NUCLEIC ACIDS OBJECTIVES Identify recognize nucleic acid u
NUCLEIC ACIDS

OBJECTIVES Identify/ recognize nucleic acid u Components in nucleic acid – monosaccharide, nucleobases, phosphoric acid u Differentiate - between 2 types of nucleic acids, DNA and RNA - between nucleotide and nucleoside u Definition – nucleotide, nucleoside, DNA and RNA u
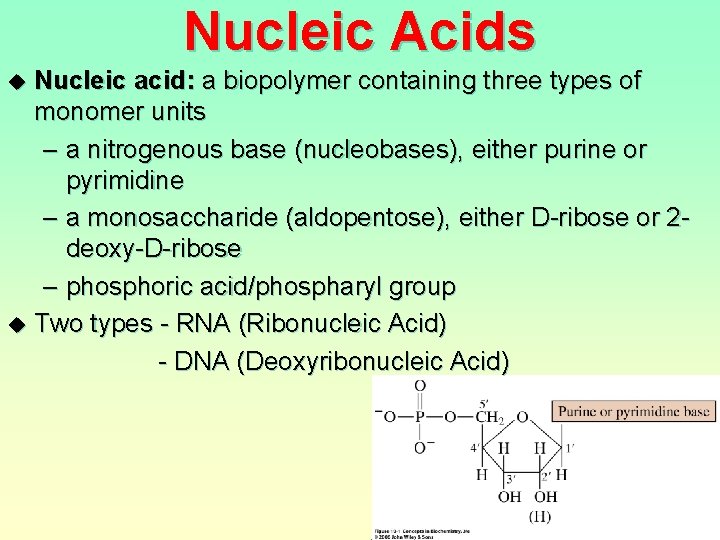
Nucleic Acids u u Nucleic acid: a biopolymer containing three types of monomer units – a nitrogenous base (nucleobases), either purine or pyrimidine – a monosaccharide (aldopentose), either D-ribose or 2 deoxy-D-ribose – phosphoric acid/phospharyl group Two types - RNA (Ribonucleic Acid) - DNA (Deoxyribonucleic Acid)
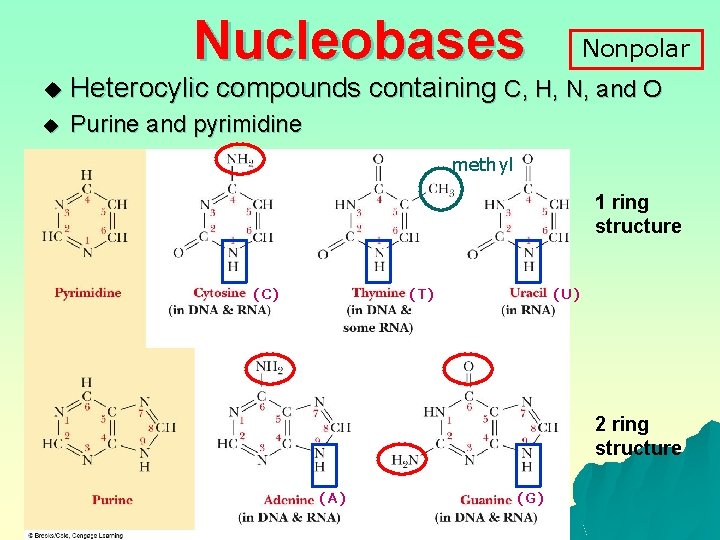
Nucleobases Nonpolar u Heterocylic compounds containing C, H, N, and O u Purine and pyrimidine methyl 1 ring structure (C) (T) (U) 2 ring structure (A) (G)
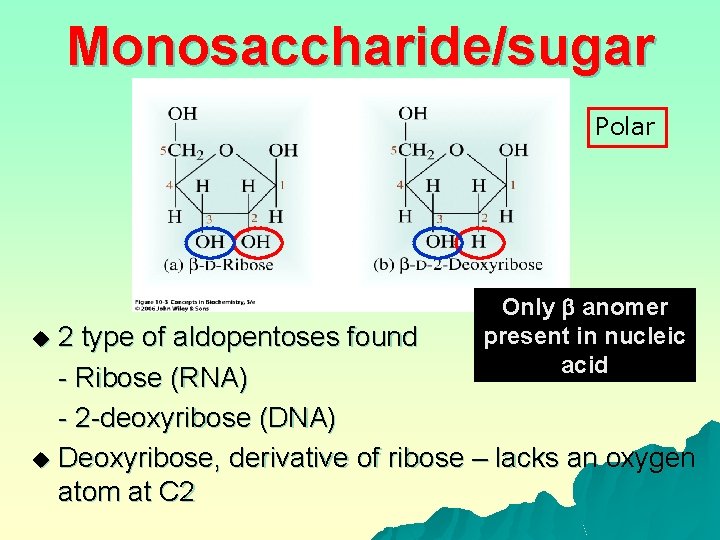
Monosaccharide/sugar Polar Only anomer present in nucleic acid 2 type of aldopentoses found - Ribose (RNA) - 2 -deoxyribose (DNA) u Deoxyribose, derivative of ribose – lacks an oxygen atom at C 2 u
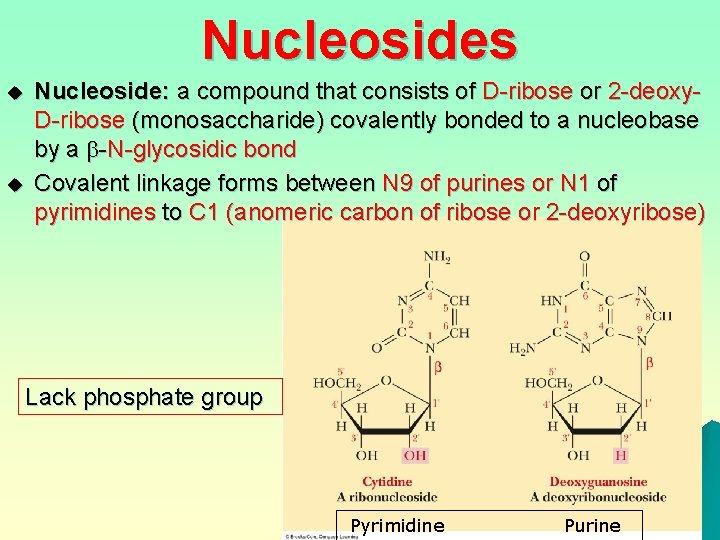
Nucleosides u u Nucleoside: a compound that consists of D-ribose or 2 -deoxy. D-ribose (monosaccharide) covalently bonded to a nucleobase by a -N-glycosidic bond Covalent linkage forms between N 9 of purines or N 1 of pyrimidines to C 1 (anomeric carbon of ribose or 2 -deoxyribose) Lack phosphate group Pyrimidine Purine

