Nucleotide Catabolism and Salvage Membrane Signaling Andy Howard

Nucleotide Catabolism and Salvage; Membrane Signaling Andy Howard Biochemistry Lectures, Spring 2019 Thursday 2 May 2019

Membranes and Metabolism n n Nucleotides can be broken down, but they’re often recycled instead Signal transduction is a critical part of what membranes do 05/02/2019 Nucleotide Catabolism; Signaling p. 2 of 63

What we’ll discuss n n n Ribonucleotide reductase Thymidylate synthesis Nucleotide catabolism and salvage n n Reactions Medical significance 05/02/2019 n Membrane signaling n n n G proteins Adenylyl cyclase PIP 2 pathways Sphingolipidbased signaling Tyrosine kinases Nucleotide Catabolism; Signaling p. 3 of 63
i. Clicker question #1 1. Which of the following is an energyproviding cosubstrate in converting IMP to adenylosuccinate on the way to making AMP? 05/02/2019 n n n (a) ATP (b) ITP (c) CTP (d) GTP (e) none of the above. Nucleotide Catabolism; Signaling p. 4 of 63
i. Clicker question #2 2. P-loop proteins are typically n n 05/02/2019 (a) CTP-binding proteins (b) GTP-binding proteins (c) ATP-binding proteins (d) none of the above. Nucleotide Catabolism; Signaling p. 5 of 63

Making deoxyribonucleotides n Conversions of nucleotides to deoxynucleotides n n occurs at the diphosphate level Reichard showed that most organisms have a single ribonucleotide reductase that converts ADP, GDP, CDP, UDP to d. ADP, d. GDP, d. CDP, and d. UDP NADPH is the reducing agent 05/02/2019 Nucleotide Catabolism; Signaling p. 6 of 63
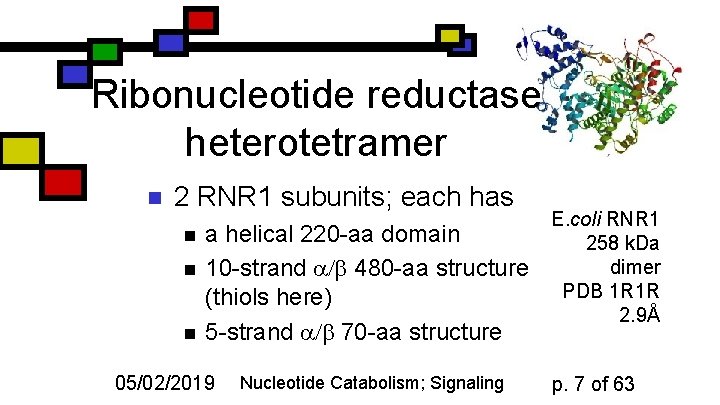
Ribonucleotide reductase heterotetramer n 2 RNR 1 subunits; each has n n n a helical 220 -aa domain 10 -strand 480 -aa structure (thiols here) 5 -strand 70 -aa structure 05/02/2019 Nucleotide Catabolism; Signaling E. coli RNR 1 258 k. Da dimer PDB 1 R 1 R 2. 9Å p. 7 of 63

RNR 2 n 2 RNR 2 subunits; each has n n A diferric ion center A stable tyrosyl free radical 05/02/2019 E. coli RNR 2 82 k. Da dimer EC 1. 17. 4. 1 PDB 1 PJ 0, 1. 9Å Nucleotide Catabolism; Signaling p. 8 of 63
Mechanism of RNR n n n Y 122 in RNR 2 is converted to stable free radical Radical transmitted to RNR 1 cys 439 Cys 439 reacts with substrate 3’-OH to form free radical at C 3’ Substrate dehydrates to carbonyl at C 3’ and free radical at C 2’; S- formed at Cys 462 Disulfide formed between Cys 462, Cys 225; radical regenerated at Cys 439 05/02/2019 Nucleotide Catabolism; Signaling p. 9 of 63
RNR : control I n n ATP, d. TTP, and d. GTP act as allosteric modulators by binding to two regulatory sites on the enzyme Activity site (A) regulates activity of catalytic site n When ATP binds at A, activity goes up n When d. ATP binds at A, activity inhibited overall 05/02/2019 Nucleotide Catabolism; Signaling p. 10 of 63

RNR Control II n Specificity site (S) controls which substrates can be turned over n n n ATP at A + ATP or d. ATP at S : pyrimidines only d. TTP at S : activates reduction of GDP d. GTP at S : activates reduction of ADP 05/02/2019 Nucleotide Catabolism; Signaling p. 11 of 63

d. UDP to d. UMP n d. UMP derived three ways en route to d. TMP: n n d. UDP + ADP d. UMP + ATP d. UDP + ATP d. UTP + ADP d. UTP + H 2 O d. UMP + PPi d. CMP + H 2 O d. UMP + NH 4+ d. TMP formed at monophosphate level (from d. UMP) 05/02/2019 Nucleotide Catabolism; Signaling p. 12 of 63

