Nucleic Acids Deoxyribonucleic acid DNA There are two
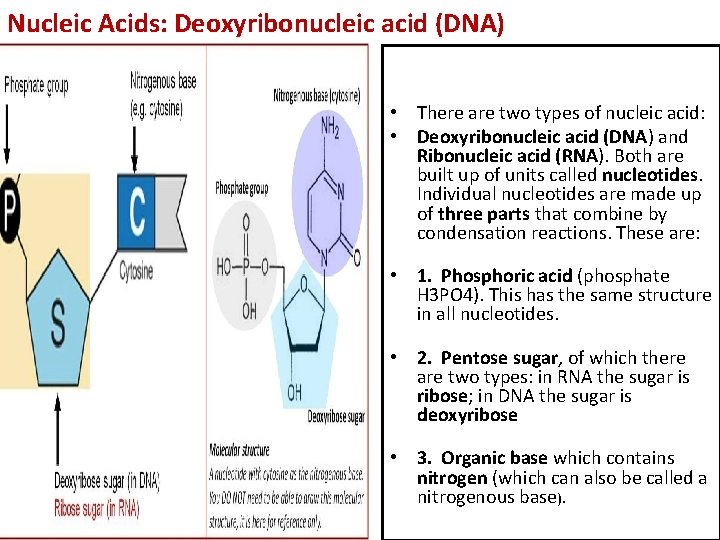
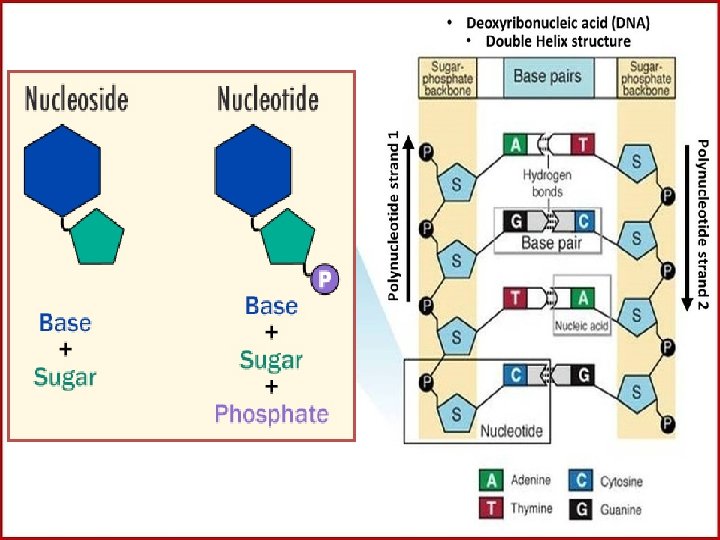
Nucleic Acids: Deoxyribonucleic acid (DNA) • There are two types of nucleic acid: • Deoxyribonucleic acid (DNA) and Ribonucleic acid (RNA). Both are built up of units called nucleotides. Individual nucleotides are made up of three parts that combine by condensation reactions. These are: • 1. Phosphoric acid (phosphate H 3 PO 4). This has the same structure in all nucleotides. • 2. Pentose sugar, of which there are two types: in RNA the sugar is ribose; in DNA the sugar is deoxyribose • 3. Organic base which contains nitrogen (which can also be called a nitrogenous base).
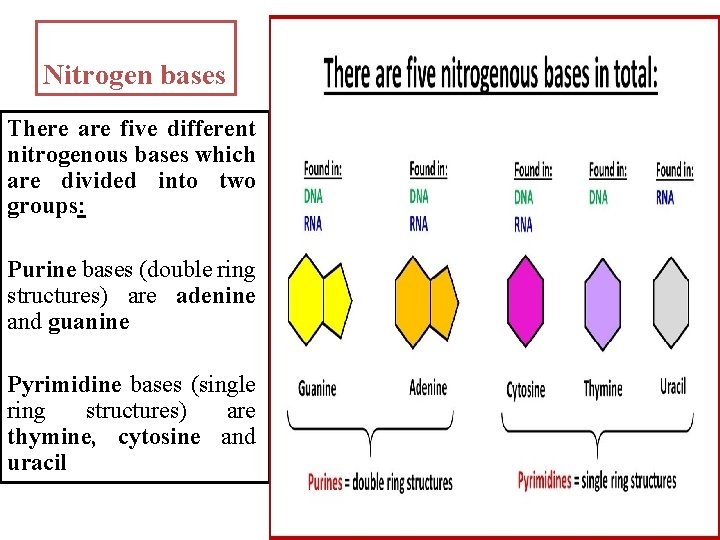
Nitrogen bases There are five different nitrogenous bases which are divided into two groups: Purine bases (double ring structures) are adenine and guanine Pyrimidine bases (single ring structures) are thymine, cytosine and uracil
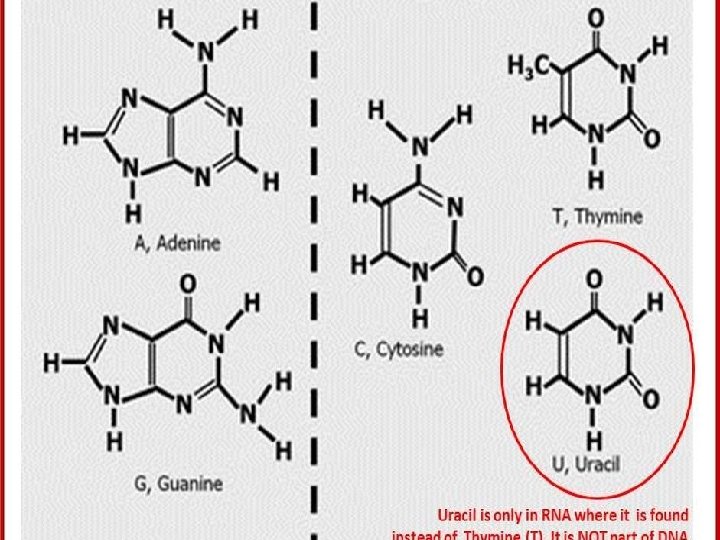
Structure of Purine and Pyrimidine
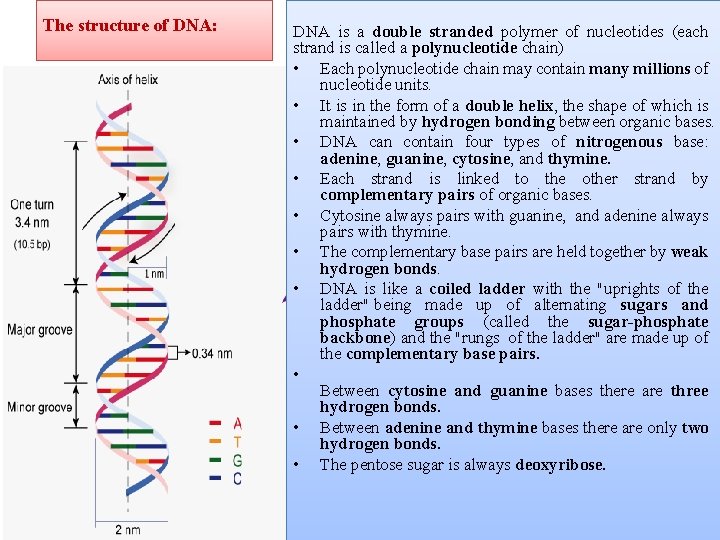
The structure of DNA: DNA is a double stranded polymer of nucleotides (each strand is called a polynucleotide chain) • Each polynucleotide chain may contain many millions of nucleotide units. • It is in the form of a double helix, the shape of which is maintained by hydrogen bonding between organic bases. • DNA can contain four types of nitrogenous base: adenine, guanine, cytosine, and thymine. • Each strand is linked to the other strand by complementary pairs of organic bases. • Cytosine always pairs with guanine, and adenine always pairs with thymine. • The complementary base pairs are held together by weak hydrogen bonds. • DNA is like a coiled ladder with the "uprights of the ladder" being made up of alternating sugars and phosphate groups (called the sugar-phosphate backbone) and the "rungs of the ladder" are made up of the complementary base pairs. • Between cytosine and guanine bases there are three hydrogen bonds. • Between adenine and thymine bases there are only two hydrogen bonds. • The pentose sugar is always deoxyribose.
