Nucleic Acids Chemistry Structure Andy Howard Introductory Biochemistry

Nucleic Acids: Chemistry & Structure Andy Howard Introductory Biochemistry 8 October 2013 Biochemistry: Nucleic Acids II 10/08/2013
DNA & RNA structure & function n DNA and RNA are dynamic molecules, but understanding their structural realities helps us understand how they work 10/08/2013 Biochemistry: Nucleic Acids II p. 2 of 51
What we’ll discuss n n n DNA duplexes and helicity DNA sequencing DNA structure n n Characterizations B, A, and Z-DNA n DNA, continued n n n Dynamics Function RNA: structure & types n n m. RNA t. RNA r. RNA Small RNAs 10/08/2013 Biochemistry: Nucleic Acids II p. 3 of 51
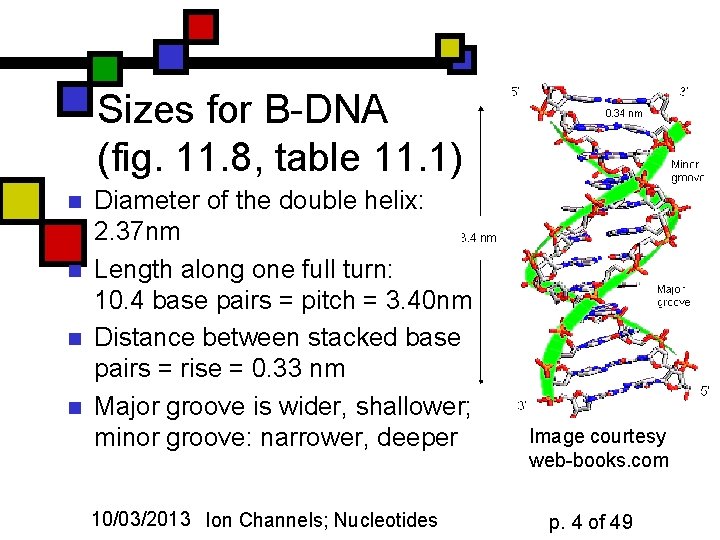
Sizes for B-DNA (fig. 11. 8, table 11. 1) n n Diameter of the double helix: 2. 37 nm Length along one full turn: 10. 4 base pairs = pitch = 3. 40 nm Distance between stacked base pairs = rise = 0. 33 nm Major groove is wider, shallower; minor groove: narrower, deeper 10/03/2013 Ion Channels; Nucleotides Image courtesy web-books. com p. 4 of 49
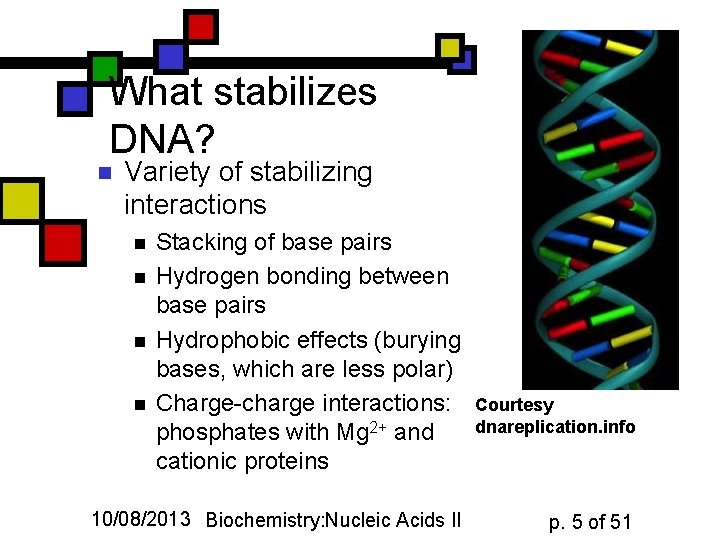
What stabilizes DNA? n Variety of stabilizing interactions n n Stacking of base pairs Hydrogen bonding between base pairs Hydrophobic effects (burying bases, which are less polar) Charge-charge interactions: phosphates with Mg 2+ and cationic proteins 10/08/2013 Biochemistry: Nucleic Acids II Courtesy dnareplication. info p. 5 of 51

How close to instability is it? n n Pretty close. Heating DNA makes it melt: fig. 11. 14 p. H > 10 separates strands too The more GC pairs, the harder it is to melt DNA thermally n n Weaker stacking interactions in A-T One more H-bond per GC than per AT 10/08/2013 Biochemistry: Nucleic Acids II p. 6 of 51
Supercoiling n n Refers to levels of organization of DNA beyond the immediate double-helix We describe circular DNA as relaxed if the closed double helix could lie flat It’s underwound or overwound if the ends are broken, twisted, and rejoined. Supercoils restore 10. 4 bp/turn relation upon rejoining 10/08/2013 Biochemistry: Nucleic Acids II p. 7 of 51