Thymidylate synthase 5, 10 methylene THF n n 05/02/2019 d. UMP + 5, 10 -methylene. THF d. TMP + 7, 8 -dihydrofolate Unusual THF reaction in that cofactor gets oxidized as well as giving up a carbon n CH 2 from 5, 10 -methylene group n extra H from C 6 Nucleotide Catabolism; Signaling p. 13 of 63
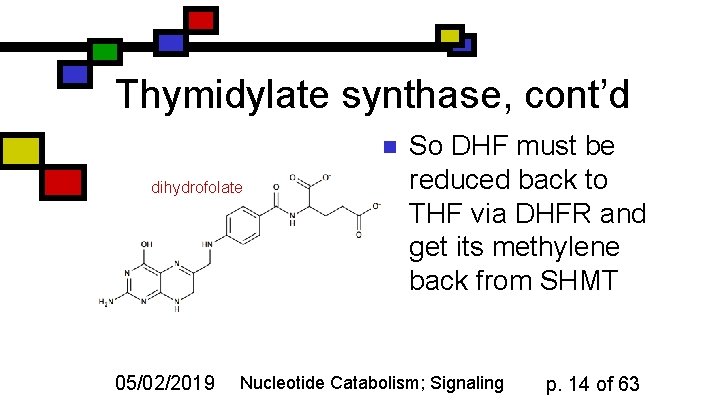
Thymidylate synthase, cont’d n dihydrofolate 05/02/2019 So DHF must be reduced back to THF via DHFR and get its methylene back from SHMT Nucleotide Catabolism; Signaling p. 14 of 63
Thymidylate synthase n n n Generally the controlling step in DNA synthesis because [d. TTP] < other [deoxynucleoside triphosphates] Therefore a target for cancer chemotherapy and otherapies that target rapidly-dividing cells Enzyme is a 2 -layer sandwich 05/02/2019 Nucleotide Catabolism; Signaling E. coli TS 64 k. Da dimer with d. UMP + GA 9 EC 2. 1. 1. 45 PDB 2 A 9 W, 1. 7Å p. 15 of 63
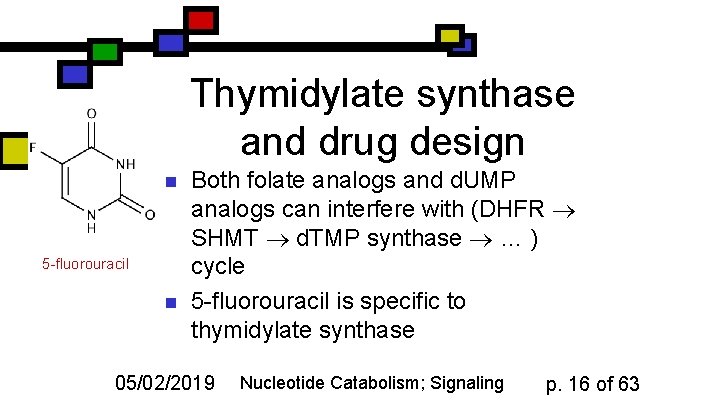
Thymidylate synthase and drug design n 5 -fluorouracil n Both folate analogs and d. UMP analogs can interfere with (DHFR SHMT d. TMP synthase … ) cycle 5 -fluorouracil is specific to thymidylate synthase 05/02/2019 Nucleotide Catabolism; Signaling p. 16 of 63

DHFR as a drug target n n n Human DHFR inhibition: kill cancer cells faster than healthy cells Eukaryotic DHFR also catalyzes folate dihydrofolate Prokaryotic DHFR doesn’t; n n DHF derived by another mechanism in bacteria So there antibacterials based on the structural differences between prokaryotic and eukaryotic DHFR 05/02/2019 Nucleotide Catabolism; Signaling p. 17 of 63

Special case: protozoan enzyme n Bifunctional enzyme: n n Thymidylate synthase Dihydrofolate reductase Plasmodium falciparum DHFR-TS EC 1. 5. 1. 3, 2. 1. 1. 45 104 k. Da dimer PDB 1 J 3 K, 2. 1Å Nucleotide Catabolism; Signaling p. 18 of 63 Presumably some entropic advantage Maybe electrostatics too, allowing the negative charges on DHF to tunnel through; but cf. Atreya et al (2003) J. Biol. Chem. 278: 28901. 05/02/2019

