Nuclear Magnetic Resonance A Introduction Nuclear Magnetic Resonance















- Slides: 85

Nuclear Magnetic Resonance A. ) Introduction: Nuclear Magnetic Resonance (NMR) measures the absorption of electromagnetic radiation in the radio-frequency region (~4 -900 MHz) - nuclei (instead of outer electrons) are involved in absorption process - sample needs to be placed in magnetic field to cause different energy states NMR was first experimentally observed by Bloch and Purcell in 1946 (received Nobel Prize in 1952) and quickly became commercially available and widely used. Probe the Composition, Structure, Dynamics and Function of the Complete Range of Chemical Entities: from small organic molecules to large molecular weight polymers and proteins. NMR is routinely and widely used as the preferred technique to rapidly elucidate the chemical structure of most organic compounds. One of the MOST Routinely used Analytical Techniques

NMR History 1937 1946 1953 1966 1975 1985 Rabi predicts and observes nuclear magnetic resonance Bloch, Purcell first nuclear magnetic resonance of bulk sample Overhauser NOE (nuclear Overhauser effect) Ernst, Anderson Fourier transform NMR Jeener, Ernst 2 D NMR Wüthrich first solution structure of a small protein (BPTI) from NOE derived distance restraints 1987 3 D NMR + 13 C, 15 N isotope labeling of recombinant proteins (resolution) 1990 pulsed field gradients (artifact suppression) 1996/7 new long range structural parameters: - residual dipolar couplings from partial alignment in liquid crystalline media - projection angle restraints from cross-correlated relaxation TROSY (molecular weight > 100 k. Da) Nobel prizes 1944 Physics Rabi (Columbia) 1952 Physics Bloch (Stanford), Purcell (Harvard) 1991 Chemistry Ernst (ETH) 2002 Chemistry Wüthrich (ETH) 2003 Medicine Lauterbur (University of Illinois in Urbana ), Mansfield (University of Nottingham)

NMR History First NMR Spectra on Water 1 H NMR spectra of water Bloch, F. ; Hansen, W. W. ; Packard, M. The nuclear induction experiment. Physical Review (1946), 70 474 -85.

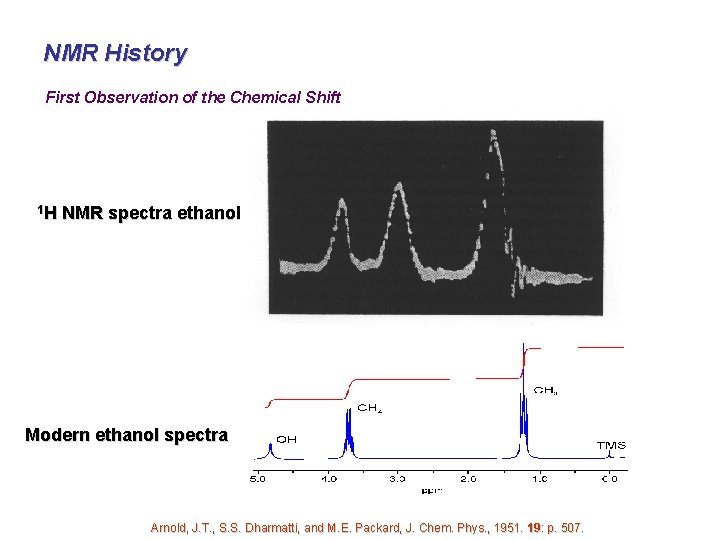
NMR History First Observation of the Chemical Shift 1 H NMR spectra ethanol Modern ethanol spectra Arnold, J. T. , S. S. Dharmatti, and M. E. Packard, J. Chem. Phys. , 1951. 19: p. 507.

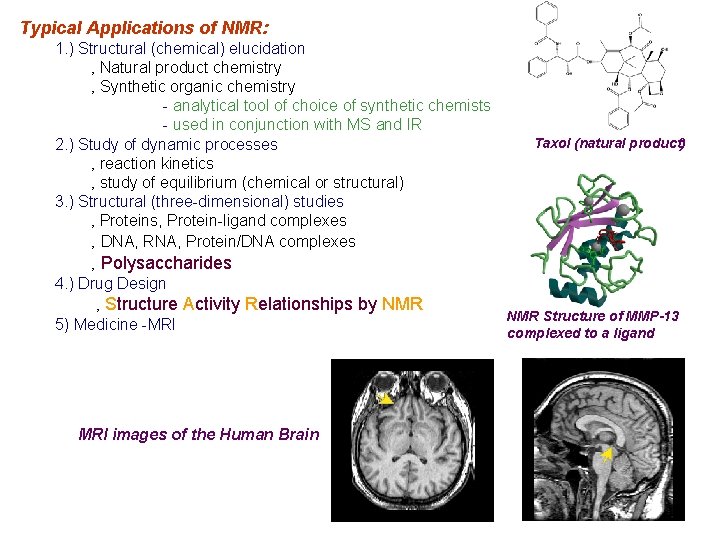
Typical Applications of NMR: 1. ) Structural (chemical) elucidation ‚ Natural product chemistry ‚ Synthetic organic chemistry - analytical tool of choice of synthetic chemists - used in conjunction with MS and IR 2. ) Study of dynamic processes ‚ reaction kinetics ‚ study of equilibrium (chemical or structural) 3. ) Structural (three-dimensional) studies ‚ Proteins, Protein-ligand complexes ‚ DNA, RNA, Protein/DNA complexes ‚ Polysaccharides 4. ) Drug Design ‚ Structure Activity Relationships by NMR 5) Medicine -MRI images of the Human Brain Taxol (natural product) NMR Structure of MMP-13 complexed to a ligand

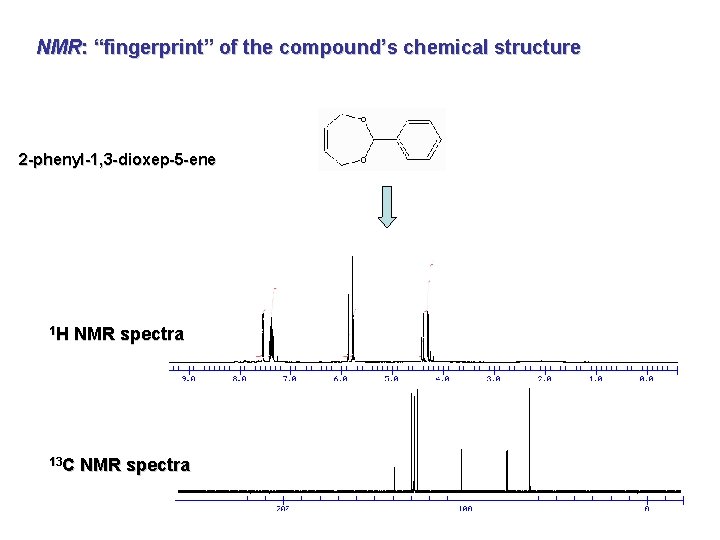
NMR: “fingerprint” of the compound’s chemical structure 2 -phenyl-1, 3 -dioxep-5 -ene 1 H NMR spectra 13 C NMR spectra

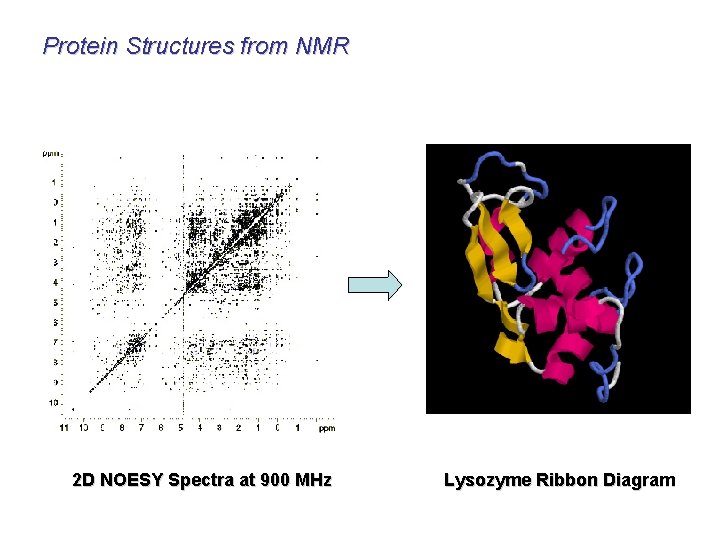
Protein Structures from NMR 2 D NOESY Spectra at 900 MHz Lysozyme Ribbon Diagram

Some Suggested NMR References “Spin Dynamics – Basics of Nuclear Magnetic Resonance” M. H. Levitt “Tables of Spectral Data for Structure Determination of Organic Compounds” Pretsch, Clerc, Seibl and Simon “Basic One- and Two-Dimensional NMR Spectroscopy” Horst Friebolin “Modern NMR Techniques for Chemistry Research” Andrew E. Derome “NMR and Chemistry- an introduction to modern NMR spectroscopy” J. W. Akitt “Nuclear Magnetic Resonance Spectroscopy” R. K Harris “Protein NMR Spectroscopy: Principals and Practice” John Cavanagh, Arthur Palmer, Nicholas J. Skelton, Wayne Fairbrother “Biomolecular NMR Spectroscopy” J. N. S. Evans “NMR of Proteins and Nucleic Acids” Kurt Wuthrich “Spectrometric Identification of Organic Compounds” Silverstein, Bassler and Morrill

