Object Oried Data Analysis Last Time HDLSS Asymptotics

Object Orie’d Data Analysis, Last Time HDLSS Asymptotics • Studied from Dual Viewpoint NCI 60 Data • Visualization – found (DWD) directions that showed clusters of cancer types • Investigated with Di. Pro. Perm test HDLSS hypothesis testing

HDLSS Asymptotics Interesting Idea from Travis Gaydos: Interpret from viewpoint of dual space Recall from Aug. 25: • Distance to origin: • Pairwise Distance: • Angle from origin: for

HDLSS Asymptotics – Dual View Would be interesting to try: • Study (i. e. explore conditions for): – Consistency – Strong Inconsistency for PCA direction vectors, from this viewpoint Perhaps other things as well…

NCI 60 Data Recall from: • Aug. 28 • Aug. 30 NCI 60 Cancer Cell Lines Microarray Data • Explored Data Combination • c. DNA & Affymetrix Measurements • Right answer is known

Real Clusters in NCI 60 Data Simple Visual Approach: • Randomly relabel data (Cancer Types) • Recompute DWD dir’ns & visualization • Get heuristic impression from this Deeper Approach • Formal Hypothesis Testing (Done later)

Real Clusters in NCI 60 Data? From Aug. 30: Simple Visual Approach: n Randomly relabel data (Cancer Types) n Recompute DWD dir’ns & visualization n Get heuristic impression from this n Some types appeared signif’ly different n Others did not Deeper Approach: Formal Hypothesis Testing

HDLSS Hypothesis Testing Approach: Di. Pro. Perm Test DIrection – PROjection – PERMutation Ideas: n Find an appropriate Direction vector n Project data into that 1 -d subspace n Construct a 1 -d test statistic n Analyze significance by Permutation

Di. Pro. Perm Simple Example 1, Totally Separate Results: n Random relabelling gives much smaller Ts n Quantiles (over 1000 sim’s) give p-val of 0 n I. e. Strongly conclusive n Conclude sub-populations are different

Needed final verification of Cross-platform Normal’n • Is statistical power actually improved? • Is there benefit to data combo by DWD? • More data more power? • Will study later now

Needed final verification of Cross-platform Normal’n Summary of Results: • P-values – Combined Results Better for 7 out of 8 cases – Combined signficant, Affy not, in 3 cases (Lukemia, NSCLC, Renal) – Shows combining platforms often worthwhile (because more data gives more power) Comparison with previous heuristics…
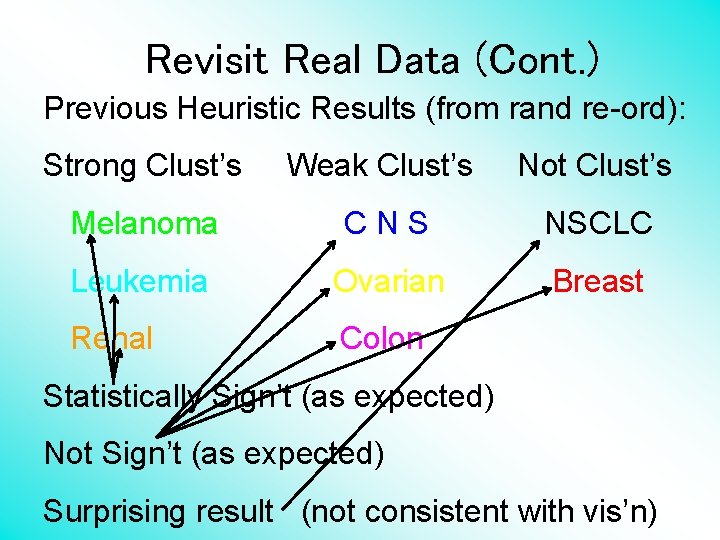
Revisit Real Data (Cont. ) Previous Heuristic Results (from rand re-ord): Strong Clust’s Weak Clust’s Not Clust’s Melanoma CNS NSCLC Leukemia Ovarian Breast Renal Colon Statistically Sign’t (as expected) Not Sign’t (as expected) Surprising result (not consistent with vis’n)

Revisit Real Data (Cont. ) Sungkyu Jung Question: How are those results driven by sample size? Add sample size to above table….

Revisit Real Data (Cont. ) Previous Heuristic Results (from rand re-ord): Strong Clust’s Weak Clust’s Not Clust’s Melanoma - 18 C N S – 12 NSCLC - 18 Leukemia - 12 Ovarian - 8 Renal - 14 Breast - 12 Colon - 12 Statistically Sign’t (as expected) Not Sign’t (as expected) Surprising result (not consistent with vis’n)

Revisit Real Data (Cont. ) Sungkyu Jung Question: How are those results driven by sample size? Add sample size to above table…. Good idea: Surprising result perhaps indeed due to larger sample size

Di. Pro. Perm Test Particulate Matter Data Consulting Class Project, for: Lindsay Whicher, Penn Watkinson, EPA Analysis by: Chihoon Lee

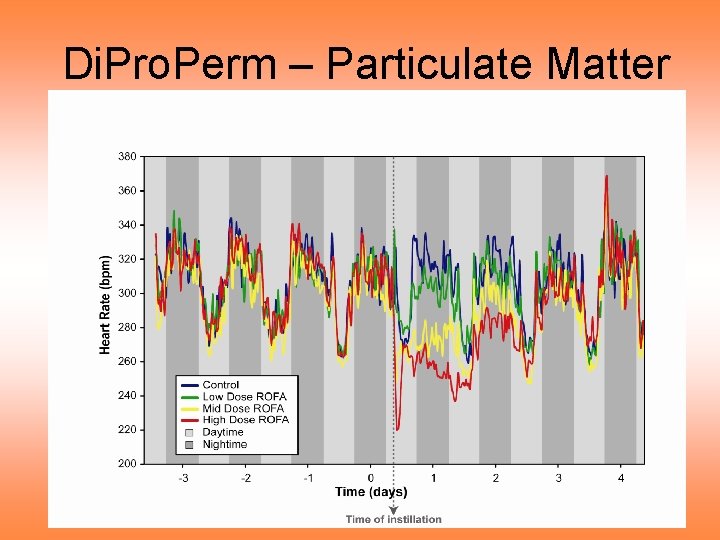
Di. Pro. Perm – Particulate Matter Data: • Measure Heart Rate of Rats • Over time (several days) • Treat with Particulate Matter • Study effect • See differences between treatments? • Statistically significant?
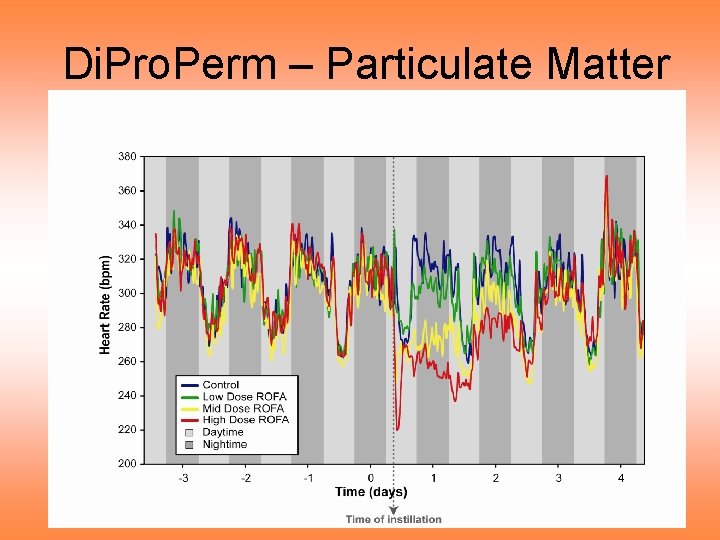
Di. Pro. Perm – Particulate Matter