Recovery pathway to d. TMP n n Deoxythymidine can be phosphorylated by thymidine kinase: Herpes simplex deoxythymidine + ATP thymidine kinase 73 k. Da monomer d. TMP + ADP EC 2. 7. 1. 21 Labeled thymidine is convenient for PDB 1 E 2 K monitoring intracellular synthesis of DNA because thymidine enters cells easily 05/02/2019 Nucleotide Catabolism; Signaling p. 19 of 63

i. Clicker question #3 3. Thymidylate synthase is particularly significant because n n 05/02/2019 (a) Excess thymidine is toxic (b) We cannot obtain thymidine from the diet (c) [d. TTP] < [other nucleoside triphosphates] (d) none of the above. Nucleotide Catabolism; Signaling p. 20 of 63

Fates of polynucleotides n n Poly- and oligonucleotides hydrolyzed to mononucleotides via nucleases Mononucleotides are dephosphorylated via nucleotidases and phosphatases Resulting nucleosides are deglycosylated Resulting bases are sent either into salvage pathways or get degraded and excreted 05/02/2019 Nucleotide Catabolism; Signaling p. 21 of 63

Salvage pathways n n n Nucleotides can be fully broken down and excreted, but often they get recycled—especially purines Typically the free base reacts with PRPP to form x. MP and PPi, and the x. MP is used; enzymes are phosphoribosyltransferases Deficiencies in salvage pathways often result in severe problems 05/02/2019 Nucleotide Catabolism; Signaling p. 22 of 63

Salvage pathways: Why? n We can describe them, and we will: but why do they matter so much? n n n They provide energy savings relative to de novo synthesis (think of all the ATP we used in making IMP!) Considerable medical significance to interference with these pathways Intracellular nucleic acid bases are usually recycled; dietary bases are usually broken down and excess nitrogen excreted 05/02/2019 Nucleotide Catabolism; Signaling p. 23 of 63

Orotate phosphoribosyl transferase n n n Principal salvage enzyme for pyrimidines Orotate + PRPP OMP + PPi OMP can then reenter UMP synthetic pathway (decarboxylation to UMP, then form UDP and CDP) 05/02/2019 Nucleotide Catabolism; Signaling Yeast OPT 50 k. Da dimer EC 2. 4. 2. 10 PDB 2 PS 1, 1. 75Å p. 24 of 63

OPT, continued n Same enzyme can act on other pyrimidines to make nucleotides: Pyr + PRPP Pyr. MP + PPi 05/02/2019 Nucleotide Catabolism; Signaling p. 25 of 63

Pyrimidine interconversions n n n All phosphorylations & dephosphorylations can and do happen UTP can be aminated to CTP CDP and UDP can be reduced to d. CDP and d. UDP d. CMP can deaminate to d. UMP Cytidine can be converted to uridine d. UMP can be methylated to d. TMP 05/02/2019 Nucleotide Catabolism; Signaling p. 26 of 63
CMP and cytidine n n CMP’s phosphate can be hydrolyzed off That’s followed by deamination of cytidine to make uridine n n Catalyzed by cytidine deaminase Another sandwich protein 05/02/2019 Nucleotide Catabolism; Signaling Mouse cytidine deaminase 66 k. Da tetramer EC 3. 5. 4. 5 PDB 2 FR 5, 1. 48Å p. 27 of 63

Uracil to acetyl Co. A; thymine to succinyl Co. A n n Reduced to dihydrouracil and dihydrothymine Thermus Hydrated and ring-opened to Dihydro-pyrimidinase ureidopropionate or ureidoisobutyrate EC 3. 5. 2. 2 Eliminate bicarbonate and ammonium to 302 k. Da hexamer PDB 1 GKP, 1. 29Å yield -alanine or -aminoisobutyrate Several reactions from there to acetyl Co. A and succinyl Co. A 05/02/2019 Nucleotide Catabolism; Signaling p. 28 of 63
Hydrolysis of U, d. U & d. T n Glycosidic bond in uridine or thymidine is hydrolyzed by phosphate: n n Uridine + Pi -D-ribose-1 -P + uracil Enyzme is uridine phosphorylase Similar enzyme handles deoxyuridine Similar reaction using thymidine phosphorylase yields thymine + -Ddeoxyribose-1 -P 05/02/2019 Nucleotide Catabolism; Signaling E. coli Uridine phosphorylase EC 2. 4. 2. 3 167 k. Da hexamer Dimer shown PDB 1 RXY, 1. 7Å p. 29 of 63

Purine salvage n n n Two phosphoribosyl transferases convert adenine, guanine, and hypoxanthine to AMP, GMP, and IMP Adenine phosphoribosyl transferase is Toxoplasma gondii HGPRT specific EC 2. 4. 2. 8 HGPRT accepts both hypoxanthine 102 k. Da tetramer and guanine dimer shown PDB 1 FSG, 1. 05Å 05/02/2019 Nucleotide Catabolism; Signaling p. 30 of 63

