Nucleic Acids Nucleic Acids Are Essential For Information























- Slides: 56

Nucleic Acids


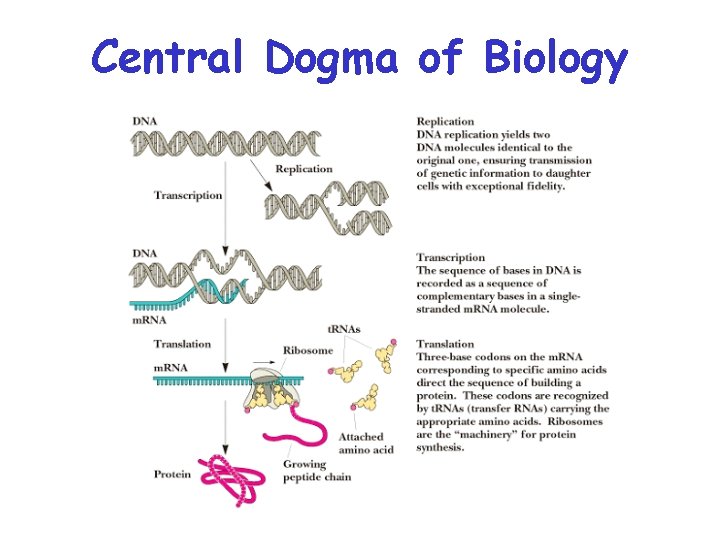
Nucleic Acids Are Essential For Information Transfer in Cells • Information encoded in a DNA molecule is transcribed via synthesis of an RNA molecule • The sequence of the RNA molecule is "read" and is translated into the sequence of amino acids in a protein.

Central Dogma of Biology

Nucleic Acids • First discovered in 1869 by Miescher. • Found as a precipitate that formed when extracts from nuclei were treated with acid. • Compound contained C, N, O, and high amount of P. • Was an acid compound found in nuclei therefore named nucleic acid

Nucleic Acids • 1944 Oswald, Avery, Mac. Leod and Mc. Carty demonstrated that DNA is the molecule that carrier genetic information. • 1953 Watson and Crick proposed the double helix model for the structure of DNA

Nucleic Acids • Nucleic acids are long polymers of nucleotides. • Nucleotides contain a 5 carbon sugar, a weakly basic nitrogenous compound (base), one or more phosphate groups. • Nucleosides are similar to nucleotides but have no phosphate groups.

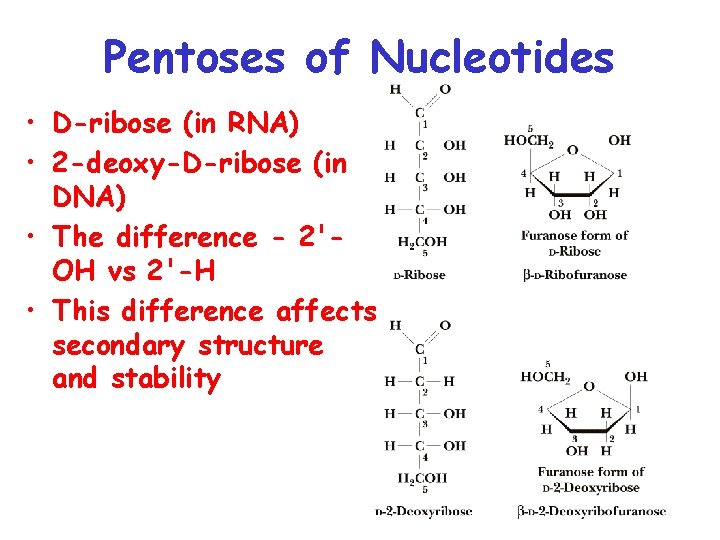
Pentoses of Nucleotides • D-ribose (in RNA) • 2 -deoxy-D-ribose (in DNA) • The difference - 2'OH vs 2'-H • This difference affects secondary structure and stability

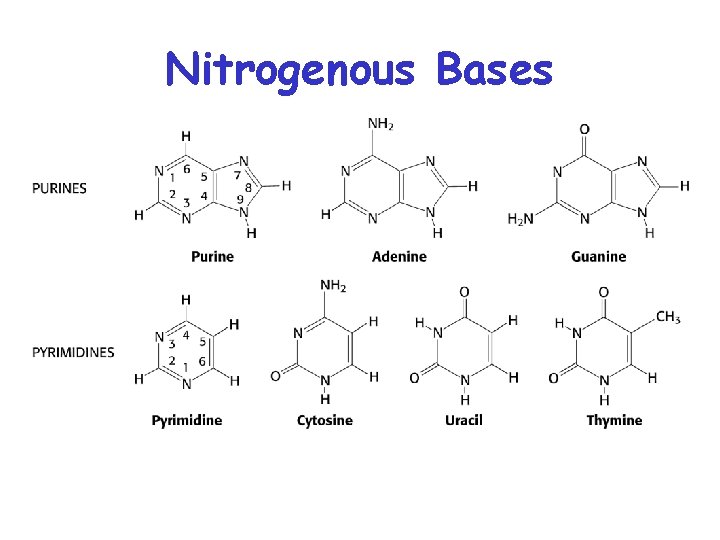
Nitrogenous Bases

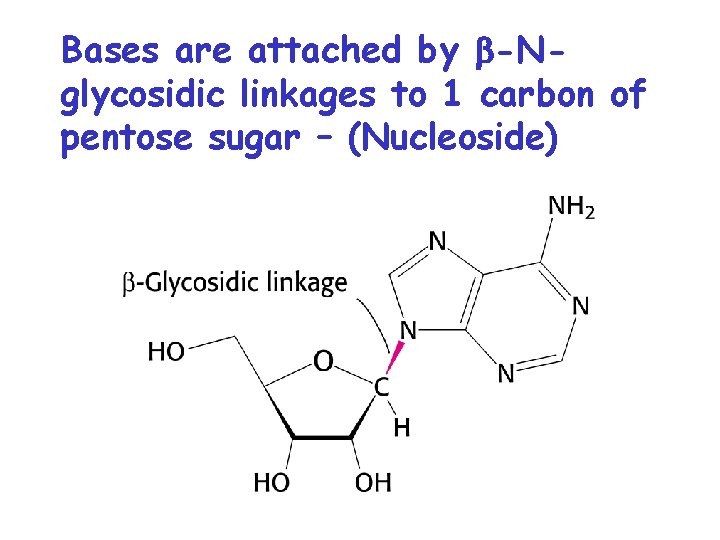
Bases are attached by b-Nglycosidic linkages to 1 carbon of pentose sugar – (Nucleoside)