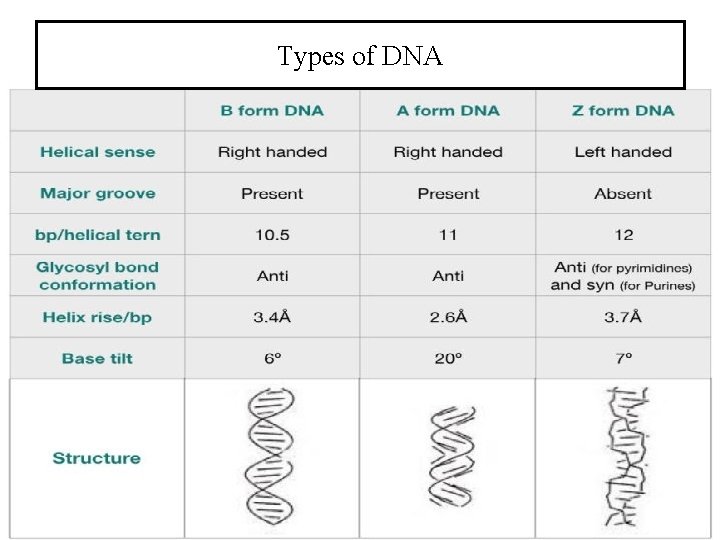
Types of DNA
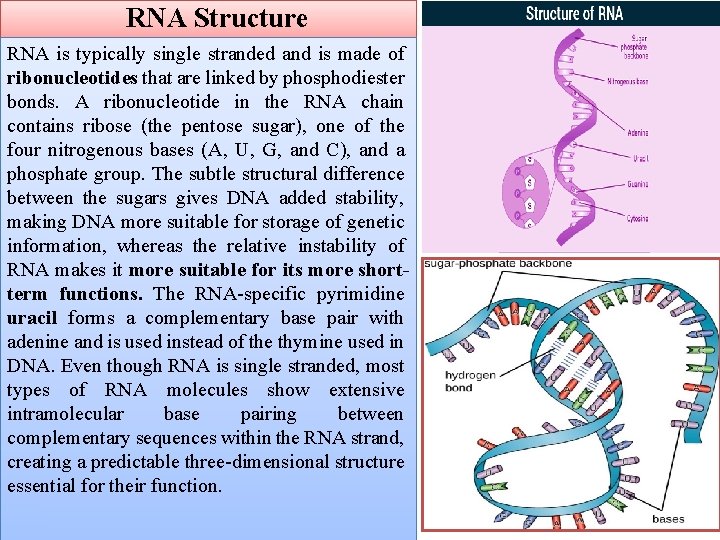
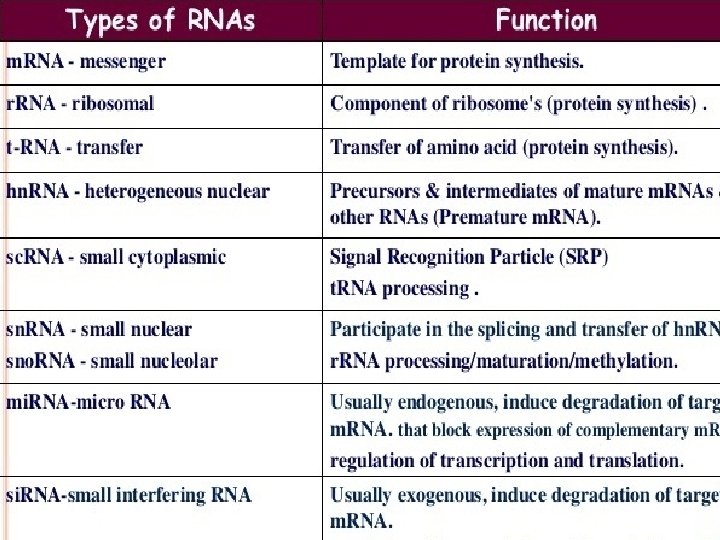
RNA Structure RNA is typically single stranded and is made of ribonucleotides that are linked by phosphodiester bonds. A ribonucleotide in the RNA chain contains ribose (the pentose sugar), one of the four nitrogenous bases (A, U, G, and C), and a phosphate group. The subtle structural difference between the sugars gives DNA added stability, making DNA more suitable for storage of genetic information, whereas the relative instability of RNA makes it more suitable for its more shortterm functions. The RNA-specific pyrimidine uracil forms a complementary base pair with adenine and is used instead of the thymine used in DNA. Even though RNA is single stranded, most types of RNA molecules show extensive intramolecular base pairing between complementary sequences within the RNA strand, creating a predictable three-dimensional structure essential for their function.

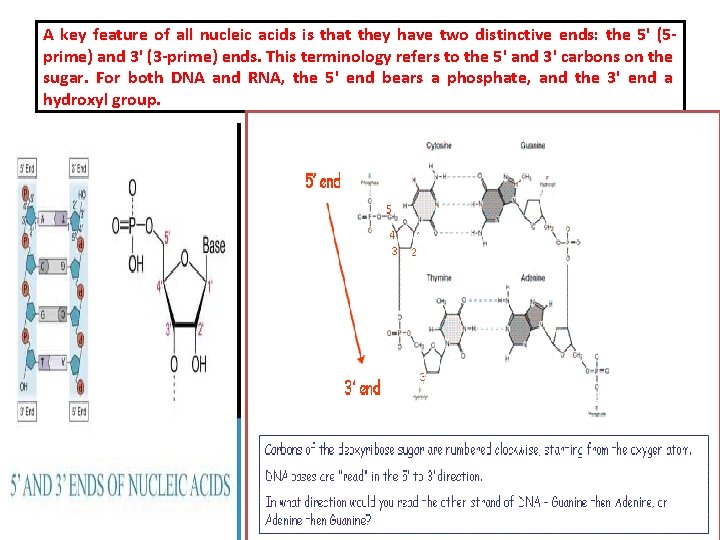
A key feature of all nucleic acids is that they have two distinctive ends: the 5' (5 prime) and 3' (3 -prime) ends. This terminology refers to the 5' and 3' carbons on the sugar. For both DNA and RNA, the 5' end bears a phosphate, and the 3' end a hydroxyl group.