Nucleotides u u Nucleotide: a nucleoside in which a molecule of phosphoric acid/phosphoryl group is esterified with an -OH of the monosaccharide, at the 5’-OH As constituents of cofactors, Coenzyme A (Co. A), flavin adenine dinucleotide (FAD) & nicotinamide adenine dinucleotides (NAD) Nucleobase, aldopentose sugar and phosphoryl group Phosphoric acid - polar
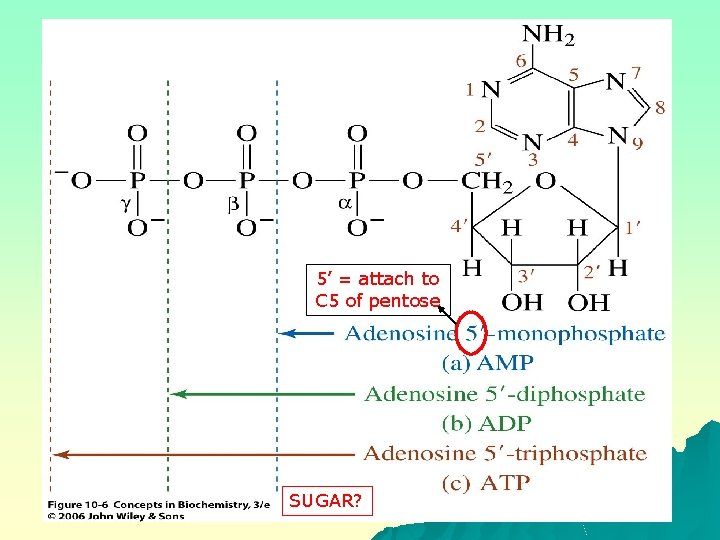
5’ = attach to C 5 of pentose SUGAR?

NOMENCLATURE of Nucleotide Based on the nucleoside, plus the phosphate group

Nucleotide Sequence u Gene: Sequence of nucleotides that encodes a polypeptide, eventually forming a functional protein u Gene: a discrete unit of hereditary information consisting of a specific nucleotide sequence in DNA (RNA in some viruses) u The nucleotide sequence is depending on the bases (nucleobases) present

Nucleic Acid: DNA Nucleoside 1. Bases = ATGC 2. Aldopentose = Ribose 3. Phosphoryl group Naming of nucleotide: if Base adenine Deoxyadenosine 5’ monophosphate Biopolymer, nucleotide as monomer RNA 1. Bases = AUGC 2. Aldopentose = Deoxyribose 3. Phosphoryl group Naming of nucleotide: if Base adenine Adenosine 5’monophosphate

Nucleic Acid - DNA and RNA u DNA stands for deoxyribonucleic acid. It is the genetic code molecule for most organisms. u RNA stands for ribonucleic acid. RNA molecules are involved in converting the genetic information in DNA into proteins. In retroviruses, RNA is the genetic material. NUCLEIC ACIDS ARE POLYMERS OF NUCLEOTIDES

