Nucleic Acid Structure II Andy Howard Introductory Biochemistry

Nucleic Acid Structure II Andy Howard Introductory Biochemistry 22 October 2014 Nucleic Acid Structure II 10/22/2014
DNA structure informs its functions n We will revisit several aspects of nucleic acid structure that help us understand how it operates. 10/22/2014 Nucleic Acid Structure II p. 2 of 43
What we’ll discuss n RNA n n Types Functions Splicing DNA tertiary structure n n n Nucleosomes Higher-level structures Bacterial organization 10/22/2014 Nucleic Acid Structure II p. 3 of 43
Ribonucleic acid n n We’re done with DNA for the moment. Let’s discuss RNA is generally, but not always, single-stranded The regions where localized basepairing occurs (local double-stranded regions) often are of functional significance 10/22/2014 Nucleic Acid Structure II p. 4 of 43
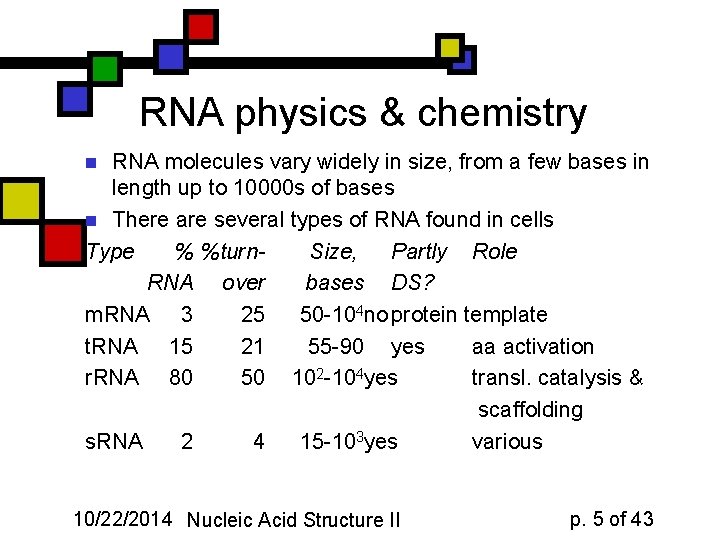
RNA physics & chemistry RNA molecules vary widely in size, from a few bases in length up to 10000 s of bases n There are several types of RNA found in cells Type % %turn. Size, Partly Role RNA over bases DS? m. RNA 3 25 50 -104 no protein template t. RNA 15 21 55 -90 yes aa activation r. RNA 80 50 102 -104 yes transl. catalysis & scaffolding s. RNA 2 4 15 -103 yes various n 10/22/2014 Nucleic Acid Structure II p. 5 of 43
Messenger RNA n n m. RNA: transcription vehicle DNA 5’-d. Ad. Cd. Gd. Td. Ad. Td. G-3’ RNA 3’- U G G C A U A C-5’ typical protein is ~500 amino acids; 3 m. RNA bases/aa: 1500 bases (after splicing) Additional noncoding regions (see later) brings it up to ~4000 bases = 4000*300 Da/base=1, 200, 000 Da Only about 3% of cellular RNA but unstable! 10/22/2014 Nucleic Acid Structure II p. 6 of 43