Some NMR Web Sites The Basics of NMR Hypertext based NMR course http: //www. cis. rit. edu/htbooks/nmr-main. htm Integrated Spectral Data Base System for Organic Compounds http: //www. aist. go. jp/RIODB/SDBS/menu-e. html Educational NMR Software All kinds of NMR software http: //www. york. ac. uk/depts/chem/services/nmr/edusoft. html NMR Knowledge Base A lot of useful NMR links http: //www. spectroscopynow. com/ NMR Information Server News, Links, Conferences, Jobs http: //www. spincore. com/nmrinfo/ Technical Tidbits Useful source for the art of shimming http: //www. acornnmr. com/nmr_topics. htm BMRB (Bio. Mag. Res. Bank) http: //www. bmrb. wisc. edu/ Database of NMR resonance assignments

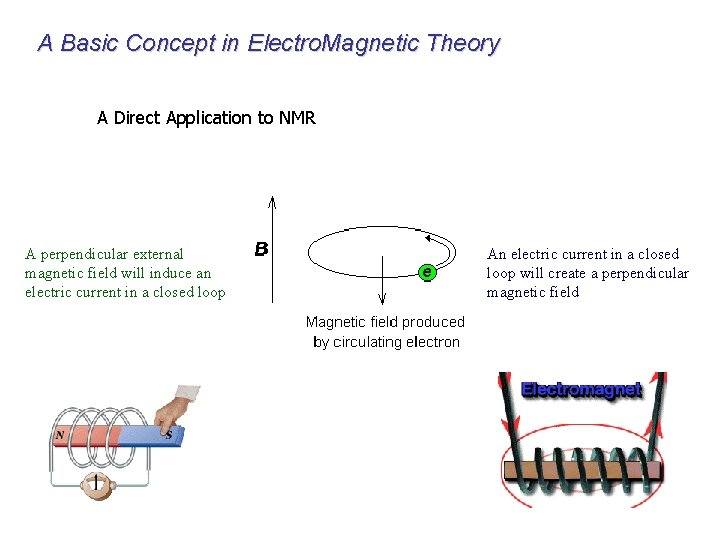
A Basic Concept in Electro. Magnetic Theory A Direct Application to NMR A perpendicular external magnetic field will induce an electric current in a closed loop An electric current in a closed loop will create a perpendicular magnetic field

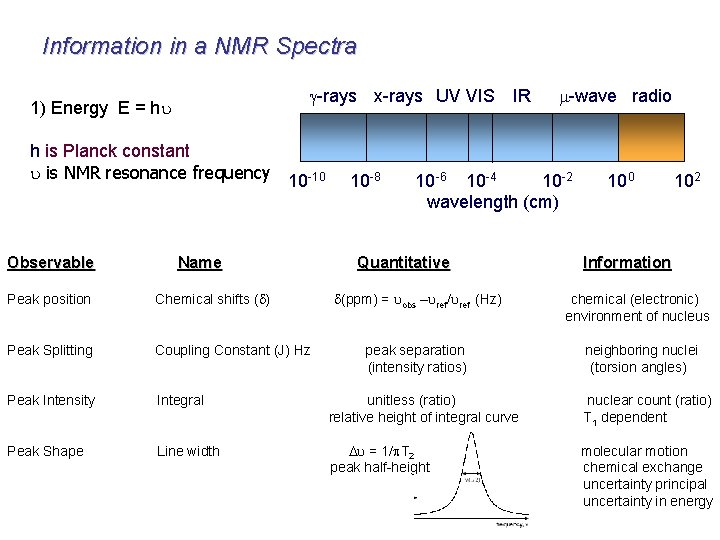
Information in a NMR Spectra g-rays x-rays UV VIS 1) Energy E = hu h is Planck constant u is NMR resonance frequency 10 -10 Observable Name 10 -8 IR m-wave radio 10 -6 10 -4 10 -2 wavelength (cm) Quantitative 100 102 Information d(ppm) = uobs –uref/uref (Hz) chemical (electronic) environment of nucleus peak separation (intensity ratios) neighboring nuclei (torsion angles) Peak position Chemical shifts (d) Peak Splitting Coupling Constant (J) Hz Peak Intensity Integral unitless (ratio) relative height of integral curve nuclear count (ratio) T 1 dependent Peak Shape Line width Du = 1/p. T 2 peak half-height molecular motion chemical exchange uncertainty principal uncertainty in energy

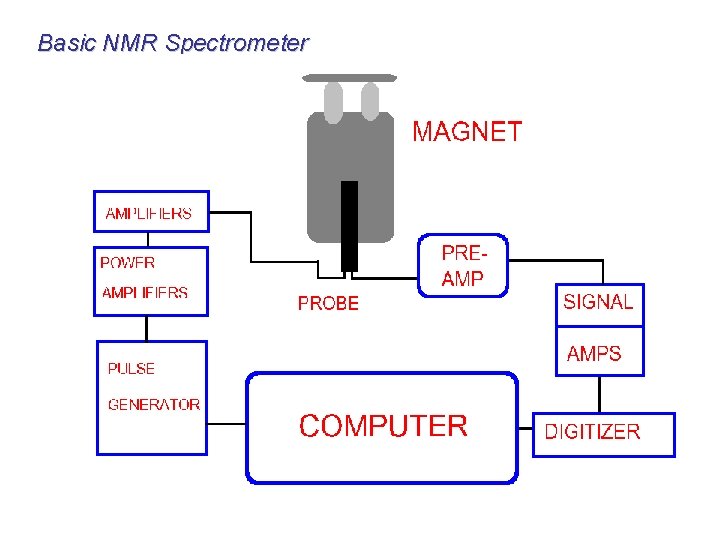
Basic NMR Spectrometer

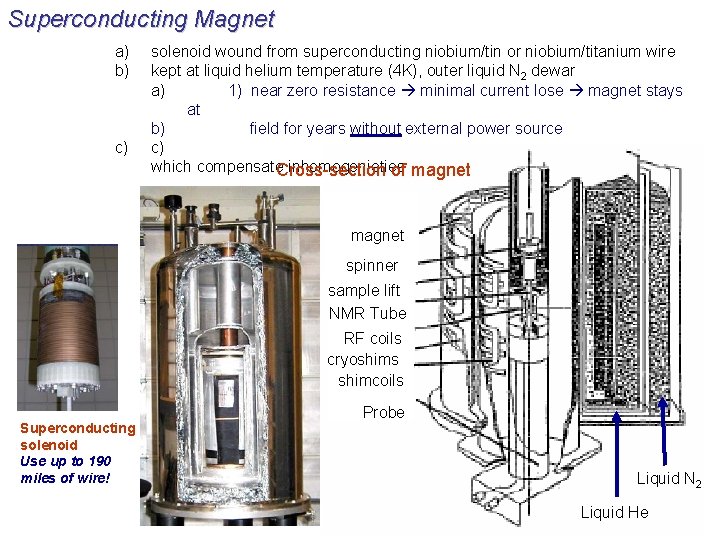
Superconducting Magnet a) b) c) solenoid wound from superconducting niobium/tin or niobium/titanium wire kept at liquid helium temperature (4 K), outer liquid N 2 dewar a) 1) near zero resistance minimal current lose magnet stays at b) field for years without external power source c) which compensate. Cross-section inhomogenieties of magnet spinner sample lift NMR Tube RF coils cryoshims shimcoils Superconducting solenoid Use up to 190 miles of wire! Probe Liquid N 2 Liquid He

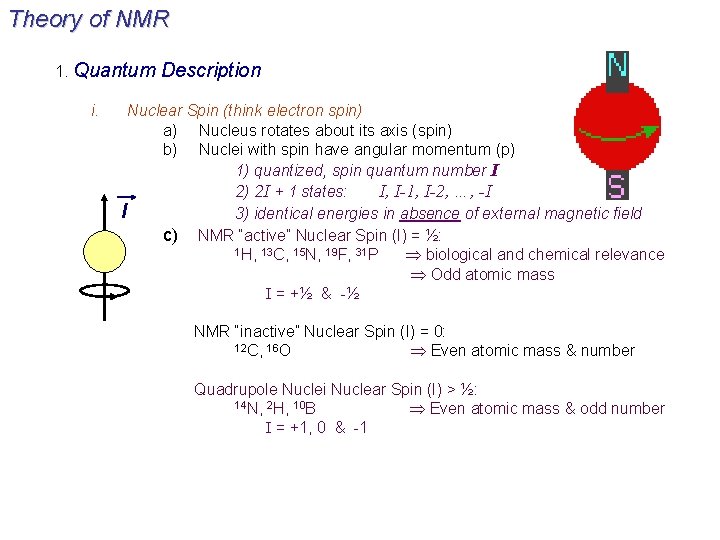
Theory of NMR 1. Quantum i. Description Nuclear Spin (think electron spin) a) Nucleus rotates about its axis (spin) b) Nuclei with spin have angular momentum (p) 1) quantized, spin quantum number I 2) 2 I + 1 states: I, I-1, I-2, …, -I l 3) identical energies in absence of external magnetic field c) NMR “active” Nuclear Spin (I) = ½: 1 H, 13 C, 15 N, 19 F, 31 P biological and chemical relevance Odd atomic mass I = +½ & -½ NMR “inactive” Nuclear Spin (I) = 0: 12 C, 16 O Even atomic mass & number Quadrupole Nuclei Nuclear Spin (I) > ½: 14 N, 2 H, 10 B Even atomic mass & odd number I = +1, 0 & -1