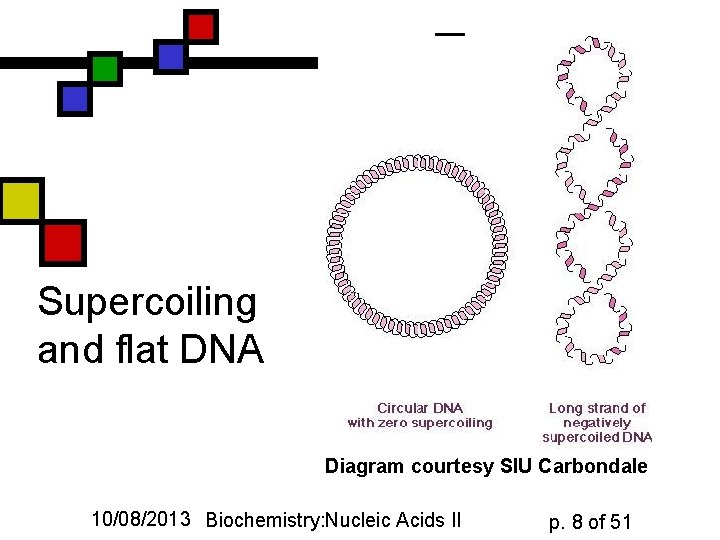
Supercoiling and flat DNA Diagram courtesy SIU Carbondale 10/08/2013 Biochemistry: Nucleic Acids II p. 8 of 51
i. Clicker quiz, 1 st question 1. Which type of ion-channel gating have we not discussed? n (a) ligand-gating n (b) substrate-gating n (c) magnetic-field gating n (d) voltage-gating n (e) we discussed all of these 10/08/2013 Biochemistry: Nucleic Acids II p. 9 of 51
i. Clicker quiz, question 2 2. We would expect the central pore of a cation channel to be lined with n (a) hydrophobic residues n (b) asp and glu residues n (c) lys and arg residues n (d) cofactors n (e) none of the above. 10/08/2013 Biochemistry: Nucleic Acids II p. 10 of 51
i. Clicker quiz, question 3 3. Which of the following atom types are absent in nucleotides? n (a) P and S n (b) S and K n (c) N and O n (d) P and H n (e) all of these atom types are found in nucleotides. 10/08/2013 Biochemistry: Nucleic Acids II p. 11 of 51
i. Clicker quiz, 4 th question 4. What positions of a pair of aromatic rings leads to stabilizing interactions? n n n (a) Parallel to one another (b) Perpendicular to one another (c) At a 45º angle to one another (d) Both (a) and (b) (e) All three: (a), (b), and ( c) 10/08/2013 Biochemistry: Nucleic Acids II p. 12 of 51

i. Clicker question 5 n 5. Which has the highest molecular mass among the compounds listed? n n n (a) cytidylate (b) thymidylate (c) adenylate (d) adenosine triphosphate (e) they’re all the same MW 10/08/2013 Biochemistry: Nucleic Acids II p. 13 of 51
Sanger dideoxy method n n Incorporates DNA replication as an analytical tool for determining sequence Uses short primer that attaches to the 3’ end of the ss. DNA, after which a specially engineered DNA polymerase replicates it Each vial includes one dideoxy. XTP and 3 ordinary d. XTPs; the dideoxy. XTP will be incorporated but will halt synthesis because the 3’ position is blocked. See figs. 11. 3 & 11. 4 for how these are read out 10/08/2013 Biochemistry: Nucleic Acids II p. 14 of 51