Relative quantities n n Note that we said there wasn’t much m. RNA around at any given moment The amount synthesized is much greater because it has a much shorter lifetime than the others Ribonucleases act more avidly on it We need a mechanism for eliminating it because the cell wants to control concentrations of specific proteins 10/22/2014 Nucleic Acid Structure II p. 7 of 43
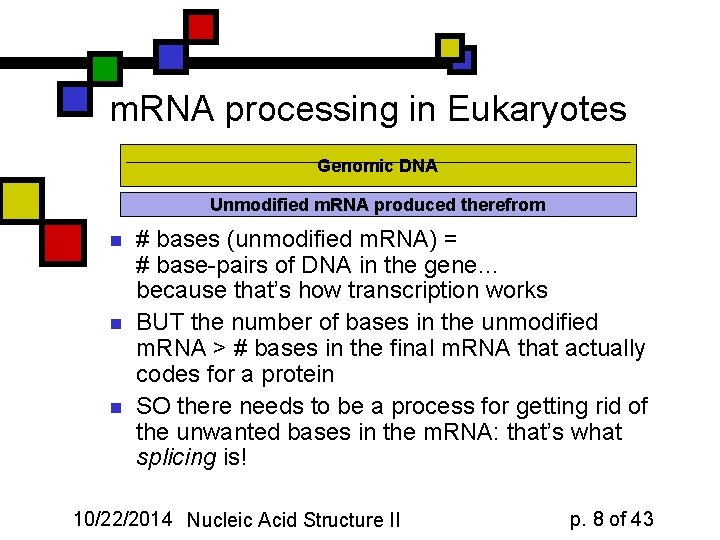
m. RNA processing in Eukaryotes Genomic DNA Unmodified m. RNA produced therefrom n n n # bases (unmodified m. RNA) = # base-pairs of DNA in the gene… because that’s how transcription works BUT the number of bases in the unmodified m. RNA > # bases in the final m. RNA that actually codes for a protein SO there needs to be a process for getting rid of the unwanted bases in the m. RNA: that’s what splicing is! 10/22/2014 Nucleic Acid Structure II p. 8 of 43
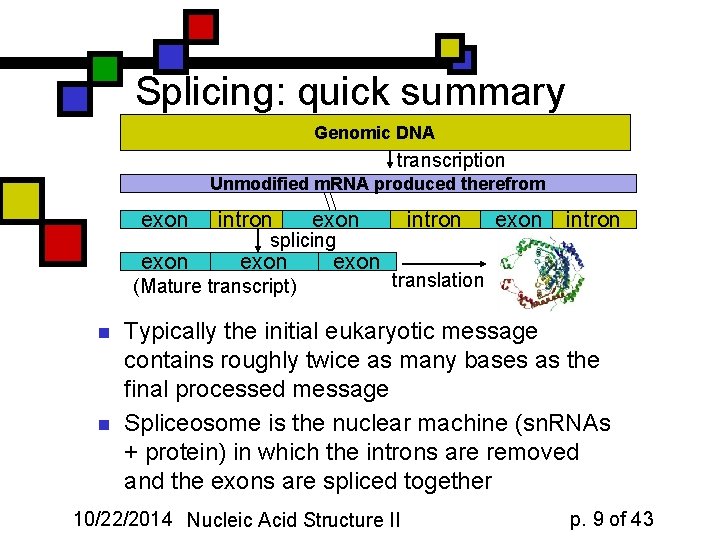
Splicing: quick summary Genomic DNA transcription Unmodified m. RNA produced therefrom exon intron splicing exon (Mature transcript) n exon intron translation Typically the initial eukaryotic message contains roughly twice as many bases as the final processed message Spliceosome is the nuclear machine (sn. RNAs + protein) in which the introns are removed and the exons are spliced together 10/22/2014 Nucleic Acid Structure II p. 9 of 43

Heterogeneity via spliceosomal flexibility n n Specific RNA sequences in the initial m. RNA signal where to start and stop each intron, but with some flexibility That flexibility enables a single gene to code for multiple mature RNAs and therefore multiple proteins 10/22/2014 Nucleic Acid Structure II p. 10 of 43
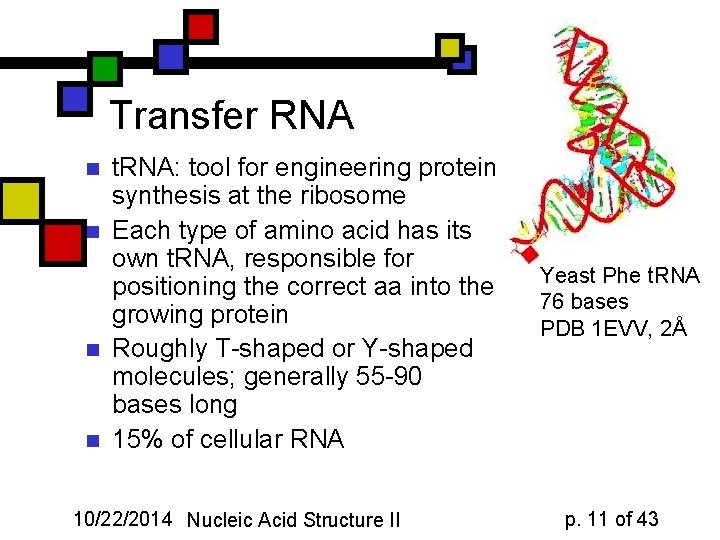