Di. Pro. Perm – Particulate Matter Notes on curve view of data: • Clear day – night effect • Apparent changes after treatment • Stronger effect for higher dose • Effect diminishes over time • Statistically significant differences? • How does “signal” compare to “noise”?

Di. Pro. Perm – Particulate Matter Alternate view of data: • Each curve is a “data point” • Study distribution of these “points” • Show replicates as points • To indicate “signal” vs. “noise” issues
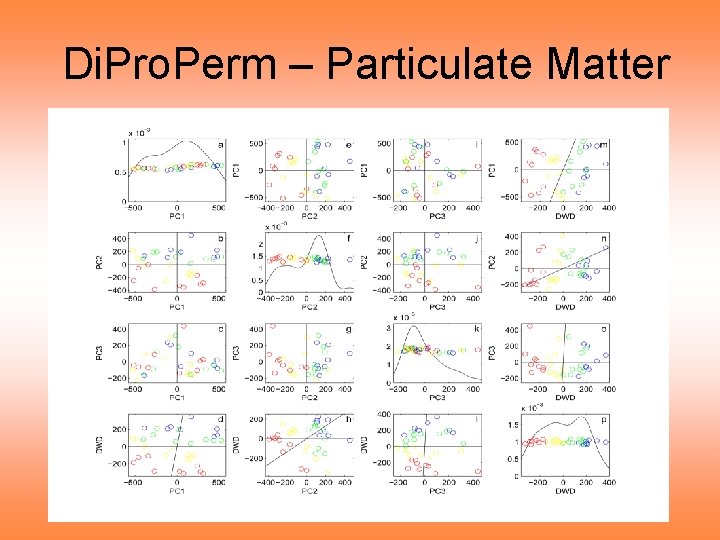
Di. Pro. Perm – Particulate Matter

Di. Pro. Perm – Particulate Matter Notes on PCA & DWD dir’n scatterplots: • Dose effect looks strong (PC 2 direction) • Systematic Pattern of colors • Ordered by doses • Suggests important differences • Statistically significant differences? • How does “signal” compare to “noise”? Address by Di. Pro. Perm tests

Di. Pro. Perm – Particulate Matter Look for differences over 48 hours: • Run Di. Pro. Perm • Test Control vs. High Dose • Study difference over long time scale
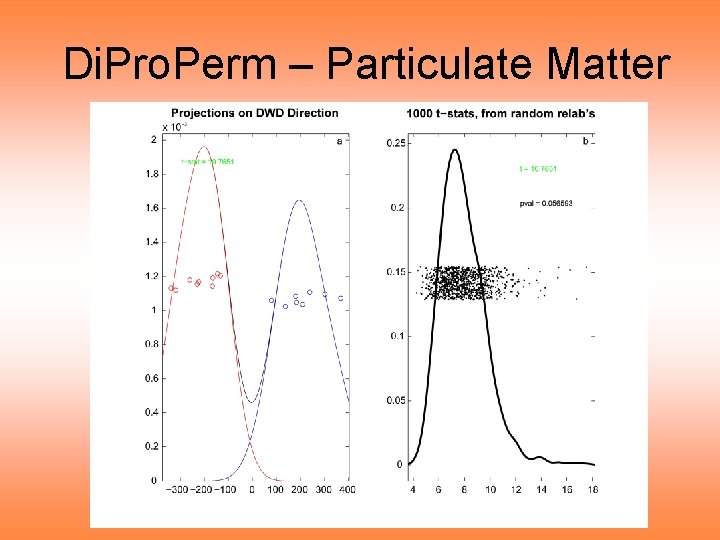
Di. Pro. Perm – Particulate Matter

Di. Pro. Perm – Particulate Matter Di. Pro. Perm Results: • P-value = 0. 056 • Not quite significant • “Noise” just overtakes “signal” • Perhaps Interval of 48 hours is too long • So try smaller interval Day 0, 9 AM – 3 PM
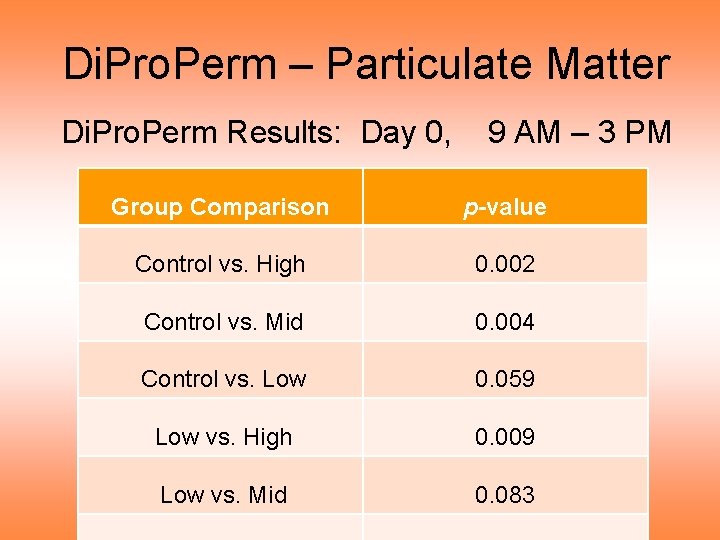
Di. Pro. Perm – Particulate Matter Di. Pro. Perm Results: Day 0, 9 AM – 3 PM Group Comparison p-value Control vs. High 0. 002 Control vs. Mid 0. 004 Control vs. Low 0. 059 Low vs. High 0. 009 Low vs. Mid 0. 083

Di. Pro. Perm – Particulate Matter Di. Pro. Perm Results: Day 0, 9 AM – 3 PM • Results consistent with data curves: – C vs. H strongly different – C vs. M & L vs. H significantly different – Others not quite significant • For more, related, results, see Wichers, Lee, et al (2007)
Di. Pro. Perm – Particulate Matter