Purine Interconnections n n n n All phosphorylations and dephosphorylations can and do occur ADP and GDP can be reduced to d. ADP and d. GDP AMP can deaminated to IMP (new) IMP can be aminated to AMP IMP can oxidized to XMP can be aminated to GMP Guanine, adenine phosphoribosylated to GMP and AMP 05/02/2019 Nucleotide Catabolism; Signaling p. 31 of 63
Purine catabolism n n Uric acid n n Nucleoside or deoxynucleoside + phosphate base + (D)-ribose 1 -P Hypoxanthine and guanine both lead to uric acid as a product Uric acid is final excreted nitrogenous compound in primates, birds, some reptiles Other organisms catabolize it further 05/02/2019 Nucleotide Catabolism; Signaling p. 32 of 63
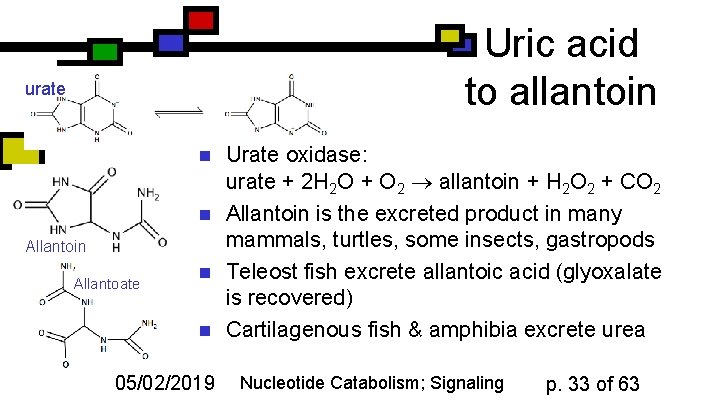
Uric acid to allantoin urate n n Allantoin Allantoate n n 05/02/2019 Urate oxidase: urate + 2 H 2 O + O 2 allantoin + H 2 O 2 + CO 2 Allantoin is the excreted product in many mammals, turtles, some insects, gastropods Teleost fish excrete allantoic acid (glyoxalate is recovered) Cartilagenous fish & amphibia excrete urea Nucleotide Catabolism; Signaling p. 33 of 63
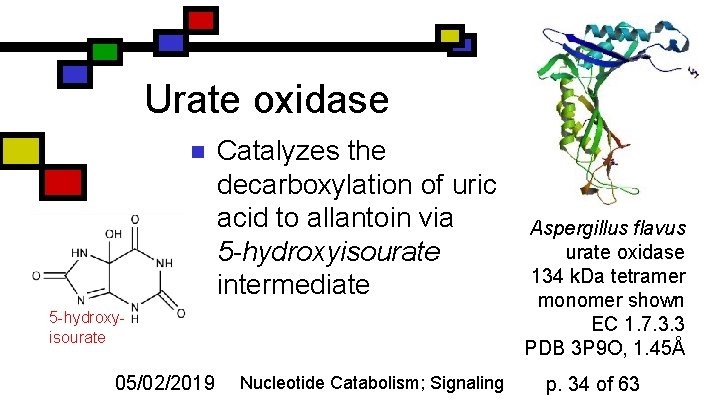
Urate oxidase n Catalyzes the decarboxylation of uric acid to allantoin via 5 -hydroxyisourate intermediate 5 -hydroxyisourate 05/02/2019 Nucleotide Catabolism; Signaling Aspergillus flavus urate oxidase 134 k. Da tetramer monomer shown EC 1. 7. 3. 3 PDB 3 P 9 O, 1. 45Å p. 34 of 63

Lesch-Nyhan syndrome n n Michael Lesch William Nyhan Complete lack of hypoxanthine-guanine phosphoribosyl transferase So hypoxanthine and guanine are degraded to uric acid rather than being built back up into IMP & GMP Leads to dangerous buildup of uric acid in nervous tissue Neurological effects are severe and poorly understood 05/02/2019 Nucleotide Catabolism; Signaling p. 35 of 63

Gout n n Sodium urate Accumulation of sodium urate and uric acid, both of which Benjamin Franklin are only moderately soluble (celebrated Arises from inadequate gout sufferer) (~10%) functionality of HGPRT, so that urate accumulates in peripheral tissues, particularly the feet Sodium urate crystals accumulating 05/02/2019 Nucleotide Catabolism; Signaling p. 36 of 63
i. Clicker question #4 4. Gout could be regarded as 05/02/2019 n n n (a) A disease only suffered by the rich (b) a mild version of Lesch-Nyhan syndrome (c) a severe version of Lesch. Nyhan syndrome (d) an entirely preventable disease n Nucleotide Catabolism; Signaling p. 37 of 63