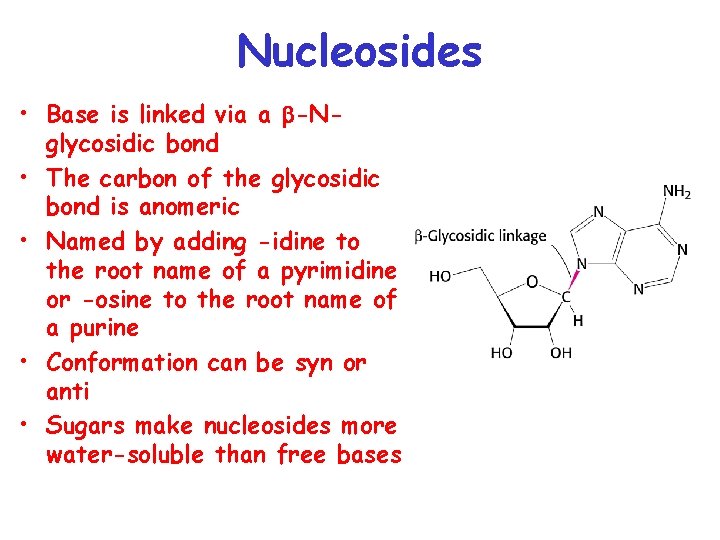
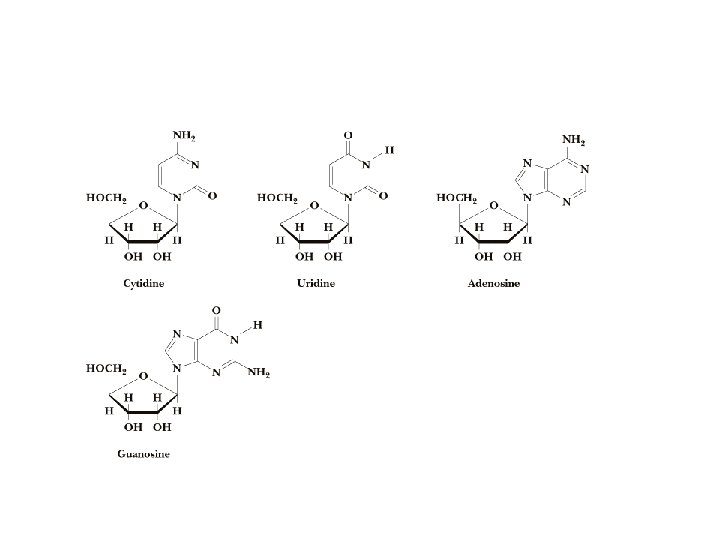
Nucleosides • Base is linked via a b-Nglycosidic bond • The carbon of the glycosidic bond is anomeric • Named by adding -idine to the root name of a pyrimidine or -osine to the root name of a purine • Conformation can be syn or anti • Sugars make nucleosides more water-soluble than free bases


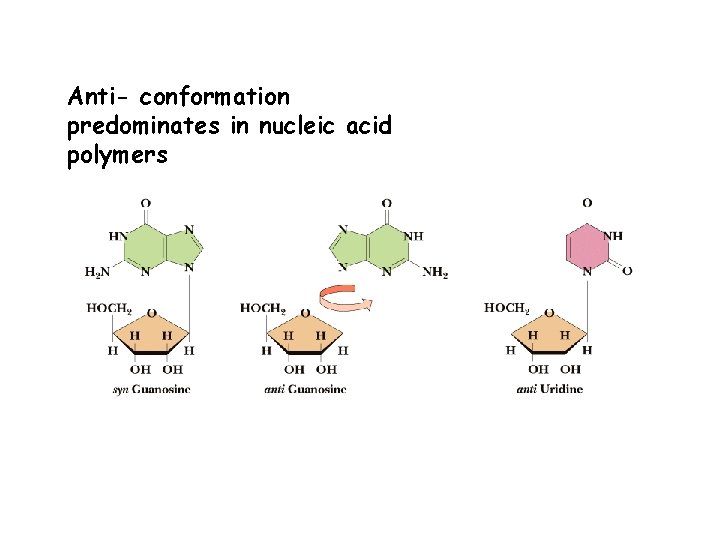
Anti- conformation predominates in nucleic acid polymers

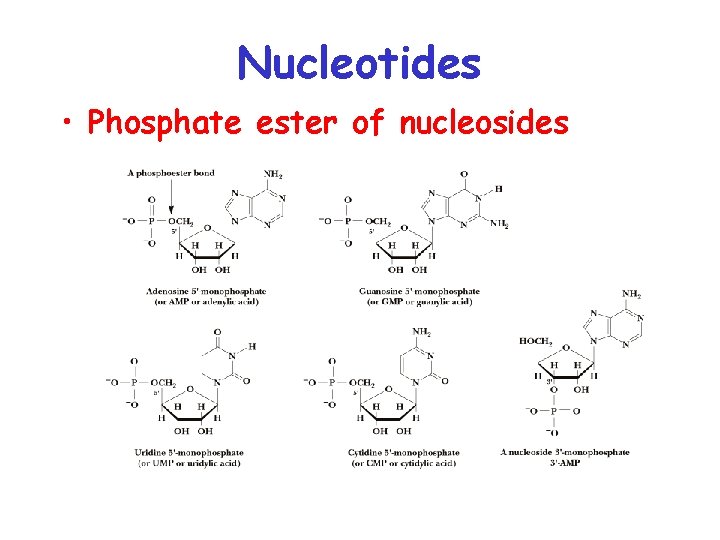
Nucleotides • Phosphate ester of nucleosides

