 74% success rate for library singletons Genome Sciences Centre BC Cancer Agency

Summary • The majority of singletons represent bona fide transcriptional elements • We have identified novel transcripts • Evidence of differentially regulated variants resulting in different protein • Data providing functional annotation Genome Sciences Centre BC Cancer Agency

… and motifs… • The transcription of a gene is dependent on at least – 1) the DNA binding factors present in the nucleus at a given time and – 2) the DNA sequences, or cisregulatory motifs, present in the gene region to which these factors can bind Our goal is to attempt to identify the regulatory motifs Genome Sciences Centre BC Cancer Agency

Project Goals • High quality in-silico discovery of gene regulatory elements on a genome wide scale Approach based on: • Overrepresentation of similar DNA motifs in upstream sequences of genes with the same regulatory control Genome Sciences Centre BC Cancer Agency

Our Method • Use orthologous genes i. e. the equivalent genes different organisms. • Use regions from genes which display strong coexpression (infer coregulation). Genome Sciences Centre BC Cancer Agency
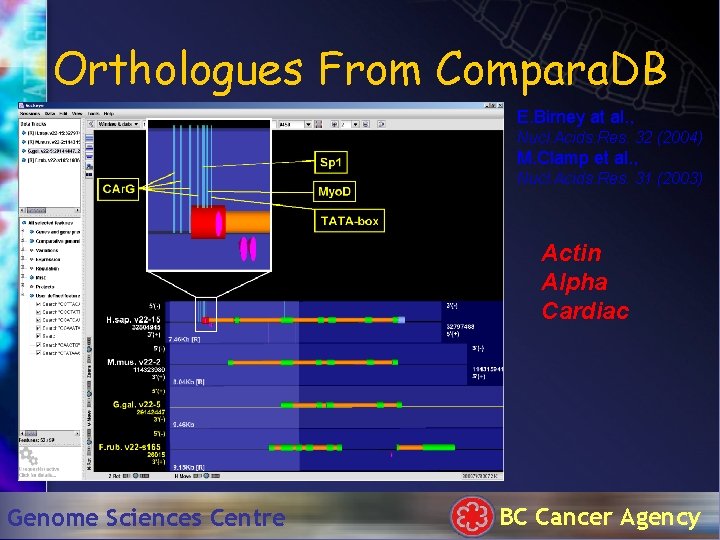
Orthologues From Compara. DB E. Birney at al. , Nucl. Acids. Res. 32 (2004) M. Clamp et al. , Nucl. Acids. Res. 31 (2003) Actin Alpha Cardiac Genome Sciences Centre BC Cancer Agency
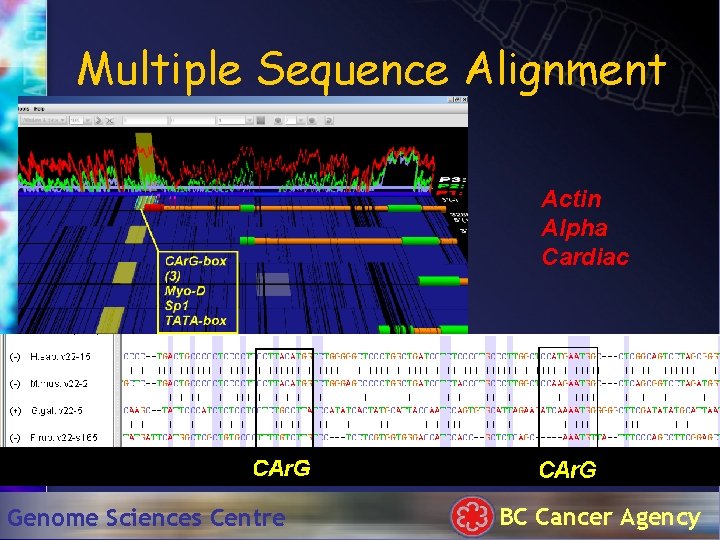
Multiple Sequence Alignment Actin Alpha Cardiac Genome Sciences Centre BC Cancer Agency
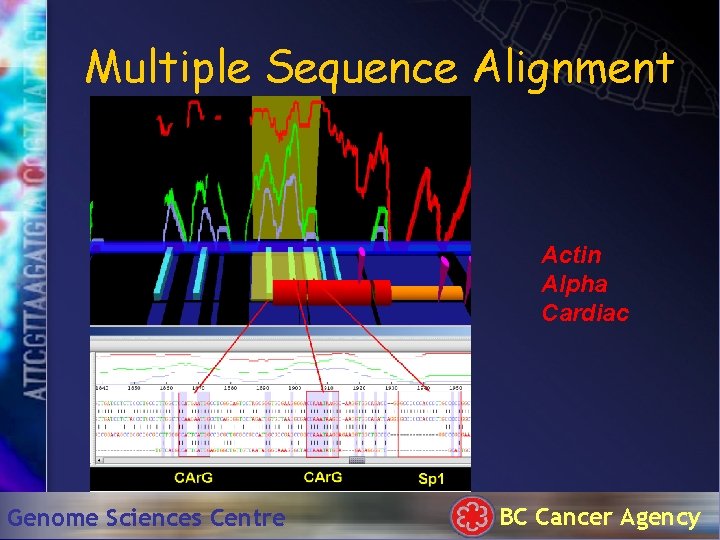
Multiple Sequence Alignment Actin Alpha Cardiac Genome Sciences Centre BC Cancer Agency