HDLSS Hypothesis Testing – Di. Pro. Perm test Chuck Perou’s 500 Breast Cancer data Based on data Merging (using DWD): • UNC – Geo, # = 102 – Unp, # = 93 • NKI – Pub, # = 220 – 97, # = 97 # = 512
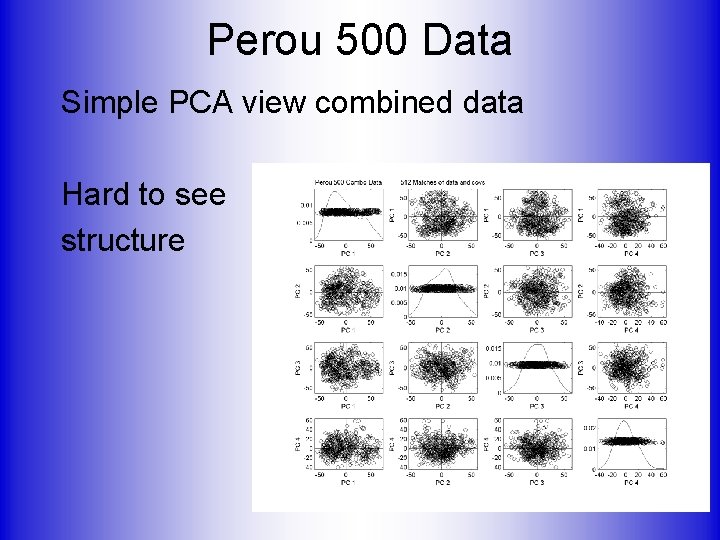
Perou 500 Data Simple PCA view combined data Hard to see structure
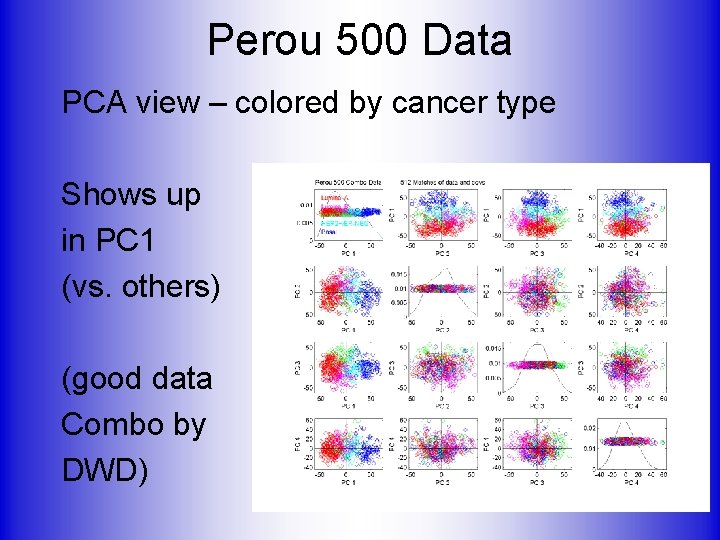
Perou 500 Data PCA view – colored by cancer type Shows up in PC 1 (vs. others) (good data Combo by DWD)
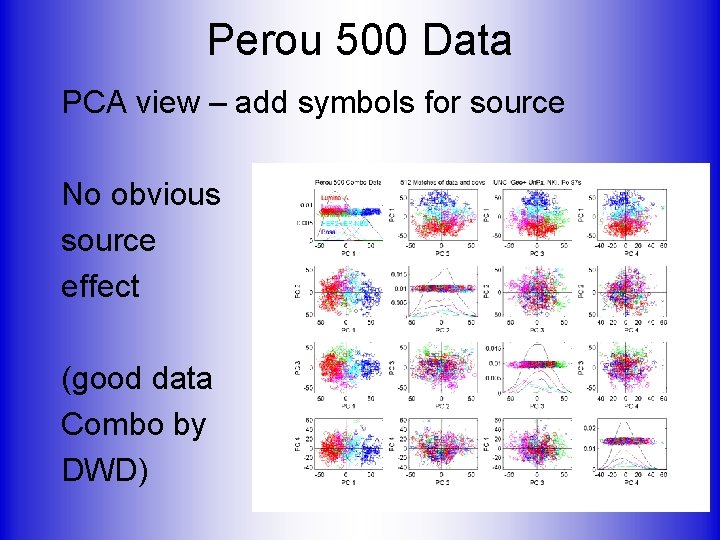
Perou 500 Data PCA view – add symbols for source No obvious source effect (good data Combo by DWD)
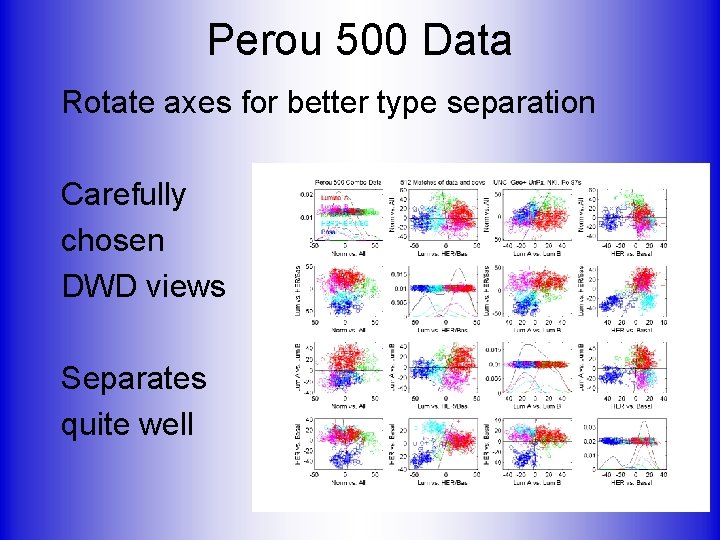
Perou 500 Data Rotate axes for better type separation Carefully chosen DWD views Separates quite well

Perou 500 Data How distinct are classes? Compare “signal” vs. “noise” Measure statistical significance Using Di. Pro. Perm test
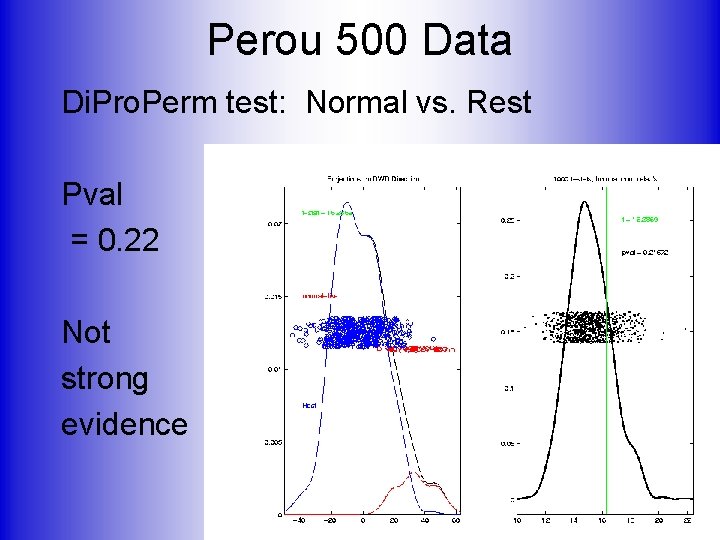
Perou 500 Data Di. Pro. Perm test: Normal vs. Rest Pval = 0. 22 Not strong evidence

Perou 500 Data Di. Pro. Perm test: Normal vs. Rest Pval = 0. 22, Not strong evidence OK, since “normal” means: biopsy missed tumor But mostly from cancer patients Instead compare with “true normals”

Perou 500 Data Di. Pro. Perm test: True Normal vs. Rest Pval = 2. 30 E-06 , Makes sense. Very strong evidence

Perou 500 Data Di. Pro. Perm test: Cancer classes • Luminals vs {Her 2 & Basals}, pval = 0 • Her 2 vs Basals, pval = 0 • Lum A vs Lum b, pval = 0. 0068 All strongly conclusive Adds statistical significance to early results

Perou 500 Data Interesting questions: • Was the DWD combination essential? • Were individual groups sign’t anyway? • What was value of DWD combo?