Automating dideoxy sequencing n n n Laser fluorescence detection allows for primer identification in real time An automated sequencing machine can handle 4500 bases/hour That’s one of the technologies that has made large-scale sequencing projects like the human genome project possible 10/08/2013 Biochemistry: Nucleic Acids II p. 15 of 51
![Base composition for DNA n n As noted, [A]=[T], [C]=[G] because of base pairing Base composition for DNA n n As noted, [A]=[T], [C]=[G] because of base pairing](http://slidetodoc.com/presentation_image_h/32d3bee04480ff3c39dbbfdc3ef205cc/image-16.jpg)
Base composition for DNA n n As noted, [A]=[T], [C]=[G] because of base pairing [A]/[C] etc. not governed by base pairing n n Can vary considerably (table 10. 3) E. coli : [A], [C] about equal Mycobacterium tuberculosis: [C] > 2*[A] Mammals: [C] < 0. 74*[A] 10/08/2013 Biochemistry: Nucleic Acids II p. 16 of 51
DNA secondary structures n If double-stranded DNA were simply a straightlegged ladder: n n n Base pairs would be 0. 6 nm apart Watson-Crick base-pairs have very uniform dimensions because the H-bonds are fixed lengths But water could get to the apolar bases So, in fact, the ladder gets twisted into a helix. The most common helix is B-DNA, but there are others. B-DNA’s properties include: n n Sugar-sugar distance is still 0. 6 nm Helix repeats itself every 3. 4 nm, i. e. 10 bp 10/08/2013 Biochemistry: Nucleic Acids II p. 17 of 51
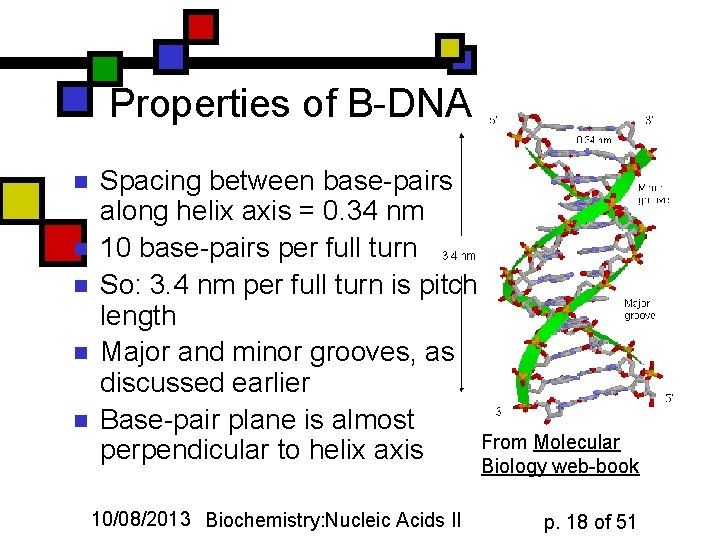
Properties of B-DNA n n n Spacing between base-pairs along helix axis = 0. 34 nm 10 base-pairs per full turn So: 3. 4 nm per full turn is pitch length Major and minor grooves, as discussed earlier Base-pair plane is almost From Molecular perpendicular to helix axis Biology web-book 10/08/2013 Biochemistry: Nucleic Acids II p. 18 of 51
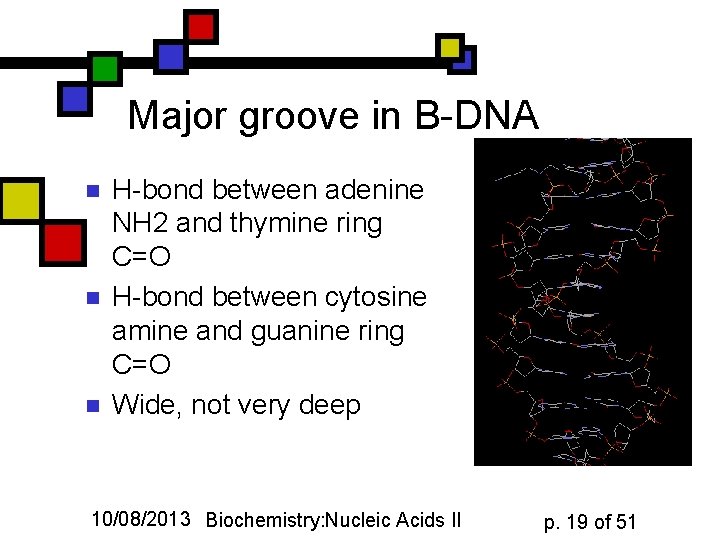
Major groove in B-DNA n n n H-bond between adenine NH 2 and thymine ring C=O H-bond between cytosine amine and guanine ring C=O Wide, not very deep 10/08/2013 Biochemistry: Nucleic Acids II p. 19 of 51
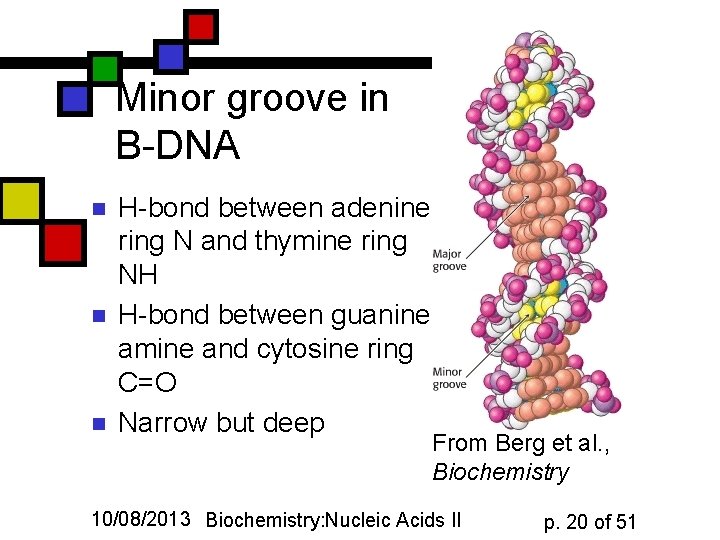
