Nucleotide Metabolism Pathways in nucleotide metabolism De novo

Nucleotide Metabolism

Pathways in nucleotide metabolism • De novo and salvage pathways • Nucleic acid degradation and the importance of nucleotide salvage • PRPP

Biosynthetic Routes: De novo and salvage pathways • Most organisms can synthesize purine and pyrimidrne nucleotides from low molecular weight precursors in amounts sufficient for their needs. These so called de novo pathways are essentially identical throughout the biological world. • Salvage pathways involve the utilization of preformed purme and pyrimidine compounds that would be otherwise lost to biodegradation. Salvage pathways represent important sites for manipulation of biological systems.
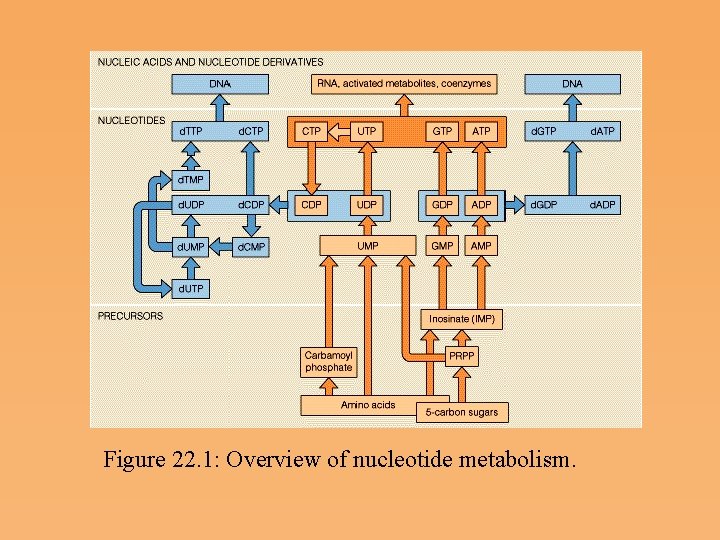
Figure 22. 1: Overview of nucleotide metabolism.
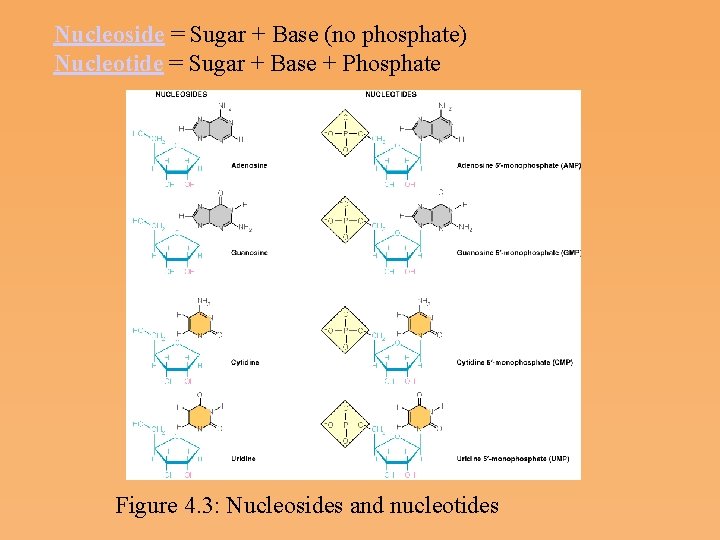
Nucleoside = Sugar + Base (no phosphate) Nucleotide = Sugar + Base + Phosphate Figure 4. 3: Nucleosides and nucleotides
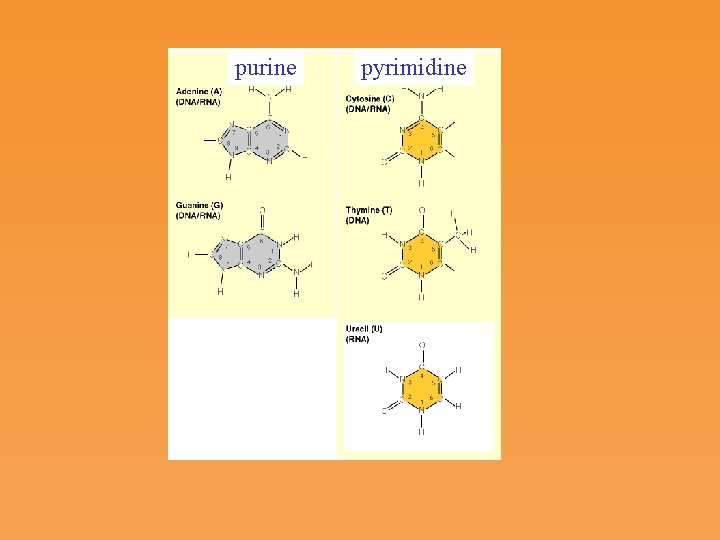
purine pyrimidine

Nucleic Acid Degradation and the Importance of Nucleotide Salvage • The salvage, or reuse, of purine and pyrimidine bases involves molecules released by nucleic acid degradation • Degradation can occur intracellularly, as the result of cell death, or, in animals, through digestion of nucleic acids ingested in the diet. • In animals, the extracellular hydrolysis of ingested nucleic acids represents the major route by which bases and nucleosides become available. Catalysis occurs by endonucleases, which function to digest nucleic acids in the small intestine. The products are mononucleotides. • If bases or nucleosides are not reused for nucleic acid synthesis via salvage pathways, the purine and pyrimidine bases are further degraded to uric acid or b ureidopropionate.
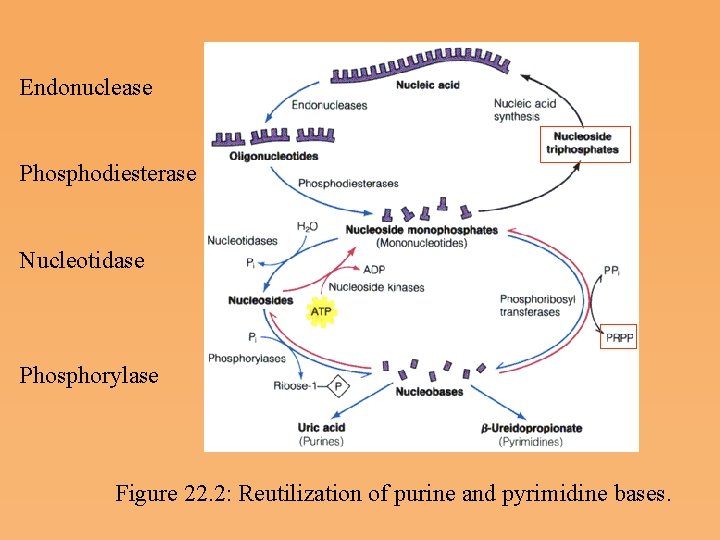
Endonuclease Phosphodiesterase Nucleotidase Phosphorylase Figure 22. 2: Reutilization of purine and pyrimidine bases.
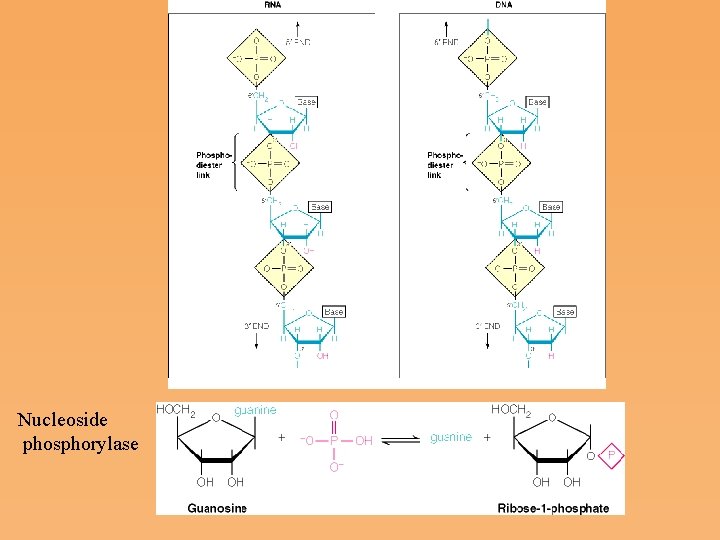
Nucleoside phosphorylase
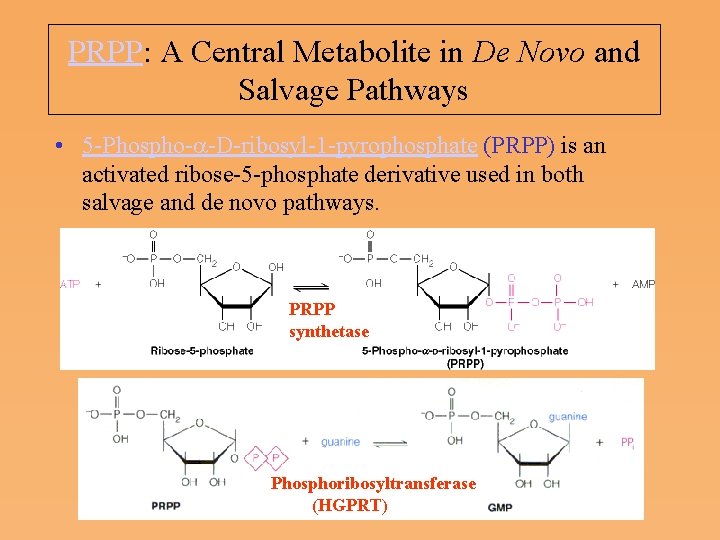
PRPP: A Central Metabolite in De Novo and Salvage Pathways • 5 Phospho a D ribosyl 1 pyrophosphate (PRPP) is an activated ribose 5 phosphate derivative used in both salvage and de novo pathways. PRPP synthetase Phosphoribosyltransferase (HGPRT)