DNA u u DNA and RNA are polymers whose monomer units are nucleotides = polynucleotides Polynucleotide = DNA and RNA Hydrolysis – break bond Condensation – form bond Deoxyribonucleic acids, DNA: a biopolymer that consists of a backbone of alternating units of 2 deoxy-D-ribose and phosphoryl group – the 3’-OH of one nucleotide is joined to the 5’ P of the next nucleotide by a phosphodiester bond 3’ 5’ -phosphodiester bond
DNA structure u Levels of structure – 1° structure: the order of bases on the polynucleotide sequence; the order of bases specifies the genetic code – 2° structure: the three-dimensional conformation of the polynucleotide backbone = double helix structure – 3° structure: supercoiling – 4° structure: interaction between DNA and proteins
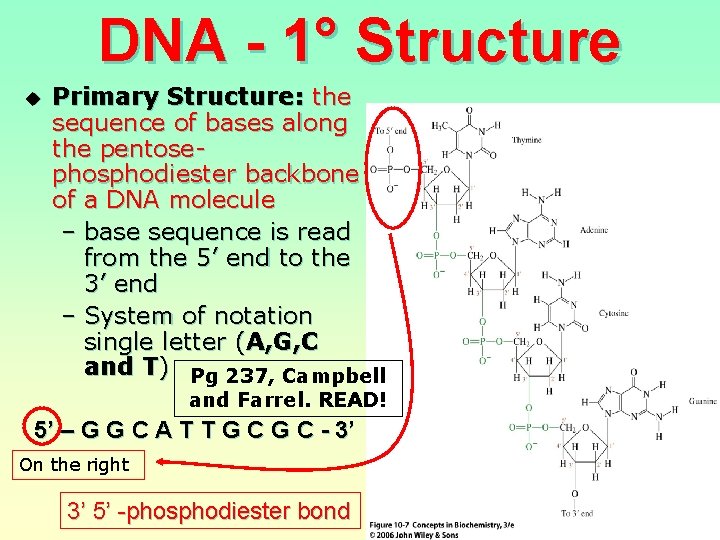
DNA - 1° Structure u Primary Structure: the sequence of bases along the pentosephosphodiester backbone of a DNA molecule – base sequence is read from the 5’ end to the 3’ end – System of notation single letter (A, G, C and T) Pg 237, Campbell and Farrel. READ! 5’ – G G C A T T G C - 3’ On the right 3’ 5’ -phosphodiester bond
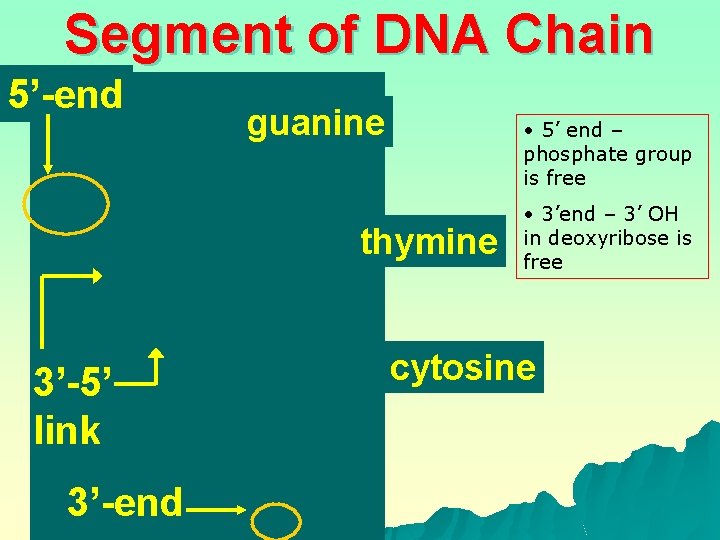
Segment of DNA Chain 5’-end guanine • 5’ end – phosphate group is free thymine 3’-5’ link 3’-end • 3’end – 3’ OH in deoxyribose is free cytosine
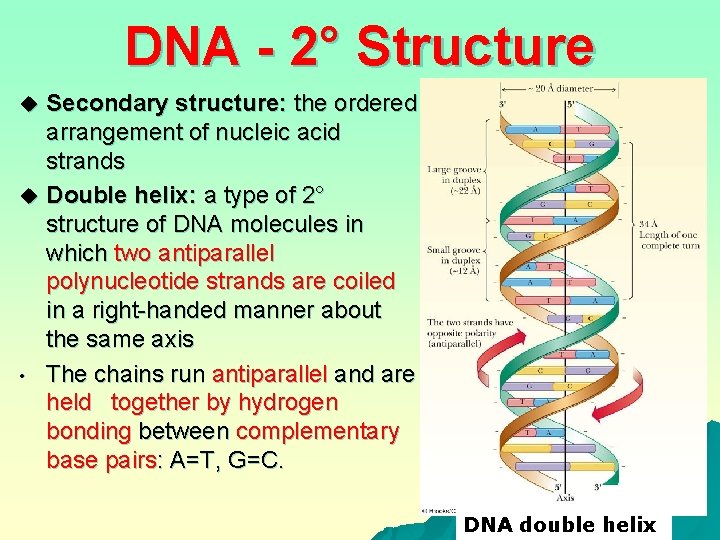
DNA - 2° Structure Secondary structure: the ordered arrangement of nucleic acid strands u Double helix: a type of 2° structure of DNA molecules in which two antiparallel polynucleotide strands are coiled in a right-handed manner about the same axis • The chains run antiparallel and are held together by hydrogen bonding between complementary base pairs: A=T, G=C. u DNA double helix
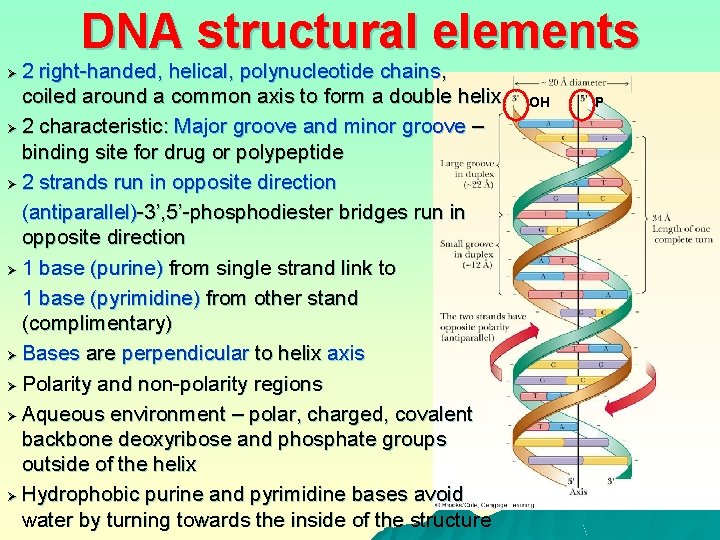
DNA structural elements 2 right-handed, helical, polynucleotide chains, coiled around a common axis to form a double helix Ø 2 characteristic: Major groove and minor groove – binding site for drug or polypeptide Ø 2 strands run in opposite direction (antiparallel)-3’, 5’-phosphodiester bridges run in opposite direction Ø 1 base (purine) from single strand link to 1 base (pyrimidine) from other stand (complimentary) Ø Bases are perpendicular to helix axis Ø Polarity and non-polarity regions Ø Aqueous environment – polar, charged, covalent backbone deoxyribose and phosphate groups outside of the helix Ø Hydrophobic purine and pyrimidine bases avoid water by turning towards the inside of the structure Ø OH P