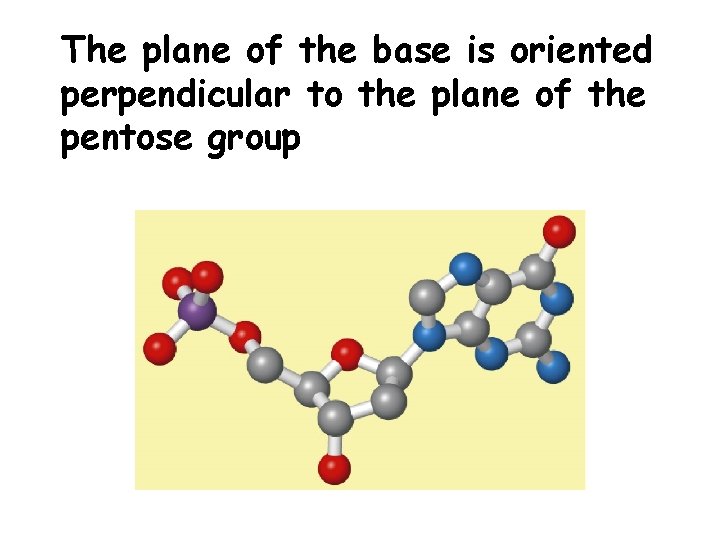
The plane of the base is oriented perpendicular to the plane of the pentose group

Other Functions of Nucleotides • Nucleoside 5'-triphosphates are carriers of energy • Bases serve as recognition units • Cyclic nucleotides are signal molecules and regulators of cellular metabolism and reproduction • ATP is central to energy metabolism • GTP drives protein synthesis • CTP drives lipid synthesis • UTP drives carbohydrate metabolism

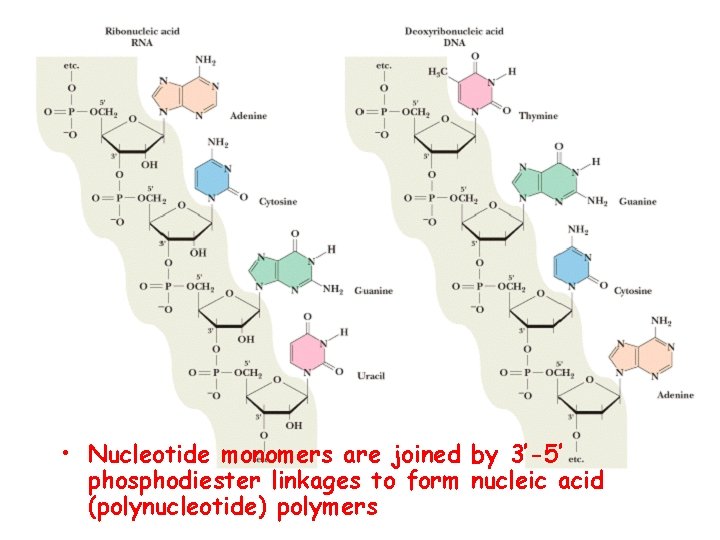
• Nucleotide monomers are joined by 3’-5’ phosphodiester linkages to form nucleic acid (polynucleotide) polymers

Nucleic Acids • Nucleic acid backbone takes on extended conformation. • Nucleotide residues are all oriented in the same direction (5’ to 3’) giving the polymer directionality. • The sequence of DNA molecules is always read in the 5’ to 3’ direction

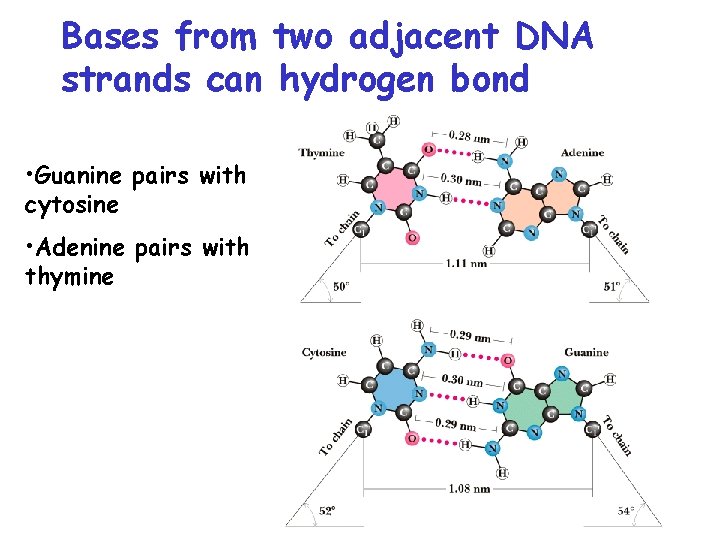
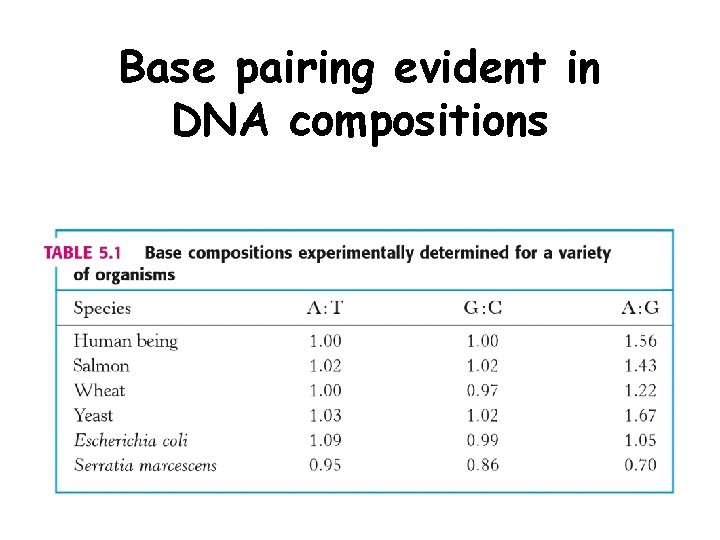
Bases from two adjacent DNA strands can hydrogen bond • Guanine pairs with cytosine • Adenine pairs with thymine