De novo biosynthesis of purine nucleotides • Purine synthesis from PRPP to inosinic acid • Synthesis of ATP and GTP from inosinic acid • Utilization of adenine nucleotides in coenzyme biosynthesis
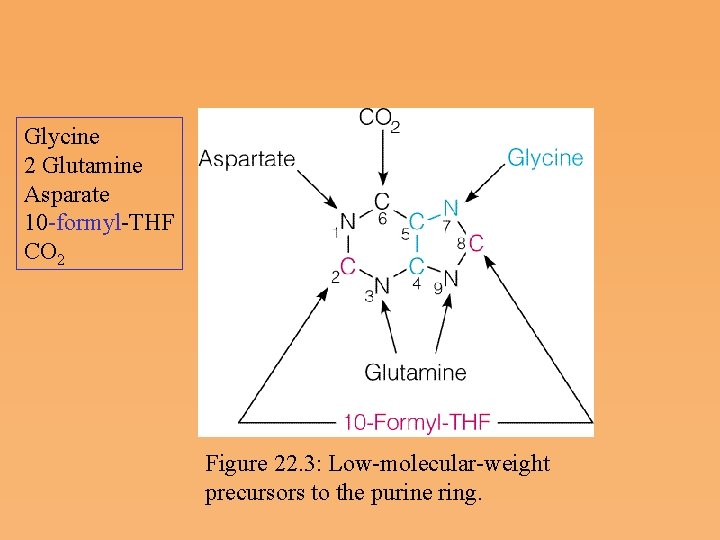
Glycine 2 Glutamine Asparate 10 formyl THF CO 2 Figure 22. 3: Low molecular weight precursors to the purine ring.

Purine synthesis from PRPP to inosinic acid • Purines are synthesized at the nucleotide level, starting with PRPP conversion to phosphoribosylamine and purine ring assembly on the amino group. • Vertebrate cells have several multifunctional enzymes involved in these processes. • Control over the biosynthesis of inosinic acid is provided through feedback regulation of early steps in purine nucleotide synthesis. PRPP synthetase is inhibited by various purine nucleotides, particularly AMP, ADP, and GDP, and PRPP amidotransferase is also inhibited by AMP, ADP, GMP, and GDP.
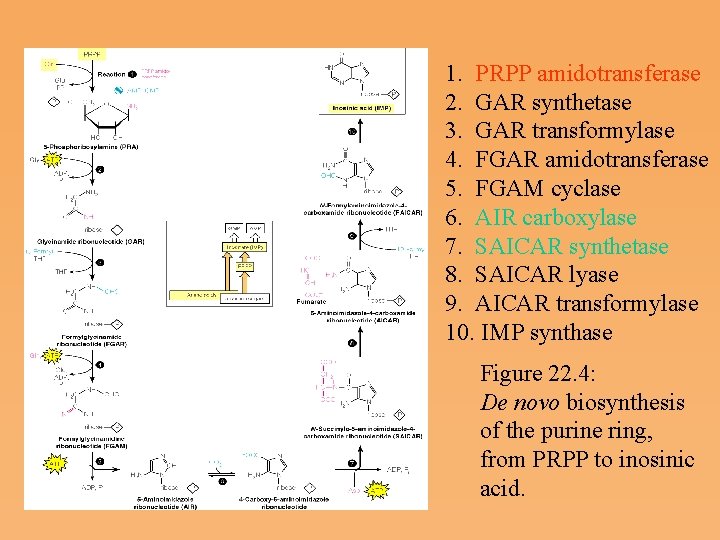
1. PRPP amidotransferase 2. GAR synthetase 3. GAR transformylase 4. FGAR amidotransferase 5. FGAM cyclase 6. AIR carboxylase 7. SAICAR synthetase 8. SAICAR lyase 9. AICAR transformylase 10. IMP synthase Figure 22. 4: De novo biosynthesis of the purine ring, from PRPP to inosinic acid.