Transducing signals n n Plasma membranes contain proteins called receptors that allow the cell to respond to chemical stimuli that can’t cross the membrane Example: Bacteria can detect chemicals. If something useful comes along, a signal is passed from the receptor to the flagella, enabling the bacterium to swim toward the source 05/02/2019 Nucleotide Catabolism; Signaling p. 38 of 63
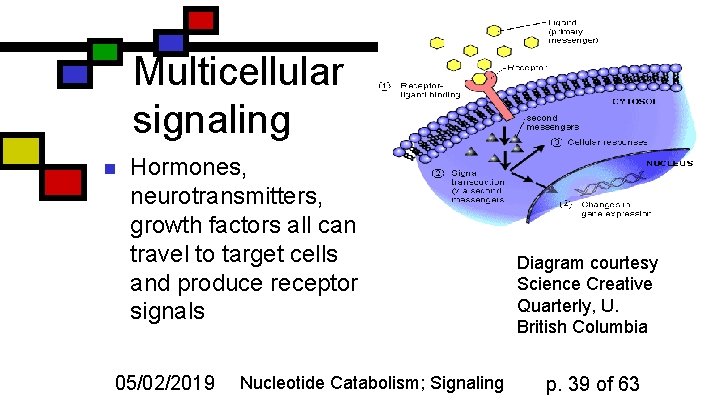
Multicellular signaling n Hormones, neurotransmitters, growth factors all can travel to target cells and produce receptor signals 05/02/2019 Nucleotide Catabolism; Signaling Diagram courtesy Science Creative Quarterly, U. British Columbia p. 39 of 63
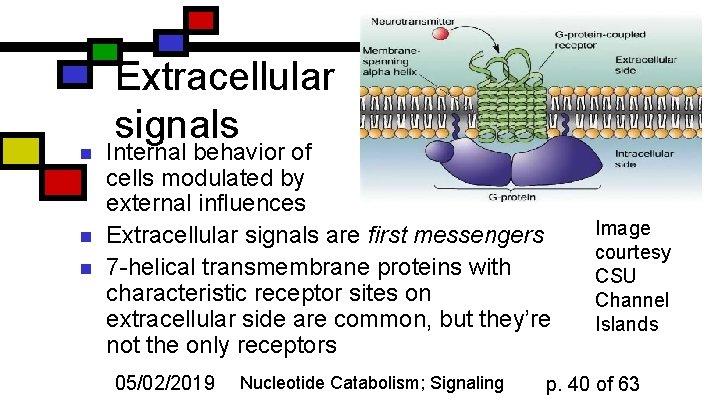
n n n Extracellular signals Internal behavior of cells modulated by external influences Extracellular signals are first messengers 7 -helical transmembrane proteins with characteristic receptor sites on extracellular side are common, but they’re not the only receptors 05/02/2019 Nucleotide Catabolism; Signaling Image courtesy CSU Channel Islands p. 40 of 63

G-protein coupled receptors n n n Not all of these transmembrane receptors interact with heterotrimeric G-proteins, but a huge percentage do These receptors are known as G-protein coupled receptors, or GPCRs Studies of GPCRs earned Robert Lefkowitz and Brian Kobilka the 2012 chemistry Nobel Prize 05/02/2019 Nucleotide Catabolism; Signaling p. 41 of 63

Internal results of signals n n Intracellular: heterotrimeric G-proteins are the transducers: they receive signal from receptor, hydrolyze GTP, and emit small molecules called second messengers Second messengers diffuse to target organelle or portion of cytoplasm Big picture: many signals, many receptors, relatively few second messengers Often there is amplification involved 05/02/2019 Nucleotide Catabolism; Signaling p. 42 of 63

Roles of these systems n n n n Response to sensory stimuli Response to hormones Response to growth factors Response to some neurotransmitters Metabolite transport Immune response This stuff gets complicated, because the kinds of signals are so varied! 05/02/2019 Nucleotide Catabolism; Signaling p. 43 of 63

G proteins n n n Transducers of external signals into the inside of the cell These are GTPases (GTP GDP + Pi) GTP-bound protein transduces signals; GDP-bound protein doesn’t Rat Gi, α subunit EC 3. 6. 5. 1 41 k. Da monomer PDB 1 SVK, 2Å 05/02/2019 p. 44 of 63 Nucleotide Catabolism; Signaling