Perou 500 Data Di. Pro. Perm Luminals vs {Her 2 & Basals}: • All Combined, p-val = 0 • UNC Combo, p-val = 0 • NKI Combo, p-val = 0 • UNC GEO, p-val = 0 • UNC Un. P, p-val = 3 e-14 • NKI Pub, p-val = 1 e-11 • NKI 97, p-val = 0. 00078

Perou 500 Data Di. Pro. Perm Luminal A vs Luminal B: • All Combined, p-val = 0. 0068 • UNC Combo, p-val = 0. 214 • NKI Combo, p-val = 0. 014 • UNC GEO, p-val = 0. 298 • UNC Un. P, p-val = 0. 396 • NKI Pub, p-val = 0. 052 • NKI 97, p-val = 0. 246 (shows clear value to combining)

Perou 500 Data Di. Pro. Perm Her 2 vs Basals: • All Combined, p-val = 0 • UNC Combo, p-val = 0 • NKI Combo, p-val = 0 • UNC GEO, p-val = 0 • UNC Un. P, p-val = 0. 02 • NKI Pub, p-val = 0. 00008 • NKI 97, p-val = 0. 246

Perou 500 Data Draw back to Di. Pro. Perm here: • Classes found by clustering • Different from e. g. NCI 60 classes • So maybe not surprising they are Different from random • I. e. find significant differences • Does this really mean: Cluster is really there? ? ? Needs deeper thought…

HDLSS Hypothesis Testing – Di. Pro. Perm test Many Open Questions on Di. Pro. Perm Test: n Which Direction is “Best”? n Which 1 -d Projected test statistic? n Permutation vs. alternatives (bootstrap? )? ? ? n How do these interact? n What are asymptotic properties?

Clustering Idea: Given data • Assign each object to a class • Of similar objects • Completely data driven • I. e. assign labels to data • “Unsupervised Learning” Contrast to Classification (Discrimination) • With predetermined classes • “Supervised Learning”

Clustering Important References: • Mc. Queen (1967) • Hartigan (1975) • Kaufman and Rousseeuw (2005),

K-means Clustering Main Idea: for data Partition indices among classes Given index sets • that partition • represent clusters by “class mean”

K-means Clustering Given index sets Measure how well clustered, using Within Class Sum of Squares Weak point: not very interpretable
![K-means Clustering Common Variation: Put on scale of proportions (i. e. in [0, 1]) K-means Clustering Common Variation: Put on scale of proportions (i. e. in [0, 1])](http://slidetodoc.com/presentation_image_h2/36340e4a709f455df40f446952cc8d05/image-48.jpg)
K-means Clustering Common Variation: Put on scale of proportions (i. e. in [0, 1]) By dividing “within class SS” by “overall SS” Gives Cluster Index:

K-means Clustering Notes on Cluster Index: • CI = 0 when all data at cluster means • CI small when gives tight clustering (within SS contains little variation) • CI big when gives poor clustering (within SS contains most of variation) • CI = 1 when all cluster means are same