Gln Glu, PPi PRPP amidotransferase AMP, GMP
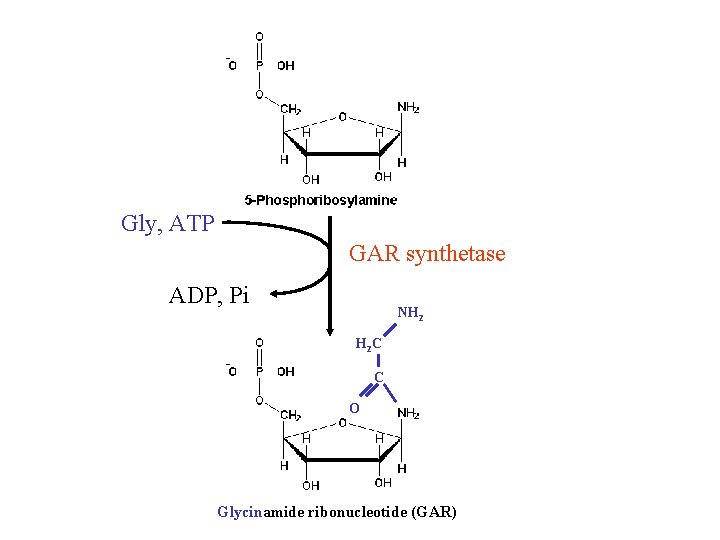
Gly, ATP GAR synthetase ADP, Pi NH 2 H 2 C C O Glycinamide ribonucleotide (GAR)
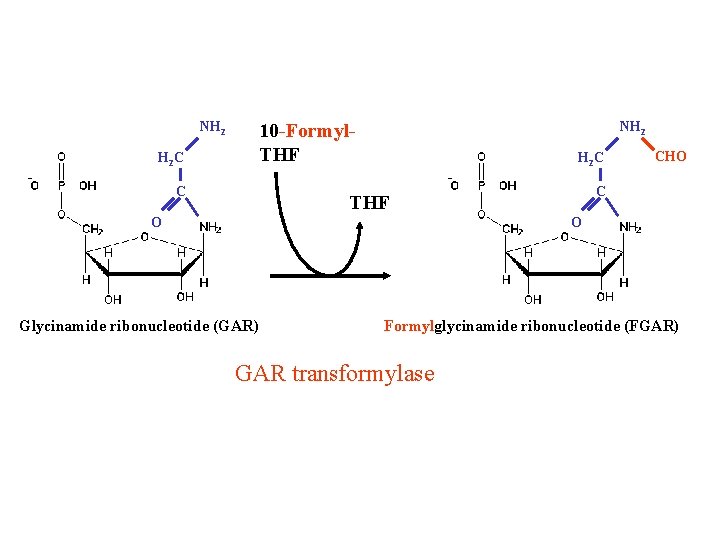
NH 2 10 -Formyl. THF H 2 C C THF O CHO O Glycinamide ribonucleotide (GAR) Formylglycinamide ribonucleotide (FGAR) GAR transformylase
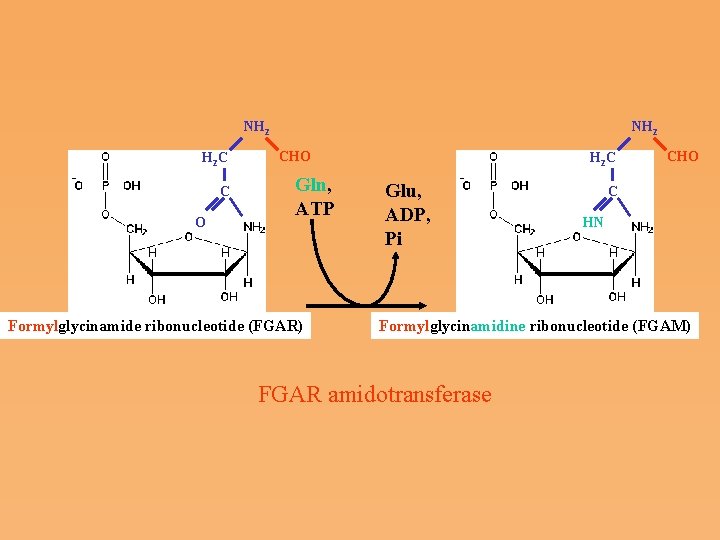
NH 2 H 2 C C O NH 2 CHO Gln, ATP Formylglycinamide ribonucleotide (FGAR) H 2 C Glu, ADP, Pi CHO C HN Formylglycinamidine ribonucleotide (FGAM) FGAR amidotransferase
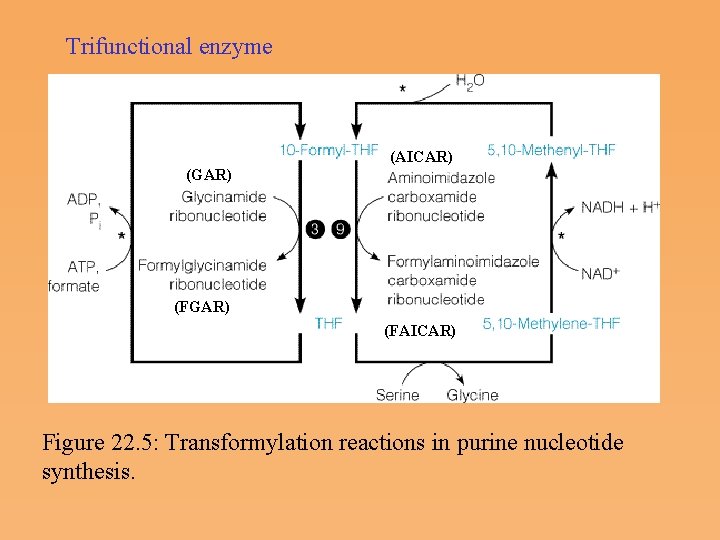
Trifunctional enzyme (AICAR) (GAR) (FAICAR) Figure 22. 5: Transformylation reactions in purine nucleotide synthesis.

Potent inhibitors of purine nucleotide synthesis structural analogs of glutamine amidotransferases

Synthesis of ATP and GTP from inosinic acid • IMP is the first fully formed purine nucleotide and is a branch point between adenine and guanine nucleotide biosynthesis. • The energy to drive the aspartate transfer reaction comes not from ATP but from GTP. • GTP accumulation would tend to promoter the pathway toward adenine nucleotide. Also accumulation of ATP could promote guanine nucleotide synthesis. • The enzyme catalyzing the pathway to make AMP is inhibited by AMP and the enzyme catalyzing the pathway to make GMP is inhibited by GMP

IMP dehydrogenase GMP AMP Inosine Monophosphate (IMP) H XMP aminase Adenylosuccinate synthetase Adenylosuccinate lyase Figure 22. 6: Pathways from inosinic acid to GMP and AMP.

• Nucleotides are active in metabolism primarily as the nucleoside triphosphates. GMP and AMP are converted to their corresponding triphosphates through two successive phosphorylation reactions. Conversion to the diphosphates involves specific ATP dependent kinases. GMP + ATP AMP + ATP Guanylate kinase Adenylate kinase GDP + ADP 2 ADP

• Phosphorylation of ADP to ATP occurs through energy metabolism, by oxidative phosphorylation, or by substrate level phosphorylations. ATP can also be formed from ADP through the action of adenylate kinase, acting in the reverse of the direction. • ATP is the phosphate donor for conversion of GDP (and other nucleotide diphosphate) to the triphosphate level through the action of nudeoside diphosphokinase. GDP + ATP GTP +ADP • Nucleoside diphosphokinase is an equilibrium driven enzyme that transfers phosphate from ATP in the synthesis of all other nucleoside triphosphates.