Complementarity of DNA • • • Complementarity of DNA strands in a double helix make it possible to use one strand as a template to construct the other. This principle plays an important role in DNA replication, setting the foundation of heredity by explaining how genetic information can be passed down to the next generation. Complementarity is also utilized in DNA transcription, which generates an RNA strand from a DNA template. DNA repair mechanisms such as proof reading are complementarity based and allow for error correction during DNA replication by removing mismatched nucleobases. Nucleic acids strands may also form hybrids in which single stranded DNA may readily anneal with complementary DNA or RNA. This principle is the basis of commonly performed laboratory techniques such as the polymerase chain reaction, PCR. Two strands of complementary sequence are referred to as sense and anti-sense. The sense strand is, generally, the transcribed sequence of DNA or the RNA that was generated in transcription. While the anti-sense strand is the strand that is complementary to the sense sequence.
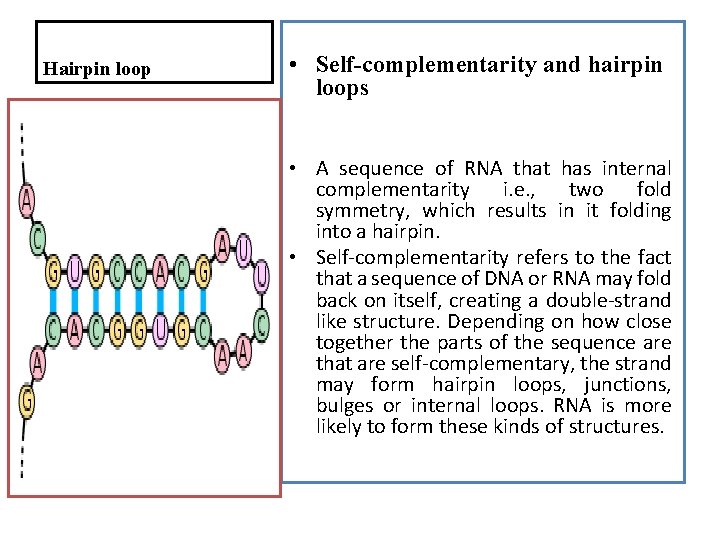
Hairpin loop • Self-complementarity and hairpin loops • A sequence of RNA that has internal complementarity i. e. , two fold symmetry, which results in it folding into a hairpin. • Self-complementarity refers to the fact that a sequence of DNA or RNA may fold back on itself, creating a double-strand like structure. Depending on how close together the parts of the sequence are that are self-complementary, the strand may form hairpin loops, junctions, bulges or internal loops. RNA is more likely to form these kinds of structures.
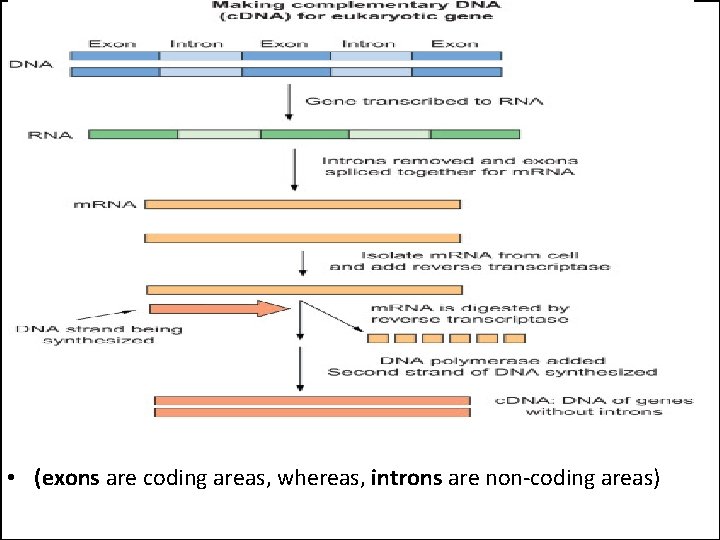
• (exons are coding areas, whereas, introns are non-coding areas)
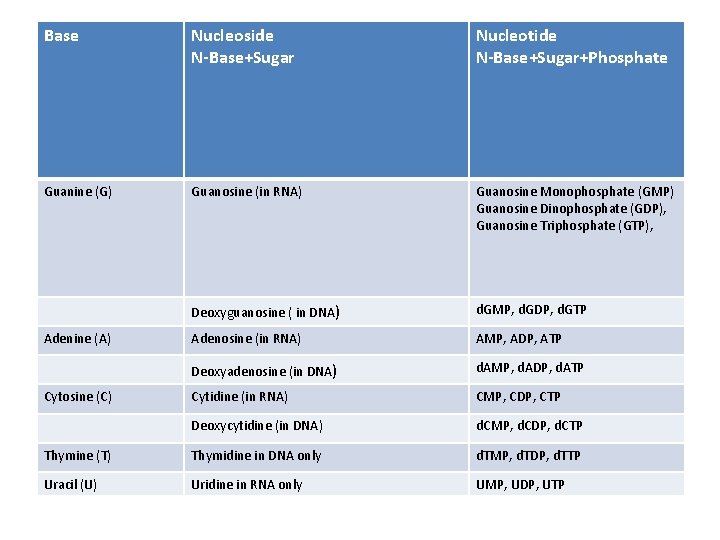
Base Nucleoside N-Base+Sugar Nucleotide N-Base+Sugar+Phosphate Guanine (G) Guanosine (in RNA) Guanosine Monophosphate (GMP) Guanosine Dinophosphate (GDP), Guanosine Triphosphate (GTP), Deoxyguanosine ( in DNA) d. GMP, d. GDP, d. GTP Adenosine (in RNA) AMP, ADP, ATP Deoxyadenosine (in DNA) d. AMP, d. ADP, d. ATP Cytidine (in RNA) CMP, CDP, CTP Deoxycytidine (in DNA) d. CMP, d. CDP, d. CTP Thymine (T) Thymidine in DNA only d. TMP, d. TDP, d. TTP Uracil (U) Uridine in RNA only UMP, UDP, UTP Adenine (A) Cytosine (C)

Biosynthesis of Purine Nucleotide 2 ways nucleotides are synthesized in our cells. De novo synthesis: The Major Pathway: Synthesis from simple molecules including CO 2, C 6 H 12 O 6, NH 3, bicarbonate, amino acids. Salvage pathway: Nucleotides are synthesized from intermediates in the degradative pathway for nucleotides and are formed during degradation of RNA and DNA. Liver synthesizes purine in cytosol. Metabolic pathway associated with pentose phosphate pathway of carbohydrate metabolism which produce Ribose-5 -Phosphote (R-5 -P).