Subunits of the G protein n Heterotrimeric proteins; association of and g subunits with subunit is disrupted by complexation with ligand-receptor complex, allowing departure of GDP & binding of GTP 05/02/2019 Nucleotide Catabolism; Signaling Alfred Gilman Martin Rodbell p. 45 of 63
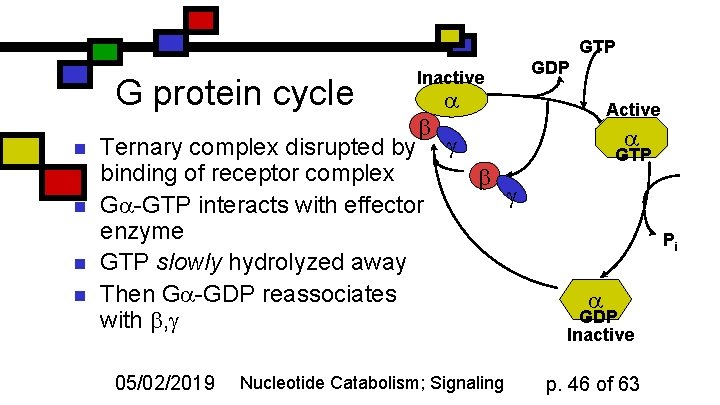
GTP G protein cycle n n Inactive Ternary complex disrupted by g binding of receptor complex g G -GTP interacts with effector enzyme GTP slowly hydrolyzed away Then G -GDP reassociates with , g 05/02/2019 GDP Nucleotide Catabolism; Signaling Active GTP Pi GDP Inactive p. 46 of 63
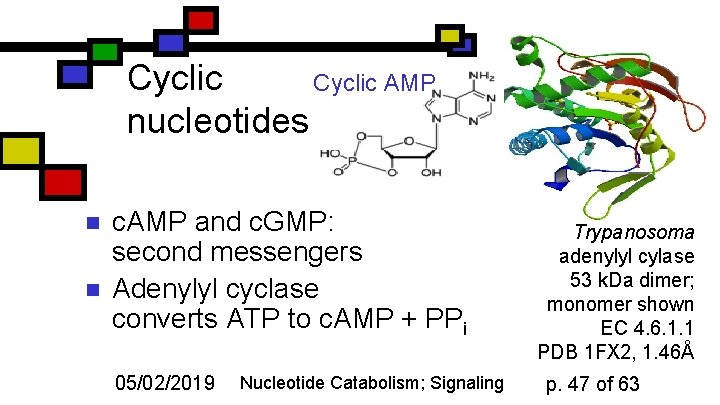
Cyclic AMP nucleotides n n c. AMP and c. GMP: second messengers Adenylyl cyclase converts ATP to c. AMP + PPi 05/02/2019 Nucleotide Catabolism; Signaling Trypanosoma adenylyl cylase 53 k. Da dimer; monomer shown EC 4. 6. 1. 1 PDB 1 FX 2, 1. 46Å p. 47 of 63

Adenylyl cyclase n n n Integral membrane enzyme; active site faces cytosol c. AMP diffuses from membrane surface through cytosol, activates protein kinase A PKA phosphorylates ser, thr in target enzymes; action reversed by specific phosphatases 05/02/2019 Nucleotide Catabolism; Signaling p. 48 of 63

Modulators of c. AMP n n Ordinarily c. AMP is shortlived (15 min): it is hydrolyzed via c. AMP phosphodiesterase, which catalyzes c. AMP + H 2 O → AMP Caffeine, theophylline inhibit c. AMP phosphodiesterase, prolonging c. AMP’s stimulatory effects on protein kinase A 05/02/2019 Nucleotide Catabolism; Signaling p. 49 of 63