K-means Clustering Goal: • Given data • Choose classes • To miminize

2 -means Clustering Study CI, using simple 1 -d examples • Varying Standard Deviation
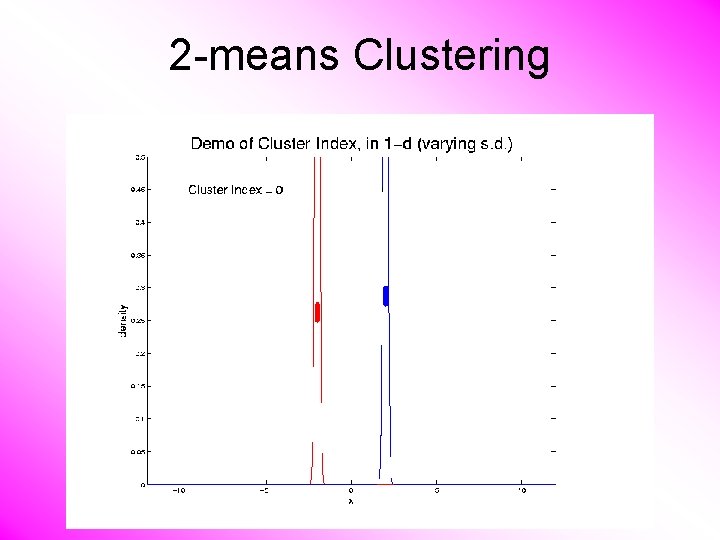
2 -means Clustering
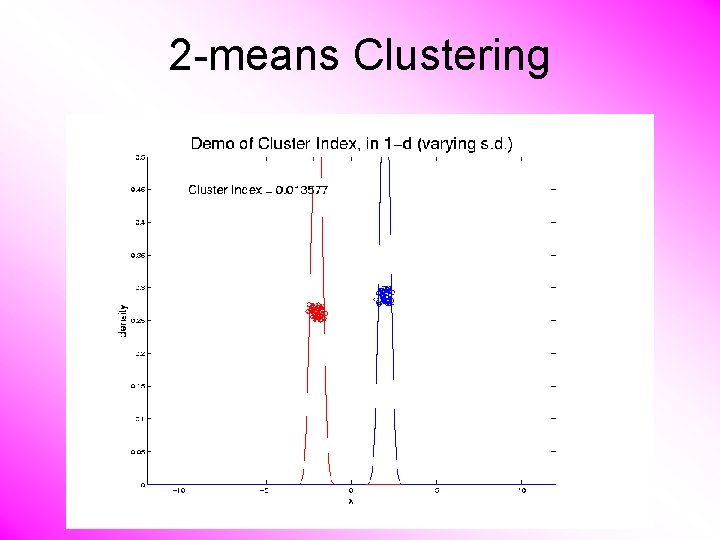
2 -means Clustering
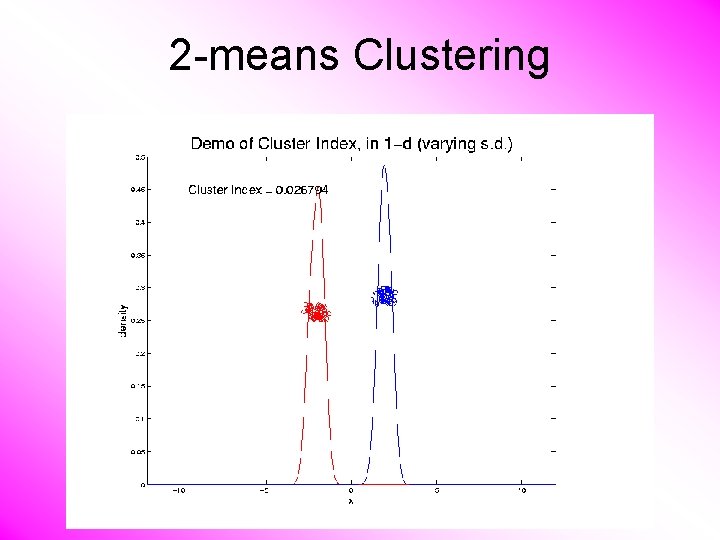
2 -means Clustering
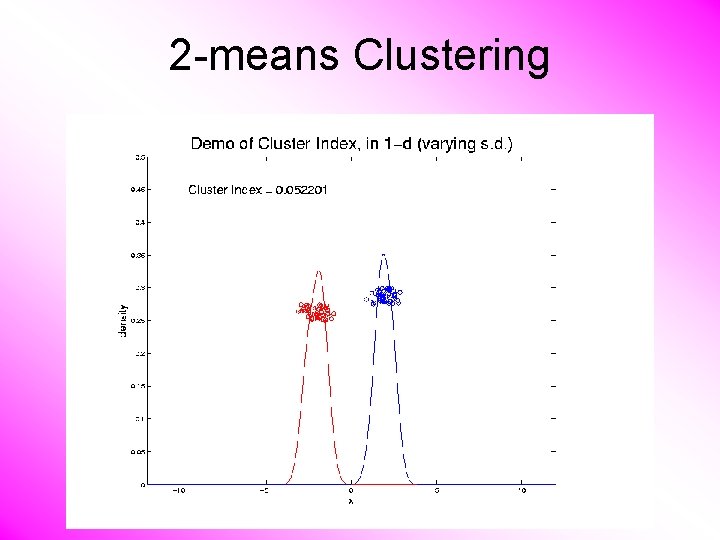
2 -means Clustering
