Numerical Modeling of Biodegradation Analytical and Numerical Methods

Numerical Modeling of Biodegradation Analytical and Numerical Methods By Philip B. Bedient

Modeling Biodegradation • Three main methods for modeling biodegradation ² Monod kinetics ² First-order decay ² Instantaneous reaction • Methods can be used where appropriate for aerobic, anaerobic, hydrocarbon, or chlorinated

• Region 1: Lag phase ² Microbial Growth microbes are adjusting to the new substrate (food source) • Region 2 Exponential growth phase, ² microbes have acclimated to the conditions • Region 3 Stationary phase, ² limiting substrate or electron acceptor limits the growth rate • Region 4 Decay phase, ² substrate supply has been exhausted

Monod Kinetics • The rate of biodegradation or biotransformation is generally the focus of environmental studies • Microbial growth and substrate consumption rates have often been described using ‘Monod kinetics’ • • C is the substrate concentration [mg/L] Mt is the biomass concentration [mg/ L] µmax is the maximum substrate utilization rate [sec-1] KC is the half-saturation coefficient [mg/L]
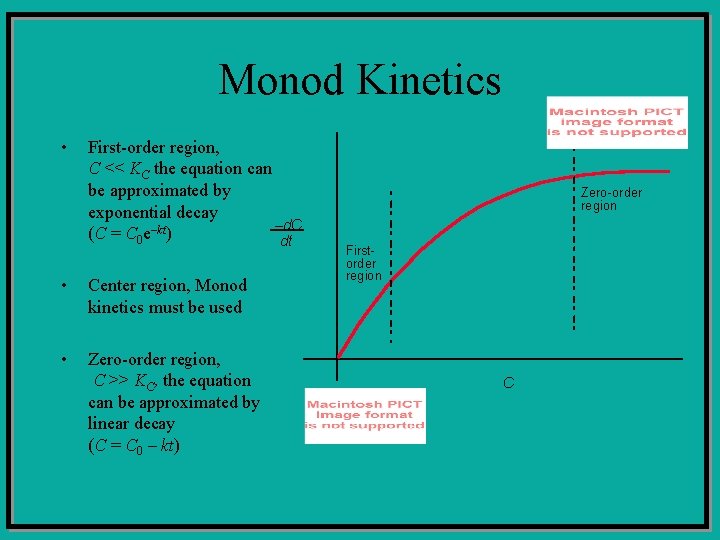
Monod Kinetics • First-order region, C << KC the equation can be approximated by exponential decay –d. C (C = C 0 e–kt) dt • Center region, Monod kinetics must be used • Zero-order region, C >> KC, the equation can be approximated by linear decay (C = C 0 – kt) Zero-order region Firstorder region C

Modeling Monod Kinetics • Reduction of concentration expressed as: • • • Mt µmax mass KC ∆t C = total microbial concentration = maximum contaminant utilization rate per of microorganisms = contaminant half-saturation constant = model time step size = concentration of contaminant

Bioplume II Equation - Monod • Including the previous equation for reaction results in this advection-dispersion-reaction equation:

Multi-Species Monod Kinetics • For multiple species, one must track the species together, and the rate is dependent on the concentrations of both species

Multi-Species • Adding these equations to the advection-dispersion equation results in one equation for each component (including microbes) • BIOPLUME III doesn’t model microbes

Modeling First-Order Decay • Cn+1 = Cn e–k∆t • Generally assumes nothing about limiting substrates or electron acceptors • Degradation rate is proportional to the concentration • Generally used as a fitting parameter, encompassing a number of uncertain parameters • BIOPLUME III can limit first-order decay to the available electron acceptors (this option has bugs)

Modeling Instantaneous Biodegradation • Excess Hydrocarbon: Hn > On/F • On+1 = 0 Hn+1 = Hn - On/F • Excess Oxygen: Hn < On/F • On+1 = On - Hn. F Hn+1 = 0 • All available substrate is biodegraded, limited only by the availability of terminal electron acceptors • First used in BIOPLUME II - 1987

Sequential Electron Acceptor Models • Newer models, such as BIOPLUME III, RT 3 D, and SEAM 3 D allow a sequential process - 1998 • After O 2 is depleted, begin using NO 3– • Continue down the list in this order O 2 ––> NO 3– ––> Fe 3+ ––> SO 42– ––> CO 2

Superposition of Components • Electron donor and acceptor are each modeled separately (advection/dispersion/sorption) • The reaction step is performed on the resulting plumes • Each cell is treated independently • Technique is called Operator Splitting

Principle of Superposition

Oxygen Utilization of Substrates • Benzene: C 6 H 6 + 7. 5 O 2 ––> 6 CO 2 + 3 H 2 O • Stoichiometric ratio (F) of oxygen to benzene • Each mg/L of benzene consumes 3. 07 mg/L of O 2

Biodegradation in BIOPLUME II

Initial Contaminant Plume

Model Parameters

Biodegrading Plume Original Plume Concentration Plume after two years Extraction Only - No Added O 2

Plume Concentrations Plume after two years O 2 Injected at 20 mg/L O 2 Injected at 40 mg/L

Biodegradation Models • • • Bioscreen -GSI Biochlor - GSI BIOPLUME II and III - Bedient & Rifai RT 3 D - Clement MT 3 D MS • SEAM 3 D

Biodegradation Models

Dehalogenation of PCE • PCE (perchloroethylene or tetrachloroethylene) • TCE (trichloroethylene) • DCE (cis-, trans-, and 1, 1 -dichloroethylene • VC (vinyl chloride)

Dehalogenation • Dehalogenation refers to the process of stripping halogens (generally Chlorine) from an organic molecule • Dehalogenation is generally an anaerobic process, and is often referred to as reductive dechlorination R–Cl + 2 e– + H+ ––> R–H + Cl– • Can occur via dehalorespiration or cometabolism • Some rare cases show cometabolic dechlorination in an aerobic environment

Chlorinated Hydrocarbons • Multiple pathways • • • Electron donor – similar to hydrocarbons Electron acceptor – depends on human-added electron donor Cometabolic • Mechanisms hard to define • First-order decay often used due to uncertainties in mechanism

Modeling Dechlorination • Few models specifically designed to simulate dechlorination • Some general models can accommodate dechlorination • Dechlorination is generally modeled as a firstorder biodegradation process • Often, the first dechlorination step results in a second compound that must also be dechlorinated

Sequential Dechlorination • Models the series of dechlorination steps between a parent compound a non-hazardous product • Each compound will have a unique decay constant • For example, the reductive dechlorination of PCE requires at least four constants • • PCE TCE DCE VC –k 1–> –k 2–> –k 3–> –k 4–> TCE DCE VC Ethene
- Slides: 27