De novo Synthesis of Purine • The starter molecule is Ribose, the sugar molecule. Which needs to be delivered in an active form. • Ribose 5 -Phosphate is produced in carbohydrate metabolism from pentose phosphate pathway (PPP). • It is the starting material for purine nucleotide synthesis. • It reacts with ATP to form Phosphoribosyl Pyrophosphate. • Parent purine nucleotide first synthesised is Inosine Mono Phosphate (IMP), a nucleotide composed of hypoxanthine +Ribose + Phosphate. • From IMP other purine nucleotides are synthesized, like Adenosine Monophosphate (AMP), Guanosine Mono Phosphate (GMP). • Nitrogen bases come from amino acids like Glysine, aspartic acid, glutamine.
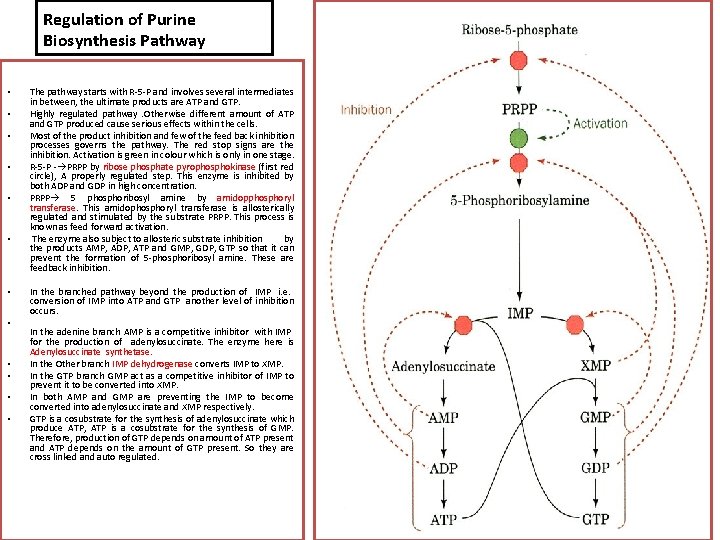
Regulation of Purine Biosynthesis Pathway • • • The pathway starts with R-5 -P and involves several intermediates in between, the ultimate products are ATP and GTP. Highly regulated pathway. Otherwise different amount of ATP and GTP produced cause serious effects within the cells. Most of the product inhibition and few of the feed back inhibition processes governs the pathway. The red stop signs are the inhibition. Activation is green in colour which is only in one stage. R-5 -P - PRPP by ribose phosphate pyrophosphokinase (first red circle), A properly regulated step. This enzyme is inhibited by both ADP and GDP in high concentration. PRPP 5 phosphoribosyl amine by amidopphosphoryl transferase. This amidophosphoryl transferase is allosterically regulated and stimulated by the substrate PRPP. This process is known as feed forward activation. The enzyme also subject to allosteric substrate inhibition by the products AMP, ADP, ATP and GMP, GDP, GTP so that it can prevent the formation of 5 -phosphoribosyl amine. These are feedback inhibition. In the branched pathway beyond the production of IMP i. e. conversion of IMP into ATP and GTP another level of inhibition occurs. In the adenine branch AMP is a competitive inhibitor with IMP for the production of adenylosuccinate. The enzyme here is Adenylosuccinate synthetase. In the Other branch IMP dehydrogenase converts IMP to XMP. In the GTP branch GMP act as a competitive inhibitor of IMP to prevent it to be converted into XMP. In both AMP and GMP are preventing the IMP to become converted into adenylosuccinate and XMP respectively. GTP is a cosubstrate for the synthesis of adenylosuccinate which produce ATP, ATP is a cosubstrate for the synthesis of GMP. Therefore, production of GTP depends on amount of ATP present and ATP depends on the amount of GTP present. So they are cross linked and auto regulated.

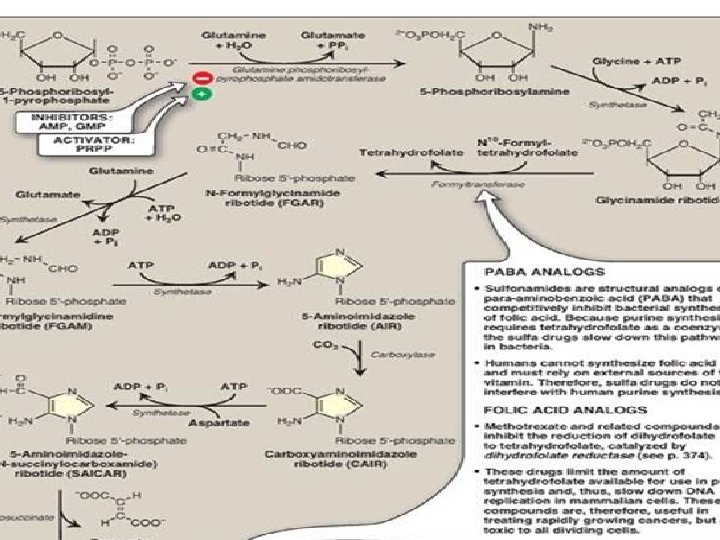
De novo Synthesis of Purine • • • • • • Ribose-5 -P (from PPP) (Enzymes are shown in red colour) PRPP synthetase 5 Phosphoribosyl 1 pyrophosphate-PRPP Glutamine PRPP amidotransferase/ amidopphosphoryl transferase 5 Phosphoribosyl beta amine GAR synthetase Glycinamide ribonucleotide (GAR) GAR transformylase Formylglycinamide ribonucleotide(FGAR) FGAM synthetase FGAM Formylglycinamidine AIR synthetase Aminoimidazole ribonucleotide (AIR) AIR carboxylase carboxyaminoimidazole ribonucleotide (CAIR) SAICAR synthetase Succinylo aminoimidazole carboxamide ribonucleotide (SAICAR) Adenylosuccinatase Aminoimidazole carboxamide ribonucleotide (AICAR) AICAR transformylase Formylaminoimidazole carboxamide ribonucleotide (FAICAR) IMP synthase Inosine Mono Phosphate (IMP)

De novo synthesis of Purine • Ribose-5 -P+ATP—PRPP synthetase- 5 Phosphoribosyl 1 pyrophosphate. PRPP----Glutamine PRPP amidotransferase Phosphoribosyl beta amine • Phosphoribosyl beta amine----GAR synthetase Glycinamide ribonucleotide (GAR) • GAR--GAR transformylase Formylglycinamide ribonucleotide(FGAR) • FGAR—FGAM synthetase FGAM Formylglycinamidine • FGAM—AIR synthetase Aminoimidazole ribonucleotide (AIR) • AIR—AIR carboxylase carboxyaminoimidazole ribonucleotide (CAIR) • CAIR—SAICAR synthetase Succinylo aminoimidazole carboxamide ribonucleotide • SAICAR—Adenylosuccinatase Aminoimidazole carboxamide ribonucleotide (AICAR) • AICAR—AICAR transformylase Formylaminoimidazole carboxamide ribonucleotide (FAICAR) • FAICAR—IMP synthase Inosine Mono Phosphate (IMP) • (This is the same material as in previous slide but in different form)

Synthesis of 5 Phosphoribosyl-1 pyrophosphate (PRPP) Metabolic pathway associated with pentose phosphate pathway of carbohydrate metabolism which produce Ribose-5 -Phosphote (R-5 -P). 5 -Phosphoribosyl-1 pyrophosphate (PRPP) is an “activated pentose” that participates in the synthesis of purine and pyrimidines. Synthesis of PRPP from ATP and ribose -5 -phosphate is catalysed by PRPP synthetase.