Minor groove in B-DNA n n n H-bond between adenine ring N and thymine ring NH H-bond between guanine amine and cytosine ring C=O Narrow but deep From Berg et al. , Biochemistry 10/08/2013 Biochemistry: Nucleic Acids II p. 20 of 51
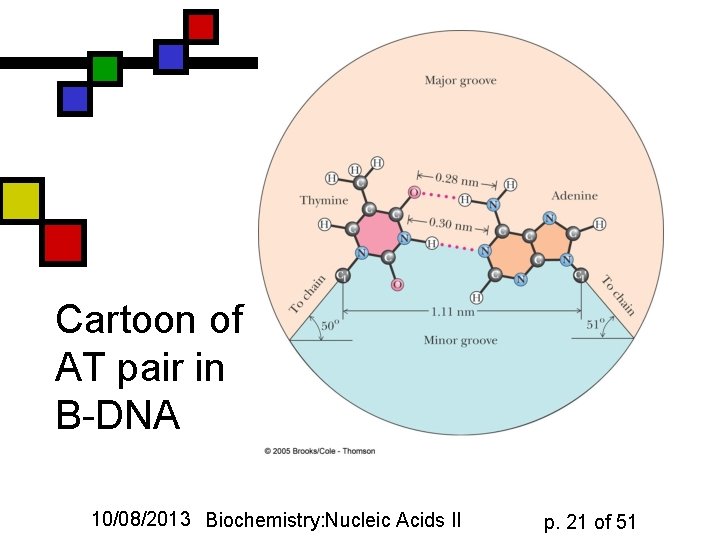
Cartoon of AT pair in B-DNA 10/08/2013 Biochemistry: Nucleic Acids II p. 21 of 51
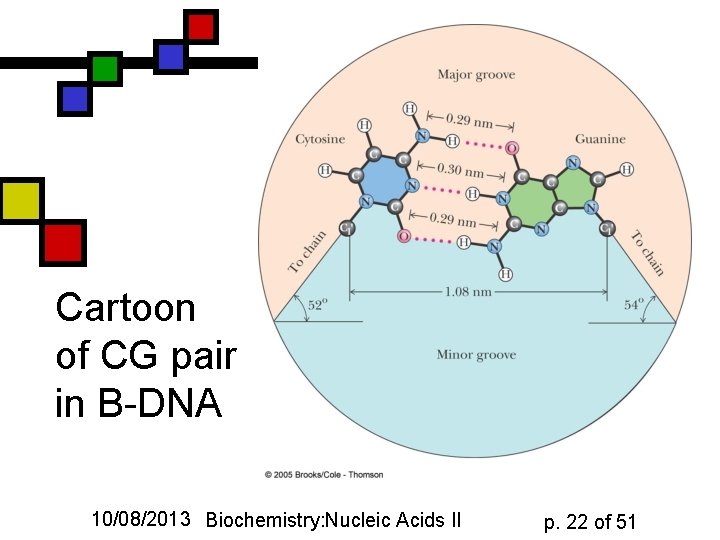
Cartoon of CG pair in B-DNA 10/08/2013 Biochemistry: Nucleic Acids II p. 22 of 51
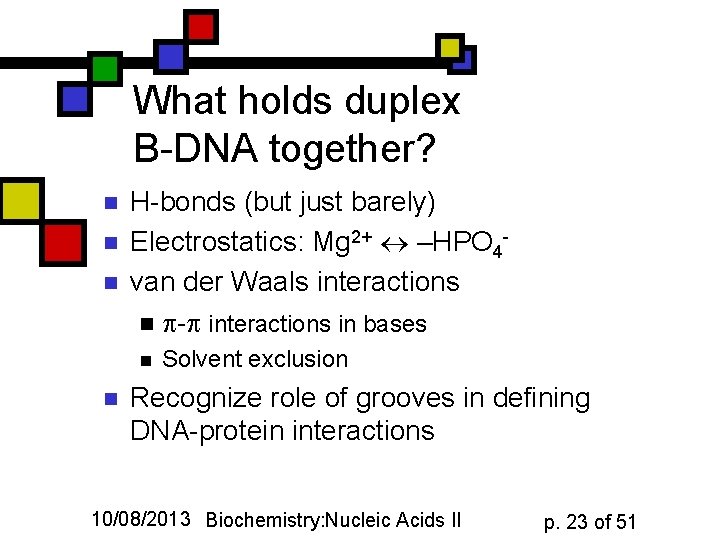
What holds duplex B-DNA together? n n n H-bonds (but just barely) Electrostatics: Mg 2+ –HPO 4 van der Waals interactions n - interactions in bases n n Solvent exclusion Recognize role of grooves in defining DNA-protein interactions 10/08/2013 Biochemistry: Nucleic Acids II p. 23 of 51
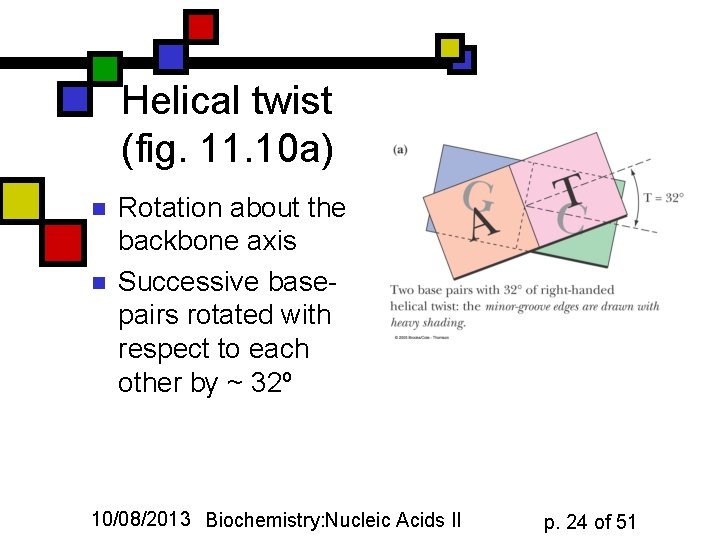
Helical twist (fig. 11. 10 a) n n Rotation about the backbone axis Successive basepairs rotated with respect to each other by ~ 32º 10/08/2013 Biochemistry: Nucleic Acids II p. 24 of 51
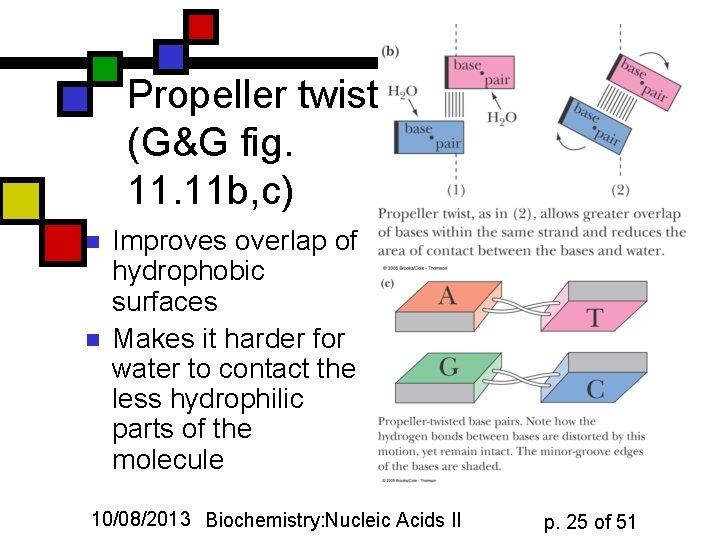
Propeller twist (G&G fig. 11 b, c) n n Improves overlap of hydrophobic surfaces Makes it harder for water to contact the less hydrophilic parts of the molecule 10/08/2013 Biochemistry: Nucleic Acids II p. 25 of 51
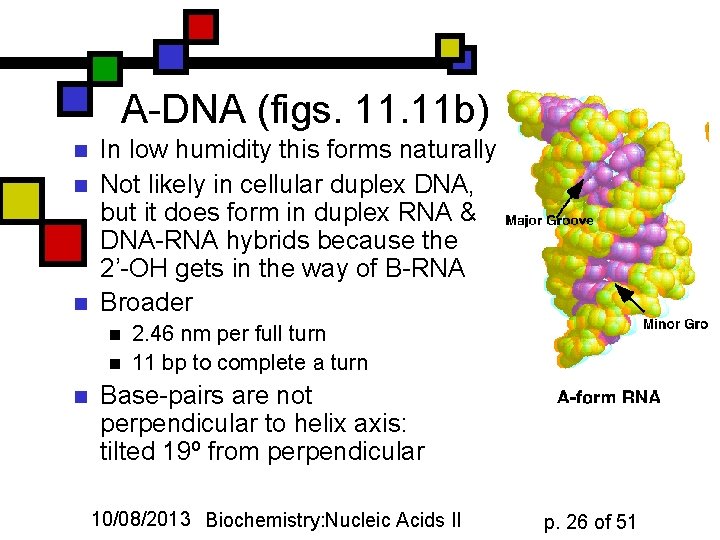
A-DNA (figs. 11 b) n n n In low humidity this forms naturally Not likely in cellular duplex DNA, but it does form in duplex RNA & DNA-RNA hybrids because the 2’-OH gets in the way of B-RNA Broader n n n 2. 46 nm per full turn 11 bp to complete a turn Base-pairs are not perpendicular to helix axis: tilted 19º from perpendicular 10/08/2013 Biochemistry: Nucleic Acids II p. 26 of 51
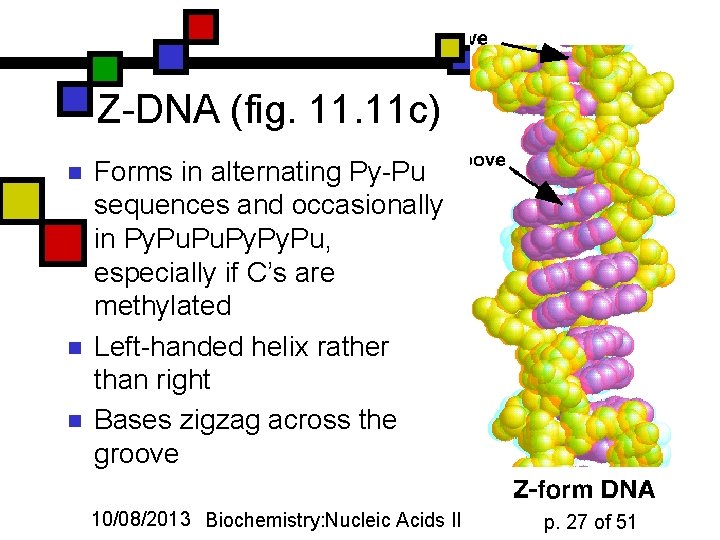
Z-DNA (fig. 11 c) n n n Forms in alternating Py-Pu sequences and occasionally in Py. Pu, especially if C’s are methylated Left-handed helix rather than right Bases zigzag across the groove 10/08/2013 Biochemistry: Nucleic Acids II p. 27 of 51
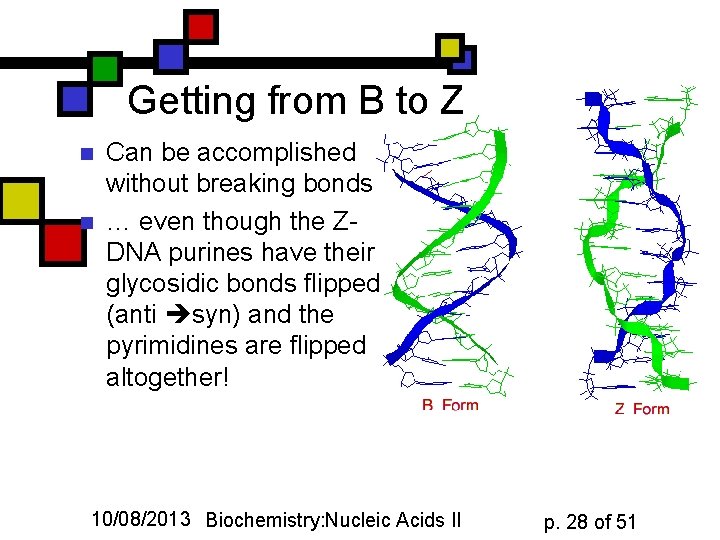