Base pairing evident in DNA compositions

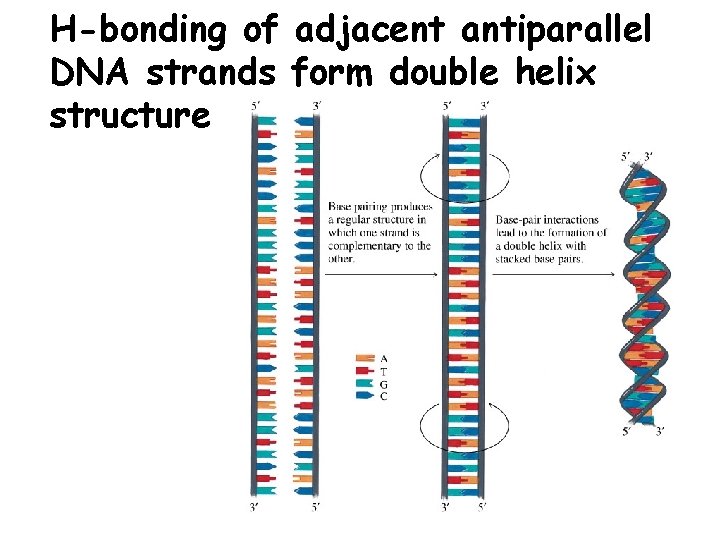
H-bonding of adjacent antiparallel DNA strands form double helix structure

Properties of DNA Double Helix • Distance between the 2 sugar-phosphate backbones is always the same, give DNA molecule a regular shape. • Plane of bases are oriented perpendicular to backbone • Hydrophillic sugar phosphate backbone winds around outside of helix • Noncovalent interactions between upper and lower surfaces of base-pairs (stacking) forms a closely packed hydrophobic interior. • Hydrophobic environment makes H-bonding between bases stronger (no competition with water) • Cause the sugar-phosphate backbone to twist.

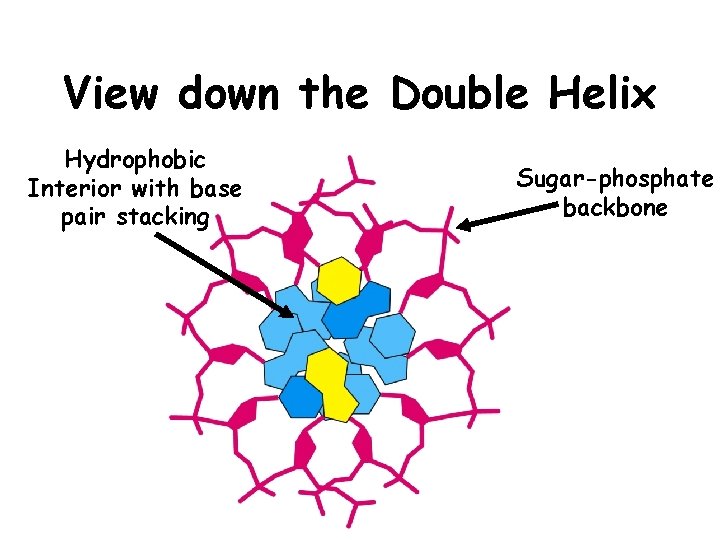
View down the Double Helix Hydrophobic Interior with base pair stacking Sugar-phosphate backbone

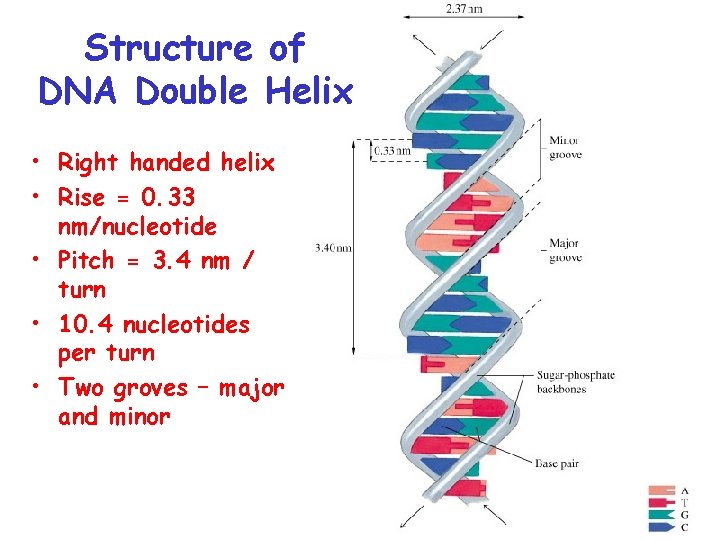
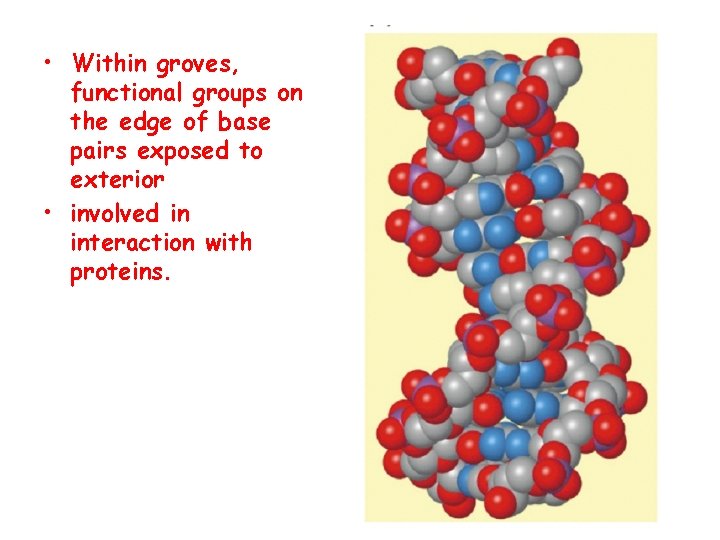
Structure of DNA Double Helix • Right handed helix • Rise = 0. 33 nm/nucleotide • Pitch = 3. 4 nm / turn • 10. 4 nucleotides per turn • Two groves – major and minor

• Within groves, functional groups on the edge of base pairs exposed to exterior • involved in interaction with proteins.