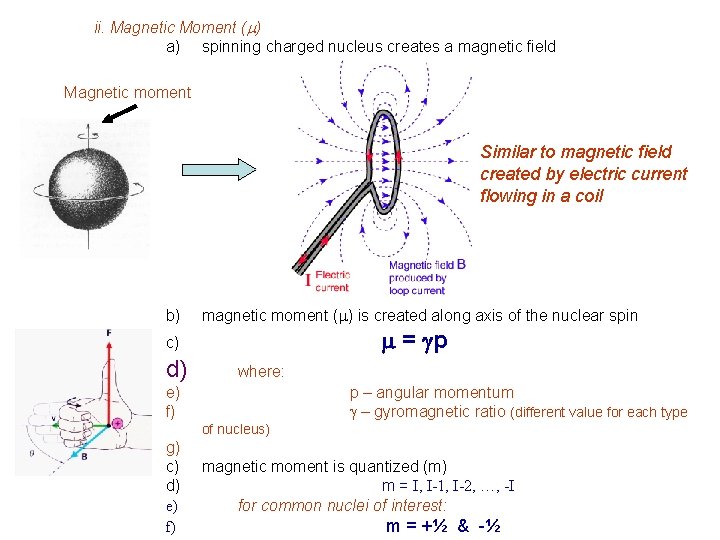
ii. Magnetic Moment (m) a) spinning charged nucleus creates a magnetic field Magnetic moment Similar to magnetic field created by electric current flowing in a coil b) magnetic moment (m) is created along axis of the nuclear spin m = gp c) d) where: e) f) p – angular momentum g – gyromagnetic ratio (different value for each type of nucleus) g) c) d) e) f) magnetic moment is quantized (m) m = I, I-1, I-2, …, -I for common nuclei of interest: m = +½ & -½

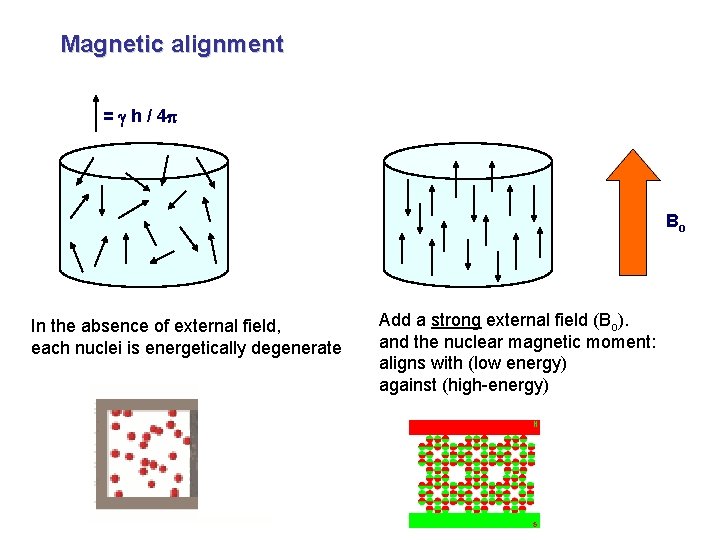
Magnetic alignment = g h / 4 p Bo In the absence of external field, each nuclei is energetically degenerate Add a strong external field (Bo). and the nuclear magnetic moment: aligns with (low energy) against (high-energy)

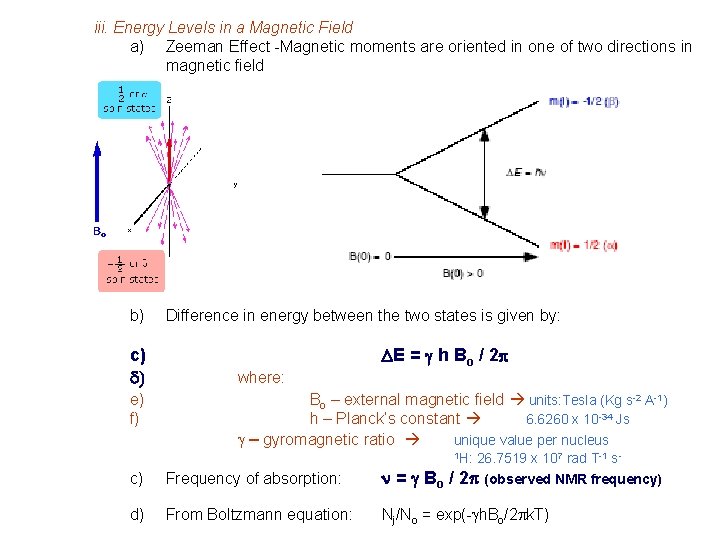
iii. Energy Levels in a Magnetic Field a) Zeeman Effect -Magnetic moments are oriented in one of two directions in magnetic field b) c) d) e) f) Difference in energy between the two states is given by: DE = g h Bo / 2 p where: Bo – external magnetic field units: Tesla (Kg s-2 A-1) h – Planck’s constant 6. 6260 x 10 -34 Js g – gyromagnetic ratio unique value per nucleus 1 H: 26. 7519 x 107 rad T-1 s 2 p (observed NMR frequency) c) Frequency of absorption: n = g Bo / d) From Boltzmann equation: Nj/No = exp(-gh. Bo/2 pk. T)

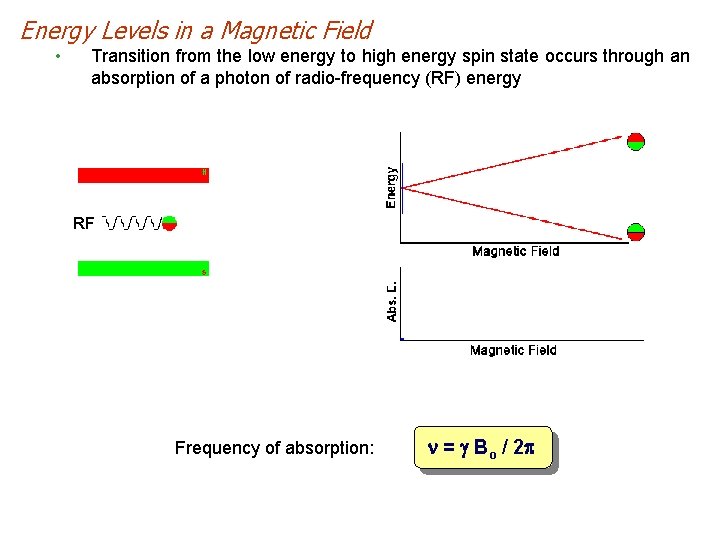
Energy Levels in a Magnetic Field • Transition from the low energy to high energy spin state occurs through an absorption of a photon of radio-frequency (RF) energy RF Frequency of absorption: n = g Bo / 2 p

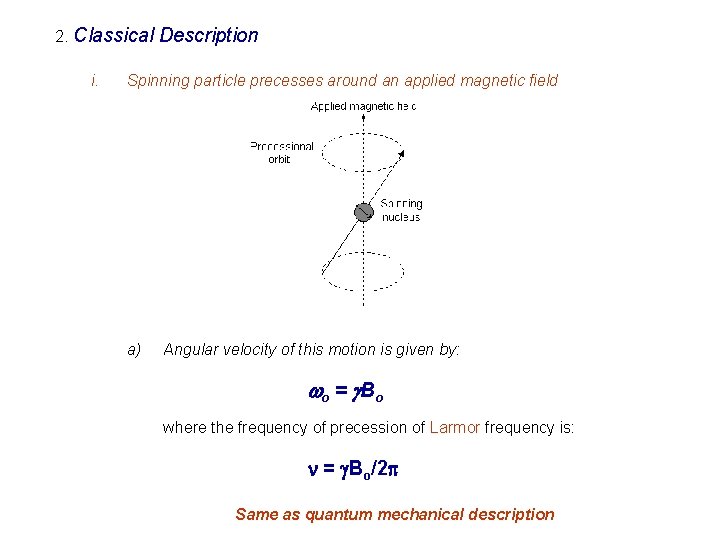
2. Classical i. Description Spinning particle precesses around an applied magnetic field a) Angular velocity of this motion is given by: wo = g B o where the frequency of precession of Larmor frequency is: n = g. Bo/2 p Same as quantum mechanical description