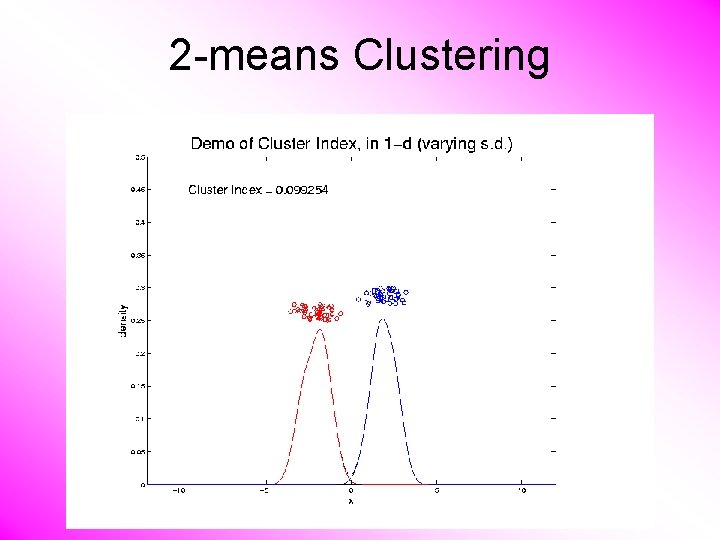
2 -means Clustering
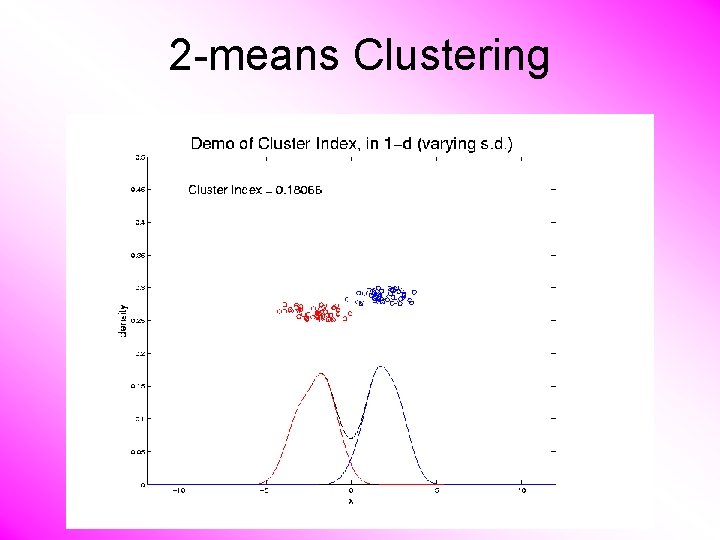
2 -means Clustering
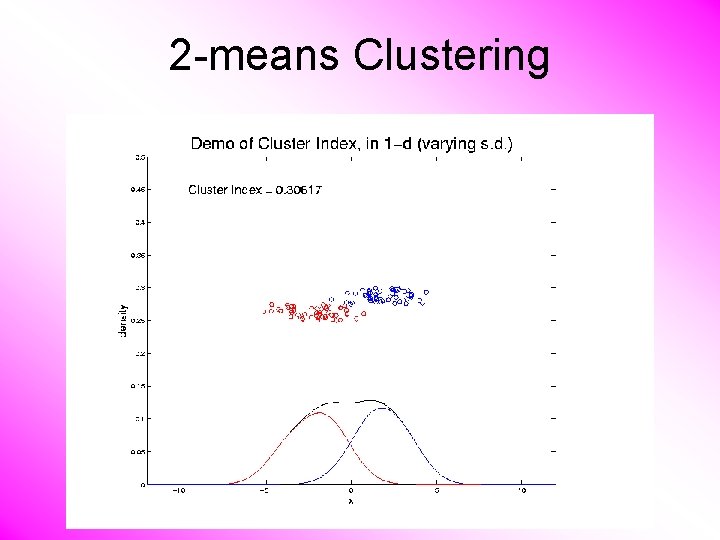
2 -means Clustering
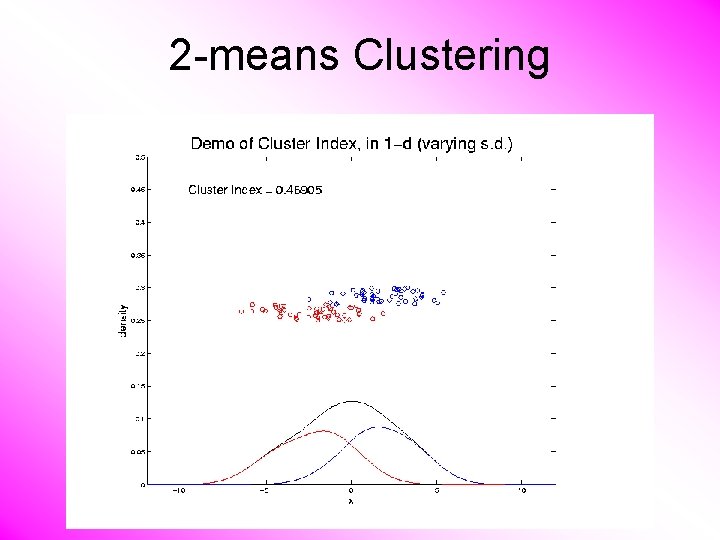
2 -means Clustering
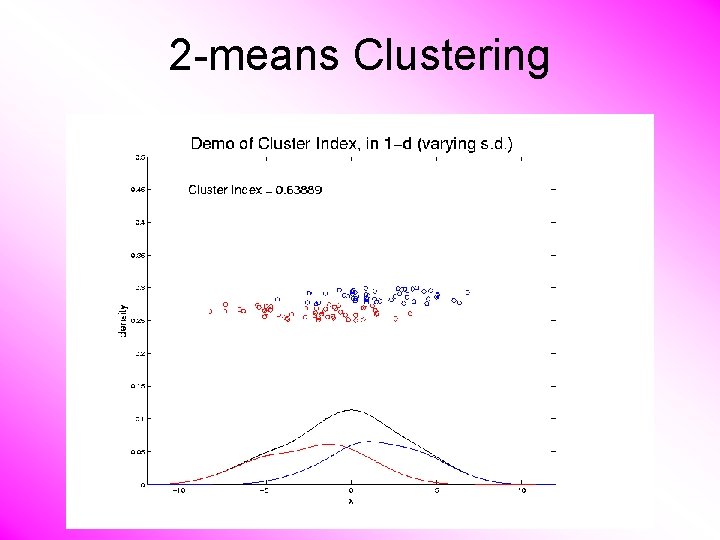
2 -means Clustering
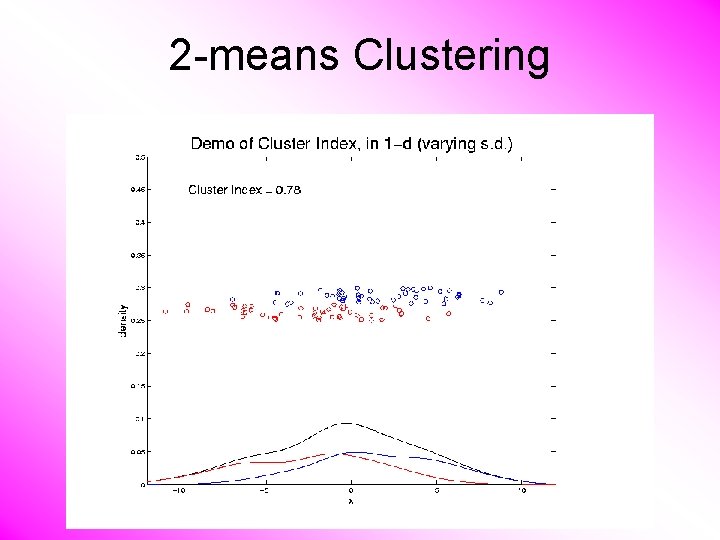
2 -means Clustering
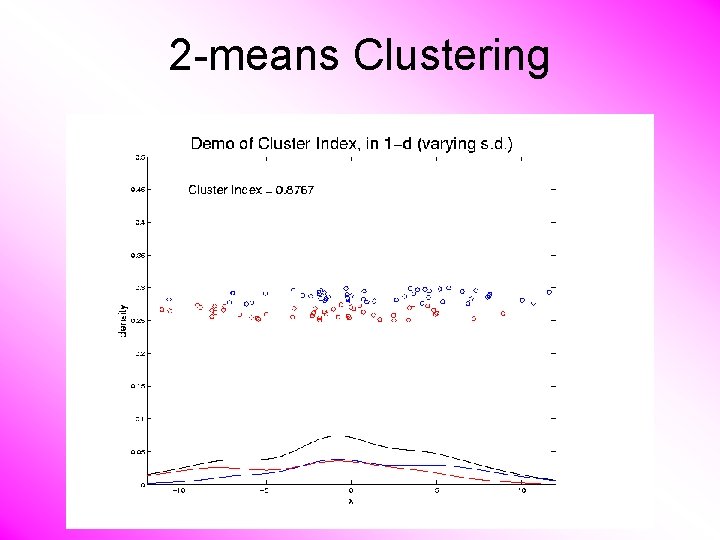
2 -means Clustering