Factors stabilizing DNA double Helix • Hydrophobic interactions – burying hydrophobic purine and pyrimidine rings in interior • Stacking interactions – van der Waals interactions between stacked bases. • Hydrogen Bonding – H-bonding between bases • Charge-Charge Interactions – Electrostatic repulsions of negatively charged phosphate groups are minimized by interaction with cations (e. g. Mg 2+)

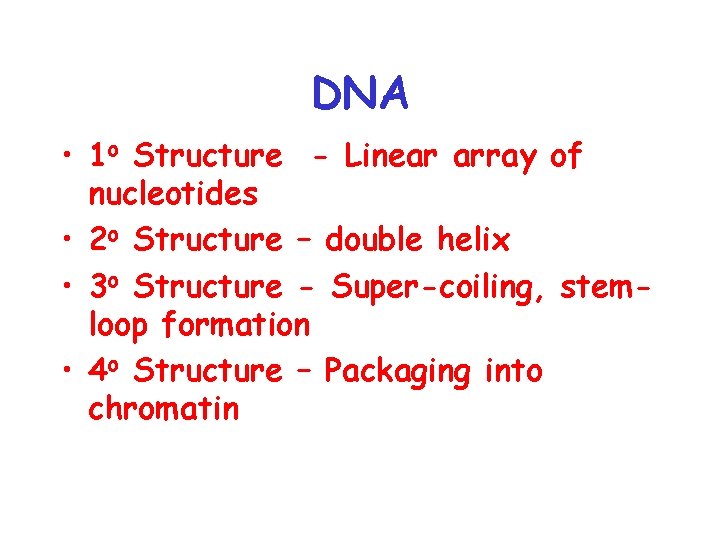
DNA • 1 o Structure - Linear array of nucleotides • 2 o Structure – double helix • 3 o Structure - Super-coiling, stemloop formation • 4 o Structure – Packaging into chromatin

Determination of the DNA 1 o Structure (DNA Sequencing) • Can determine the sequence of DNA base pairs in any DNA molecule • Chain-termination method developed by Sanger • Involves in vitro replication of target DNA • Technology led to the sequencing of the human genome

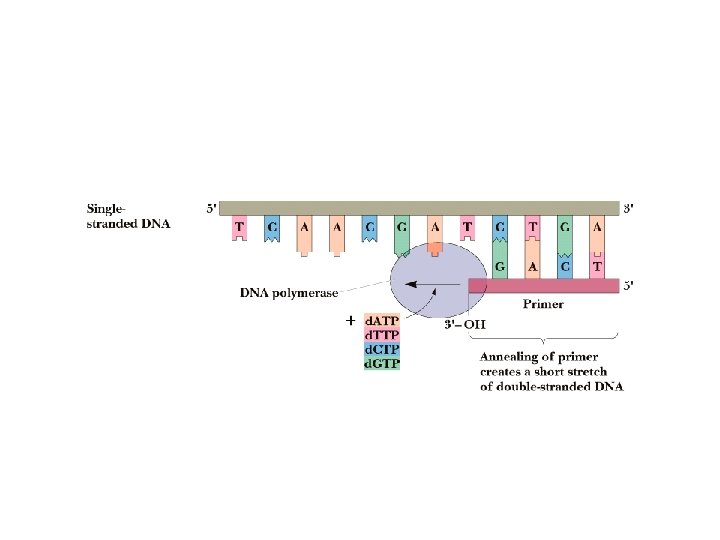
DNA Replication • DNA is a double-helical molecule • Each strand of the helix must be copied in complementary fashion by DNA polymerase • Each strand is a template for copying • DNA polymerase requires template and primer • Primer: an oligonucleotide that pairs with the end of the template molecule to form ds. DNA • DNA polymerases add nucleotides in 5'-3' direction


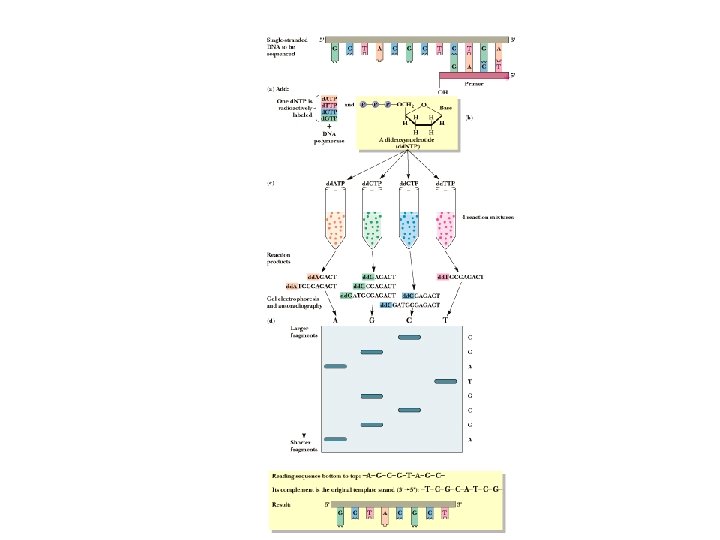
Chain Termination Method • Based on DNA polymerase reaction • 4 separate rxns • Each reaction mixture contains d. ATP, d. GTP, d. CTP and d. TTP • Each reaction also contains a small amount of one dideoxynucleotide (dd. ATP, dd. GTP, dd. CTP and dd. TTP). • Each of the 4 dideoxynucleotides are labeled with a different fluorescent dye. • Dideoxynucleotides missing 3’-OH group. Once incorporated into the DNA chain, chain elongation stops)

Chain Termination Method • Most of the time, the polymerase uses normal nucleotides and DNA molecules grow normally • Occasionally, the polymerase uses a dideoxynucleotide, which adds to the chain and then prevents further growth in that molecule • Random insertion of dd-nucleotides leaves (optimally) at least a few chains terminated at every occurrence of a given nucleotide