Transfer RNA n n t. RNA: tool for engineering protein synthesis at the ribosome Each type of amino acid has its own t. RNA, responsible for positioning the correct aa into the growing protein Roughly T-shaped or Y-shaped molecules; generally 55 -90 bases long 15% of cellular RNA 10/22/2014 Nucleic Acid Structure II Yeast Phe t. RNA 76 bases PDB 1 EVV, 2Å p. 11 of 43

Secondary and Tertiary Structure of t. RNA n n Extensive H-bonding creates four double helical domains, three capped by loops, one by a stem Only one t. RNA structure (alone) is known Phenylalanine t. RNA is "L-shaped" Many non-canonical bases found in t. RNA 10/22/2014 Nucleic Acid Structure II p. 12 of 43
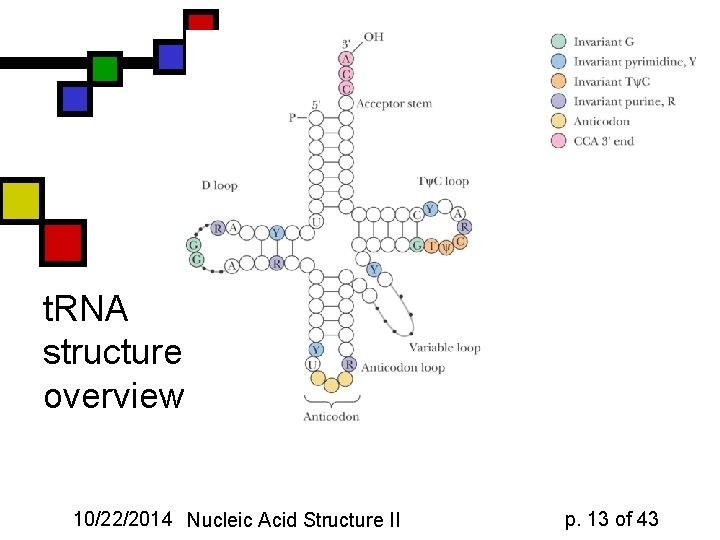
t. RNA structure: overview 10/22/2014 Nucleic Acid Structure II p. 13 of 43
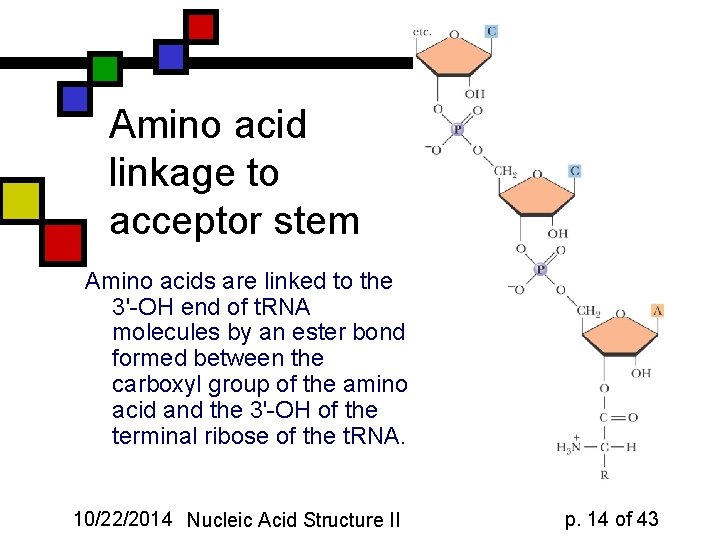
Amino acid linkage to acceptor stem Amino acids are linked to the 3'-OH end of t. RNA molecules by an ester bond formed between the carboxyl group of the amino acid and the 3'-OH of the terminal ribose of the t. RNA. 10/22/2014 Nucleic Acid Structure II p. 14 of 43
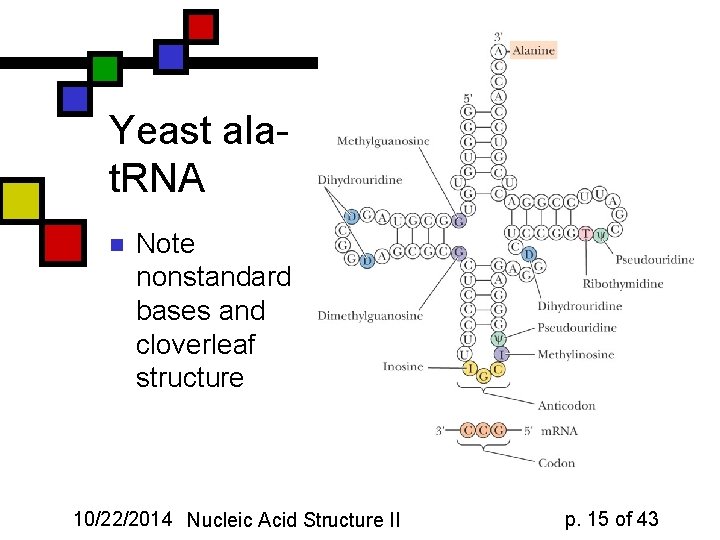
Yeast alat. RNA n Note nonstandard bases and cloverleaf structure 10/22/2014 Nucleic Acid Structure II p. 15 of 43
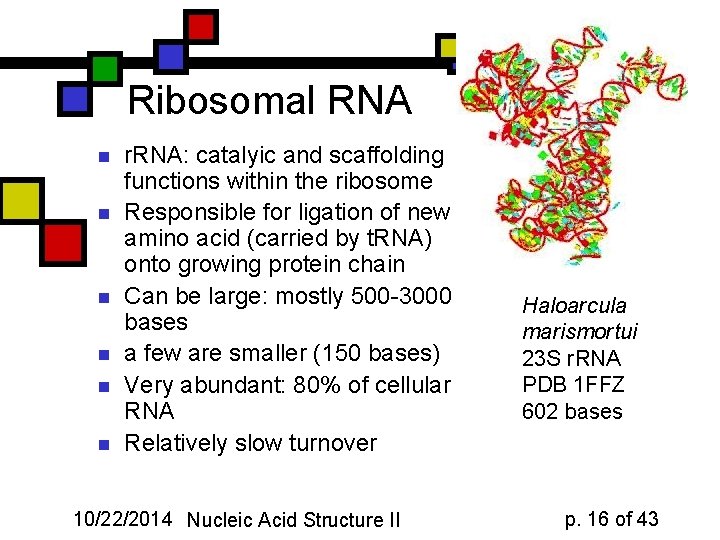
Ribosomal RNA n n n r. RNA: catalyic and scaffolding functions within the ribosome Responsible for ligation of new amino acid (carried by t. RNA) onto growing protein chain Can be large: mostly 500 -3000 bases a few are smaller (150 bases) Very abundant: 80% of cellular RNA Relatively slow turnover 10/22/2014 Nucleic Acid Structure II Haloarcula marismortui 23 S r. RNA PDB 1 FFZ 602 bases p. 16 of 43