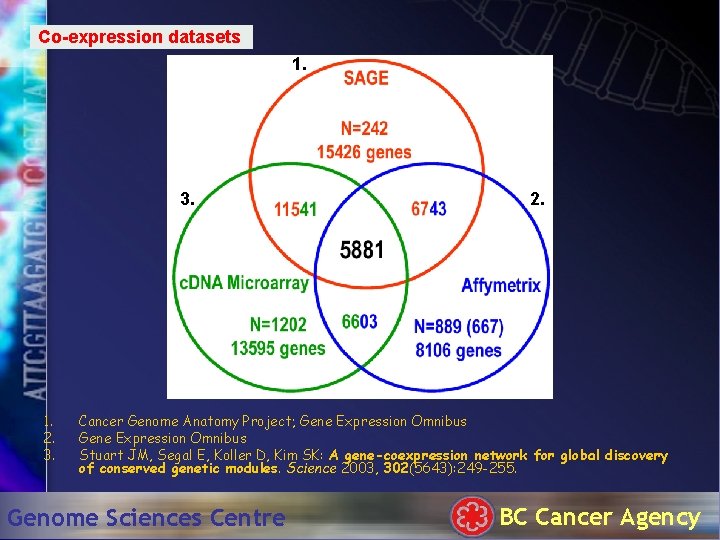
Co-expression datasets 1. 3. 1. 2. 3. 2. Cancer Genome Anatomy Project; Gene Expression Omnibus Stuart JM, Segal E, Koller D, Kim SK: A gene-coexpression network for global discovery of conserved genetic modules. Science 2003, 302(5643): 249 -255. Genome Sciences Centre BC Cancer Agency
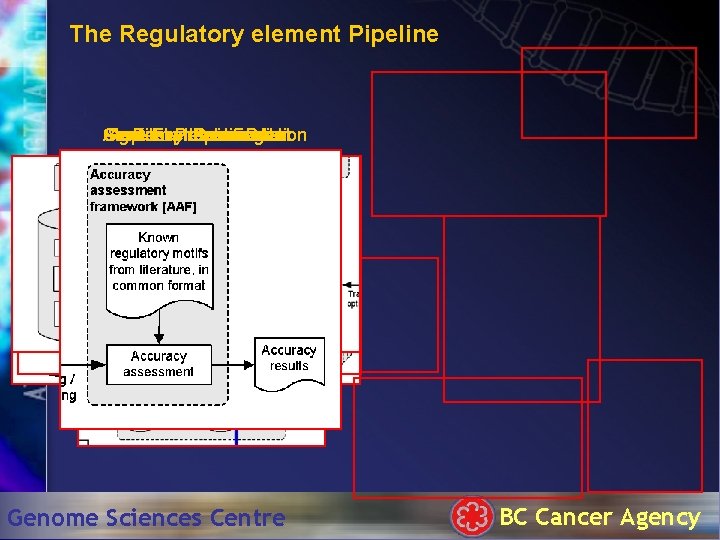
The Regulatory element Pipeline Gene Sequence Algorithm Accuracy Known Post-Processing Expression Implementation Identification Assessment Resources Data Genome Sciences Centre BC Cancer Agency
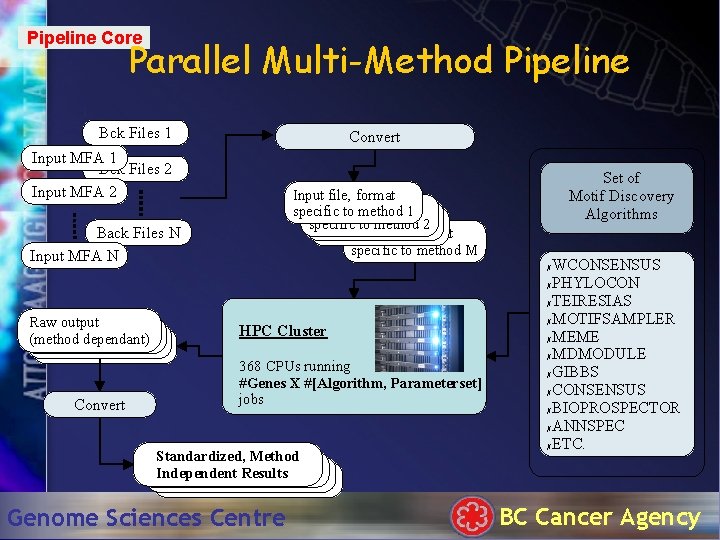
Pipeline Core Parallel Multi-Method Pipeline Bck Files 1 Convert Input MFA 1 Bck Files 2 Input MFA 2 Input file, format Input to file, format 1 specific method specific to method 2 Input file, format