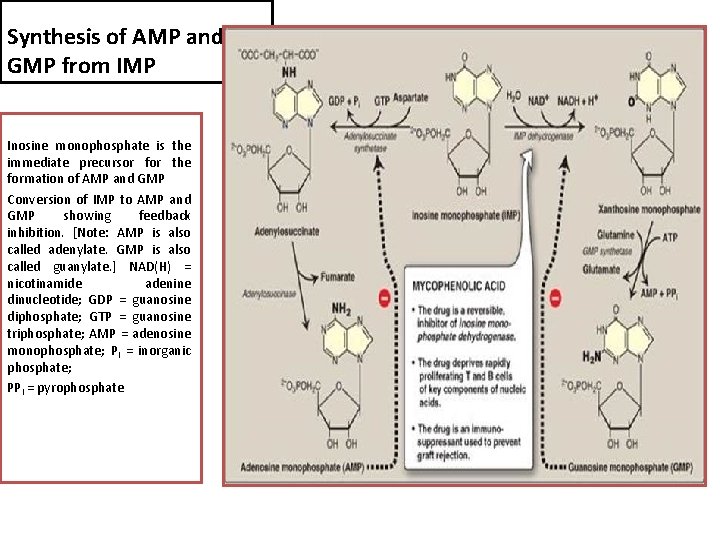
Synthesis of AMP and GMP from IMP Inosine monophosphate is the immediate precursor for the formation of AMP and GMP Conversion of IMP to AMP and GMP showing feedback inhibition. [Note: AMP is also called adenylate. GMP is also called guanylate. ] NAD(H) = nicotinamide adenine dinucleotide; GDP = guanosine diphosphate; GTP = guanosine triphosphate; AMP = adenosine monophosphate; Pi = inorganic phosphate; PPi = pyrophosphate

Conversion of nucleoside monophosphates to nucleoside diphosphates Nucleoside diphosphates are synthesized from the corresponding nucleoside monophosphates by basespecific nucleoside monophosphate kinases. [Note: These kinases do not discriminate between ribose or deoxyribose in the substrate. ] ATP is generally the source of the transferred phosphate because it is present in higher concentrations than the other nucleoside triphosphates. Adenylate kinase is particularly active in the liver and in muscle, where the turnover of energy from ATP is high. Its function is to maintain equilibrium among the adenine nucleotides (AMP, ADP, and ATP). Nucleoside diphosphates and triphosphates are interconverted by nucleoside diphosphate kinase, an enzyme that, unlike the monophosphate kinases, has broad substrate specificityhosphates and triphosphates AMP = adenosine monophosphate; ADP = adenosine diphosphate; GMP = guanosine monophosphate; GDP = guanosine diphosphate; GTP = guanosine triphosphate; CDP = cytidine diphosphate; CTP = cytidine triphosphate.
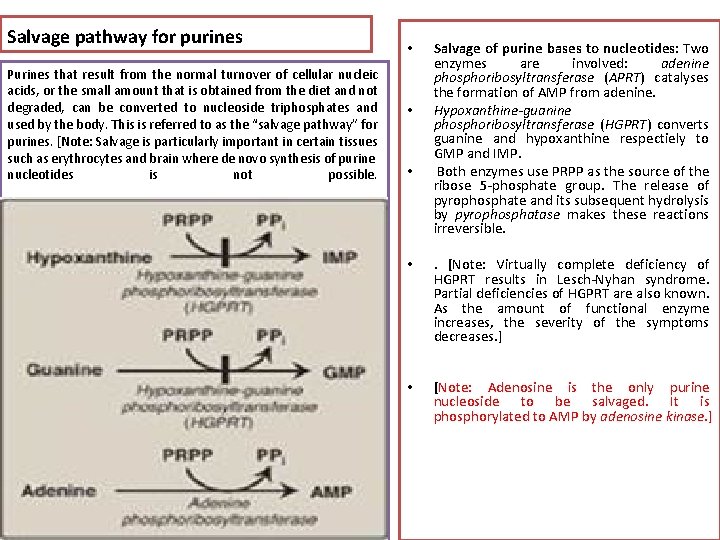
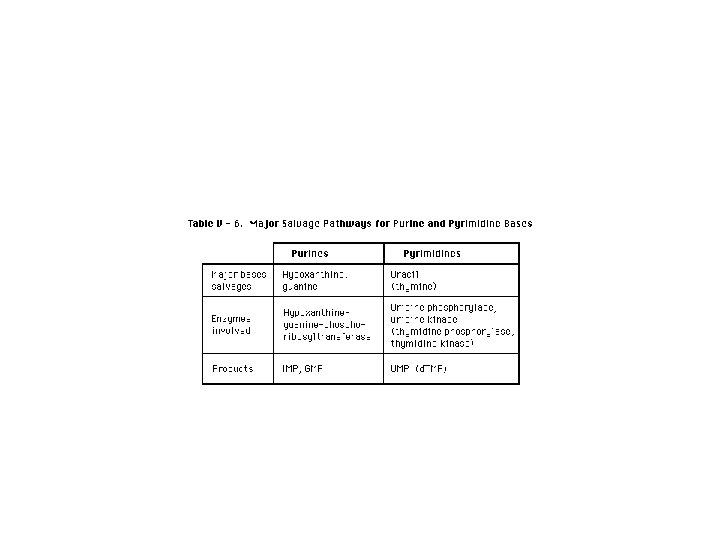
Salvage pathway for purines Purines that result from the normal turnover of cellular nucleic acids, or the small amount that is obtained from the diet and not degraded, can be converted to nucleoside triphosphates and used by the body. This is referred to as the “salvage pathway” for purines. [Note: Salvage is particularly important in certain tissues such as erythrocytes and brain where de novo synthesis of purine nucleotides is not possible. • • • Salvage of purine bases to nucleotides: Two enzymes are involved: adenine phosphoribosyltransferase (APRT) catalyses the formation of AMP from adenine. Hypoxanthine-guanine phosphoribosyltransferase (HGPRT) converts guanine and hypoxanthine respectiely to GMP and IMP. Both enzymes use PRPP as the source of the ribose 5 -phosphate group. The release of pyrophosphate and its subsequent hydrolysis by pyrophosphatase makes these reactions irreversible. • . [Note: Virtually complete deficiency of HGPRT results in Lesch-Nyhan syndrome. Partial deficiencies of HGPRT are also known. As the amount of functional enzyme increases, the severity of the symptoms decreases. ] • [Note: Adenosine is the only purine nucleoside to be salvaged. It is phosphorylated to AMP by adenosine kinase. ]