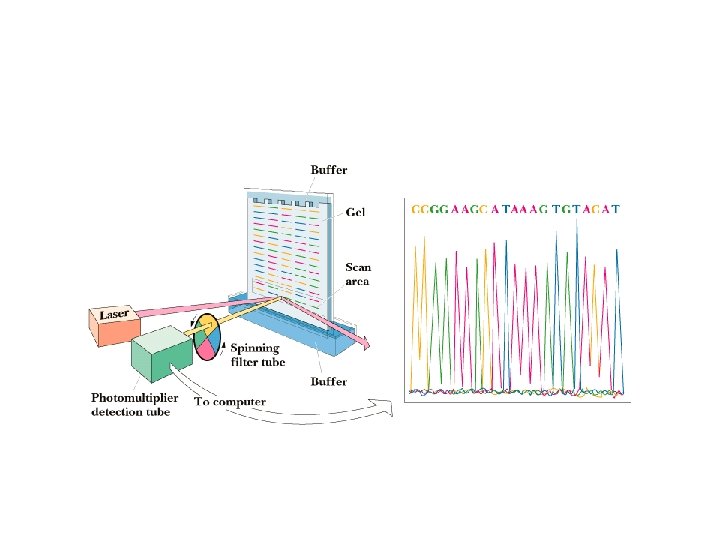
Chain Termination Method • Run each reaction mixture on electrophoresis gel • Short fragments go to bottom, long fragments on top • Read the "sequence" from bottom of gel to top • Convert this "sequence" to the complementary sequence • Now read from the other end and you have the sequence you wanted - read 5' to 3'



DNA Secondary structure • DNA is double stranded with antiparallel strands • Right hand double helix • Three different helical forms (A, B and Z DNA.

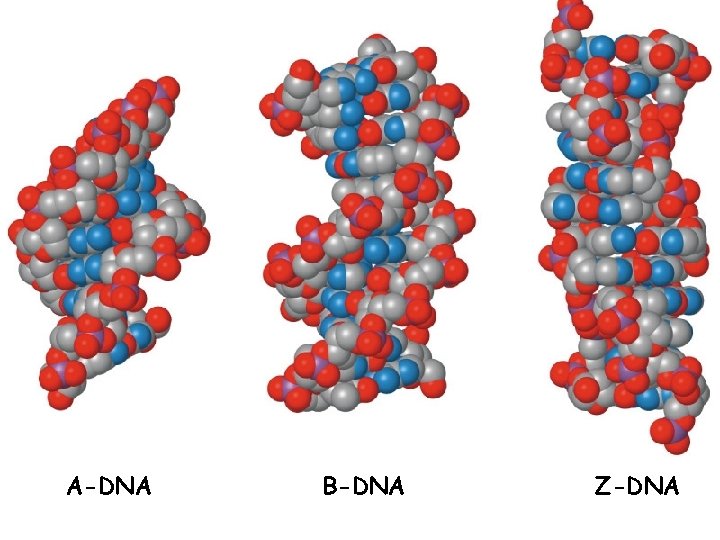
Comparison of A, B, Z DNA • A: right-handed, short and broad, 2. 3 A, 11 bp per turn • B: right-handed, longer, thinner, 3. 32 A, 10 bp per turn • Z: left-handed, longest, thinnest, 3. 8 A, 12 bp per turn

A-DNA B-DNA Z-DNA

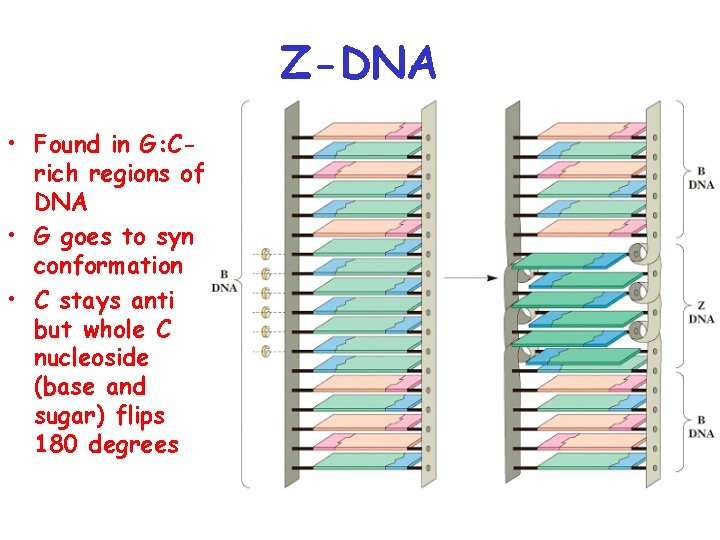
Z-DNA • Found in G: Crich regions of DNA • G goes to syn conformation • C stays anti but whole C nucleoside (base and sugar) flips 180 degrees

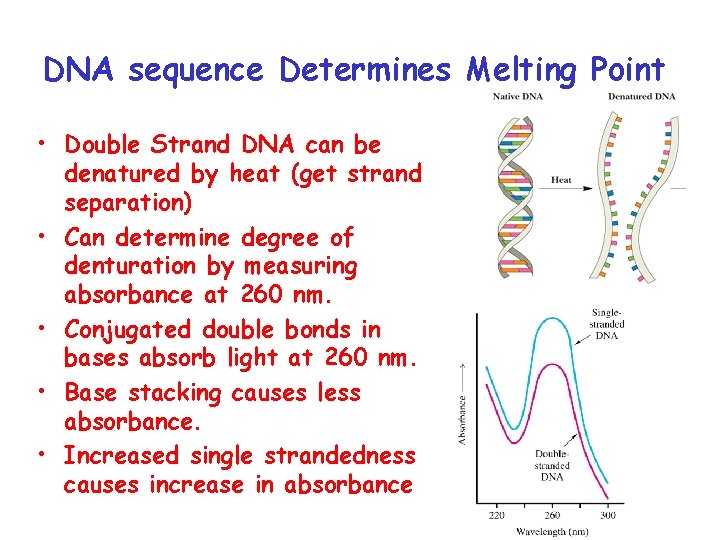
DNA sequence Determines Melting Point • Double Strand DNA can be denatured by heat (get strand separation) • Can determine degree of denturation by measuring absorbance at 260 nm. • Conjugated double bonds in bases absorb light at 260 nm. • Base stacking causes less absorbance. • Increased single strandedness causes increase in absorbance