specific to method M Back Files N Input MFA N Raw output (method dependant) Convert Set of Motif Discovery Algorithms WCONSENSUS ✗PHYLOCON ✗TEIRESIAS ✗MOTIFSAMPLER ✗MEME ✗MDMODULE ✗GIBBS ✗CONSENSUS ✗BIOPROSPECTOR ✗ANNSPEC ✗ETC. ✗ HPC Cluster 368 CPUs running #Genes X #[Algorithm, Parameterset] jobs Standardized, Method Independent Results Genome Sciences Centre BC Cancer Agency
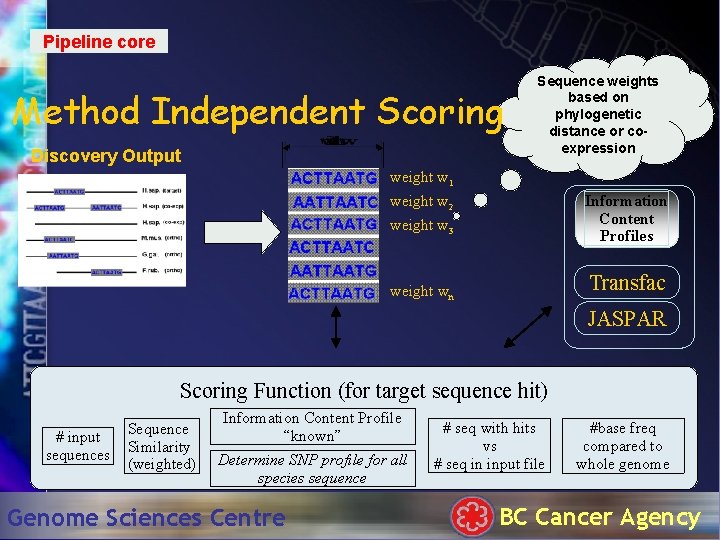
Pipeline core Method Independent Scoring Discovery Output Sequence weights based on phylogenetic distance or coexpression weight w 1 Information Content Profiles weight w 2 weight w 3 Transfac weight wn JASPAR Scoring Function (for target sequence hit) # input sequences Sequence Similarity (weighted) Information Content Profile “known” Determine SNP profile for all species sequence Genome Sciences Centre # seq with hits vs # seq in input file #base freq compared to whole genome BC Cancer Agency