T-A Base Pairing u Base pairing is complementary: A=T, G C u A major factor stabilizing the double helix is base pairing by hydrogen bonding between T-A and between C-G u T-A base pair comprised of 2 hydrogen bonds Complementary base pairing
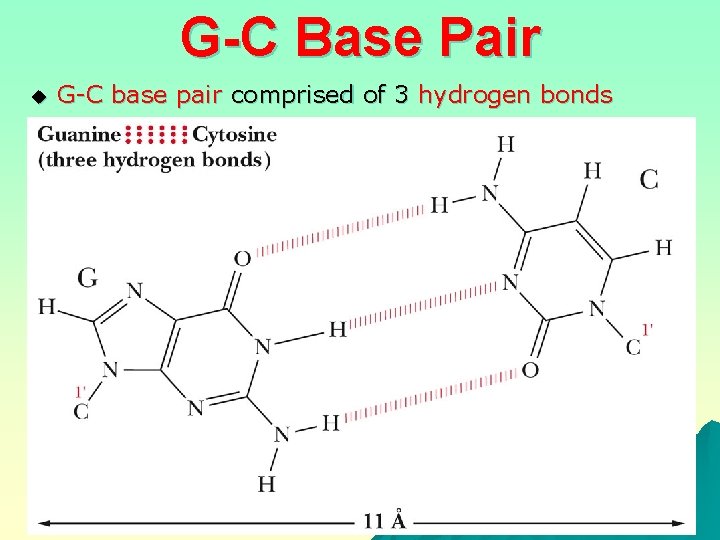
G-C Base Pair u G-C base pair comprised of 3 hydrogen bonds

Forms of DNA B-DNA – considered the physiological form – a right-handed helix, inside diameter 11Å – 10 base pairs per turn (34Å) of the helix u A-DNA – a right-handed helix, but thicker than B-DNA – 11 base pairs per turn of the helix – has not been found in vivo u Z-DNA • a left-handed double helix • may play a role in gene expression u • Z-DNA occurs in nature, usually consists of alternating purine-pyrimidine bases • Methylated cytosine found also in Z-DNA
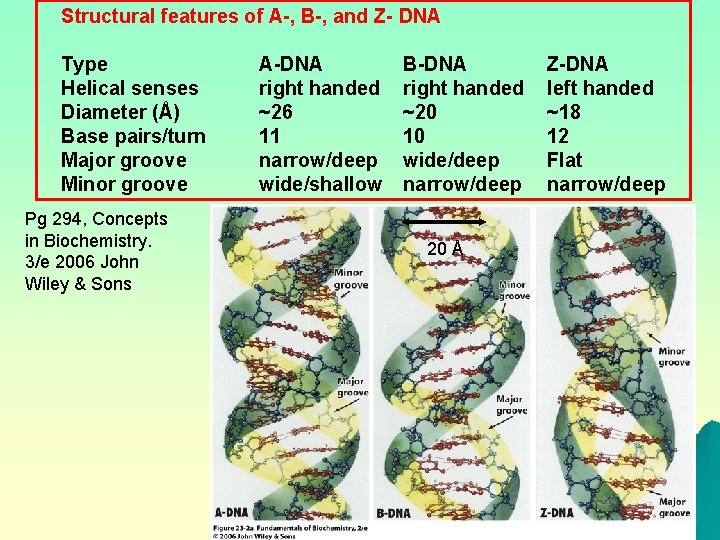
Structural features of A-, B-, and Z- DNA Type Helical senses Diameter (Å) Base pairs/turn Major groove Minor groove Pg 294, Concepts in Biochemistry. 3/e 2006 John Wiley & Sons A-DNA right handed ~26 11 narrow/deep wide/shallow B-DNA right handed ~20 10 wide/deep narrow/deep 20 Å Z-DNA left handed ~18 12 Flat narrow/deep
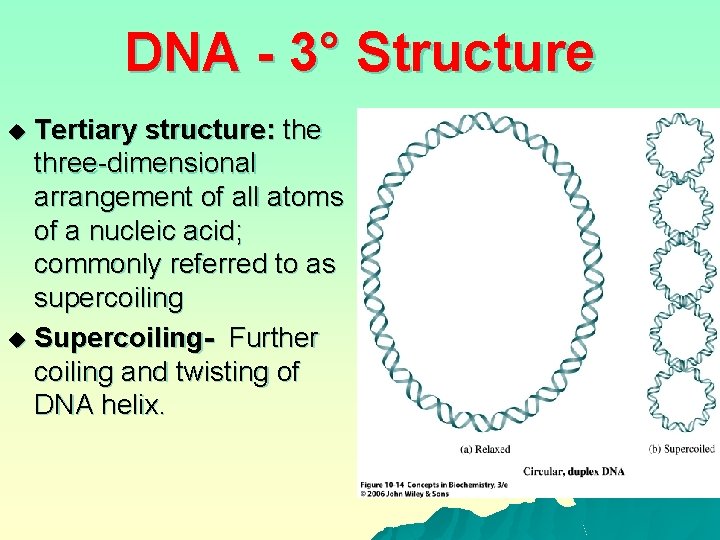
DNA - 3° Structure Tertiary structure: the three-dimensional arrangement of all atoms of a nucleic acid; commonly referred to as supercoiling u Supercoiling- Further coiling and twisting of DNA helix. u