Getting from B to Z n n Can be accomplished without breaking bonds … even though the ZDNA purines have their glycosidic bonds flipped (anti syn) and the pyrimidines are flipped altogether! 10/08/2013 Biochemistry: Nucleic Acids II p. 28 of 51
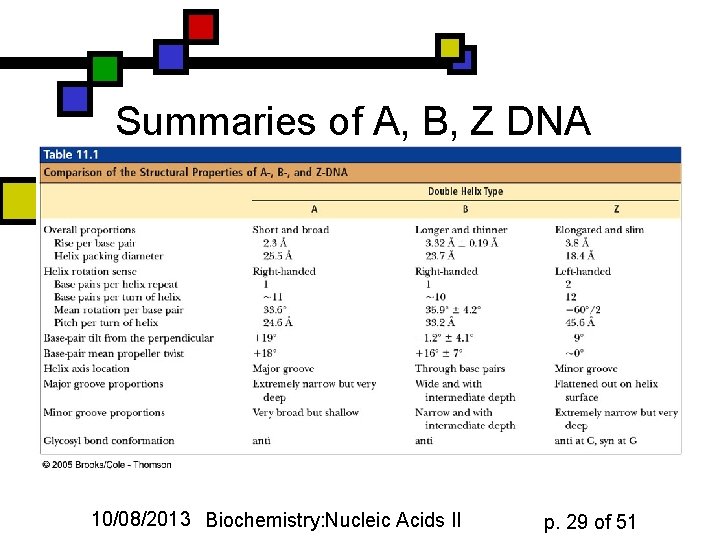
Summaries of A, B, Z DNA 10/08/2013 Biochemistry: Nucleic Acids II p. 29 of 51
DNA is dynamic n n n Don’t think of these diagrams as static The H-bonds stretch and the torsions allow some rotations, so the ropes can form roughly spherical shapes when not constrained by histones Shape is sequence-dependent, which influences protein-DNA interactions 10/08/2013 Biochemistry: Nucleic Acids II p. 30 of 51
What does DNA do? n Serve as the storehouse and the propagator of genetic information: That means that it’s made up of genes n n n Some code for m. RNAs that code for protein Others code for other types of RNA Genes contain non-coding segments (introns) But it also contains stretches that are not parts of genes at all and are serving controlling or structural roles Avoid the term junk DNA! 10/08/2013 Biochemistry: Nucleic Acids II p. 31 of 51
Ribonucleic acid n n We’re done with DNA for the moment. Let’s discuss RNA is generally, but not always, singlestranded The regions where localized base-pairing occurs (local double-stranded regions) often are of functional significance 10/08/2013 Biochemistry: Nucleic Acids II p. 32 of 51
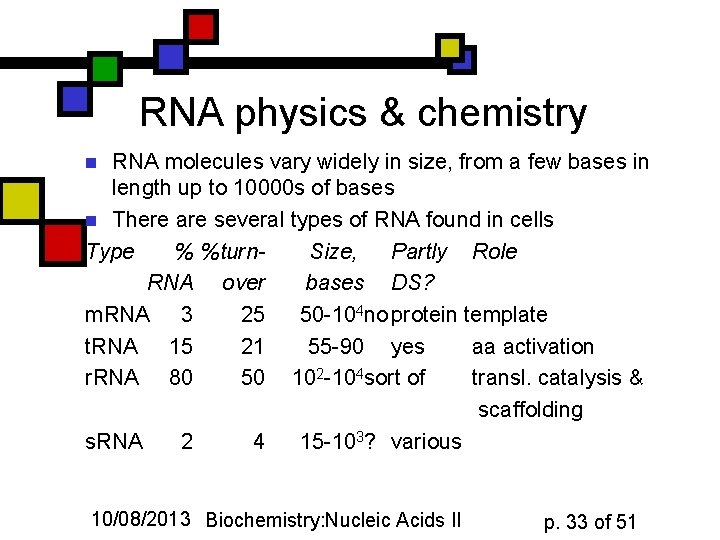
RNA physics & chemistry RNA molecules vary widely in size, from a few bases in length up to 10000 s of bases n There are several types of RNA found in cells Type % %turn. Size, Partly Role RNA over bases DS? m. RNA 3 25 50 -104 no protein template t. RNA 15 21 55 -90 yes aa activation r. RNA 80 50 102 -104 sort of transl. catalysis & scaffolding s. RNA 2 4 15 -103? various n 10/08/2013 Biochemistry: Nucleic Acids II p. 33 of 51
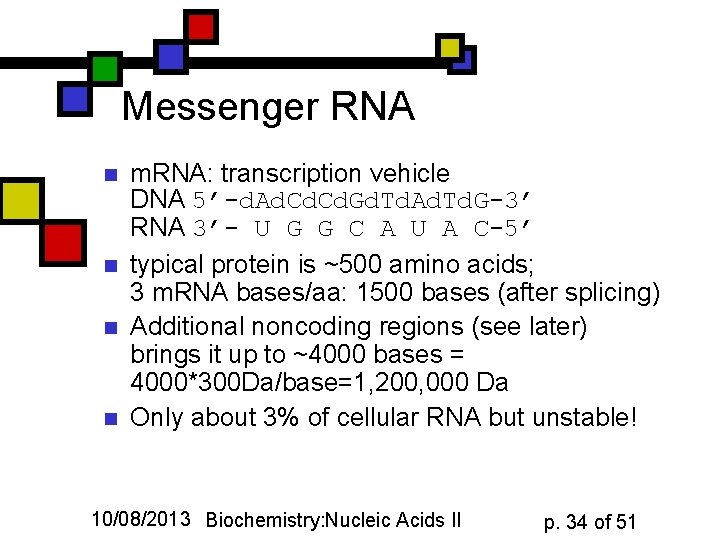
Messenger RNA n n m. RNA: transcription vehicle DNA 5’-d. Ad. Cd. Gd. Td. Ad. Td. G-3’ RNA 3’- U G G C A U A C-5’ typical protein is ~500 amino acids; 3 m. RNA bases/aa: 1500 bases (after splicing) Additional noncoding regions (see later) brings it up to ~4000 bases = 4000*300 Da/base=1, 200, 000 Da Only about 3% of cellular RNA but unstable! 10/08/2013 Biochemistry: Nucleic Acids II p. 34 of 51