Hormonal modulators of c. AMP n n Hormones that bind to stimulatory receptors activate adenylyl cyclase, raising c. AMP levels Hormones that bind to inhibitory receptors inhibit adenylyl cyclase activity via receptor interaction with the transducer Gi. 05/02/2019 Nucleotide Catabolism; Signaling p. 50 of 63
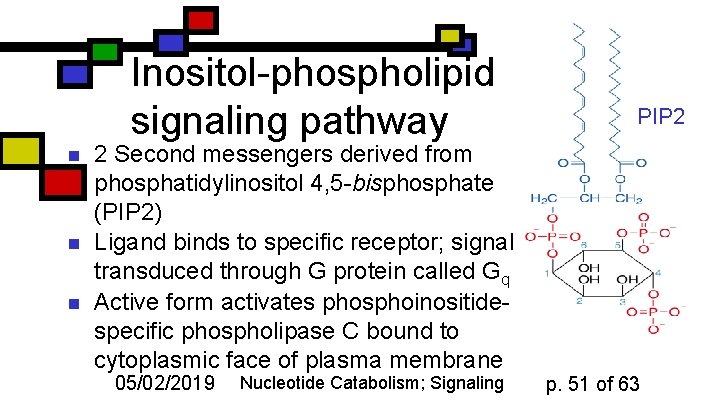
Inositol-phospholipid signaling pathway n n n PIP 2 2 Second messengers derived from phosphatidylinositol 4, 5 -bisphosphate (PIP 2) Ligand binds to specific receptor; signal transduced through G protein called Gq Active form activates phosphoinositidespecific phospholipase C bound to cytoplasmic face of plasma membrane 05/02/2019 Nucleotide Catabolism; Signaling p. 51 of 63
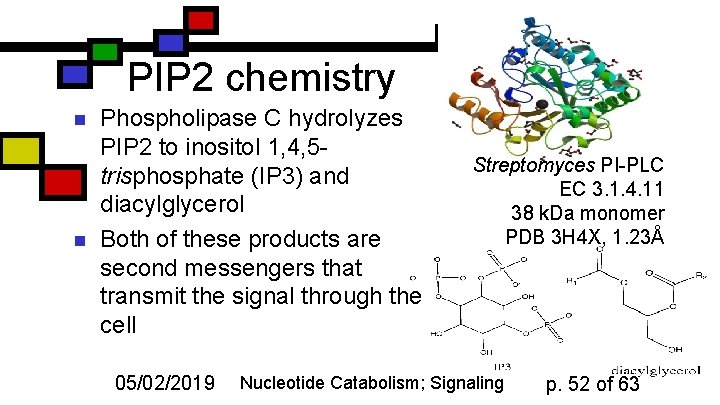
PIP 2 chemistry n n Phospholipase C hydrolyzes PIP 2 to inositol 1, 4, 5 trisphosphate (IP 3) and diacylglycerol Both of these products are second messengers that transmit the signal through the cell 05/02/2019 Streptomyces PI-PLC EC 3. 1. 4. 11 38 k. Da monomer PDB 3 H 4 X, 1. 23Å Nucleotide Catabolism; Signaling p. 52 of 63
IP 3 and calcium n n IP 3 diffuses through cytosol and binds to a calcium channel in the membrane of the endoplasmic reticulum The calcium channel opens, releasing Ca 2+ from lumen of ER into cytosol 05/02/2019 Nucleotide Catabolism; Signaling p. 53 of 63
Calcium as a 2 nd messenger n Ca 2+ is a short-lived 2 nd messenger too: it activates Ca 2+-dependent protein kinases that catalyze phosphorylation of certain proteins 05/02/2019 Nucleotide Catabolism; Signaling p. 54 of 63
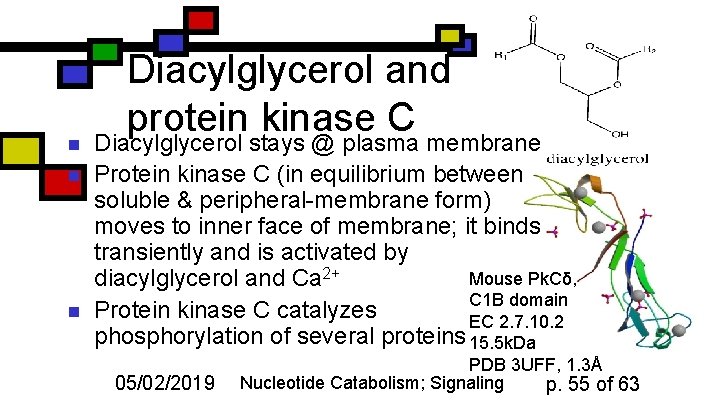
n n n Diacylglycerol and protein kinase C Diacylglycerol stays @ plasma membrane Protein kinase C (in equilibrium between soluble & peripheral-membrane form) moves to inner face of membrane; it binds transiently and is activated by Mouse Pk. Cδ, diacylglycerol and Ca 2+ C 1 B domain Protein kinase C catalyzes EC 2. 7. 10. 2 phosphorylation of several proteins 15. 5 k. Da 05/02/2019 PDB 3 UFF, 1. 3Å Nucleotide Catabolism; Signaling p. 55 of 63

Control of inositolphospholipid pathway n Figure courtesy Motifolio. com After GTP hydrolysis, Gq is inactive so it no longer stimulates phospholipase C 05/02/2019 Nucleotide Catabolism; Signaling p. 56 of 63
Activities of 2 nd messengers are transient n n n IP 3 rapidly hydrolyzed to other things Diacylglycerol is phosphorylated to form phosphatidate Calcium gets used or sequestered 05/02/2019 Nucleotide Catabolism; Signaling p. 57 of 63

Sphingolipids give rise to 2 nd messengers n n Some signals activate hydrolases that convert sphingomyelin to sphingosine, sphingosine-1 -P, and ceramides Sphingosine inhibits Protein Kinase C 05/02/2019 Nucleotide Catabolism; Signaling p. 58 of 63