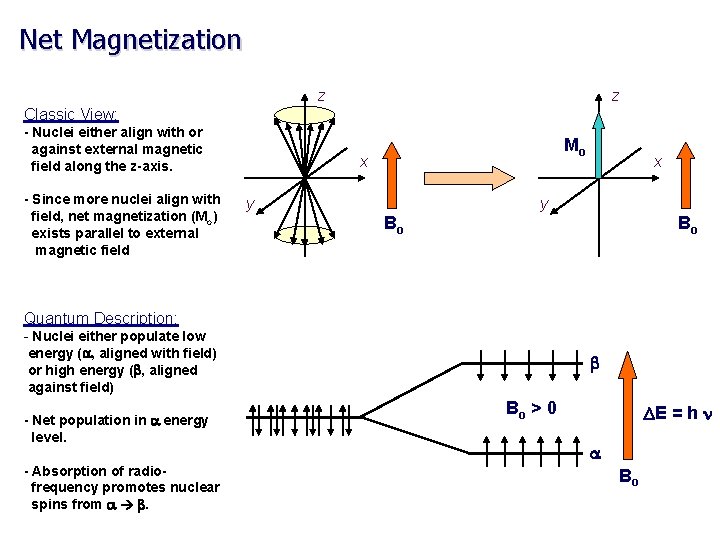
Net Magnetization z z Classic View: - Nuclei either align with or against external magnetic field along the z-axis. - Since more nuclei align with field, net magnetization (Mo) exists parallel to external magnetic field Mo x y Bo Bo Quantum Description: - Nuclei either populate low energy (a, aligned with field) or high energy (b, aligned against field) - Net population in a energy level. - Absorption of radiofrequency promotes nuclear spins from a b. b Bo > 0 DE = h n a Bo

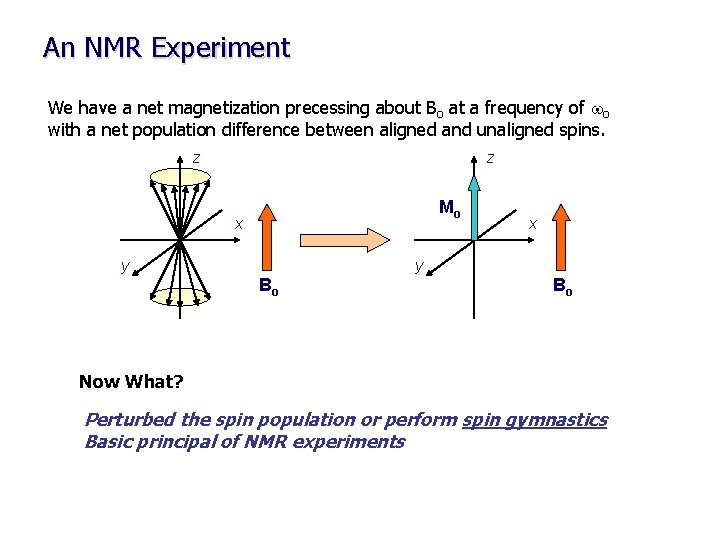
An NMR Experiment We have a net magnetization precessing about Bo at a frequency of wo with a net population difference between aligned and unaligned spins. z z Mo x y Bo Bo Now What? Perturbed the spin population or perform spin gymnastics Basic principal of NMR experiments

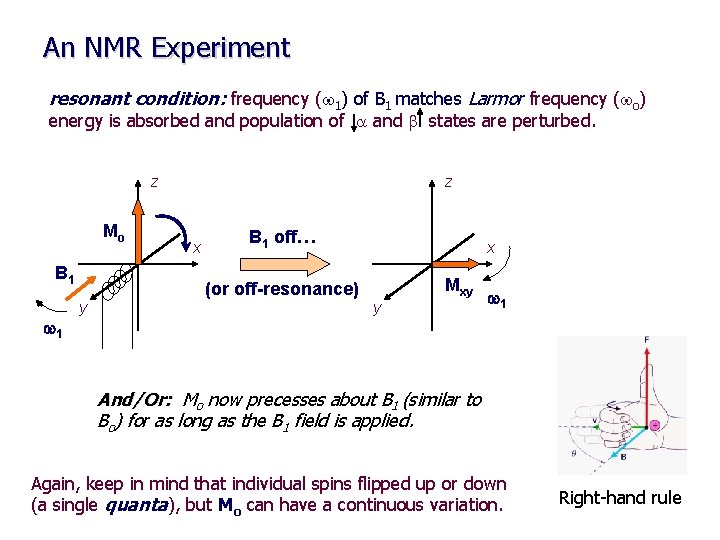
An NMR Experiment resonant condition: frequency (w 1) of B 1 matches Larmor frequency (wo) energy is absorbed and population of a and b states are perturbed. z Mo B 1 w 1 y z x B 1 off… (or off-resonance) x Mxy y w 1 And/Or: Mo now precesses about B 1 (similar to Bo) for as long as the B 1 field is applied. Again, keep in mind that individual spins flipped up or down (a single quanta), but Mo can have a continuous variation. Right-hand rule

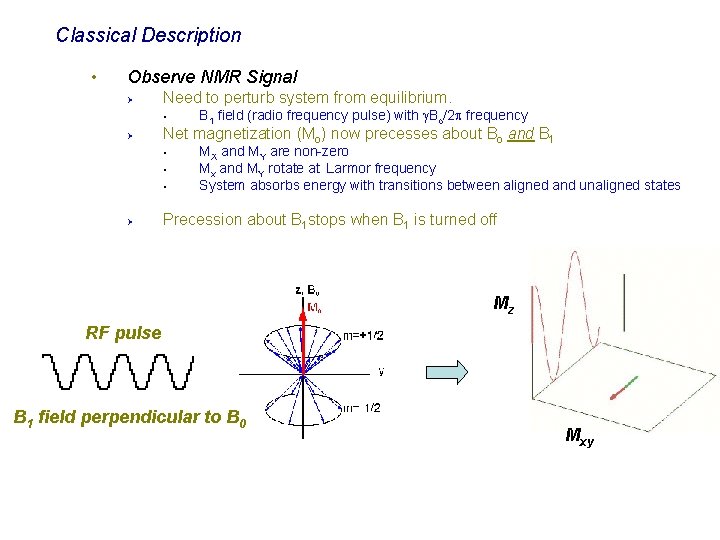
Classical Description • Observe NMR Signal Ø Need to perturb system from equilibrium. § Ø Net magnetization (Mo) now precesses about Bo and B 1 § § § Ø B 1 field (radio frequency pulse) with g. Bo/2 p frequency MX and MY are non-zero Mx and MY rotate at Larmor frequency System absorbs energy with transitions between aligned and unaligned states Precession about B 1 stops when B 1 is turned off Mz RF pulse B 1 field perpendicular to B 0 Mxy

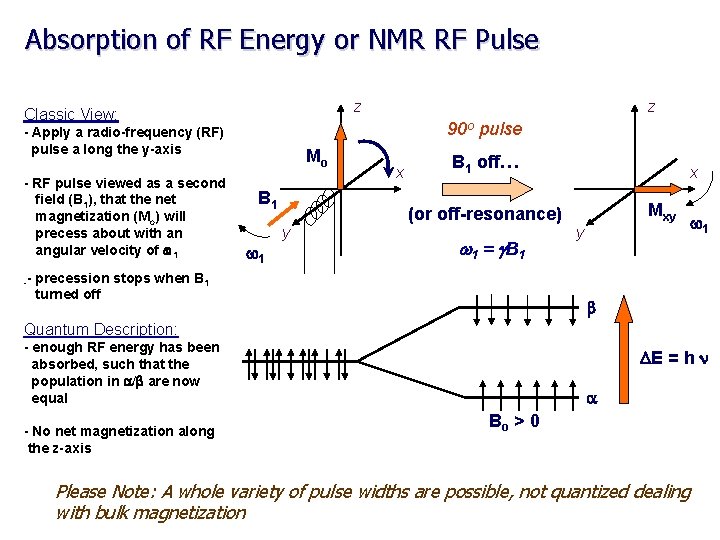
Absorption of RF Energy or NMR RF Pulse z Classic View: 90 o pulse - Apply a radio-frequency (RF) pulse a long the y-axis - RF pulse viewed as a second field (B 1), that the net magnetization (Mo) will precess about with an angular velocity of w 1 -- z Mo B 1 w 1 y x B 1 off… (or off-resonance) w 1 = g B 1 precession stops when B 1 turned off x Mxy y w 1 b Quantum Description: - enough RF energy has been absorbed, such that the population in a/b are now equal - No net magnetization along the z-axis DE = h n a Bo > 0 Please Note: A whole variety of pulse widths are possible, not quantized dealing with bulk magnetization