2 -means Clustering Study CI, using simple 1 -d examples • Varying Standard Deviation • Varying Mean
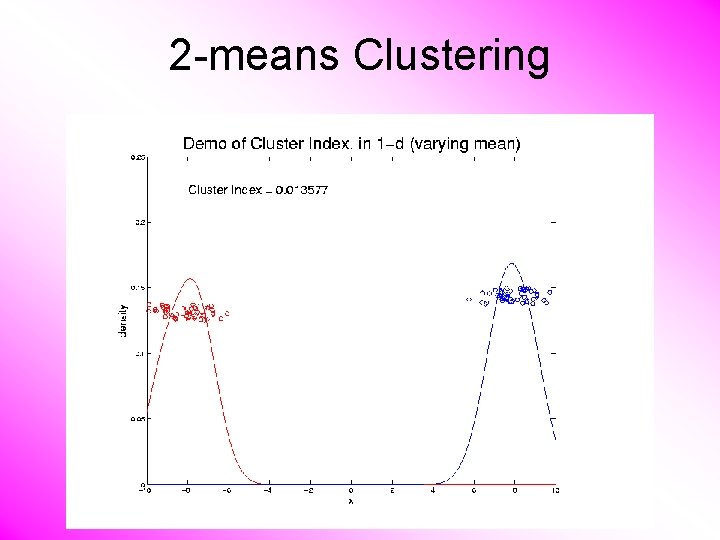
2 -means Clustering
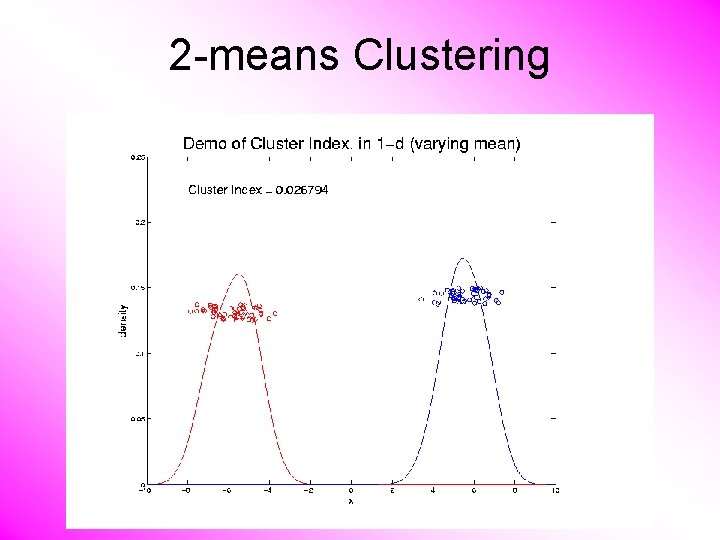
2 -means Clustering
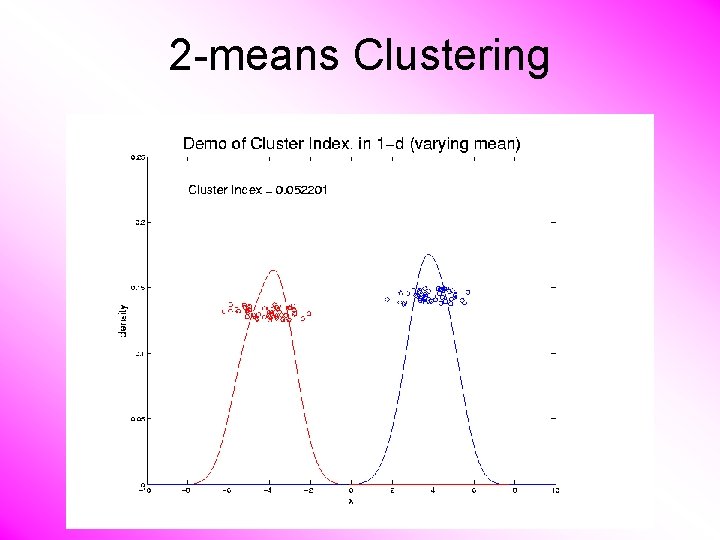
2 -means Clustering
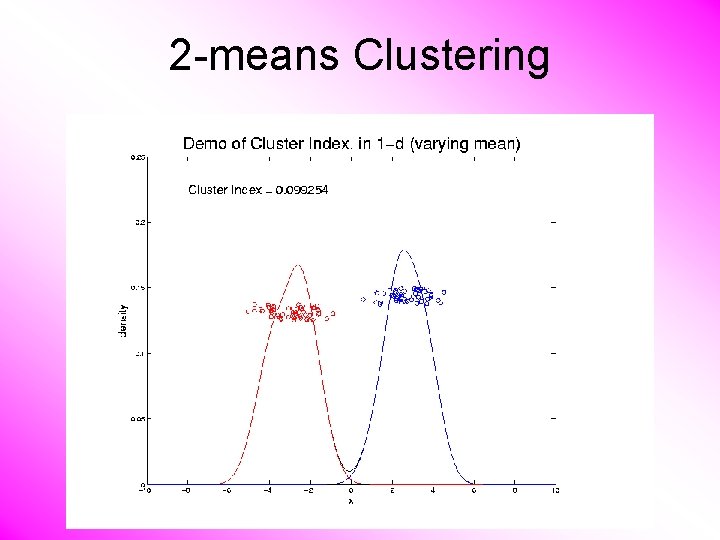
2 -means Clustering
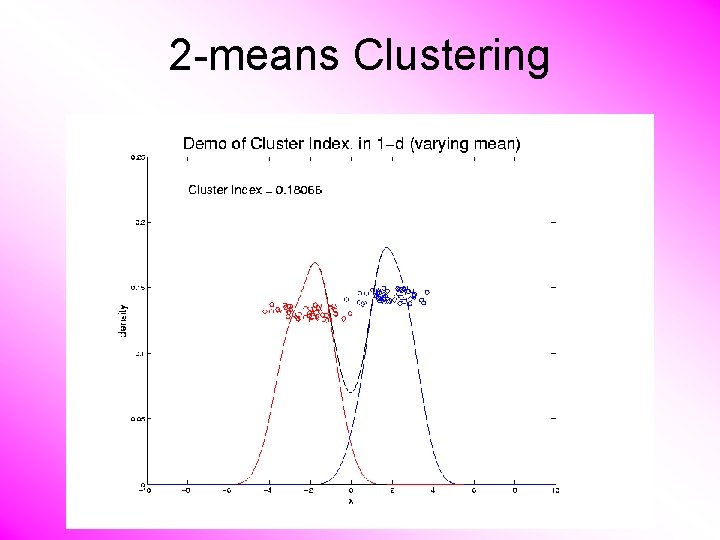
2 -means Clustering