Relative quantities n n Note that we said there wasn’t much m. RNA around at any given moment The amount synthesized is much greater because it has a much shorter lifetime than the others Ribonucleases act more avidly on it We need a mechanism for eliminating it because the cell wants to control concentrations of specific proteins 10/08/2013 Biochemistry: Nucleic Acids II p. 35 of 51
m. RNA processing in Eukaryotes Genomic DNA Unmodified m. RNA produced therefrom n n n # bases (unmodified m. RNA) = # base-pairs of DNA in the gene… because that’s how transcription works BUT the number of bases in the unmodified m. RNA > # bases in the final m. RNA that actually codes for a protein SO there needs to be a process for getting rid of the unwanted bases in the m. RNA: that’s what splicing is! 10/08/2013 Biochemistry: Nucleic Acids II p. 36 of 51
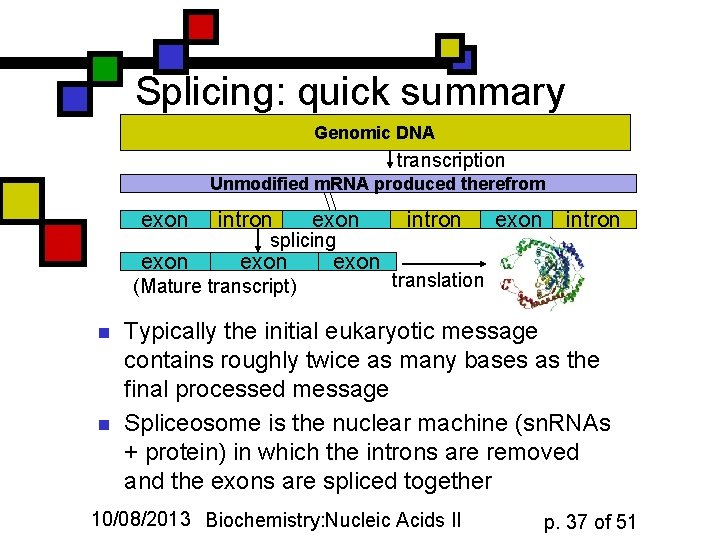
Splicing: quick summary Genomic DNA transcription Unmodified m. RNA produced therefrom exon intron splicing exon (Mature transcript) n n exon intron translation Typically the initial eukaryotic message contains roughly twice as many bases as the final processed message Spliceosome is the nuclear machine (sn. RNAs + protein) in which the introns are removed and the exons are spliced together 10/08/2013 Biochemistry: Nucleic Acids II p. 37 of 51

Heterogeneity via spliceosomal flexibility n n Specific RNA sequences in the initial m. RNA signal where to start and stop each intron, but with some flexibility That flexibility enables a single gene to code for multiple mature RNAs and therefore multiple proteins 10/08/2013 Biochemistry: Nucleic Acids II p. 38 of 51
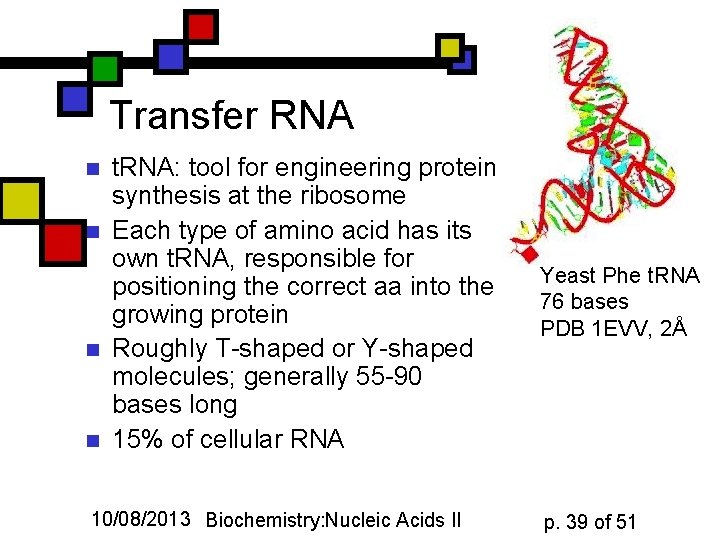
Transfer RNA n n t. RNA: tool for engineering protein synthesis at the ribosome Each type of amino acid has its own t. RNA, responsible for positioning the correct aa into the growing protein Roughly T-shaped or Y-shaped molecules; generally 55 -90 bases long 15% of cellular RNA 10/08/2013 Biochemistry: Nucleic Acids II Yeast Phe t. RNA 76 bases PDB 1 EVV, 2Å p. 39 of 51