Purine degradation and clinical disorders of purine metabolism Formation uric acid • All purine nucleotide catabolism yields uric acid. • Purine catabolism in primates ends with uric acid, which is excreted. Most other animals further oxidize the purine ring, to allantoin and then to allantoic acid, which is either excreted or further catabolized to urea or ammonia.
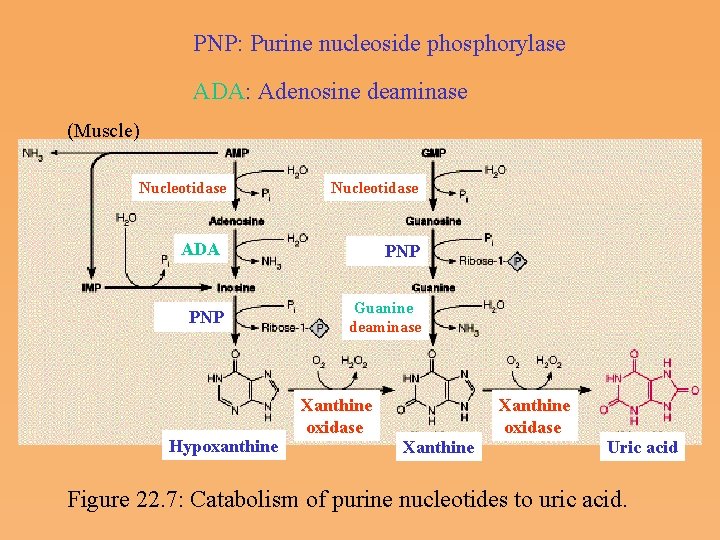
PNP: Purine nucleoside phosphorylase ADA: Adenosine deaminase (Muscle) Nucleotidase ADA PNP Guanine deaminase Hypoxanthine Xanthine oxidase Xanthine Uric acid Figure 22. 7: Catabolism of purine nucleotides to uric acid.
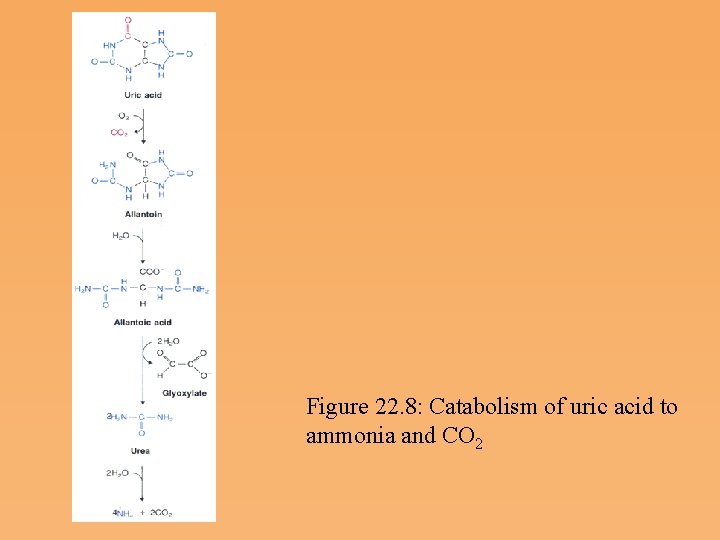
Figure 22. 8: Catabolism of uric acid to ammonia and CO 2

Excessive accumulation of uric acid: gout • Uric acid and its urate salts are very insoluble. This is an advantage to egg laying animals, because it provides a route for disposition of excess nitrogen in a closed environment. • Insolubility of urates can present difficulties in mammalian metabolism. About 3 humans in 1000 suffer from hyperuricemia, which is chronic elevation of blood uric acid levels well beyond normal levels. The biochemical reasons for this vary, but the condition goes by a single clinical name, which is gout.

• Prolonged or acute elevation of blood urate leads to precipitation, as crystals of sodium urate, in the synovial fluid of joints. These precipitates cause inflammation, resulting in painful arthritis, which can lead to severe degeneration of the joints. • Gout results from overproduction of purine nucleotides, leading to excessive uric acid synthesis, or from impaired uric acid excretion through the kidney • Several known genetic alterations in purine metabolism lead to purine oversynthesis, uric acid overproduction, and gout. Gout can also result from mutations in PRPP amidotransferase that render it less sensitive to feedback inhibition by purine nucleotides. Another cause of gout is a deficiency of the salvage enzyme hypoxanthine Guanine phosphoribosyltransferase (HGPRT).
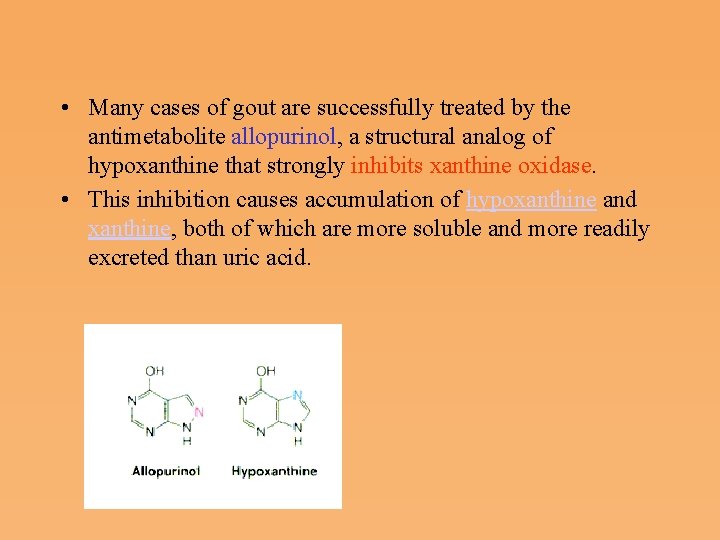
• Many cases of gout are successfully treated by the antimetabolite allopurinol, a structural analog of hypoxanthine that strongly inhibits xanthine oxidase. • This inhibition causes accumulation of hypoxanthine and xanthine, both of which are more soluble and more readily excreted than uric acid.
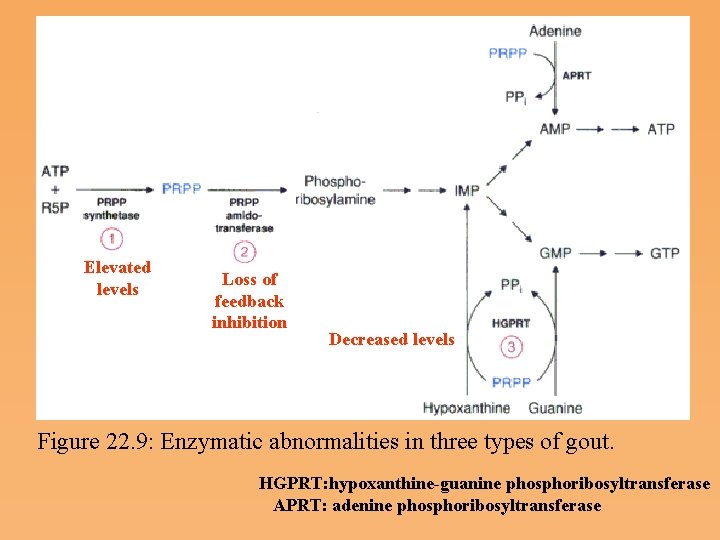
Elevated levels Loss of feedback inhibition Decreased levels Figure 22. 9: Enzymatic abnormalities in three types of gout. HGPRT: hypoxanthine-guanine phosphoribosyltransferase APRT: adenine phosphoribosyltransferase

Lesch Nyhan syndrome: HGPRT defficiency • Lesch Nyhan syndrome is a sex linked trait, because the structural gene for HGPRT is located on the X chromosome. • Patients with this condition display a severe gouty arthritis, but they also have dramatic malfunction of the nervous system, manifested as behavioral disorders, learning disabilities, and hostile or aggressive behavior, often self directed. • At present, there is no successful treatment, and afflicted individuals rarely live beyond 20 years.

Severe combined immune deficiency (SCID) • Patients with a hereditary condition called severe combined immunodeficiency syndrome are susceptible, often fatally, to infectious diseases because of an inability to mount an immune response to antigenic chanllenge. • In this condition, both B and T lymphocytes are affected. Neither class of cells can proliferate as they must if antibodies are to be synthesized. In many cases the condition is caused from a heritable lack of the degradative enzyme adenosine deaminase (ADA). • The deficiency of ADA leads to accumulation of d. ATP which is known to be a potent inhibitor of DNA replication.

• A less severe immunodeficiency results from the lack of another purine degradative enzyme, purine nuceloside phosphorylase (PNP). Decreased activity of this enzyme leads to accumulation primarily of d. GTP. This accumulation also affects DNA replication, but less severely than does excessive d. ATP. • Interestingly, the phosphorylase deficiency destroys only the T class of lymphocytes and not the B cells.