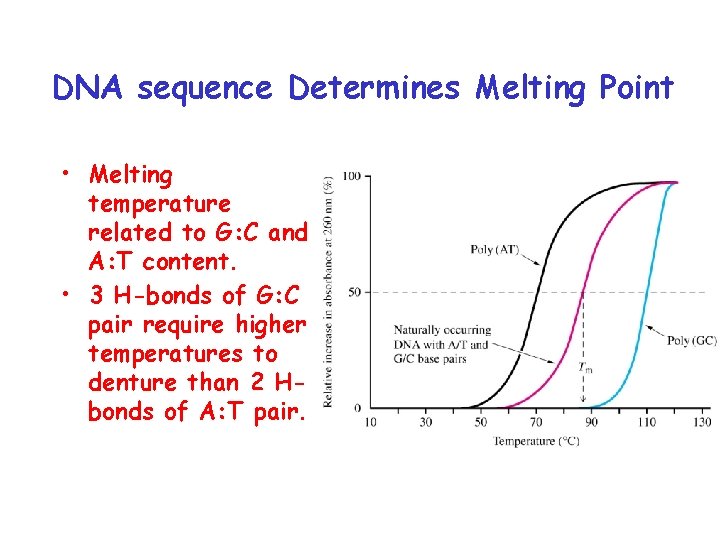
DNA sequence Determines Melting Point • Melting temperature related to G: C and A: T content. • 3 H-bonds of G: C pair require higher temperatures to denture than 2 Hbonds of A: T pair.

DNA o 3 Structure • Super coiling • Cruciform structures

Supercoils • In duplex DNA, ten bp per turn of helix (relaxed form) • DNA helix can be over-wound. • Over winding of DNA helix can be compensated by supercoiling. • Supercoiling prevalent in circular DNA molecules and within local regions of long linear DNA strands • Enzymes called topoisomerases or gyrases can introduce or remove supercoils • In vivo most DNA is negatively supercoiled. • Therefore, it is easy to unwind short regions of the molecule to allow access for enzymes

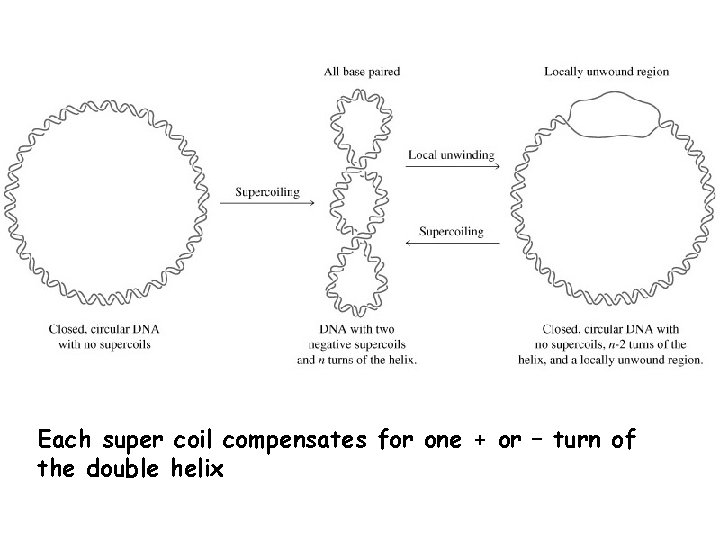
Each super coil compensates for one + or – turn of the double helix

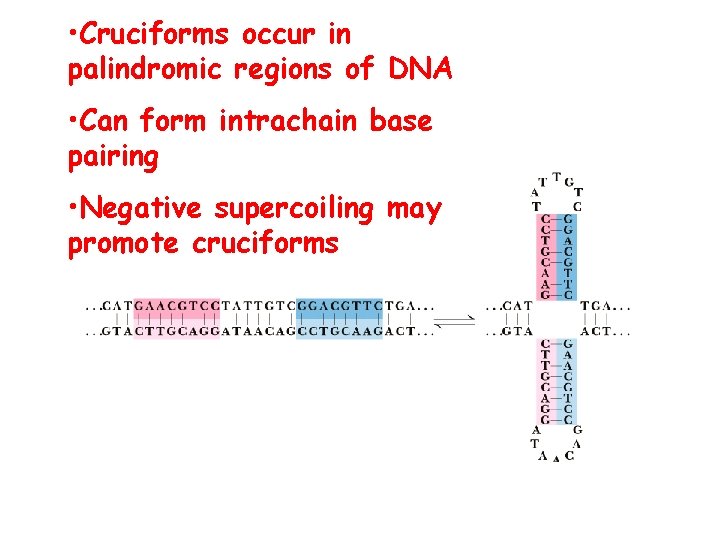
• Cruciforms occur in palindromic regions of DNA • Can form intrachain base pairing • Negative supercoiling may promote cruciforms

DNA o 4 Structure • In chromosomes, DNA is tightly associated with proteins

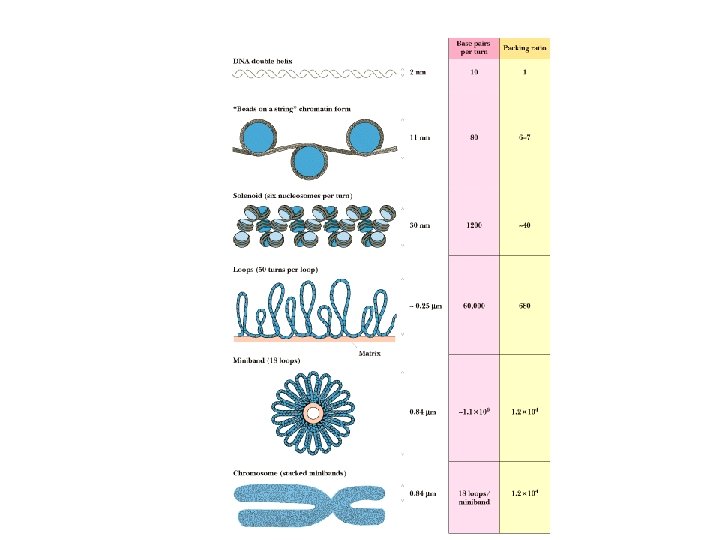
Chromosome Structure • Human DNA’s total length is ~2 meters! • This must be packaged into a nucleus that is about 5 micrometers in diameter • This represents a compression of more than 100, 000! • It is made possible by wrapping the DNA around protein spools called nucleosomes and then packing these in helical filaments