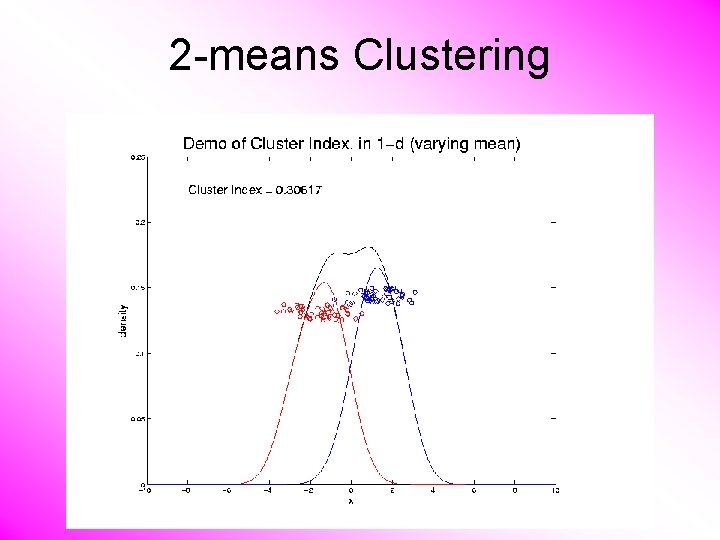
2 -means Clustering
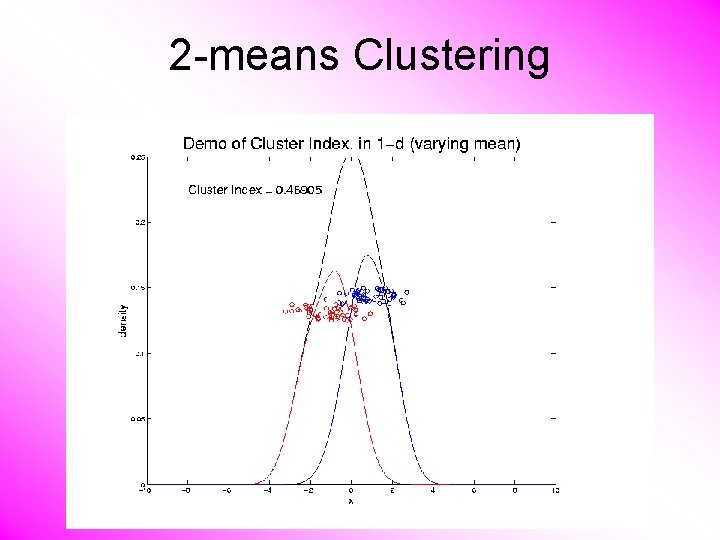
2 -means Clustering
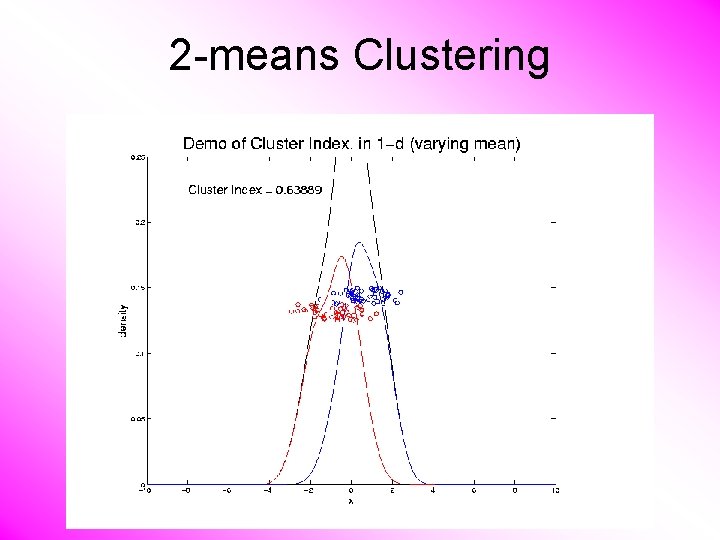
2 -means Clustering
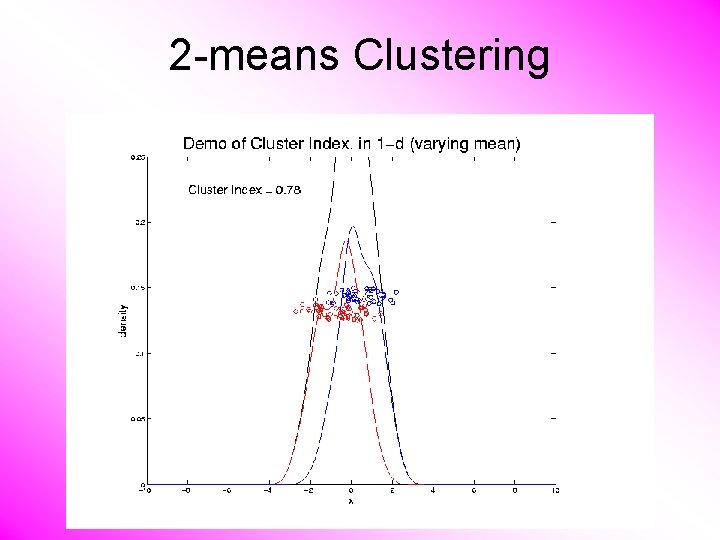
2 -means Clustering
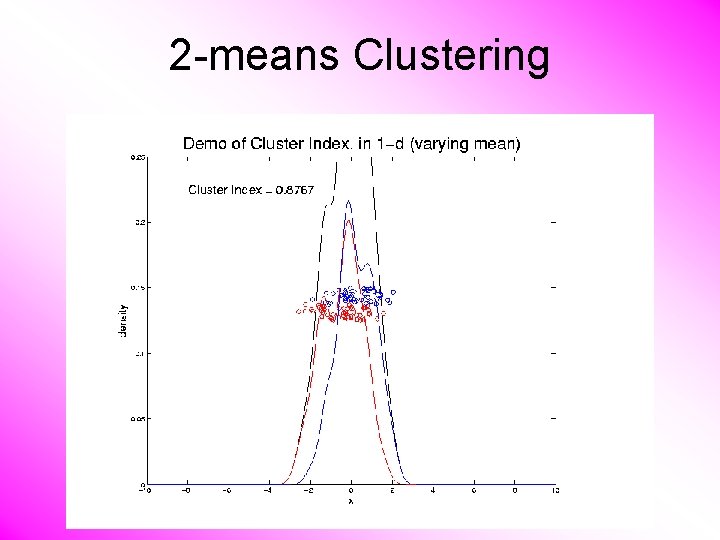
2 -means Clustering
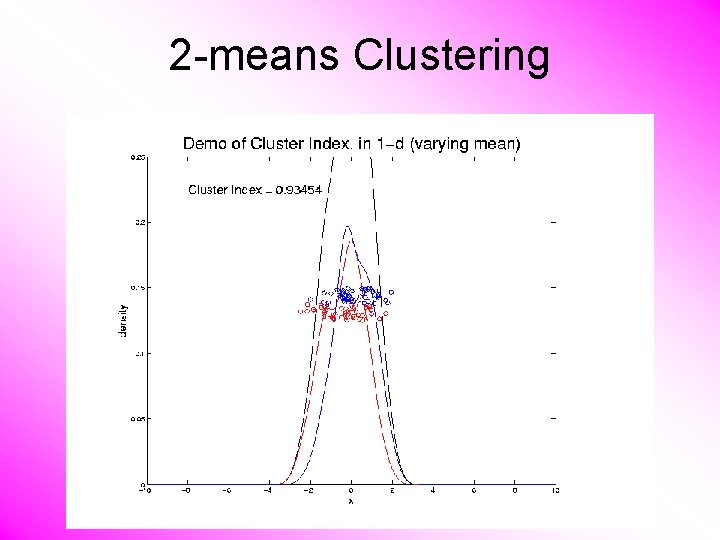
2 -means Clustering
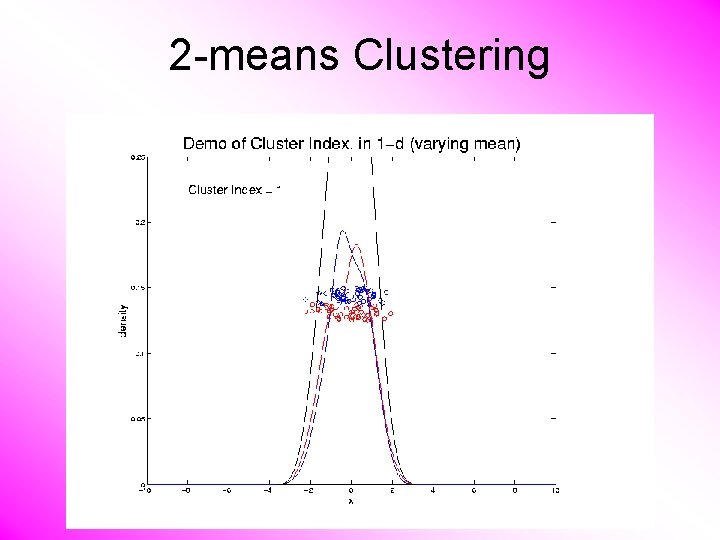
2 -means Clustering
- Slides: 75