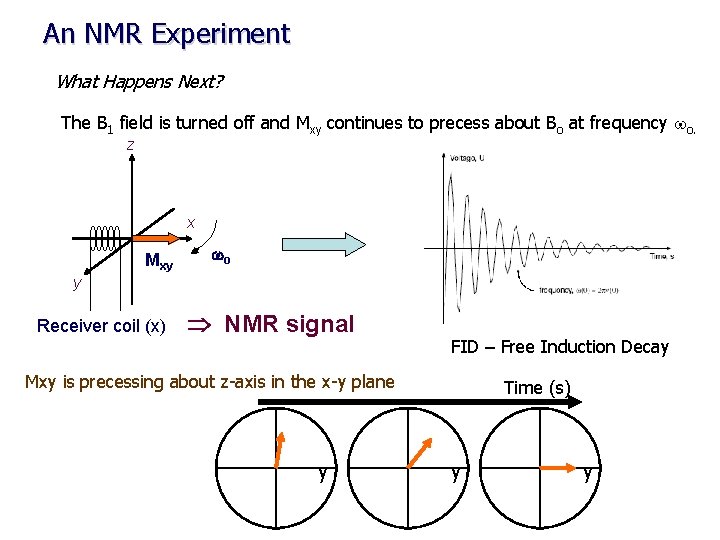
An NMR Experiment What Happens Next? The B 1 field is turned off and Mxy continues to precess about Bo at frequency wo. z x Mxy wo y Receiver coil (x) NMR signal FID – Free Induction Decay Mxy is precessing about z-axis in the x-y plane y Time (s) y y

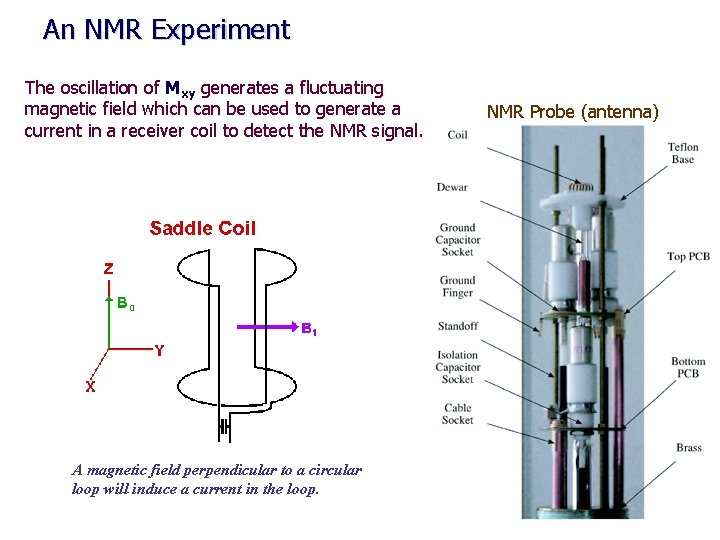
An NMR Experiment The oscillation of Mxy generates a fluctuating magnetic field which can be used to generate a current in a receiver coil to detect the NMR signal. A magnetic field perpendicular to a circular loop will induce a current in the loop. NMR Probe (antenna)

NMR Signal Detection - FID The FID reflects the change in the magnitude of Mxy as the signal is changing relative to the receiver along the y-axis Detect signal along X RF pulse along Y Again, the signal is precessing about Bo at its Larmor Frequency (wo).

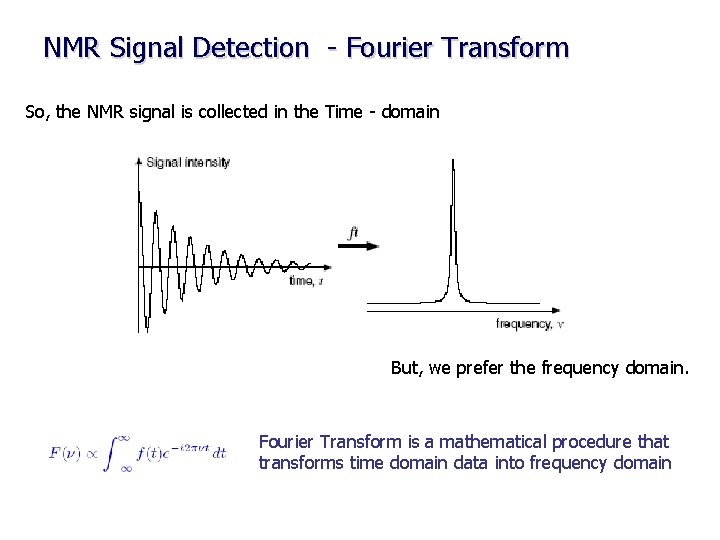
NMR Signal Detection - Fourier Transform So, the NMR signal is collected in the Time - domain But, we prefer the frequency domain. Fourier Transform is a mathematical procedure that transforms time domain data into frequency domain

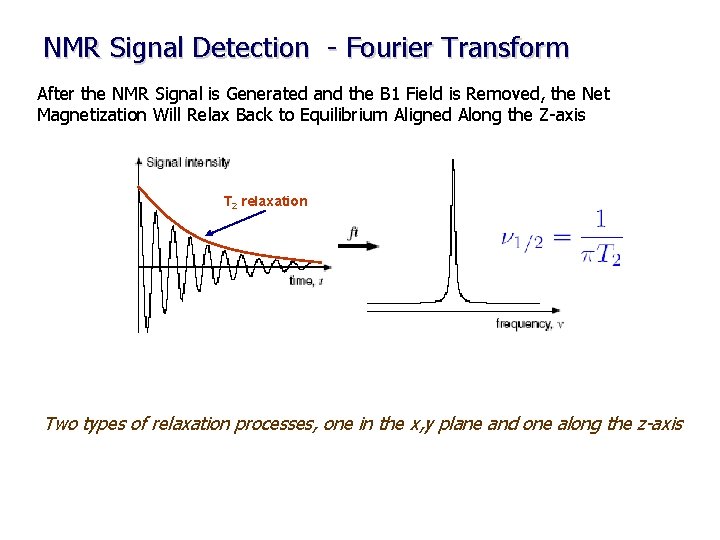
NMR Signal Detection - Fourier Transform After the NMR Signal is Generated and the B 1 Field is Removed, the Net Magnetization Will Relax Back to Equilibrium Aligned Along the Z-axis T 2 relaxation Two types of relaxation processes, one in the x, y plane and one along the z-axis

NMR Relaxation a) b) No spontaneous reemission of photons to relax down to ground state 1) Probability too low cube of the frequency Two types of NMR relaxation processes 1) spin-lattice or longitudinal relaxation (T 1) 2) i. transfer of energy to the lattice or solvent material 3) ii. coupling of nuclei magnetic field with magnetic fields created 4) by the ensemble of vibrational and rotational motion of the 5) lattice or solvent. 6) iii. results in a minimal temperature increase in sample 7) iv. Relaxation time (T 1) exponential decay Mz = M 0(1 -exp(-t/T 1)) Please Note: General practice is to wait 5 x. T 1 for the system to have fully relaxed.

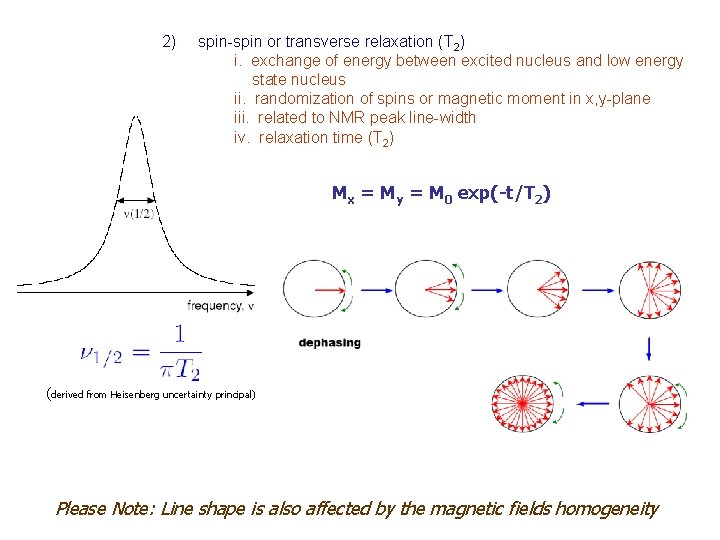
2) spin-spin or transverse relaxation (T 2) i. exchange of energy between excited nucleus and low energy state nucleus ii. randomization of spins or magnetic moment in x, y-plane iii. related to NMR peak line-width iv. relaxation time (T 2) Mx = My = M 0 exp(-t/T 2) (derived from Heisenberg uncertainty principal) Please Note: Line shape is also affected by the magnetic fields homogeneity

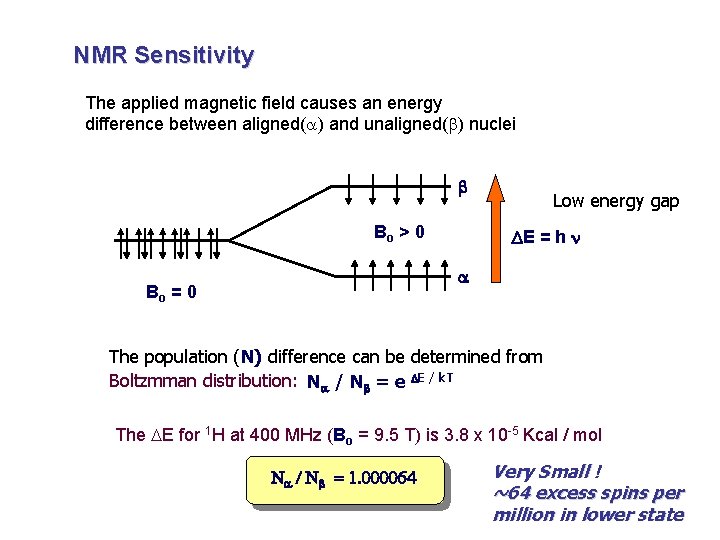
NMR Sensitivity The applied magnetic field causes an energy difference between aligned(a) and unaligned(b) nuclei b Bo > 0 Low energy gap DE = h n a Bo = 0 The population (N) difference can be determined from Boltzmman distribution: Na / Nb = e DE / k. T The DE for 1 H at 400 MHz (Bo = 9. 5 T) is 3. 8 x 10 -5 Kcal / mol Na / Nb = 1. 000064 Very Small ! ~64 excess spins per million in lower state