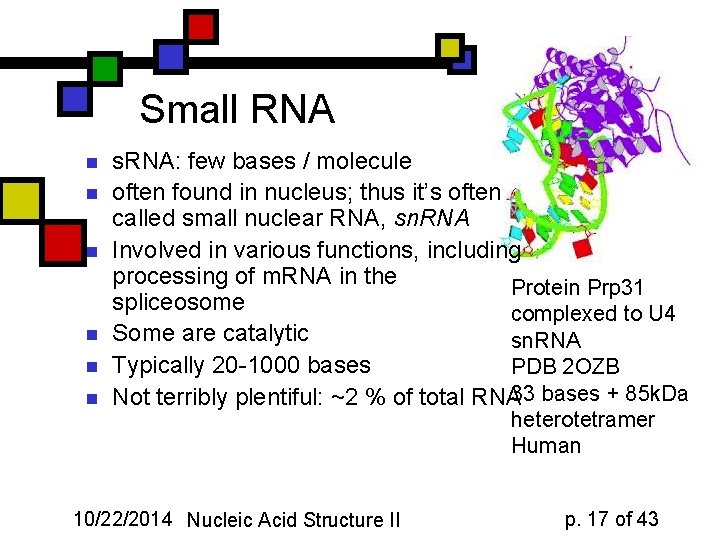
Small RNA n n n s. RNA: few bases / molecule often found in nucleus; thus it’s often called small nuclear RNA, sn. RNA Involved in various functions, including processing of m. RNA in the Protein Prp 31 spliceosome complexed to U 4 Some are catalytic sn. RNA Typically 20 -1000 bases PDB 2 OZB Not terribly plentiful: ~2 % of total RNA 33 bases + 85 k. Da 10/22/2014 Nucleic Acid Structure II heterotetramer Human p. 17 of 43
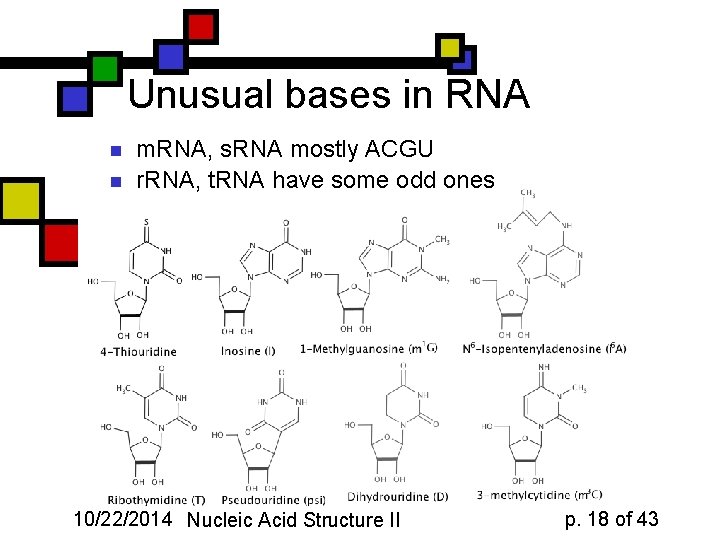
Unusual bases in RNA n n m. RNA, s. RNA mostly ACGU r. RNA, t. RNA have some odd ones 10/22/2014 Nucleic Acid Structure II p. 18 of 43
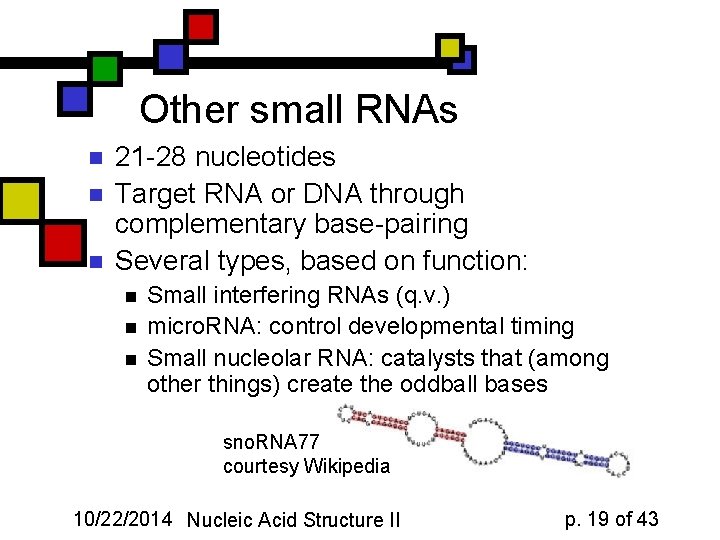
Other small RNAs n n n 21 -28 nucleotides Target RNA or DNA through complementary base-pairing Several types, based on function: n n n Small interfering RNAs (q. v. ) micro. RNA: control developmental timing Small nucleolar RNA: catalysts that (among other things) create the oddball bases sno. RNA 77 courtesy Wikipedia 10/22/2014 Nucleic Acid Structure II p. 19 of 43
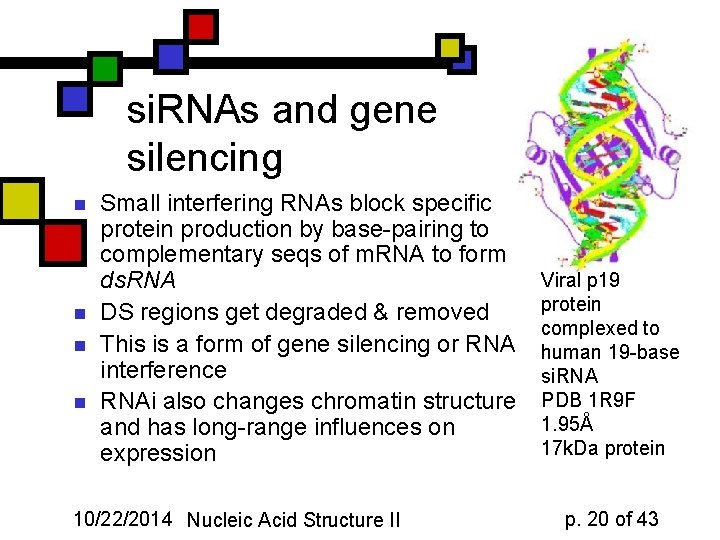
si. RNAs and gene silencing n n Small interfering RNAs block specific protein production by base-pairing to complementary seqs of m. RNA to form ds. RNA DS regions get degraded & removed This is a form of gene silencing or RNA interference RNAi also changes chromatin structure and has long-range influences on expression 10/22/2014 Nucleic Acid Structure II Viral p 19 protein complexed to human 19 -base si. RNA PDB 1 R 9 F 1. 95Å 17 k. Da protein p. 20 of 43
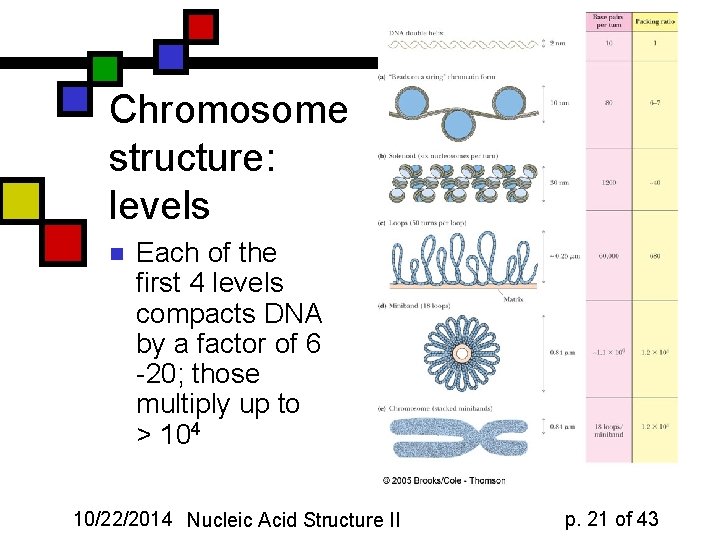
Chromosome structure: levels n Each of the first 4 levels compacts DNA by a factor of 6 -20; those multiply up to > 104 10/22/2014 Nucleic Acid Structure II p. 21 of 43