DNA can forms tertiary structure by twist into complex arrangement – supercoil u Circular DNA: a type of double-stranded DNA in which the 5’ and 3’ ends of each strand (2 polynucleotide chains) are joined by phosphodiester bonds u Can be found in microorganisms (bacteriophages, bacteria) u Circular twisted into supercoiled DNA - 3° Structure u Supercoil - results of extra twisting in the linear duplex form u
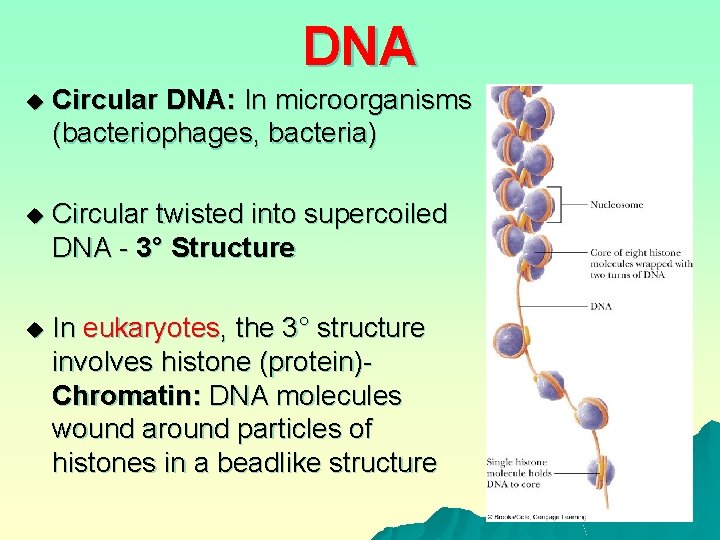
DNA u Circular DNA: In microorganisms (bacteriophages, bacteria) u Circular twisted into supercoiled DNA - 3° Structure u In eukaryotes, the 3° structure involves histone (protein)Chromatin: DNA molecules wound around particles of histones in a beadlike structure