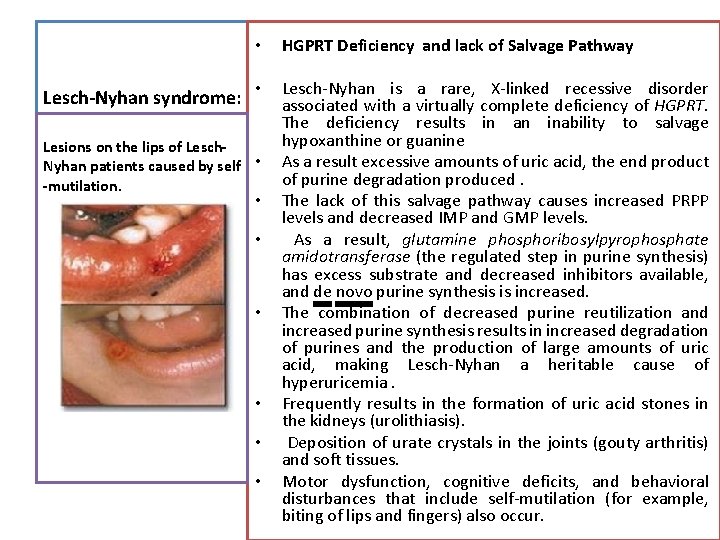
Lesch-Nyhan syndrome: • HGPRT Deficiency and lack of Salvage Pathway • Lesch-Nyhan is a rare, X-linked recessive disorder associated with a virtually complete deficiency of HGPRT. The deficiency results in an inability to salvage hypoxanthine or guanine As a result excessive amounts of uric acid, the end product of purine degradation produced. The lack of this salvage pathway causes increased PRPP levels and decreased IMP and GMP levels. As a result, glutamine phosphoribosylpyrophosphate amidotransferase (the regulated step in purine synthesis) has excess substrate and decreased inhibitors available, and de novo purine synthesis is increased. The combination of decreased purine reutilization and increased purine synthesis results in increased degradation of purines and the production of large amounts of uric acid, making Lesch-Nyhan a heritable cause of hyperuricemia. Frequently results in the formation of uric acid stones in the kidneys (urolithiasis). Deposition of urate crystals in the joints (gouty arthritis) and soft tissues. Motor dysfunction, cognitive deficits, and behavioral disturbances that include self-mutilation (for example, biting of lips and fingers) also occur. Lesions on the lips of Lesch. Nyhan patients caused by self • -mutilation. • • •

Inhibitors of Purine synthesis • Sulfonamides are the structural analogs of paraaminobenzoic acid (PABA). • These sulfur drugs can be used to inhibit the synthesis of folic acid by microorganisms. • This indirectly reduces the synthesis of purines and therefore, the nucleic acids (DNA and RNA). • Sulfonamides have no influence on humans, since folic acid is not synthesized and is supplied through diet. • The structural analogue of folic acid ( methotrexate) are widely used to control cancer. A chemotherapy agent and immune system suppressant. Cancer, autoimmune disease, ectopic pregnancy and for medical abortions. • They inhibit the synthesis of purine nucleotides and thus nucleic acids. • These inhibitors also affect the proliferation of normally growing cells. • These cause many side effects including anaemia, baldness, scaly skin etc.
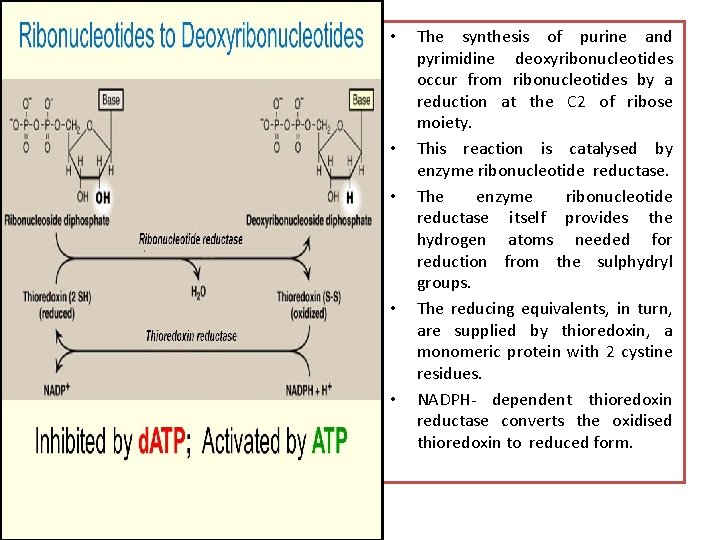
• • • The synthesis of purine and pyrimidine deoxyribonucleotides occur from ribonucleotides by a reduction at the C 2 of ribose moiety. This reaction is catalysed by enzyme ribonucleotide reductase. The enzyme ribonucleotide reductase itself provides the hydrogen atoms needed for reduction from the sulphydryl groups. The reducing equivalents, in turn, are supplied by thioredoxin, a monomeric protein with 2 cystine residues. NADPH- dependent thioredoxin reductase converts the oxidised thioredoxin to reduced form.