NMR Sensitivity NMR signal depends on: signal (s) % g 4 Bo 2 NB 1 g(u)/T 1) 2) 3) 4) 5) Number of Nuclei (N) (limited to field homogeneity and filling factor) Gyromagnetic ratio (in practice g 3) Inversely to temperature (T) External magnetic field (Bo 2/3, in practice, homogeneity) B 12 exciting field strength DE = g h Bo / 2 p Na / Nb = e DE / k. T Increase energy gap -> Increase population difference -> Increase NMR signal DE ≡ Bo ≡ g g - Intrinsic property of nucleus can not be changed. (g. H/g. C)3 1 H for 13 C is 64 x (g. H/g. N)3 for is ~ 64 x as sensitive as 13 C 15 N is 1000 x and 1000 x as sensitive as 15 N ! Consider that the natural abundance of 13 C is 1. 1% and 15 N is 0. 37% relative sensitivity increases to ~6, 400 x and ~2. 7 x 105 x !!

NMR Sensitivity • Relative sensitivity of 1 H, 13 C, 15 N and other nuclei NMR spectra depend on Ø Gyromagnetic ratio (g) Ø Natural abundance of the isotope g - Intrinsic property of nucleus can not be changed. (g. H/g. C)3 1 H for 13 C is 64 x (g. H/g. N)3 for is ~ 64 x as sensitive as 13 C 15 N is 1000 x and 1000 x as sensitive as 15 N ! Consider that the natural abundance of 13 C is 1. 1% and 15 N is 0. 37% relative sensitivity increases to ~6, 400 x and ~2. 7 x 105 x !! 1 H NMR spectra of caffeine 8 scans ~12 secs 13 C NMR spectra of caffeine 10, 000 scans ~4. 2 hours

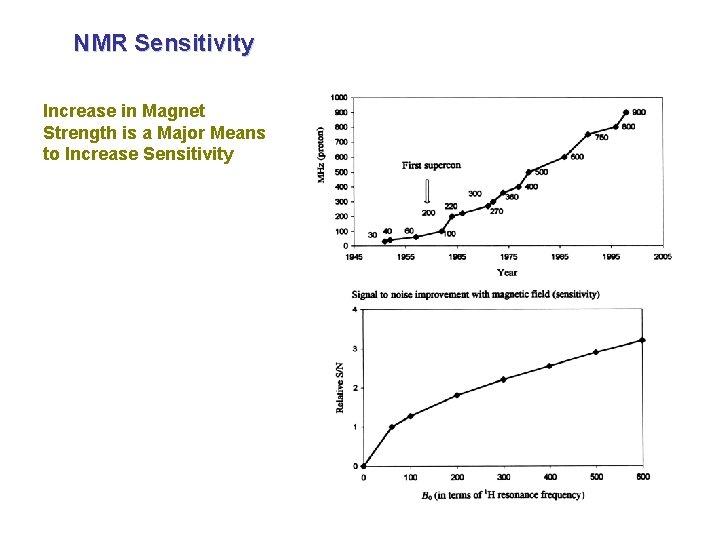
NMR Sensitivity Increase in Magnet Strength is a Major Means to Increase Sensitivity

NMR Sensitivity But at a significant cost! ~$800, 000 ~$2, 000 ~$4, 500, 000

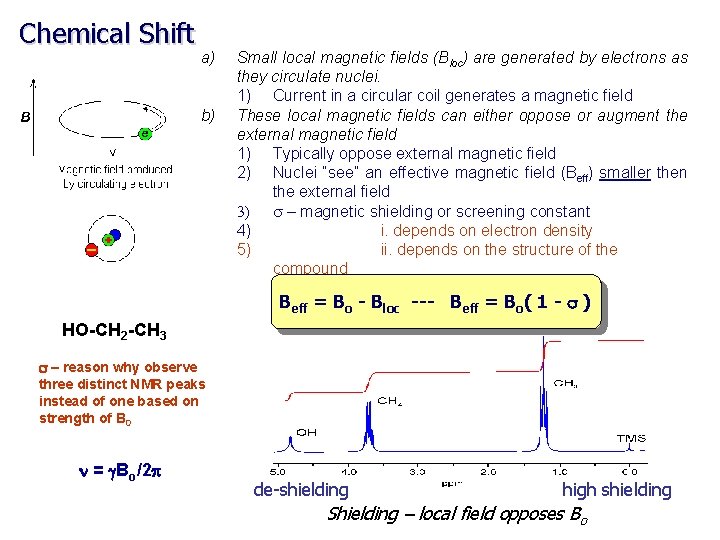
Chemical Shift Up to this point, we have been treating nuclei in general terms. Simply comparing 1 H, 13 C, 15 N etc. If all 1 H resonate at 500 MHz at a field strength of 11. 7 T, NMR would not be very interesting The chemical environment for each nuclei results in a unique local magnetic field (Bloc) for each nuclei: Beff = Bo - Bloc --- Beff = Bo( 1 - s ) s is the magnetic shielding of the nucleus

Chemical Shift a) b) Small local magnetic fields (Bloc) are generated by electrons as they circulate nuclei. 1) Current in a circular coil generates a magnetic field These local magnetic fields can either oppose or augment the external magnetic field 1) Typically oppose external magnetic field 2) Nuclei “see” an effective magnetic field (Beff) smaller then the external field 3) s – magnetic shielding or screening constant 4) i. depends on electron density 5) ii. depends on the structure of the compound Beff = Bo - Bloc --- Beff = Bo( 1 - s ) HO-CH 2 -CH 3 s – reason why observe three distinct NMR peaks instead of one based on strength of B 0 n = g. Bo/2 p de-shielding high shielding Shielding – local field opposes Bo

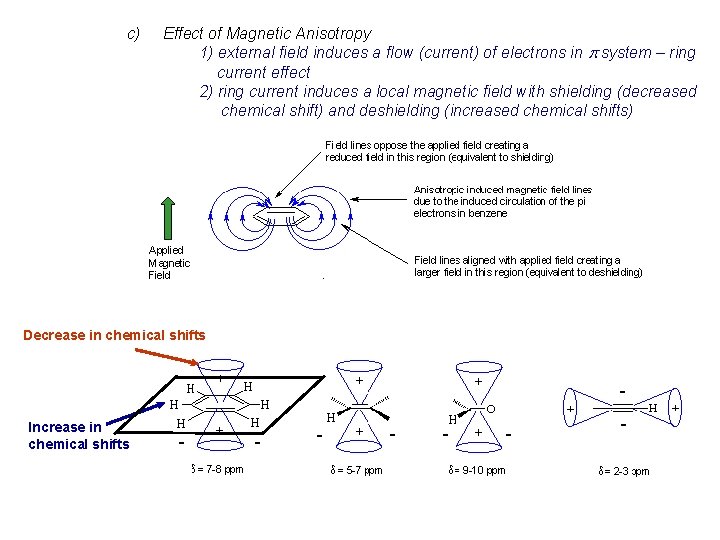
c) Effect of Magnetic Anisotropy 1) external field induces a flow (current) of electrons in p system – ring current effect 2) ring current induces a local magnetic field with shielding (decreased chemical shift) and deshielding (increased chemical shifts) Decrease in chemical shifts Increase in chemical shifts

The NMR scale (d, ppm) Bo >> Bloc -- MHz compared to Hz Comparing small changes in the context of a large number is cumbersome d= w - wref ppm (parts per million) Instead use a relative scale, and refer all signals (w) in the spectrum to the signal of a particular compound (wref). IMPORTANT: absolute frequency is field dependent (n = g Bo / 2 p) CH 3 Tetramethyl silane (TMS) is a common reference chemical H 3 C Si CH 3

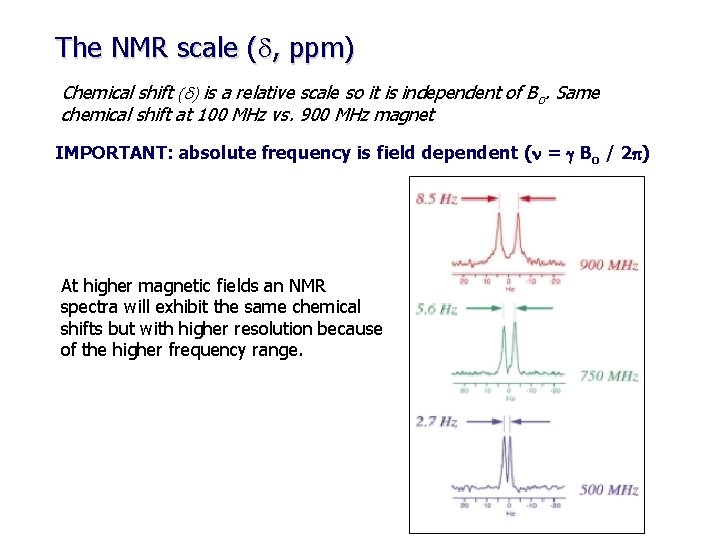
The NMR scale (d, ppm) Chemical shift (d) is a relative scale so it is independent of Bo. Same chemical shift at 100 MHz vs. 900 MHz magnet IMPORTANT: absolute frequency is field dependent (n = g Bo / 2 p) At higher magnetic fields an NMR spectra will exhibit the same chemical shifts but with higher resolution because of the higher frequency range.