Other second messengers arising from sphingolipids n n Ceramides activate a protein kinase and a protein phosphatase Sphingosine-1 -P can activate Phospholipase D, which catalyzes hydrolysis of phosphatidylcholine; products are 2 nd messengers 05/02/2019 Nucleotide Catabolism; Signaling p. 59 of 63
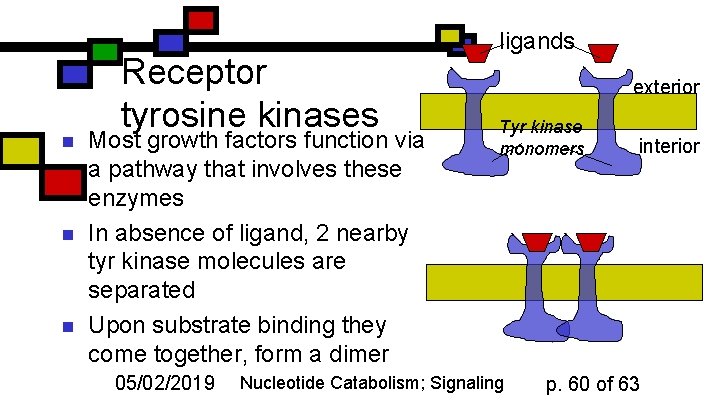
n n n Receptor tyrosine kinases Most growth factors function via a pathway that involves these enzymes In absence of ligand, 2 nearby tyr kinase molecules are separated Upon substrate binding they come together, form a dimer 05/02/2019 ligands exterior Tyr kinase monomers Nucleotide Catabolism; Signaling interior p. 60 of 63
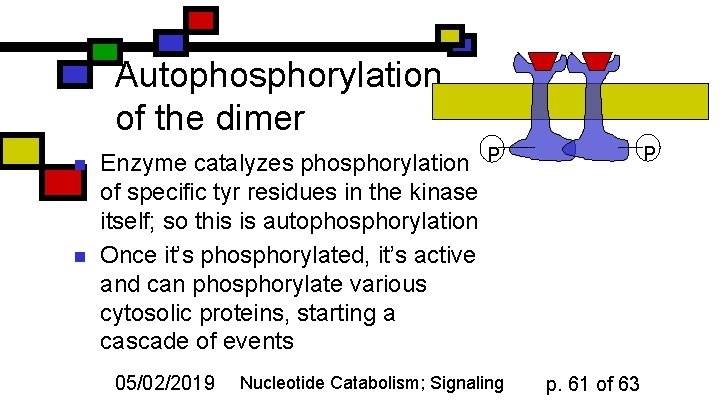
Autophosphorylation of the dimer n n P Enzyme catalyzes phosphorylation P of specific tyr residues in the kinase itself; so this is autophosphorylation Once it’s phosphorylated, it’s active and can phosphorylate various cytosolic proteins, starting a cascade of events 05/02/2019 Nucleotide Catabolism; Signaling p. 61 of 63
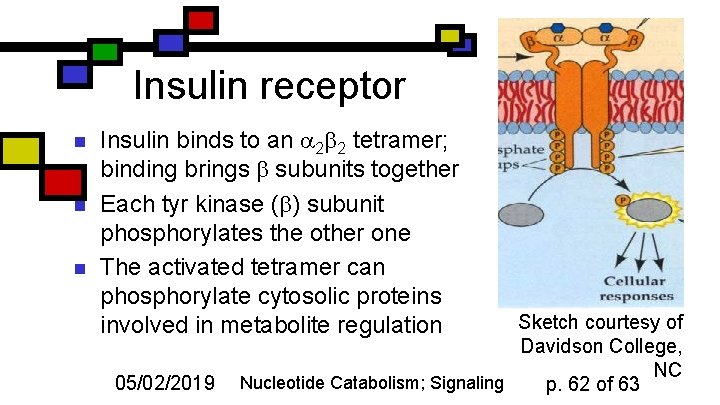
Insulin receptor n n n Insulin binds to an 2 2 tetramer; binding brings subunits together Each tyr kinase ( ) subunit phosphorylates the other one The activated tetramer can phosphorylate cytosolic proteins involved in metabolite regulation 05/02/2019 Sketch courtesy of Davidson College, NC Nucleotide Catabolism; Signaling p. 62 of 63

i. Clicker question #5 5. Which of the following have we not described as a second messenger? 05/02/2019 n n n (a) cyclic AMP (b) Na+ (c) sphingosine-1 -phosphate (d) Ca 2+ (e) all four of these are second messengers Nucleotide Catabolism; Signaling p. 63 of 63
- Slides: 63