Nucleosome Structure n n n Chromatin, the nucleoprotein complex, consists of histones and nonhistone chromosomal proteins Histone octamer structure has been solved without DNA: Moudrianakis, 1991 with DNA by Richmond Nonhistone proteins are regulators of gene expression 10/22/2014 Nucleic Acid Structure II p. 22 of 43
Histone types n n n H 2 a, H 2 b, H 3, H 4 make up core particle, associating with ~146 bp of DNA: two copies of each, so: protein octamer All histones are KR-rich, small proteins H 1 associates with ~54 bp region between two nucleosomes 10/22/2014 Nucleic Acid Structure II p. 23 of 43
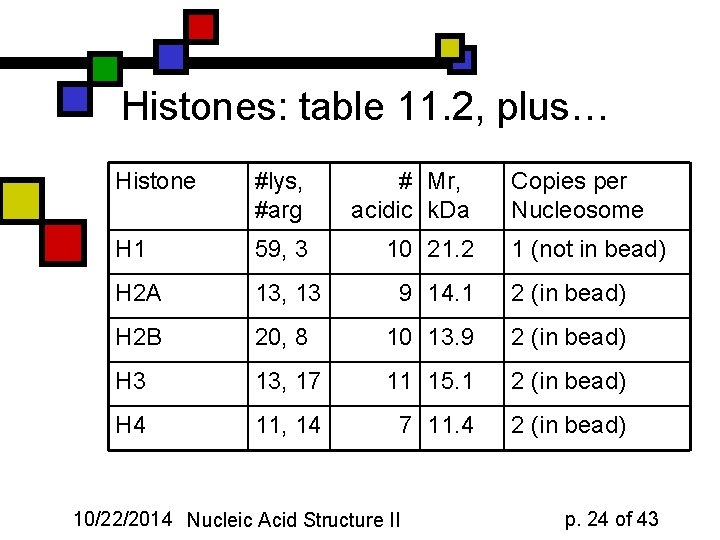
Histones: table 11. 2, plus… Histone #lys, #arg # Mr, acidic k. Da H 1 59, 3 10 21. 2 H 2 A 13, 13 9 14. 1 2 (in bead) H 2 B 20, 8 10 13. 9 2 (in bead) H 3 13, 17 11 15. 1 2 (in bead) H 4 11, 14 7 11. 4 2 (in bead) 10/22/2014 Nucleic Acid Structure II Copies per Nucleosome 1 (not in bead) p. 24 of 43

Unfolded chromatin n Treat chromatin with low ionic strength; that disrupts higher level interactions so the individual nucleosomes are strung out relative to one another like beads on a string Image courtesy U. Maine 10/22/2014 Nucleic Acid Structure II p. 25 of 43
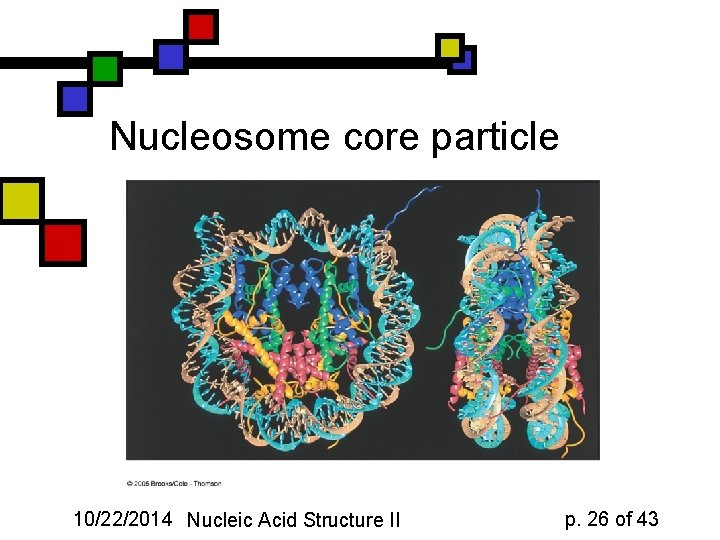
Nucleosome core particle 10/22/2014 Nucleic Acid Structure II p. 26 of 43
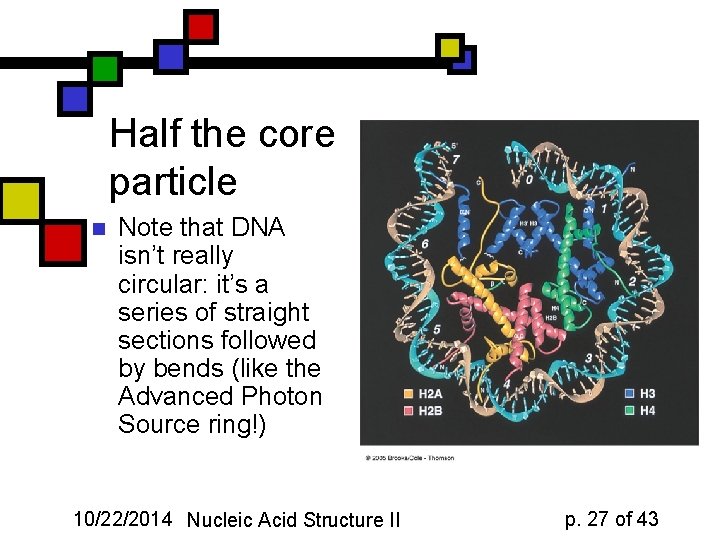
Half the core particle n Note that DNA isn’t really circular: it’s a series of straight sections followed by bends (like the Advanced Photon Source ring!) 10/22/2014 Nucleic Acid Structure II p. 27 of 43
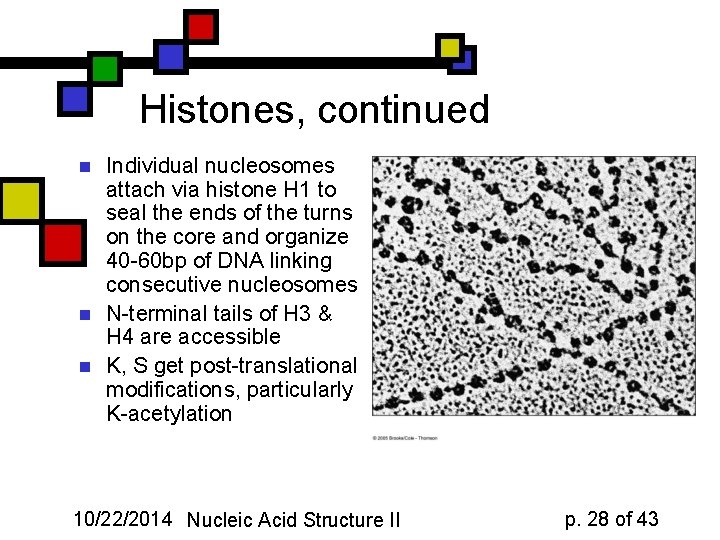