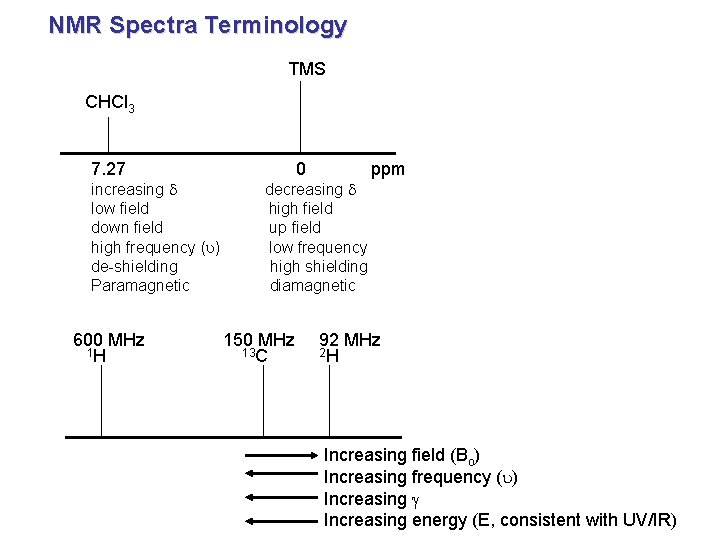
NMR Spectra Terminology TMS CHCl 3 7. 27 increasing d low field down field high frequency (u) de-shielding Paramagnetic 600 MHz 1 H 0 decreasing d high field up field low frequency high shielding diamagnetic 150 MHz 13 C ppm 92 MHz 2 H Increasing field (Bo) Increasing frequency (u) Increasing g Increasing energy (E, consistent with UV/IR)

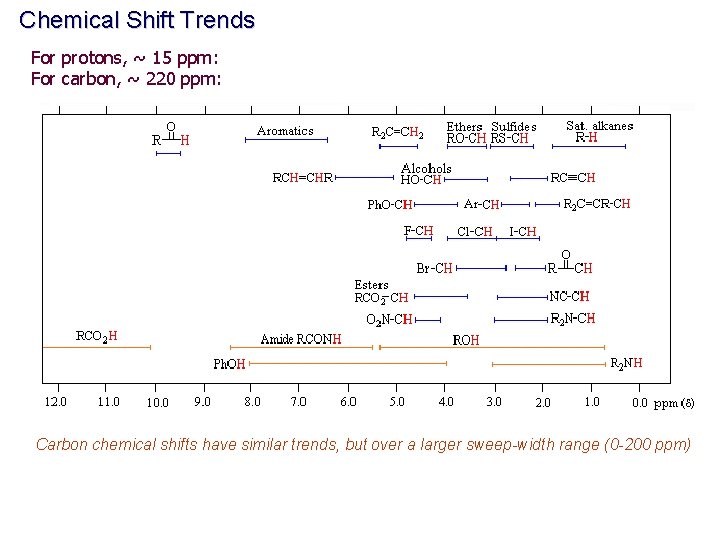
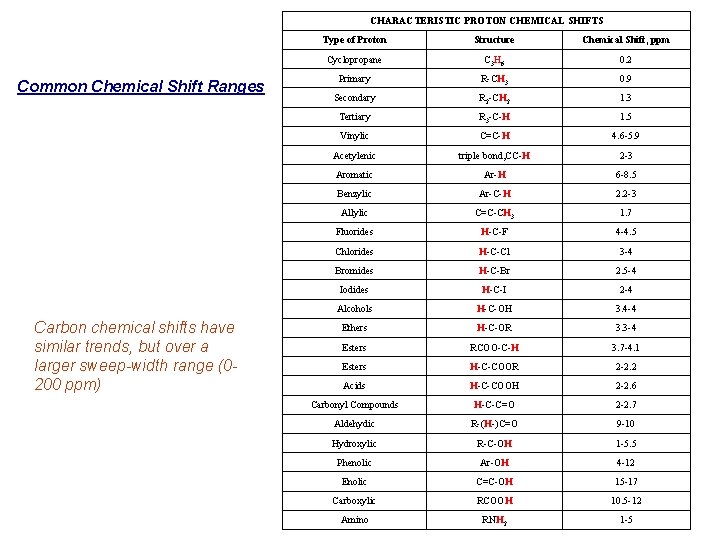
Chemical Shift Trends For protons, ~ 15 ppm: For carbon, ~ 220 ppm: Carbon chemical shifts have similar trends, but over a larger sweep-width range (0 -200 ppm)

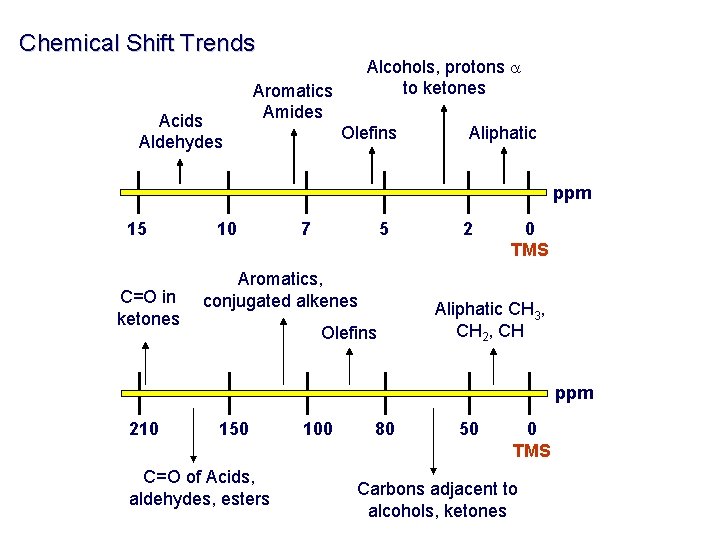
Chemical Shift Trends Acids Aldehydes Alcohols, protons a to ketones Aromatics Amides Olefins Aliphatic ppm 15 C=O in ketones 10 7 5 Aromatics, conjugated alkenes Olefins 2 0 TMS Aliphatic CH 3, CH 2, CH ppm 210 150 C=O of Acids, aldehydes, esters 100 80 50 0 TMS Carbons adjacent to alcohols, ketones

CHARACTERISTIC PROTON CHEMICAL SHIFTS Common Chemical Shift Ranges Carbon chemical shifts have similar trends, but over a larger sweep-width range (0200 ppm) Type of Proton Structure Chemical Shift, ppm Cyclopropane C 3 H 6 0. 2 Primary R-CH 3 0. 9 Secondary R 2 -CH 2 1. 3 Tertiary R 3 -C-H 1. 5 Vinylic C=C-H 4. 6 -5. 9 Acetylenic triple bond, CC-H 2 -3 Aromatic Ar-H 6 -8. 5 Benzylic Ar-C-H 2. 2 -3 Allylic C=C-CH 3 1. 7 Fluorides H-C-F 4 -4. 5 Chlorides H-C-Cl 3 -4 Bromides H-C-Br 2. 5 -4 Iodides H-C-I 2 -4 Alcohols H-C-OH 3. 4 -4 Ethers H-C-OR 3. 3 -4 Esters RCOO-C-H 3. 7 -4. 1 Esters H-C-COOR 2 -2. 2 Acids H-C-COOH 2 -2. 6 Carbonyl Compounds H-C-C=O 2 -2. 7 Aldehydic R-(H-)C=O 9 -10 Hydroxylic R-C-OH 1 -5. 5 Phenolic Ar-OH 4 -12 Enolic C=C-OH 15 -17 Carboxylic RCOOH 10. 5 -12 Amino RNH 2 1 -5

Predicting Chemical Shift Assignments Numerous Experimental NMR Data has been compiled and general trends identified • See: § “Tables of Spectral Data for Structure Determination of Organic Compounds” Pretsch, Clerc, Seibl and Simon § “Spectrometric Identification of Organic Compounds” Silverstein, Bassler and Morrill • Spectral Databases: § Aldrich/ACD Library of FT NMR Spectra § Sadtler/Spectroscopy (UV/Vis, IR, MS, GC and NMR) Ongoing effort to predict chemical shifts from first principals (quantum mechanical description of factors contributing to chemical shifts)