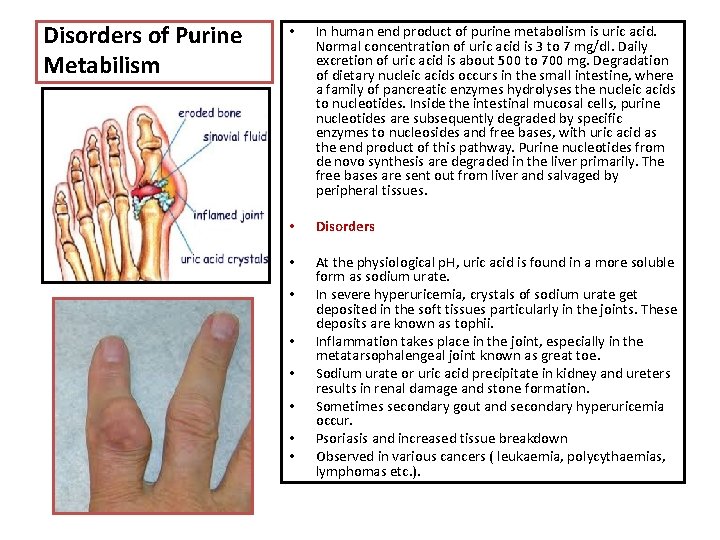
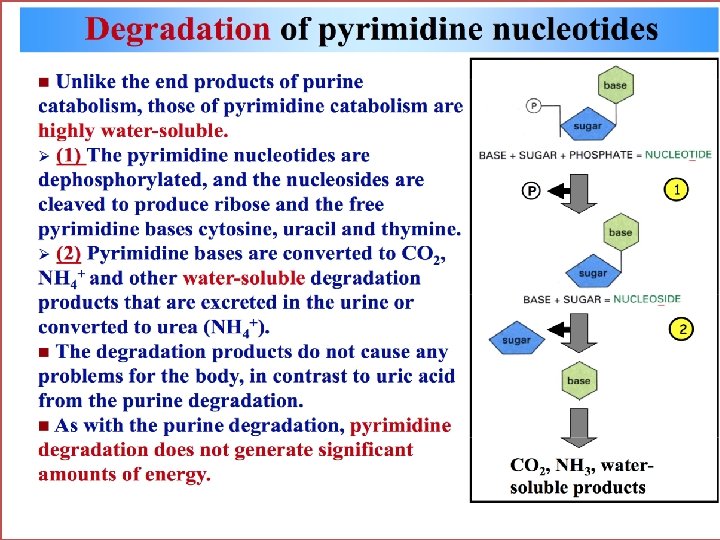
Disorders of Purine Metabilism • In human end product of purine metabolism is uric acid. Normal concentration of uric acid is 3 to 7 mg/dl. Daily excretion of uric acid is about 500 to 700 mg. Degradation of dietary nucleic acids occurs in the small intestine, where a family of pancreatic enzymes hydrolyses the nucleic acids to nucleotides. Inside the intestinal mucosal cells, purine nucleotides are subsequently degraded by specific enzymes to nucleosides and free bases, with uric acid as the end product of this pathway. Purine nucleotides from de novo synthesis are degraded in the liver primarily. The free bases are sent out from liver and salvaged by peripheral tissues. • Disorders • At the physiological p. H, uric acid is found in a more soluble form as sodium urate. In severe hyperuricemia, crystals of sodium urate get deposited in the soft tissues particularly in the joints. These deposits are known as tophii. Inflammation takes place in the joint, especially in the metatarsophalengeal joint known as great toe. Sodium urate or uric acid precipitate in kidney and ureters results in renal damage and stone formation. Sometimes secondary gout and secondary hyperuricemia occur. Psoriasis and increased tissue breakdown Observed in various cancers ( leukaemia, polycythaemias, lymphomas etc. ). • • •
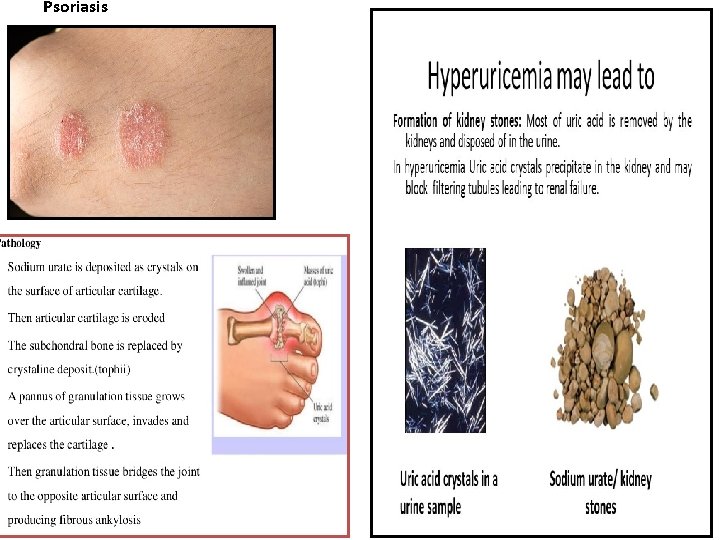
Psoriasis

Pyrimidine Synthesis • • • Synthesis of Carbamoyl Phosphate Synthesis of Carbamoyl Aspartate Ring Closure to form dihydroorotate Oxidation of Dihydro Orotate Addition of ribose Phosphate moiety De Carboxylation to form UMP
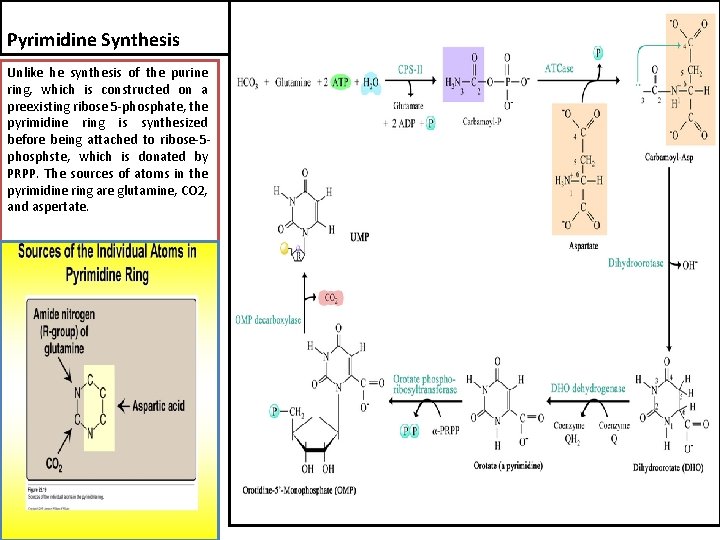
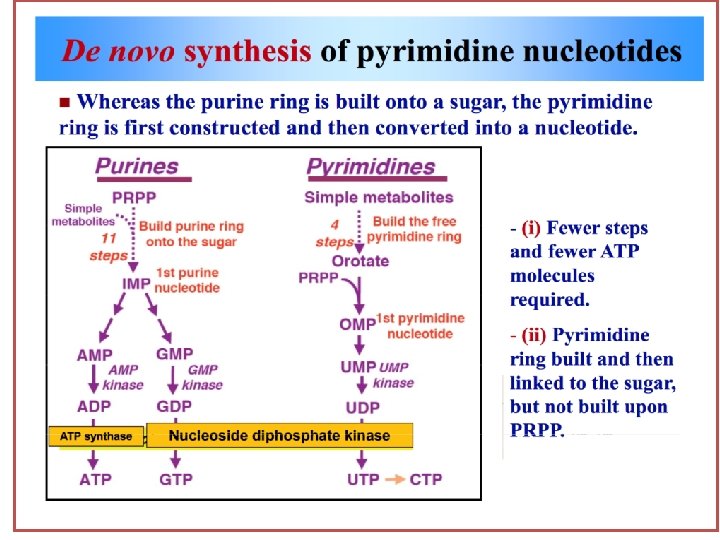
Pyrimidine Synthesis Unlike he synthesis of the purine ring, which is constructed on a preexisting ribose 5 -phosphate, the pyrimidine ring is synthesized before being attached to ribose-5 phosphste, which is donated by PRPP. The sources of atoms in the pyrimidine ring are glutamine, CO 2, and aspertate.