PROPERTIES OF SUPERCOIL u Supercoiled is less stable than the relaxed form u Compact hence it more easily stored in the cell u Play a regulatory role in DNA replication
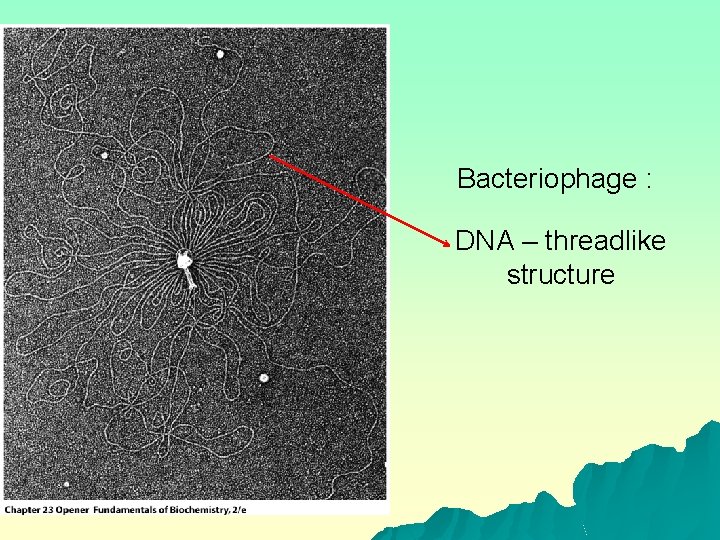
Bacteriophage : DNA – threadlike structure
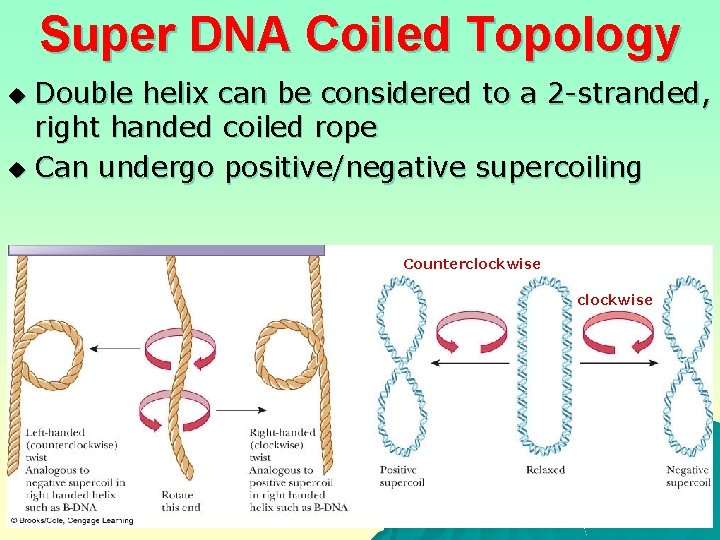
Super DNA Coiled Topology Double helix can be considered to a 2 -stranded, right handed coiled rope u Can undergo positive/negative supercoiling u Counterclockwise
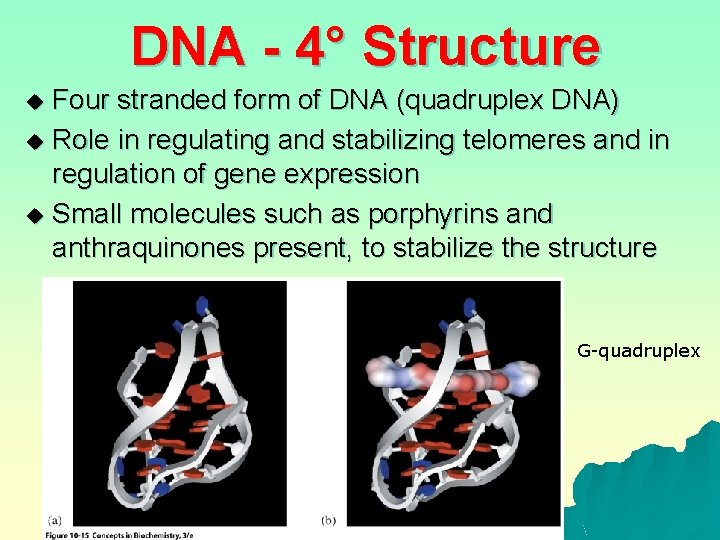
DNA - 4° Structure Four stranded form of DNA (quadruplex DNA) u Role in regulating and stabilizing telomeres and in regulation of gene expression u Small molecules such as porphyrins and anthraquinones present, to stabilize the structure u G-quadruplex

NUCLEIC ACIDS (2)

OBJECTIVES u Denaturation of DNA u Identify/ recognize RNA u Differentiate between DNA and RNA u Differentiate between m. RNA, t. RNA, r. RNA
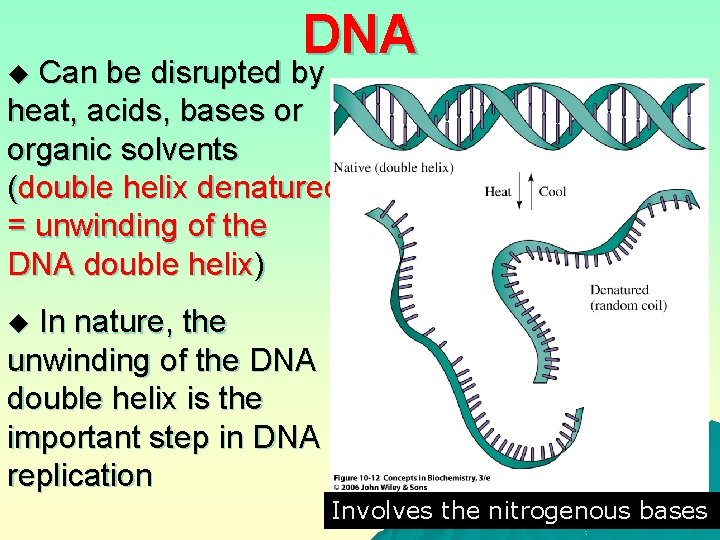
DNA Can be disrupted by heat, acids, bases or organic solvents (double helix denatured = unwinding of the DNA double helix) u In nature, the unwinding of the DNA double helix is the important step in DNA replication u Involves the nitrogenous bases