Pyrimidine nucleotide metabolism • Pyrimidine nucleotide synthesis occurs primarily at the free base level, with conversion to a nucleotide occurring later in the unbranched pathway. • Pyrimidine synthesis begins with formation of carbamoyl phosphate. The first reaction committed solely to pyrimidine synthesis is the formation of carbamoyl aspartate from carbamoyl phosphate and aspartate, catalyzed by aspartate transcarbamoylase, or ATCase. • In enteric bacteria, this enzyme represents an example of feedback control. The enzyme is inhibited by the end product CTP and activated by ATP.
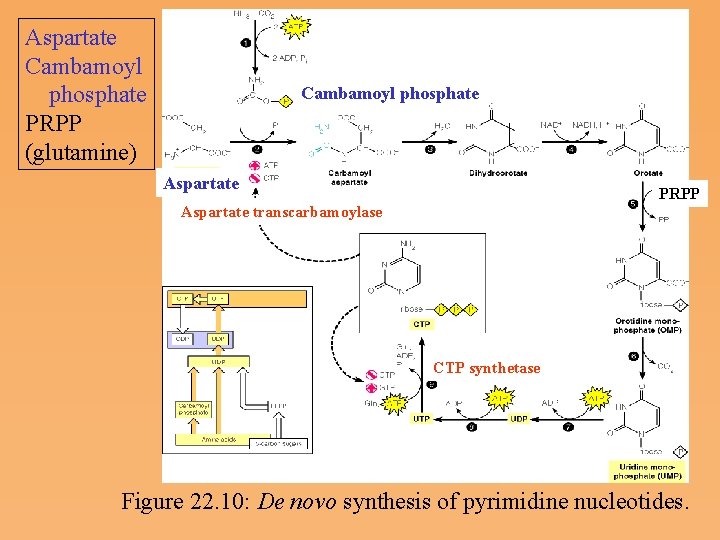
Aspartate Cambamoyl phosphate PRPP (glutamine) Cambamoyl phosphate Aspartate PRPP Aspartate transcarbamoylase CTP synthetase Figure 22. 10: De novo synthesis of pyrimidine nucleotides.

Multifunctional Enzymes in Eukaryotic Pyrimidine Synthesis • Aspartate transcarbamoylase in eukaryotes is strikingly different from the E. coli enzyme. In eukaryotes, the first three reactions of pyrimidine synthesis are catalyzed by a multifunctional enzyme, the CAD protein (Carbamoyl phosphate synthetase, Aspartate transcarbamoylase, and Dihydroorotase). • In mammalian cells, reactions 5 and 6 (see Figure 22. 10) are also catalyzed by a single protein, which is called UMP synthase. • A site for control of pyrimidine nucleotide synthesis is the amidotransferase, CTP synthetase, which converts UTP to CTP.

Salvage Synthesis and Pyrimidine Catabolism • Pyrimidine nucleotides are also synthesized by salvage pathways involving phosphorylases and kinases, comparable to those already discussed for purines. • The catabolic pathways for pyrimidines are simpler than those for purines. Because the intermediates are relatively soluble, there are few known derangements of pyrimidine breakdown. • b alanine is used in the biosynthesis of coenzyme A
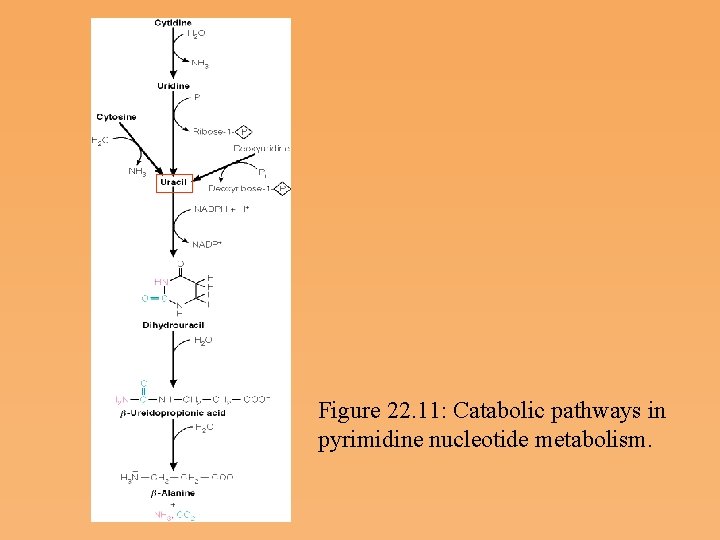
Figure 22. 11: Catabolic pathways in pyrimidine nucleotide metabolism.

Deoxyribonucleotide biosynthesis and metabolism • Most cells contain 5 to 10 times as much RNA as DNA. • The small fraction that is diverted to the synthesis of deoxyribonucleoside triphosphates (d. NTPs) is of paramount importance to the cell. • d. NTPs are used almost exclusively in the biosynthesis of DNA. There are very close regulatory relationships between DNA synthesis and d. NTP metabolism. • DNA differs chemically from RNA in the nature of the sugar and in the identity of one of the pyrimidine bases. • The deoxyribonucleotide biosynthesis on two specific processes: the conversion of ribose to deoxyribose, and the conversion of uracil to thymine. Both processes occur at the nucleotide level.
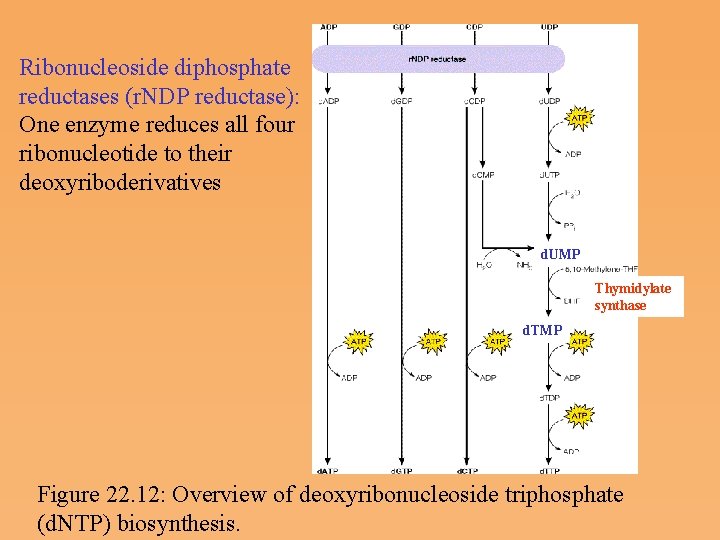
Ribonucleoside diphosphate reductases (r. NDP reductase): One enzyme reduces all four ribonucleotide to their deoxyriboderivatives d. UMP Thymidylate synthase d. TMP Figure 22. 12: Overview of deoxyribonucleoside triphosphate (d. NTP) biosynthesis.