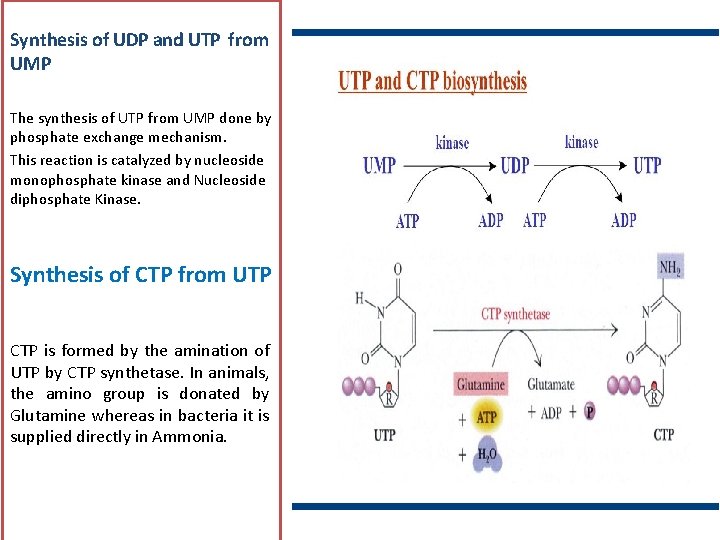
Synthesis of UDP and UTP from UMP The synthesis of UTP from UMP done by phosphate exchange mechanism. This reaction is catalyzed by nucleoside monophosphate kinase and Nucleoside diphosphate Kinase. Synthesis of CTP from UTP CTP is formed by the amination of UTP by CTP synthetase. In animals, the amino group is donated by Glutamine whereas in bacteria it is supplied directly in Ammonia.

UMP to d. UMP • Deoxyuridine monophosphate (d. UMP) is the deoxygenated form of uridine monophosphate (UMP), and is the precursor to deoxythymine monophosphate (d. TMP), a component of DNA nucleotide biosynthesis. By replacing the hydroxyl group at the 2' carbon of ribose with a hydrogen, UMP becomes deoxygenated to d. UMP.
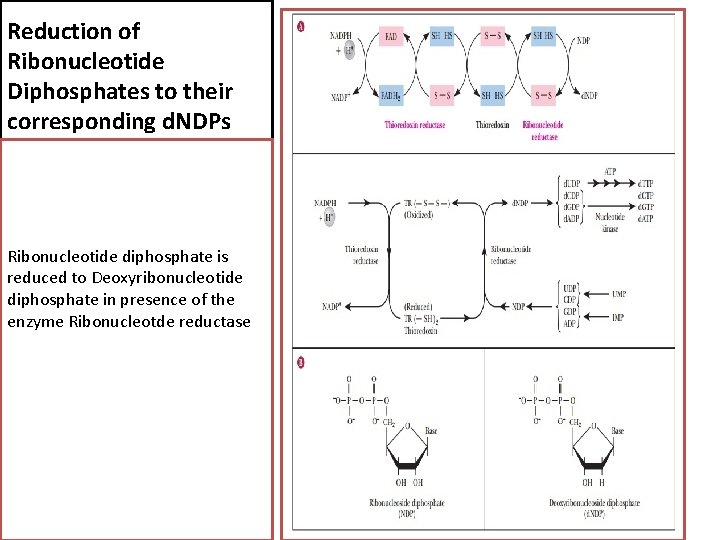
Reduction of Ribonucleotide Diphosphates to their corresponding d. NDPs Ribonucleotide diphosphate is reduced to Deoxyribonucleotide diphosphate in presence of the enzyme Ribonucleotde reductase
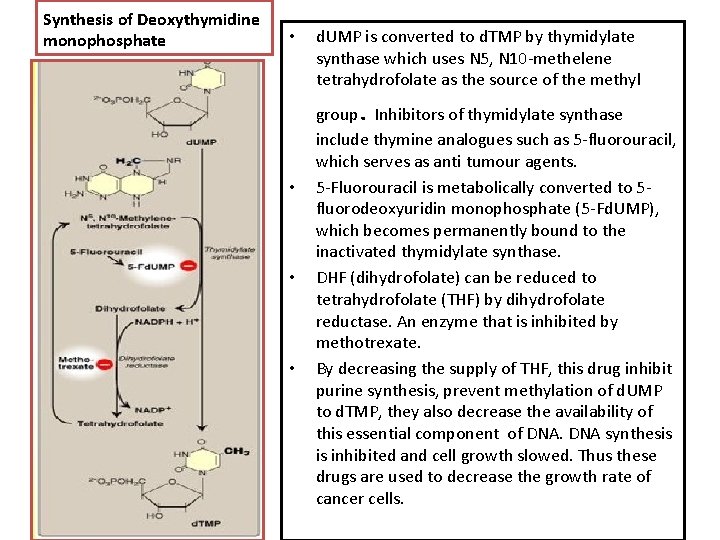
Synthesis of Deoxythymidine monophosphate • d. UMP is converted to d. TMP by thymidylate synthase which uses N 5, N 10 -methelene tetrahydrofolate as the source of the methyl . • • • group Inhibitors of thymidylate synthase include thymine analogues such as 5 -fluorouracil, which serves as anti tumour agents. 5 -Fluorouracil is metabolically converted to 5 fluorodeoxyuridin monophosphate (5 -Fd. UMP), which becomes permanently bound to the inactivated thymidylate synthase. DHF (dihydrofolate) can be reduced to tetrahydrofolate (THF) by dihydrofolate reductase. An enzyme that is inhibited by methotrexate. By decreasing the supply of THF, this drug inhibit purine synthesis, prevent methylation of d. UMP to d. TMP, they also decrease the availability of this essential component of DNA synthesis is inhibited and cell growth slowed. Thus these drugs are used to decrease the growth rate of cancer cells.

Pyrimidine Synthesis • • • ATP+Glutamine + CO 2 carbamoyl phosphate synthetase II carbamoyl phosphate + Aspertate transcarbamolase Carbamoyl aspertate Dihydroorotase--. Dihydroorotate dehydrogenase Orotate phosphoribosyl transferase Orotidine 5 Monophosphate (OMP) OMP decarboxylase Uridine 5 Monophosphate (UMP, the 1 st true pyrimidine ribonucleotide) • UMP Phosphorylation UDP and UTP • • Formation of CTP UTP CTP Synthetase CTP(Nitrogen is provided by glutamine) NDP Ribonucleotide Reductase d. NDP d. UMP Thymidylate synthase d. TMP • •

Pyrimidine base+PRPP Pyrimidine phosphoribosyl transferase Pyrimidine nucleotide+Pi


- Slides: 42