Histones, continued n n n Individual nucleosomes attach via histone H 1 to seal the ends of the turns on the core and organize 40 -60 bp of DNA linking consecutive nucleosomes N-terminal tails of H 3 & H 4 are accessible K, S get post-translational modifications, particularly K-acetylation 10/22/2014 Nucleic Acid Structure II p. 28 of 43
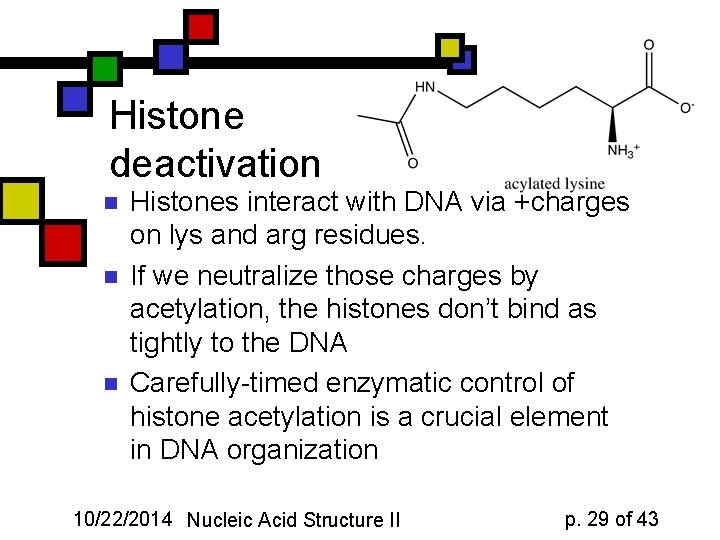
Histone deactivation n Histones interact with DNA via +charges on lys and arg residues. If we neutralize those charges by acetylation, the histones don’t bind as tightly to the DNA Carefully-timed enzymatic control of histone acetylation is a crucial element in DNA organization 10/22/2014 Nucleic Acid Structure II p. 29 of 43
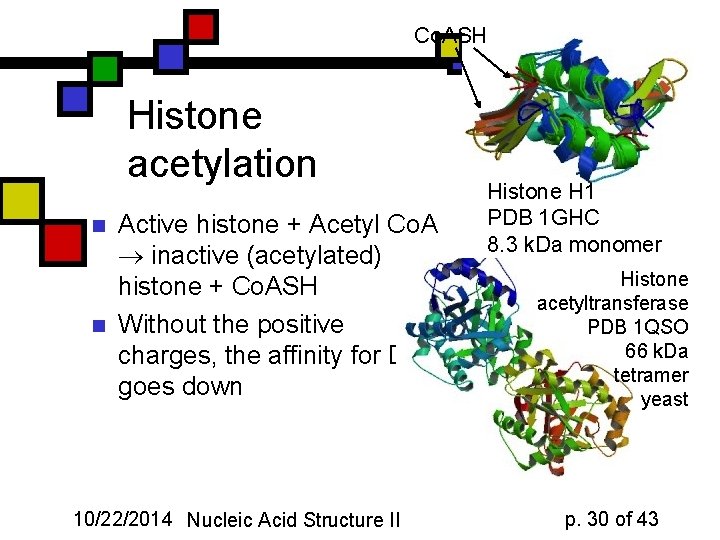
Co. ASH Histone acetylation n n Active histone + Acetyl Co. A inactive (acetylated) histone + Co. ASH Without the positive charges, the affinity for DNA goes down 10/22/2014 Nucleic Acid Structure II Histone H 1 PDB 1 GHC 8. 3 k. Da monomer Chicken Histone acetyltransferase PDB 1 QSO 66 k. Da tetramer yeast p. 30 of 43
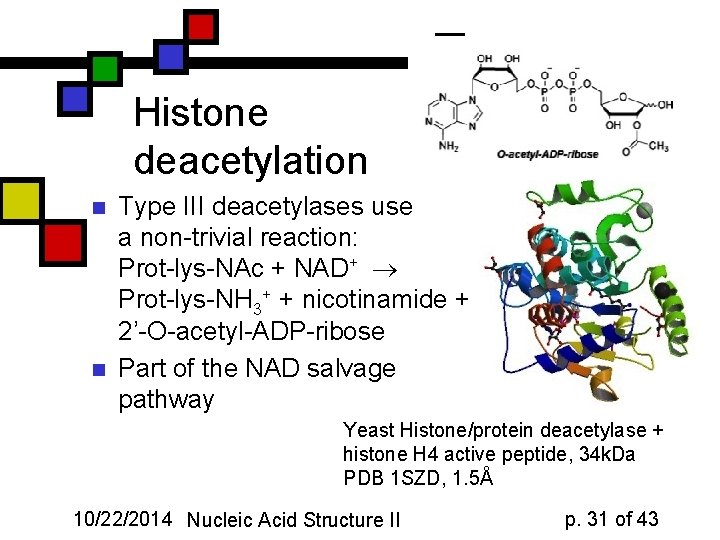
Histone deacetylation n n Type III deacetylases use a non-trivial reaction: Prot-lys-NAc + NAD+ Prot-lys-NH 3+ + nicotinamide + 2’-O-acetyl-ADP-ribose Part of the NAD salvage pathway Yeast Histone/protein deacetylase + histone H 4 active peptide, 34 k. Da PDB 1 SZD, 1. 5Å 10/22/2014 Nucleic Acid Structure II p. 31 of 43
Other histone PTM n Histones can be post-translationally modified in other ways as well n n n Methylation: e. g. lysines 4, 27 of H 3 Phosphorylation: H 2 A phosphorylated at several sites near “hinge” These are correlated with acetylation and play a role in folding and function 10/22/2014 Nucleic Acid Structure II p. 32 of 43