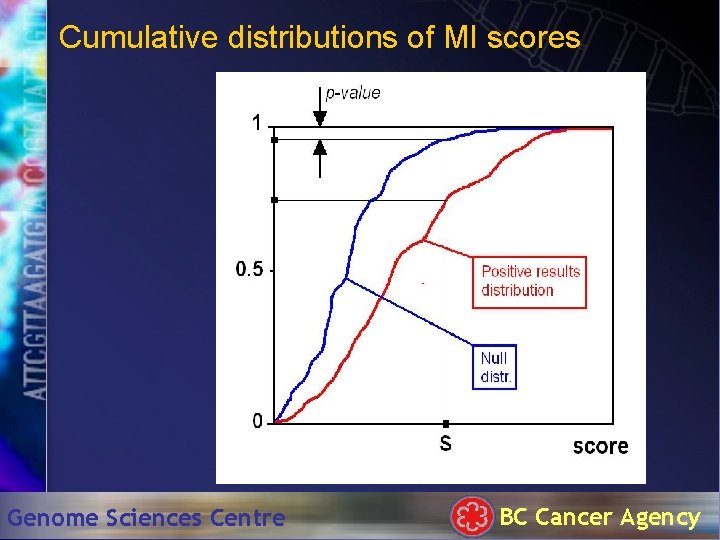
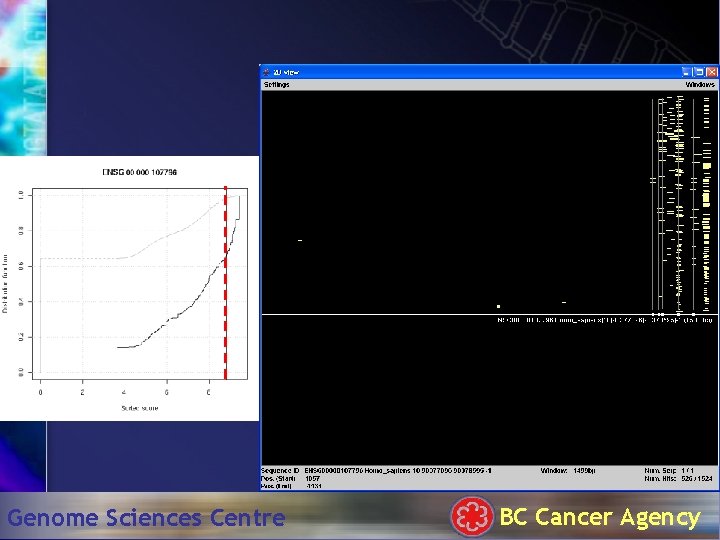
Cumulative distributions of MI scores Genome Sciences Centre BC Cancer Agency
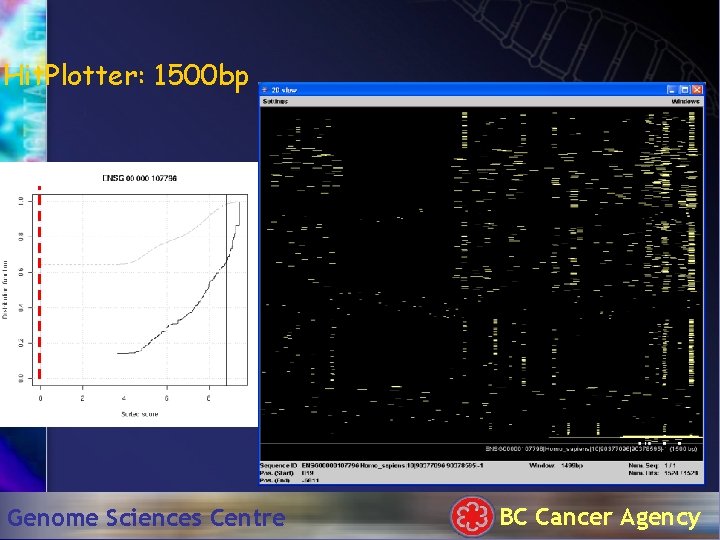
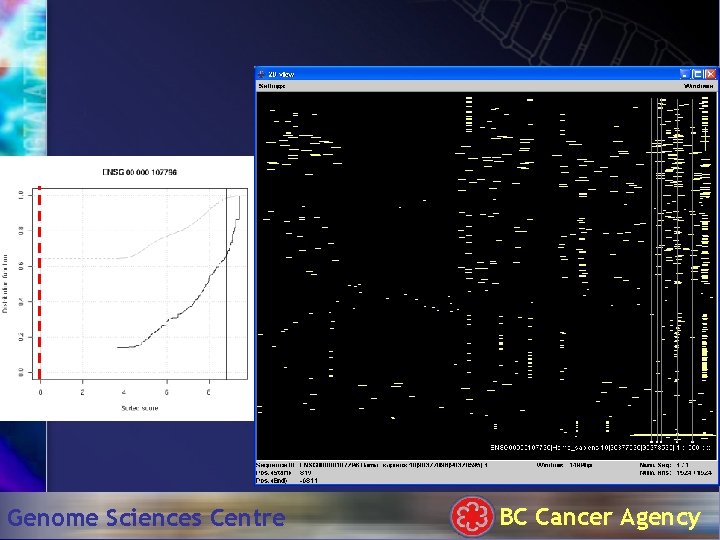
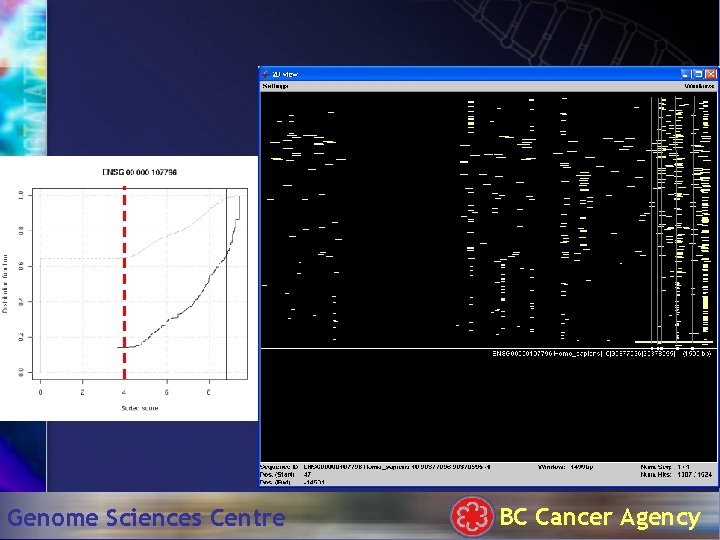
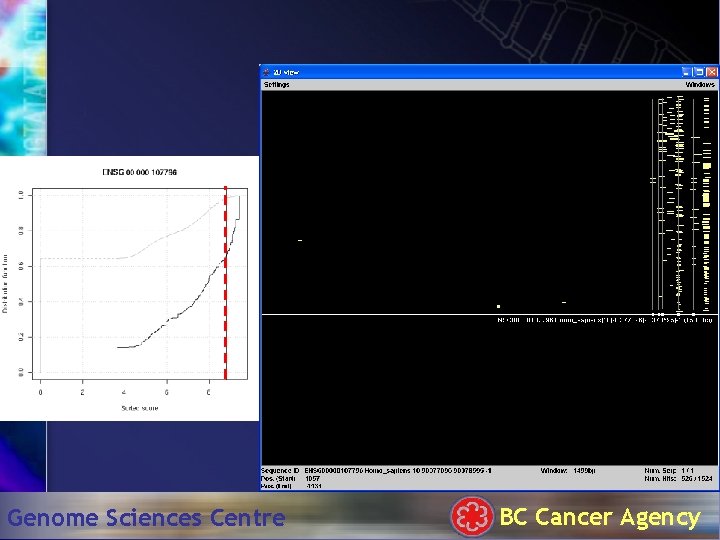
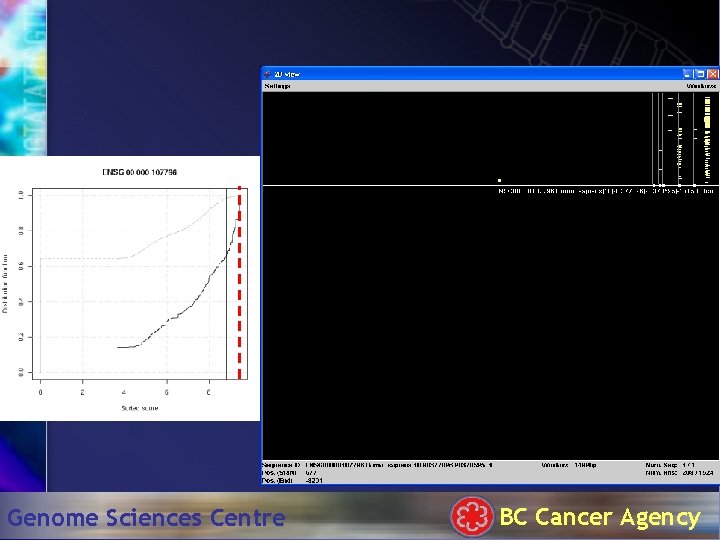
Hit. Plotter: 1500 bp Genome Sciences Centre BC Cancer Agency
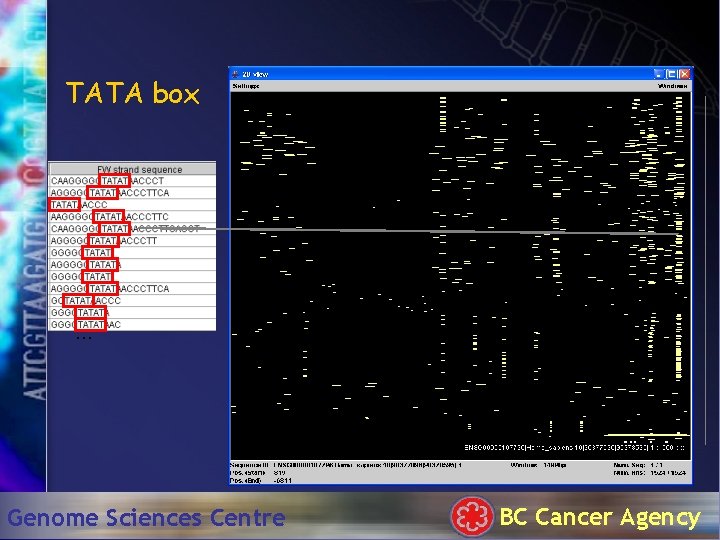
TATA box . . . Genome Sciences Centre BC Cancer Agency
Genome Sciences Centre BC Cancer Agency
Genome Sciences Centre BC Cancer Agency
Genome Sciences Centre BC Cancer Agency
Genome Sciences Centre BC Cancer Agency
Genome Sciences Centre BC Cancer Agency