u Denaturation of DNA Denaturation: disruption of 2° structure – most commonly by heat denaturation (melting- the heat At the nitrogenous bases denaturation of DNA) – as strands separate, absorbance at 260 nm increases – increase is called hyperchromicitythe wavelength of absorption does not change but the amount of light absorbed increases – midpoint of transition (melting) curve = Tm – the higher the % G-C, the higher the Tm – Renaturation/annealing is possible on slow cooling v Principal use in PCR
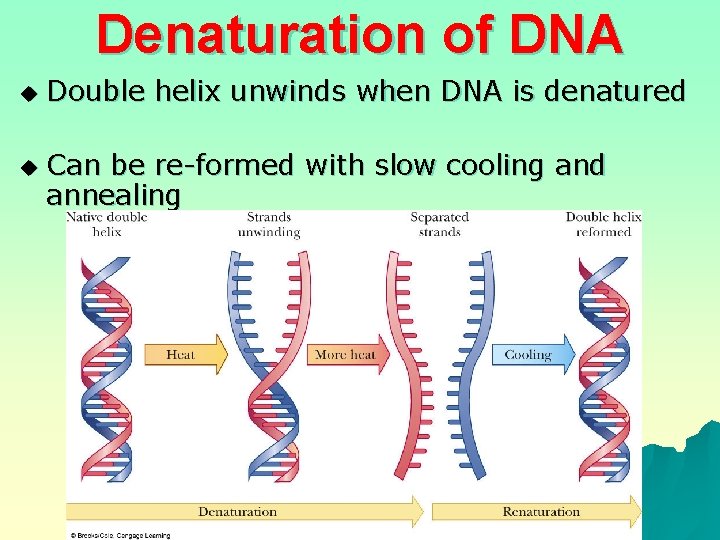
Denaturation of DNA u u Double helix unwinds when DNA is denatured Can be re-formed with slow cooling and annealing

RNA u consist of long, unbranched chains of nucleotides joined by phosphodiester bonds between the 3’-OH of one pentose and the 5’-P of the next nucleotide u the pentose unit is -D-ribose (it is 2 -deoxy-D-ribose in DNA)- the extra OH present in RNA makes this nucleotide more susceptible to hydrolysis than DNA. u the pyrimidine bases are uracil and cytosine (they are thymine and cytosine in DNA) u RNA is single stranded (DNA is double stranded) The bases sequences of all types of RNA are determined by that of DNA

RNA u RNA molecules are classified according to their structure and function
RNA structure u Levels of structure – 1° structure: the order of bases on the polynucleotide sequence; complementary to the DNA template – 2° structure: no specific 2° arrangements, but RNA is not completely lacking of regular structure – 3° structure: interaction between DNA and proteins
RNA- 1° Structure u Polymer of nucleotide u Involves single polynucleotide strand

RNA- 2° Structure u Loop back onto themselves to fold into conformation containing several different structural elements: 1 -hairpin turns 2 -right-handed double helixes 3 -internal loops All classes of RNA synthesized as single-stranded molecules 3’OH 5’P
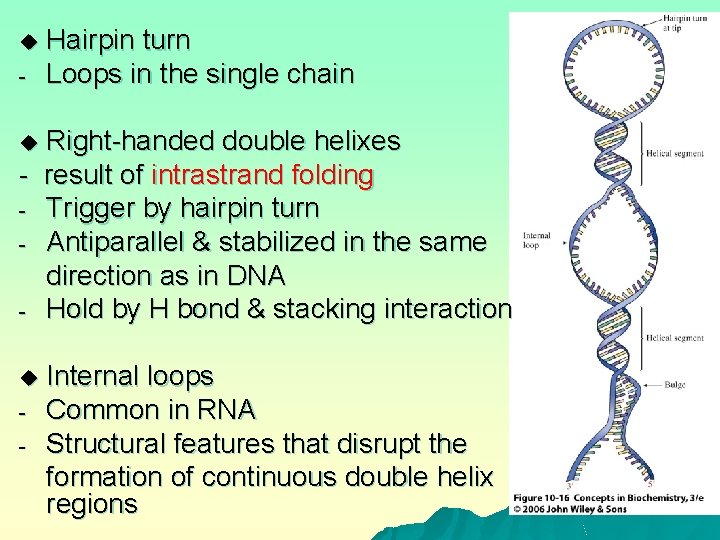
u - Hairpin turn Loops in the single chain Right-handed double helixes - result of intrastrand folding - Trigger by hairpin turn - Antiparallel & stabilized in the same direction as in DNA - Hold by H bond & stacking interaction u u - Internal loops Common in RNA Structural features that disrupt the formation of continuous double helix regions
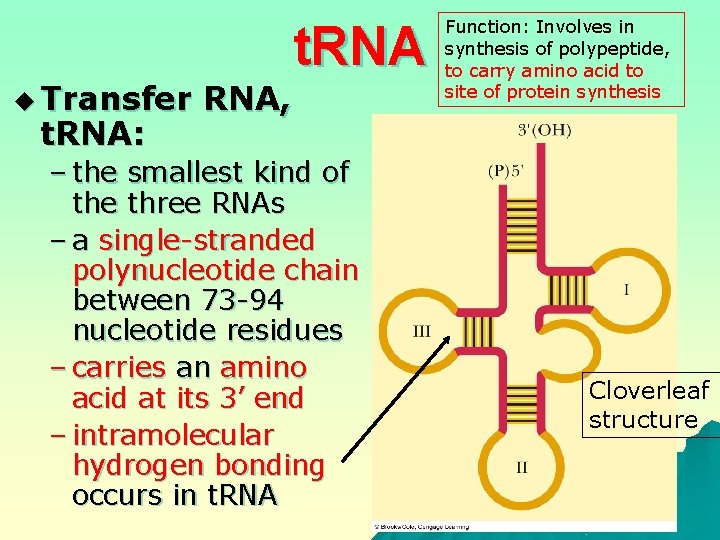
u Transfer t. RNA: RNA, t. RNA – the smallest kind of the three RNAs – a single-stranded polynucleotide chain between 73 -94 nucleotide residues – carries an amino acid at its 3’ end – intramolecular hydrogen bonding occurs in t. RNA Function: Involves in synthesis of polypeptide, to carry amino acid to site of protein synthesis Cloverleaf structure