How much does this coil up? n n n 200 bp extended would be about 50 nm The width of the core-particle disk is 5 nm So this is a tenfold reduction Nucleosomal organization corresponds to negative supercoiling … so DNA ends up supercoiled when we take away the histones 10/22/2014 Nucleic Acid Structure II p. 33 of 43
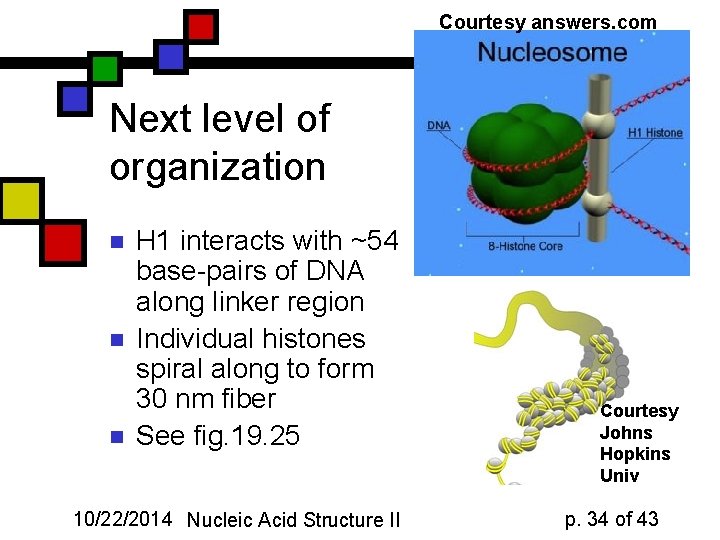
Courtesy answers. com Next level of organization n H 1 interacts with ~54 base-pairs of DNA along linker region Individual histones spiral along to form 30 nm fiber See fig. 19. 25 10/22/2014 Nucleic Acid Structure II Courtesy Johns Hopkins Univ p. 34 of 43
Even higher… n n The 30 nm fibers are attached to an RNA-protein scaffold that holds the 30 nm fibers in large loops Typical chromosome has ~200 loops Loops are attached to scaffold at their base Ends can rotate so it can be supercoiled 10/22/2014 Nucleic Acid Structure II p. 35 of 43
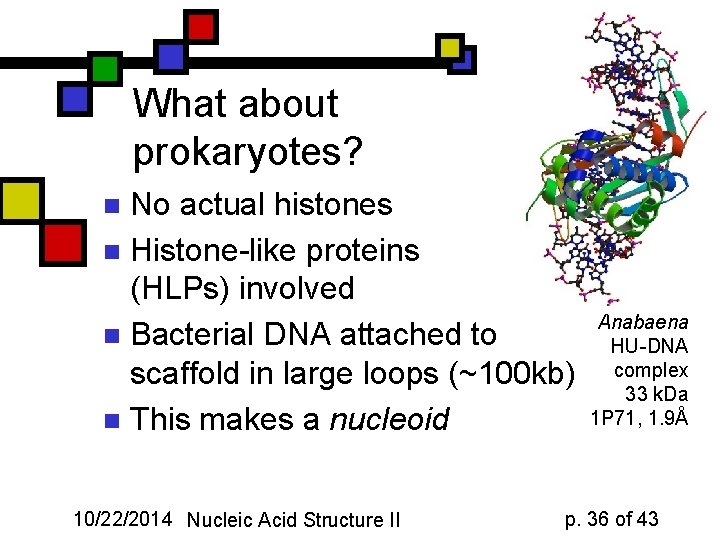
What about prokaryotes? No actual histones n Histone-like proteins (HLPs) involved n Bacterial DNA attached to scaffold in large loops (~100 kb) n This makes a nucleoid n 10/22/2014 Nucleic Acid Structure II Anabaena HU-DNA complex 33 k. Da 1 P 71, 1. 9Å p. 36 of 43
How many loops in bacteria? n n Typical bacterial genome (E. coli) has 3000 open reading frames ~ 3000 genes. Assume 500 amino acids per protein = 1500 bases per gene (ignores transcriptional elements) Then genome is 1500 bp/gene * 3000 genes = 4. 5*106 base-pairs That’s (4. 5*106 bp)/(1*105 bp/loop) = 45 loops 10/22/2014 Nucleic Acid Structure II p. 37 of 43
i. Clicker question 1 n 1. Which of the following is a potential restriction site? (a) ACTTCA n (b) AGCGCT n (c) TGGCCT n (d) AACCGG n (e) none of the above. n 10/22/2014 Nucleic Acid Structure II p. 38 of 43