Nucleosome Structure • Chromatin, the nucleoprotein complex, consists of histones and nonhistone chromosomal proteins • % major histone proteins: H 1, H 2 A, H 2 B, H 3 and H 4 • Histone octamers are major part of the “protein spools” • Nonhistone proteins are regulators of gene expression

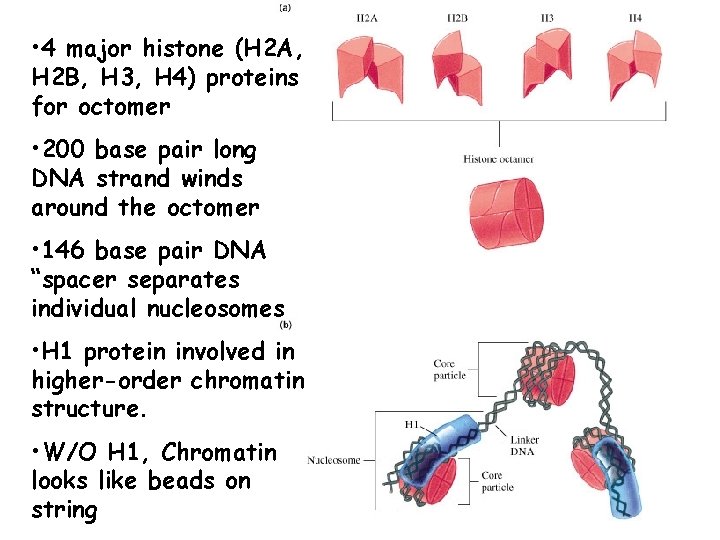
• 4 major histone (H 2 A, H 2 B, H 3, H 4) proteins for octomer • 200 base pair long DNA strand winds around the octomer • 146 base pair DNA “spacer separates individual nucleosomes • H 1 protein involved in higher-order chromatin structure. • W/O H 1, Chromatin looks like beads on string

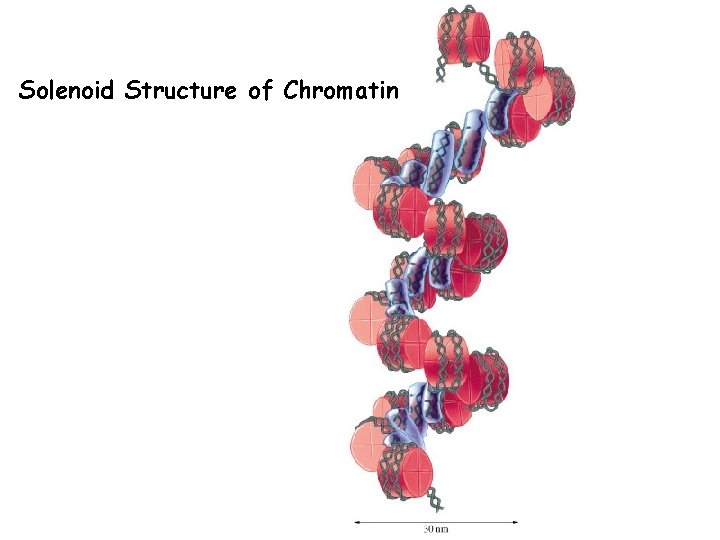
Solenoid Structure of Chromatin


RNA • Single stranded molecule • Chemically less stable than DNA • presence of 2’-OH makes RNA more susceptible to hydrolytic attack (especially form bases) • Prone to degradation by Ribonucleases (Rnases) • Has secondary structure. Can form intrachain base pairing (i. e. cruciform structures). • Multiple functions

Type of RNA • Ribosomal RNA (r. RNA) – integral part of ribosomes (very abundant) • Transfer RNA (t. RNA) – carries activated amino acids to ribosomes. • Messenger RNA (m. RNA) – endcodes sequences of amino acids in proteins. • Catalytic RNA (Ribozymes) – catalzye cleavage of specific RNA species.

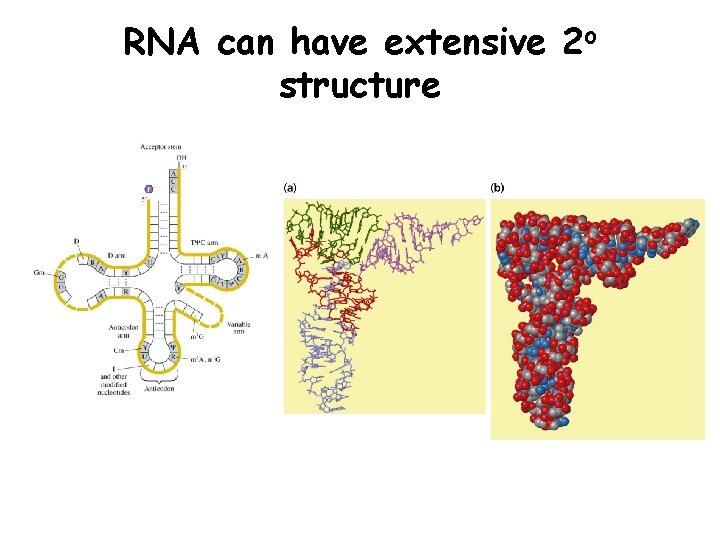
RNA can have extensive 2 o structure