Co-occurring Motifs • Red and Blue motifs co-occur in the promoter regions of these two genes • The separation of the two motifs may be constrained • Use co-occurrence motifs to define regulatory modules Genome Sciences Centre BC Cancer Agency
Putting it all together… Tissue Specific Gene Expression Patterns Tissue Specific Motifs and Modules Genome Sciences Centre BC Cancer Agency

…And best laid plans • But, Mousie, thou art no thy lane, In proving foresight may be vain; The best-laid schemes o' mice an' men Gang aft agley, An'lea'e us nought but grief an' pain, For promis'd joy! – Robert Burns Genome Sciences Centre BC Cancer Agency

The Moral of the Story • If you’re a mouse, don’t make your home in a farmer’s field – build it next to the field! • Risk Management! • What are the issues associated with running a large bioinformatics activity ? Genome Sciences Centre BC Cancer Agency

Running a bioinformatics group • • What does everyone do ? How are they doing it ? Are they talking to the right people ? Have they got the right requirements ? Is anyone waiting for information ? Are they running on schedule ? Is there an issue that needs escalating? Are there HR, training, management, coaching issues that need to be addressed ? Genome Sciences Centre BC Cancer Agency

Organizational Complexity • The organizational complexity of bioinformatics projects has increased: – Made up of larger teams – Have multiple stakeholders – Contain many organizational layers Genome Sciences Centre BC Cancer Agency

Technical Complexity • • • Number of databases increasing Number of methods increasing Body of knowledge is developing rapidly Requirements change rapidly Must be well-read in a large number of fields Genome Sciences Centre BC Cancer Agency