i. Clicker question 2 n 2. A DNA sample has Tm=94ºC. It is probably (a) AT-rich n (b) CG-rich n (c) characterized in the presence of chelators n (d) measured in pure water rather than buffer n (e) either (c) or (d) n 10/22/2014 Nucleic Acid Structure II p. 39 of 43

i. Clicker question 3 n 3. One step in gyrase activity depends on ATP hydrolysis. It is: (a) association of the gyrase with the DNA loop n (b) DNA cleavage n (c) DNA re-ligation and release of DNA from gyrase n (d) all of the above n 10/22/2014 Nucleic Acid Structure II p. 40 of 43

i. Clicker question 4 n 4. When DNA is compressed, which of the compression steps accomplishes the most substantial reduction in size? (a) helix to beads-on-a-string n (b) beads-on-a-string to solenoid n (c) solenoid to loop n (d) loop to 18 -loop miniband n (e) 18 -loop miniband to chromosome n 10/22/2014 Nucleic Acid Structure II p. 41 of 43

i. Clicker quiz, question 5 n 5. Suppose a mutation in the gene coding for histone H 1 makes it fold up incorrectly. How will this mutation influence DNA organization? n n n (a) It will prevent formation of nucleosomes (b) It will interfere with the beads-on-a-string organization between nucleosomes (c) It will interfere with higher-level organization involving assembly of solenoids into loops (d) All of the above (e) None of the above 10/22/2014 Nucleic Acid Structure II p. 42 of 43
i. Clicker question 6 n n n 6. When a histone becomes acetylated, the net charge on the protein (a) goes up (b) goes down (c) does not change (d) will go up or down depending on the circumstance 10/22/2014 Nucleic Acid Structure II p. 43 of 43
- Slides: 43