Ribonucleoside diphosphate reductase • The enzyme catalyzing the synthesis of d. NDPs from r. NDPs, reduces the hydroxyl at carbon 2’ to a hydrogen via a free radical mechanism. • Ribonucleotide reductase contains catalytic residues on each of its subunits-redox active thiols and a tyrosine free radical stabilized by an iron-Oxygen complex. • Hydroxyurea, an inhibitor of ribonucleotide reductase, destroys the free radical.
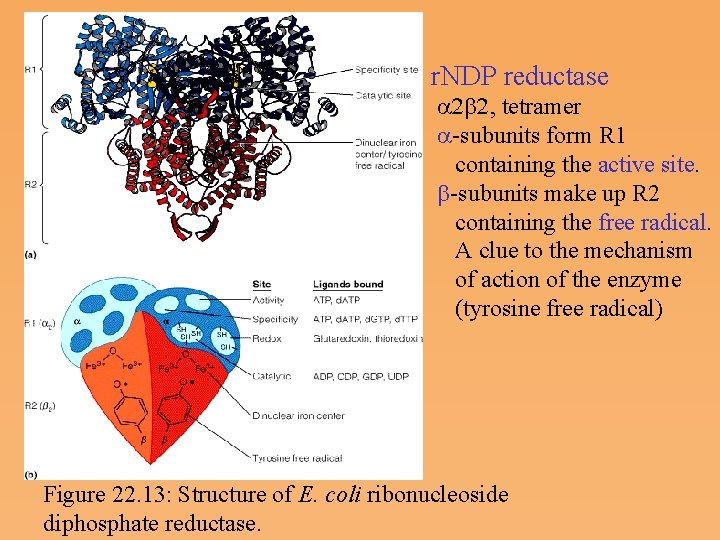
r. NDP reductase a 2 b 2, tetramer a subunits form R 1 containing the active site. b subunits make up R 2 containing the free radical. A clue to the mechanism of action of the enzyme (tyrosine free radical) Figure 22. 13: Structure of E. coli ribonucleoside diphosphate reductase.
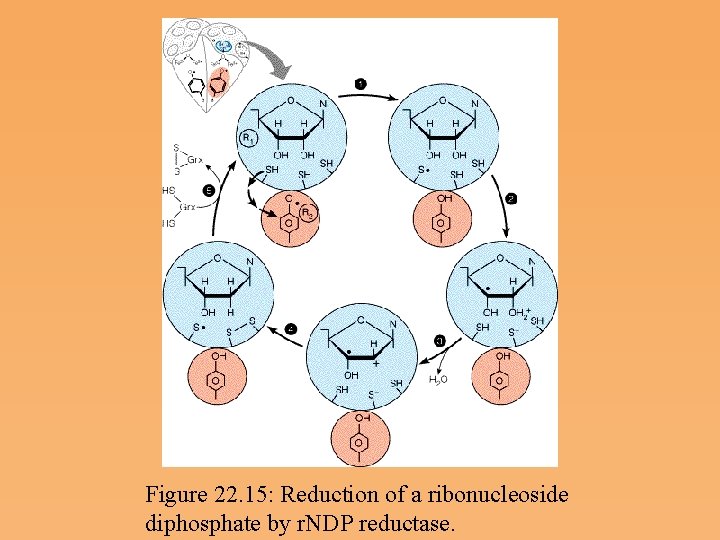
Figure 22. 15: Reduction of a ribonucleoside diphosphate by r. NDP reductase.

Source of Electrons for r. NDP Reduction • Electrons for the reduction of ribonucleotides come ultimately from NADPH, but they are shuttled to r. NDP reductase by a coenzyme that is unusual because it is itself a protein. • Ribonucleotide reductase uses a protein cofactor, thioredoxin or glutaredoxin, to provide electrons for reduction of the ribonucleotide substrate.
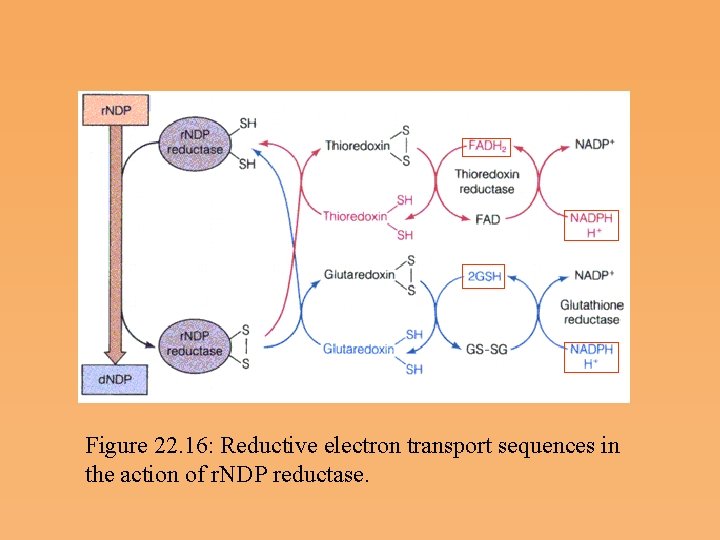
Figure 22. 16: Reductive electron transport sequences in the action of r. NDP reductase.

Table 22. 1 Biological activities of thioredoxin

Regulation of Ribonucleotide Reductase Activity • Ribonucleotide reductase has two classes of allosteric sites. Activity sites influence catalytic efficiency, and specificity sites determine specificity for one or more of the four substrates. • The activity sites bind either ATP or d. ATP with relatively low affinity, whereas the specificity sites bind ATP, d. GTP, or d. TTP with relatively high affinity. • Inhibition of DNA synthesis by thymidine or deoxyadenosine involves allosteric inhibition of ribonucleotide reductase by d. TTP or d. ATP, respectively

Table 22. 2 Regulation of Ribonucleotide Reductase Activity

Biosynthesis of Thymine Deoxyribonucleotides • d. UMP, the substrate for thymidylate synthesis, can arise either from UDP reduction and dephosphorylation or from deammation of a deoxycytidine nucleotide (d. CMP). • In the reaction catalyzed by thymidylate synthase, 5, 10 methylenetetrahydrofolate donates both a single carbon group and an electron pair to reduce that group to the methyl level.
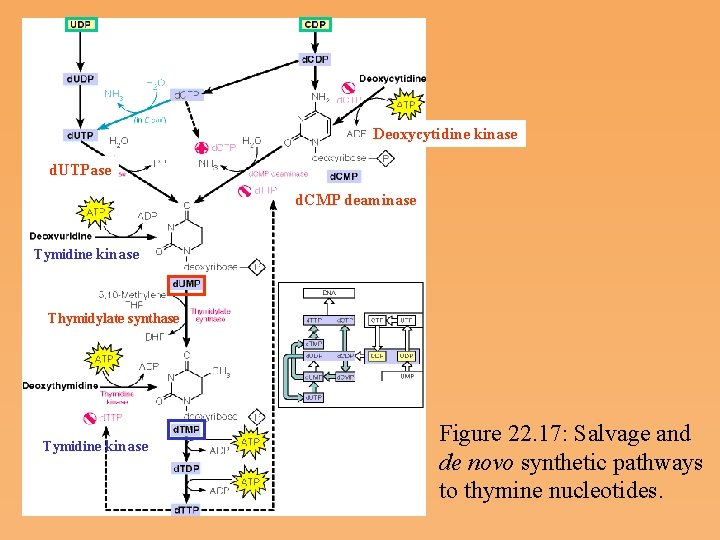
Deoxycytidine kinase d. UTPase d. CMP deaminase Tymidine kinase Thymidylate synthase Tymidine kinase Figure 22. 17: Salvage and de novo synthetic pathways to thymine nucleotides.
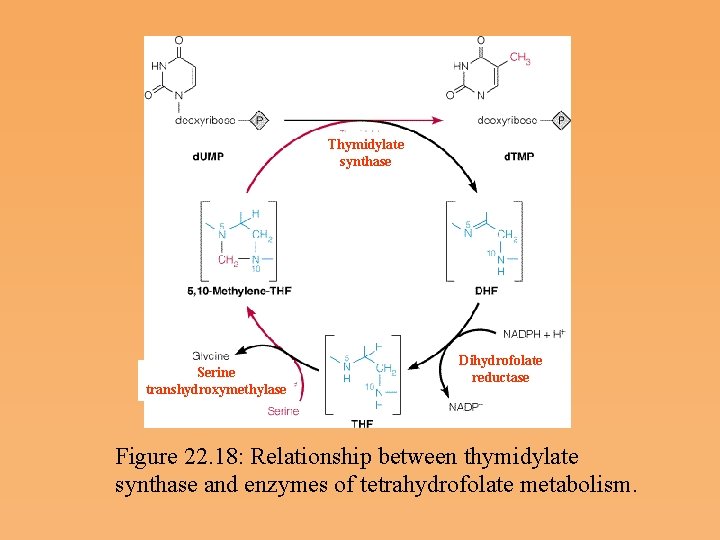
Thymidylate synthase Serine transhydroxymethylase Dihydrofolate reductase Figure 22. 18: Relationship between thymidylate synthase and enzymes of tetrahydrofolate metabolism.