t. RNA structure Smallest types of RNA u Highly structured u All t. RNAs contain between 74 and 93 nucleotides in a single chain u Structural features: hairpin turns, regions of double helix and loops (non-hydrogen bonded portions) u Carriers of specific amino acids used for protein synthesis u Reads the codon message on m. RNA and incorporates amino acid into the protein being synthesized u 20 amino acid – 20 t. RNA u
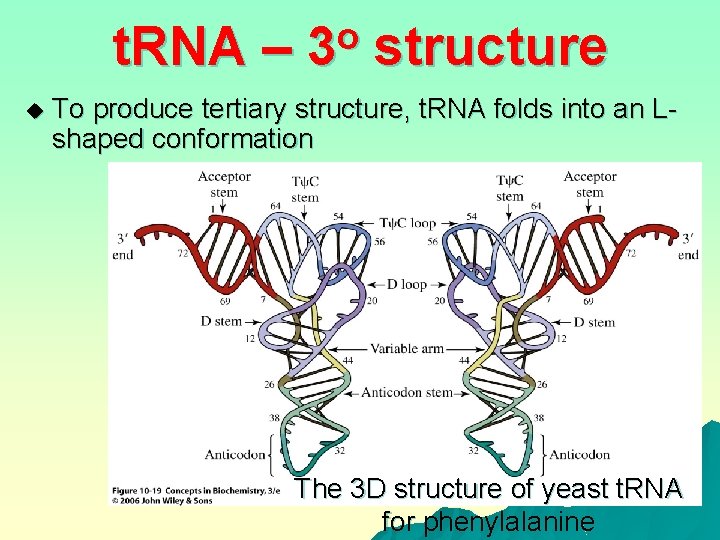
t. RNA – u o 3 structure To produce tertiary structure, t. RNA folds into an Lshaped conformation The 3 D structure of yeast t. RNA for phenylalanine

r. RNA u Ribosomal RNA, r. RNA: a ribonucleic acid found in ribosomes, the site of protein synthesis – only a few types of r. RNA exist in cells – ribosomes (protein-synthesizing organelles) consist of 60 to 65% r. RNA and 35 to 40% protein – in both prokaryotes and eukaryotes, ribosomes consist of two subunits, one larger than the other – analyzed by analytical ultracentrifugation – particles characterized by sedimentation coefficients, expressed in Svedberg units (S) – Sequencing of 16 S RNA (small subunit of bacteria r. RNA) - identification of bacteria
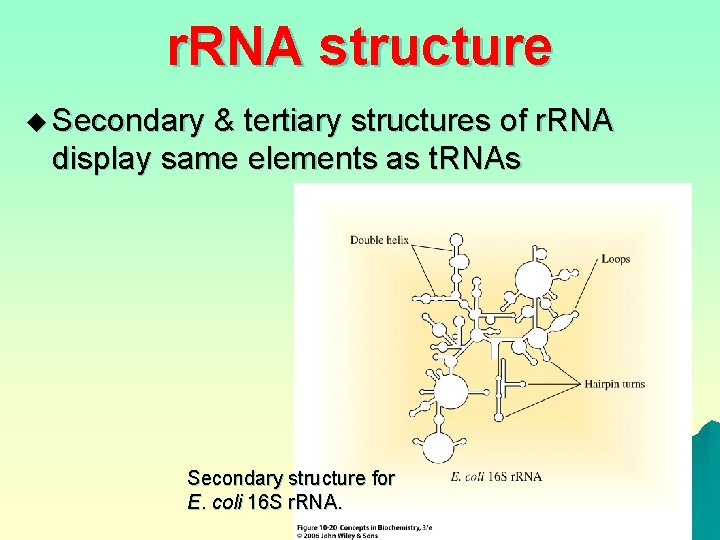
r. RNA structure u Secondary & tertiary structures of r. RNA display same elements as t. RNAs Secondary structure for E. coli 16 S r. RNA.

m. RNA u Messenger Structure: Linear polynucleotide strand RNA, m. RNA: a ribonucleic acid that carries coded genetic information from DNA to ribosomes for the synthesis of proteins – – present in cells in relatively small amounts and very short-lived (less abundant form of RNA) single stranded biosynthesis is directed by information encoded on DNA Synthesize from DNA, the nucleotide sequence in m. RNA is similar with the 5’-3’ strand of DNA, with the exception of U replacing T 5’-3’ DNA sequence is the same with RNA sequence (complementary to 3’-5’DNA) sequence
m. RNA structure Serves as a template for protein synthesis (Carries the transient message for protein synthesis from nuclear DNA to the ribosomes) u Move the information contained in DNA to the translation machinery u Each molecule carries the instruction for each gene (codes for one type of polypeptide product) u 5’ – G G C A U U G C - 3’

u Initiation - codon codes for the 1 st amino acid in all polypeptide sequences (AUG) N-formyl methionine in prokaryotes and methionine in eukaryotes u Termination - codon UAA, UAG & UGA do not code for an amino acid & thus signal the end of protein synthesis Also called stop codon or nonsense codon
- Slides: 48