Secondary and Tertiary Structure of t. RNA n n Extensive H-bonding creates four double helical domains, three capped by loops, one by a stem Only one t. RNA structure (alone) is known Phenylalanine t. RNA is "L-shaped" Many non-canonical bases found in t. RNA 10/08/2013 Biochemistry: Nucleic Acids II p. 40 of 51
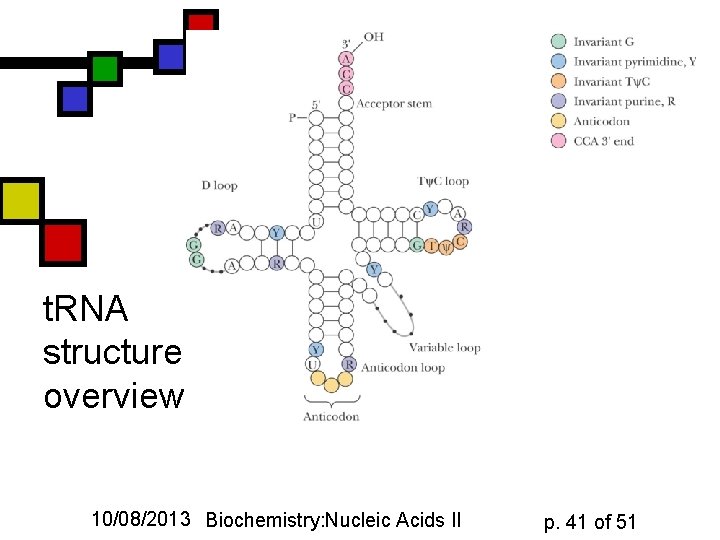
t. RNA structure: overview 10/08/2013 Biochemistry: Nucleic Acids II p. 41 of 51
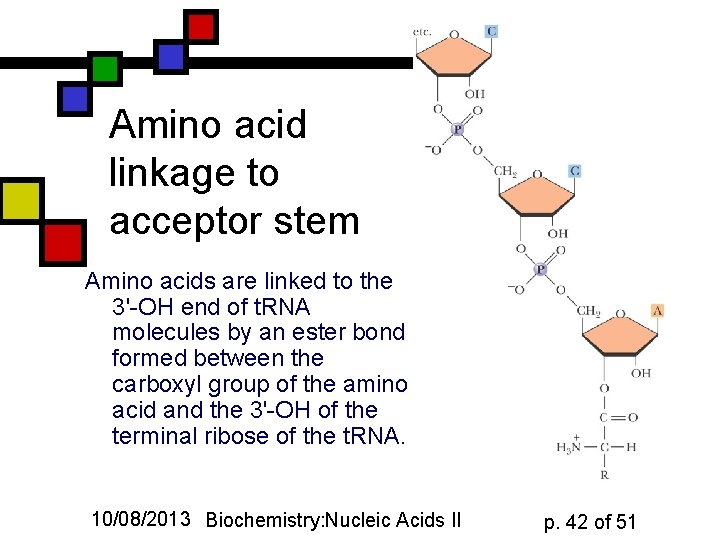
Amino acid linkage to acceptor stem Amino acids are linked to the 3'-OH end of t. RNA molecules by an ester bond formed between the carboxyl group of the amino acid and the 3'-OH of the terminal ribose of the t. RNA. 10/08/2013 Biochemistry: Nucleic Acids II p. 42 of 51
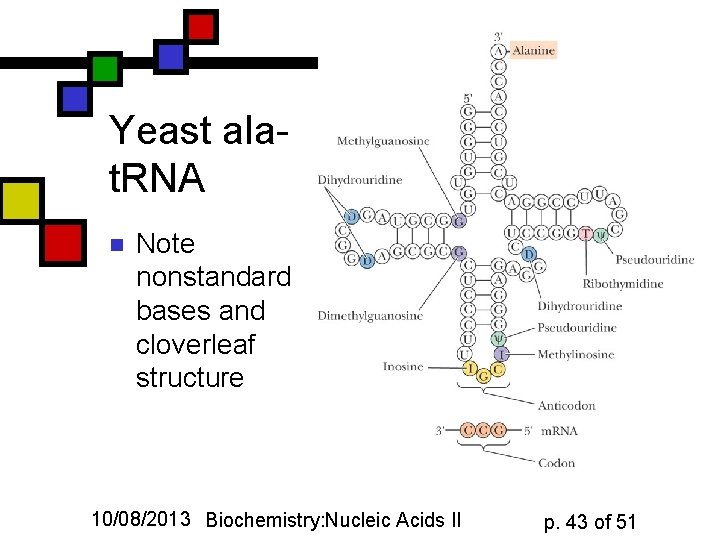
Yeast alat. RNA n Note nonstandard bases and cloverleaf structure 10/08/2013 Biochemistry: Nucleic Acids II p. 43 of 51
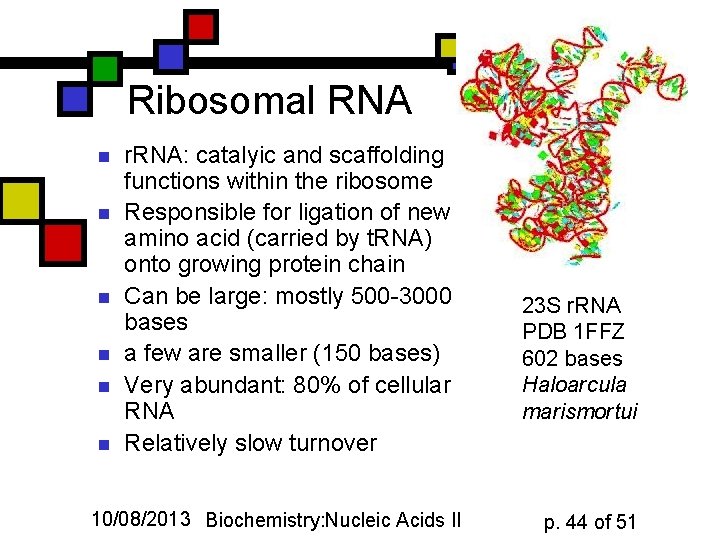
Ribosomal RNA n n n r. RNA: catalyic and scaffolding functions within the ribosome Responsible for ligation of new amino acid (carried by t. RNA) onto growing protein chain Can be large: mostly 500 -3000 bases a few are smaller (150 bases) Very abundant: 80% of cellular RNA Relatively slow turnover 10/08/2013 Biochemistry: Nucleic Acids II 23 S r. RNA PDB 1 FFZ 602 bases Haloarcula marismortui p. 44 of 51
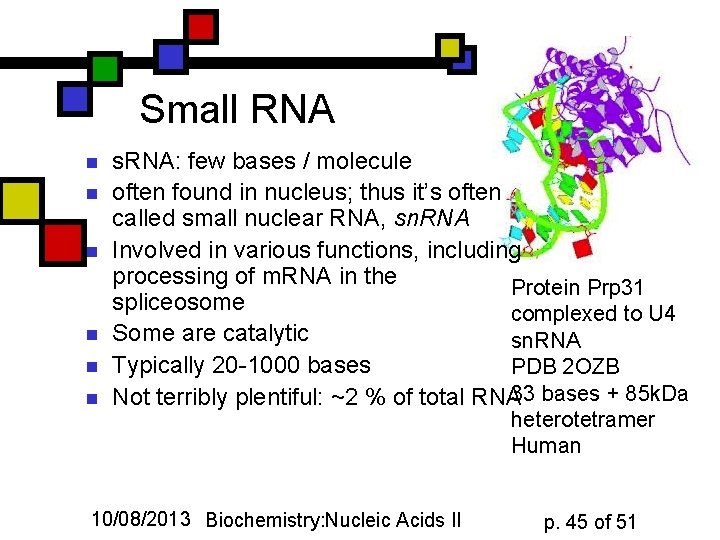
Small RNA n n n s. RNA: few bases / molecule often found in nucleus; thus it’s often called small nuclear RNA, sn. RNA Involved in various functions, including processing of m. RNA in the Protein Prp 31 spliceosome complexed to U 4 Some are catalytic sn. RNA Typically 20 -1000 bases PDB 2 OZB Not terribly plentiful: ~2 % of total RNA 33 bases + 85 k. Da 10/08/2013 Biochemistry: Nucleic Acids II heterotetramer Human p. 45 of 51
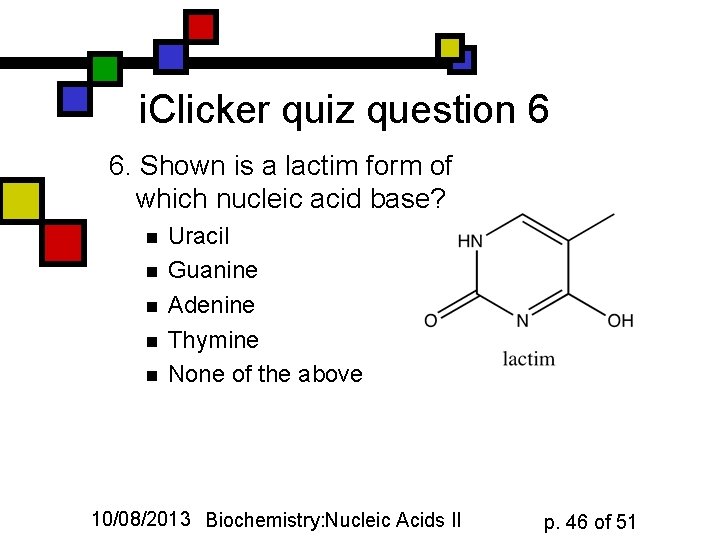