Deoxyuridine Nucleotide Metabolism • In addition to the biosynthetic function of d. UTPase in forming d. UMP for thymine nucleotide formation, the enzyme plays an important role in excluding uracil from DNA. • d. UMP residues can arise in DNA not only by d. UTP incorporation but also by the spontaneous deamination of d. CMP residues.

Salvage Routes to Deoxyribonucleotide Synthesis • Of the various deoxyribonucleoside kinases, one that merits special mention is thymidine kinase (TK). This enzyme is allosterically inhibited by d. TTP. Activity of thymidine kinase in a given cell is closely related to the proliferative state of that cell. During the cell cycle, activity of TK rises dramatically as cells enter S phase. In general, rapidly dividing cells have high levels of this enzyme. • The salvage pathway to d. TTP competes very efficiently with the thymidylate synthase mediated de novo pathway. • Deoxycytidine kinase is also a salvage enzyme that is feedback inhibited by d. CTP. This enzyme also functions as a deoxyadenosine kinase and a deoxyguanosine kinase. Unlike thymidine kinase, whose activity fluctuates over the course of the cell cycle, the activity of deoxycytidine kinase stays relatively constant.

Thymidylate synthase: a target enzyme for chemotherapy • A goal of chemotherapy is to exploit a biochemical difference between the disease process and the host tissue in order to interfere selectively with the disease process. • Many chemotherapeutic agents were discovered by chance, through testing of analogs of normal metabolites. Most of these agents are limited in their effectiveness by unanticipated side effects, incomplete selectivity, and the development of resistance to the agent. • One of the most exciting areas of modern biochemical pharmacology is drug architecture: the design of specific inhibitors based on knowledge of the molecular structure of the site to which the inhibitor will bind and the mechanism of action of the target molecule.

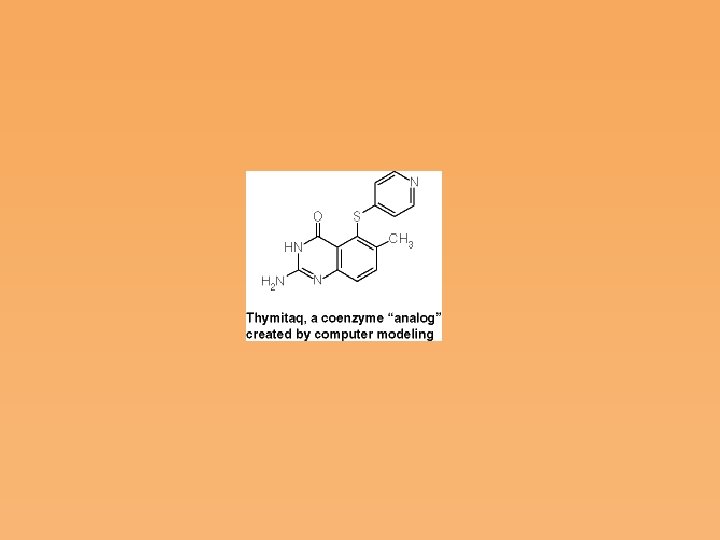
• Inhibition of thymidylate synthase is an approach to cancer chemotherapy, by causing specific inhibition of DNA synthesis. Cells that are not rapidly proliferating should be relatively immune to such agents. Thus, cancer and a wide range of infectious diseases should be amenable to treatment by this approach. • Knowing the active site structure and reaction mechanism of an enzyme allows the design of novel inhibitors, an approach used for thymidylate synthase, but applicable to many other drugs.
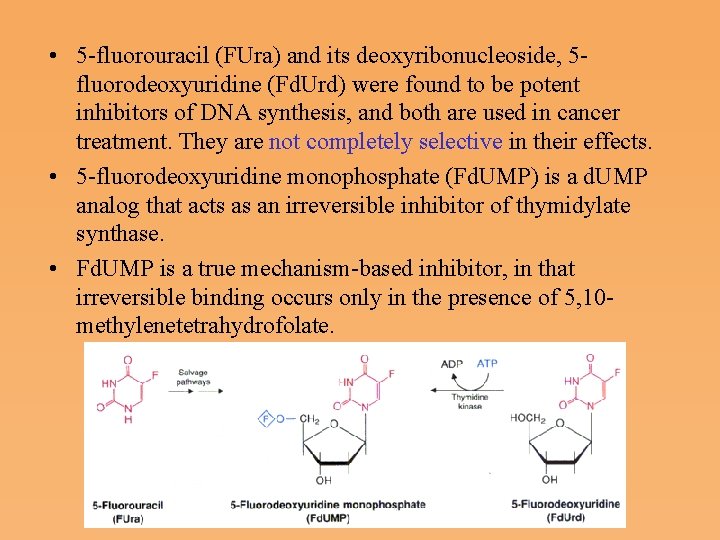
• 5 fluorouracil (FUra) and its deoxyribonucleoside, 5 fluorodeoxyuridine (Fd. Urd) were found to be potent inhibitors of DNA synthesis, and both are used in cancer treatment. They are not completely selective in their effects. • 5 fluorodeoxyuridine monophosphate (Fd. UMP) is a d. UMP analog that acts as an irreversible inhibitor of thymidylate synthase. • Fd. UMP is a true mechanism based inhibitor, in that irreversible binding occurs only in the presence of 5, 10 methylenetetrahydrofolate.
Figure 22. 20: Mechanism for the reaction catalyzed by thymidylate synthase.
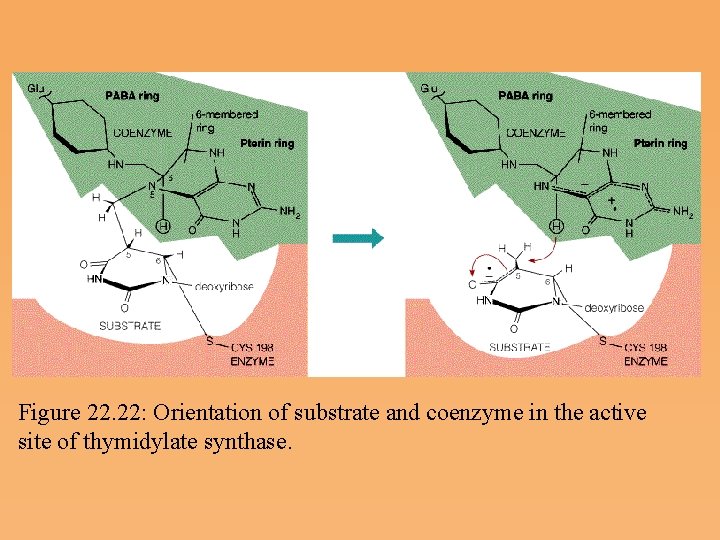
Figure 22. 22: Orientation of substrate and coenzyme in the active site of thymidylate synthase.