Common Statements – “Things change all the time - it’s impossible to plan” – “I’d like you to do some analysis” – “I don’t have time to plan” – “We’ll figure it out as we go along” – Not so common – “An ounce of prevention is worth a pound of cure” Genome Sciences Centre BC Cancer Agency

Software Engineering Management • • • Large body of knowledge Requirements engineering Architecture and Design Validation Change management Risk management Genome Sciences Centre BC Cancer Agency

Solutions at the GSC • • • CM controls Bug tracking controls Some validation controls Various levels of design and architecture Implementation of structured engineering process under way to define, track and manage work – Requirements control – Risk/Change management Genome Sciences Centre BC Cancer Agency

• ~90% of work performed by the group can be planned or have a LOE assigned • Some areas harder – finishing a genome, algorithm development, exploratory analysis • There is always a schedule and a budget Genome Sciences Centre BC Cancer Agency
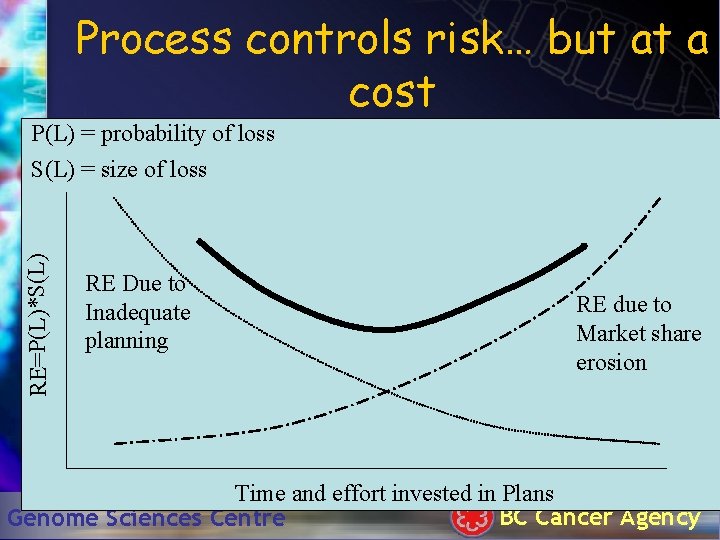
Process controls risk… but at a cost P(L) = probability of loss RE=P(L)*S(L) = size of loss RE Due to Inadequate planning RE due to Market share erosion Time and effort invested in Plans BC Cancer Agency Genome Sciences Centre

Hacky Scripts/Code have their place • Ideal for prototyping – Only prototype when you are trying to get a handle on things • Throw away the prototype, when you’re done experimenting! • …but stop and think! Genome Sciences Centre BC Cancer Agency

And standards… Genome Sciences Centre BC Cancer Agency

Mouse Atlas Pamela Hoodless Marco Marra Terry Fox Laboratory Genome Sciences Centre Jim Rupert Mona Wu Rebecca Cullum Jaswinder Khattra Allen Delaney Jennifer Asano Susanna Chan Cheryl Helgason Steven Jones Cancer Endocrinology Genome Sciences Centre Brad Hoffman Teresa Ruiz de Alagara Ida Zhang Asim Siddiqui Scott Zuyderduyn Richard Varhol Derek Leung Kevin Teague Lisa Lee Anita Landry Caroline Astell Project Manager Genome Sciences Centre Elizabeth M. Simpson CMMT Robert Xie Slavita Bohacec Byron Kuo Adrian Burke Genome. BC Gregory Riggins John Hopkins BC Cancer Agency

Cis. Red GSC Marco Marra Gordon Robertson Richard Varhol Kevin Teague Obi Griffith Erin Pleasance Debra Fulton Keven Lin Mikhail Bilenky Neil Roberston Monica Sluemer Stephen Montgomery Asim Siddiqui Genome Sciences Centre Ian Holmes, UC Berkeley Stanford University Rick Myers Nathan Trinklein Shelley Force Alldred Sarah Hartman Ewan Birney, EBI BC Cancer Agency

www. mouse. Atlas. org www. cis. Red. org Genome Sciences Centre BC Cancer Agency
- Slides: 63