i. Clicker quiz question 6 6. Shown is a lactim form of which nucleic acid base? n n n Uracil Guanine Adenine Thymine None of the above 10/08/2013 Biochemistry: Nucleic Acids II p. 46 of 51

i. Clicker quiz question 7 7. Suppose someone reports that he has characterized the genomic DNA of an organism as having 29% A and 22% T. How would you respond? n n (a) That’s a reasonable result (b) This result is unlikely because [A] ~ [T] in duplex DNA (c) That’s plausible if it’s a bacterium, but not if it’s a eukaryote (d) none of the above 10/08/2013 Biochemistry: Nucleic Acids II p. 47 of 51
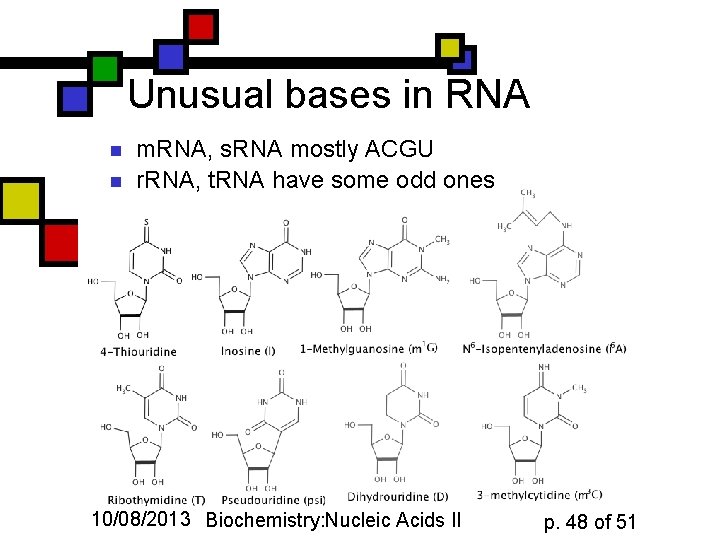
Unusual bases in RNA n n m. RNA, s. RNA mostly ACGU r. RNA, t. RNA have some odd ones 10/08/2013 Biochemistry: Nucleic Acids II p. 48 of 51
Other small RNAs n n n 21 -28 nucleotides Target RNA or DNA through complementary base-pairing Several types, based on function: n n n Small interfering RNAs (q. v. ) micro. RNA: control developmental timing Small nucleolar RNA: catalysts that (among other things) create the oddball bases sno. RNA 77 courtesy Wikipedia 10/08/2013 Biochemistry: Nucleic Acids II p. 49 of 51
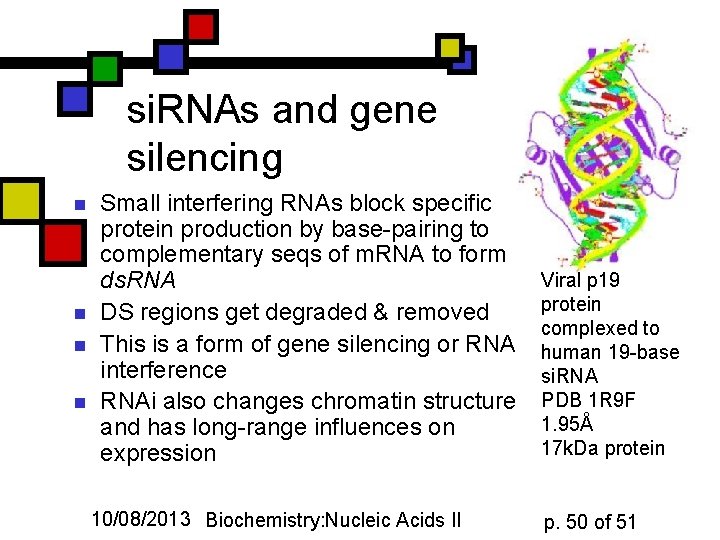
si. RNAs and gene silencing n n Small interfering RNAs block specific protein production by base-pairing to complementary seqs of m. RNA to form ds. RNA DS regions get degraded & removed This is a form of gene silencing or RNA interference RNAi also changes chromatin structure and has long-range influences on expression 10/08/2013 Biochemistry: Nucleic Acids II Viral p 19 protein complexed to human 19 -base si. RNA PDB 1 R 9 F 1. 95Å 17 k. Da protein p. 50 of 51

Do the differences between RNA and DNA matter? Yes! n n Deoxy sugars don’t degrade as readily! DNA has deoxythymidine, RNA has uridine: n n cytidine spontaneously degrades to uridine d. C spontaneously degrades to d. U The only d. U found in DNA is there because of degradation: d. T goes with d. A So when a cell finds d. U in its DNA, it knows it should replace it with d. C or else synthesize d. G opposite the d. U instead of d. A 10/08/2013 Biochemistry: Nucleic Acids II p. 51 of 51
- Slides: 51