Virus Directed Alterations of Nucleotide Metabolism • The T even bacteriophages that viruses can redirect the metabolism of their host cells.
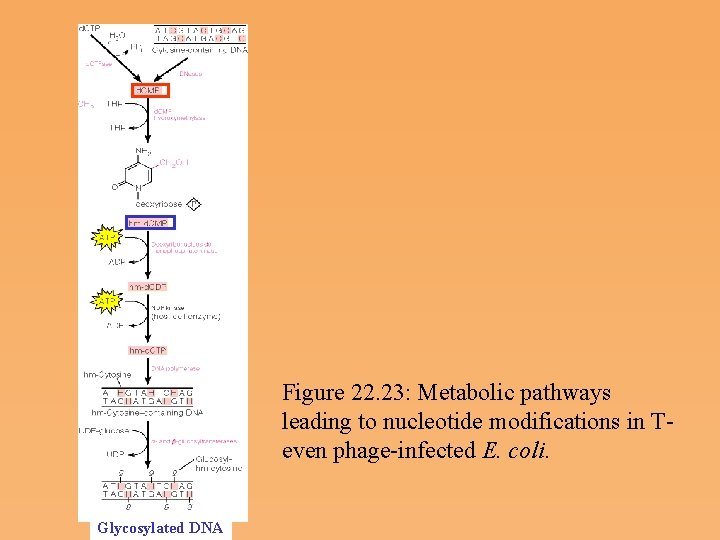
Figure 22. 23: Metabolic pathways leading to nucleotide modifications in T even phage infected E. coli. Glycosylated DNA
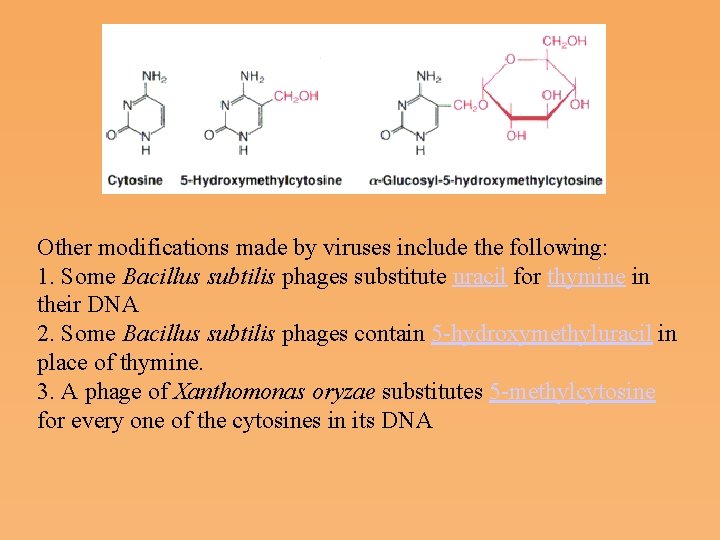
Other modifications made by viruses include the following: 1. Some Bacillus subtilis phages substitute uracil for thymine in their DNA 2. Some Bacillus subtilis phages contain 5 hydroxymethyluracil in place of thymine. 3. A phage of Xanthomonas oryzae substitutes 5 methylcytosine for every one of the cytosines in its DNA

Biological and Medical Importance of Other Nucleotide Analogs • Nucleotide Analogs as Chemotherapeutic Agents Antiviral Nucleoside Analogs Purine Salvage as a Target Folate Antagonists (Figure 22. 18) • Nucleotide Analogs and Mutagenesis (Figure 22. 24) • Nucleotide Metabolizing Enzymes as Selectable Genetic Markers

Nucleotide Analogs as Chemotherapeutic Agents • The enzymes of nucleotide synthesis have been widely studied as target sites for the action of antiviral or antimicrobial drugs. • Other analogs receiving considerable attention are those being used to combat acquired immune deficiency syndrome (AIDS). One such analog is AZT, the first drug approved in the United States for the treatment of HIV infections.
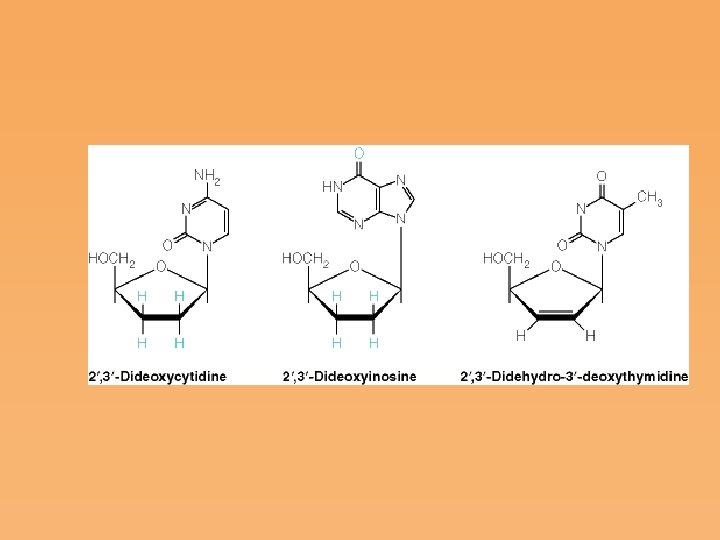
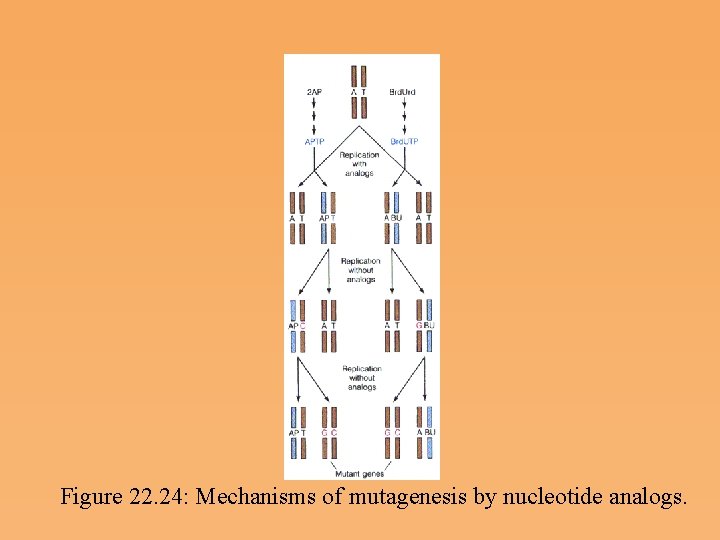
Figure 22. 24: Mechanisms of mutagenesis by nucleotide analogs.

Selectable genetic markers • Because most cells can synthesize nucleotides de novo, the enzymes of salvage synthesis are usually nonessential for cell viability. This means that nucleotide metabolizing enzymes and the genes that encode them provide selectable genetic markers, which have a variety of uses. • Separate salvage and de novo pathways allow selection for survival or death of cells with particular metabolic traits.
• Modified nucleotides can be used to select cells containing or lacking specific enzymes. Examples include the following: 6 Thioguanine selects for cells lacking an active hypoxanthine guanine phosphoribosyltransferase (HGPRT). Cells containing an active enzyme convert 6 thioguanine to a toxic compound. • 5 Bromodeoxyuridine (Brd. Urd) can be used to select cells lacking thymidine kinase, which is needed to metabolize Brd. Urd to a toxic metabolite.

• HAT Selection The compounds hypoxanthine, aminopterin, and thymidine (H, A, and T, respectively) can be used to select for cells having functional salvage pathways. • Aminopterin inhibits dihydrofolate reductase, which blocks de novo purine and thymidine synthesis. Only cells which can utilize thymidine (pyrimidine salvage) and hypoxanthine (purine salvage) can grow in this medium.
- Slides: 70