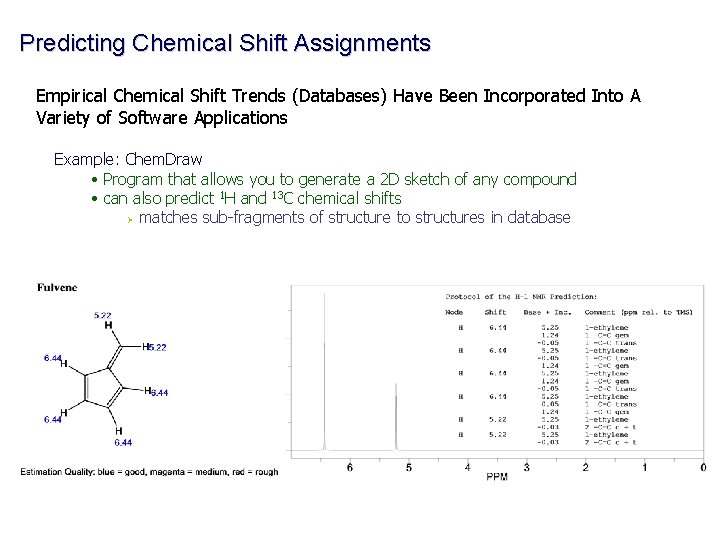
Predicting Chemical Shift Assignments Empirical Chemical Shift Trends (Databases) Have Been Incorporated Into A Variety of Software Applications Example: Chem. Draw • Program that allows you to generate a 2 D sketch of any compound • can also predict 1 H and 13 C chemical shifts Ø matches sub-fragments of structure to structures in database

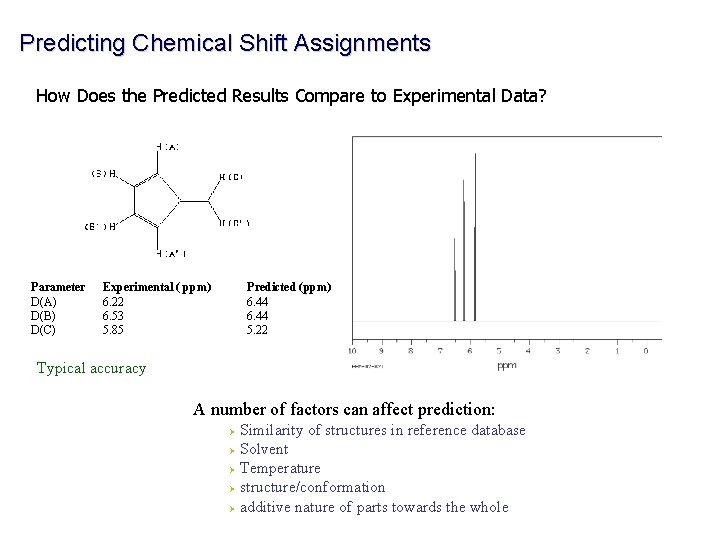
Predicting Chemical Shift Assignments How Does the Predicted Results Compare to Experimental Data? Parameter D(A) D(B) D(C) Experimental ( ppm) 6. 22 6. 53 5. 85 Predicted (ppm) 6. 44 5. 22 Typical accuracy A number of factors can affect prediction: Similarity of structures in reference database Ø Solvent Ø Temperature Ø structure/conformation Ø additive nature of parts towards the whole Ø

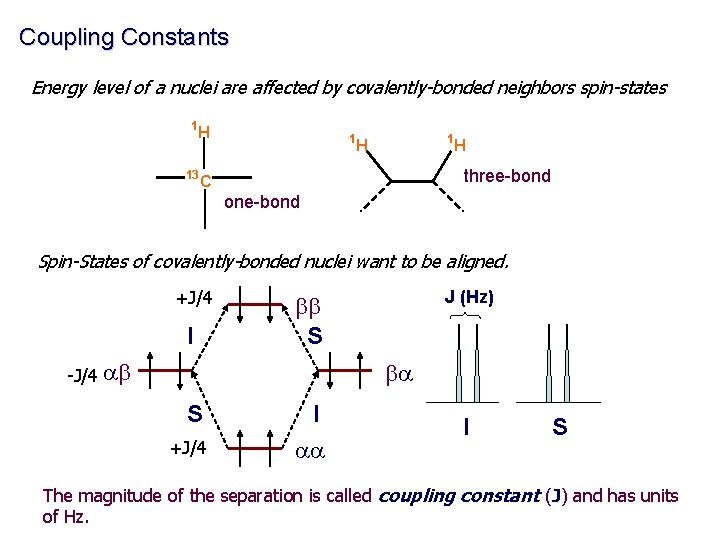
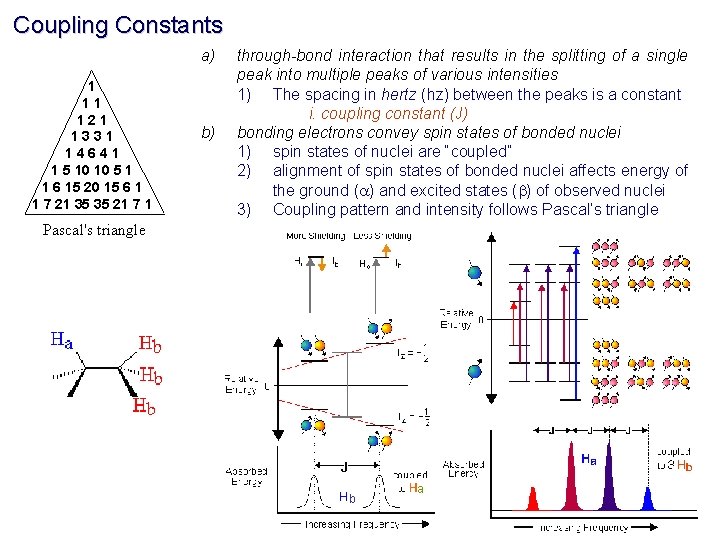
Coupling Constants Energy level of a nuclei are affected by covalently-bonded neighbors spin-states 1 H 13 1 1 H H three-bond C one-bond Spin-States of covalently-bonded nuclei want to be aligned. +J/4 I J (Hz) bb S -J/4 ab ba S +J/4 I aa I S The magnitude of the separation is called coupling constant (J) and has units of Hz.

Coupling Constants a) 1 11 121 1331 14641 1 5 10 10 5 1 1 6 15 20 15 6 1 1 7 21 35 35 21 7 1 b) through-bond interaction that results in the splitting of a single peak into multiple peaks of various intensities 1) The spacing in hertz (hz) between the peaks is a constant i. coupling constant (J) bonding electrons convey spin states of bonded nuclei 1) spin states of nuclei are “coupled” 2) alignment of spin states of bonded nuclei affects energy of the ground (a) and excited states (b) of observed nuclei 3) Coupling pattern and intensity follows Pascal’s triangle Pascal's triangle b a

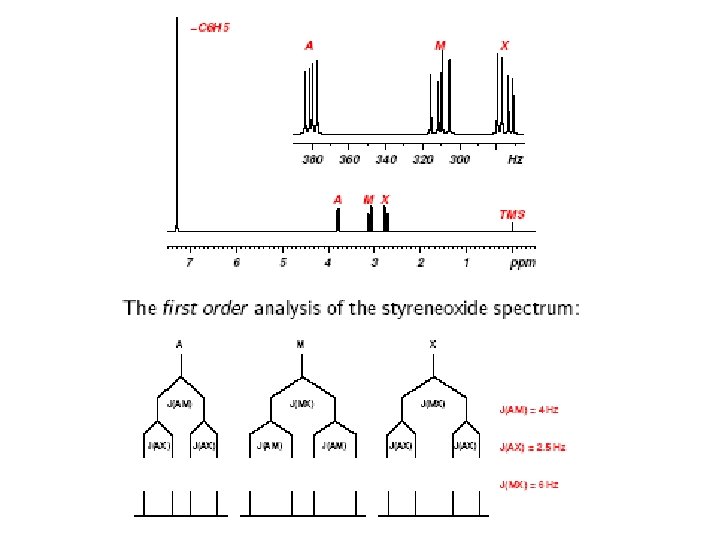
Common NMR Splitting Patterns Multiplets consist of 2 n. I + 1 lines I is the nuclear spin quantum number (usually 1/2) and n is the number of neighboring spins. singlet doublet 1: 1 triplet quartet 1: 2: 1 1: 3: 3: 1 pentet 1: 4: 6: 4: 1 Coupling Rules: 1. 2. 3. 4. 5. 6. equivalent nuclei do not interact coupling constants decreases with separation ( typically # 3 bonds) multiplicity given by number of attached equivalent protons (n+1) multiple spin systems multiplicity (na+1)(nb+1) Relative peak heights/area follows Pascal’s triangle Coupling constant are independent of applied field strength IMPORTANT: Coupling constant pattern allow for the identification of bonded nuclei.


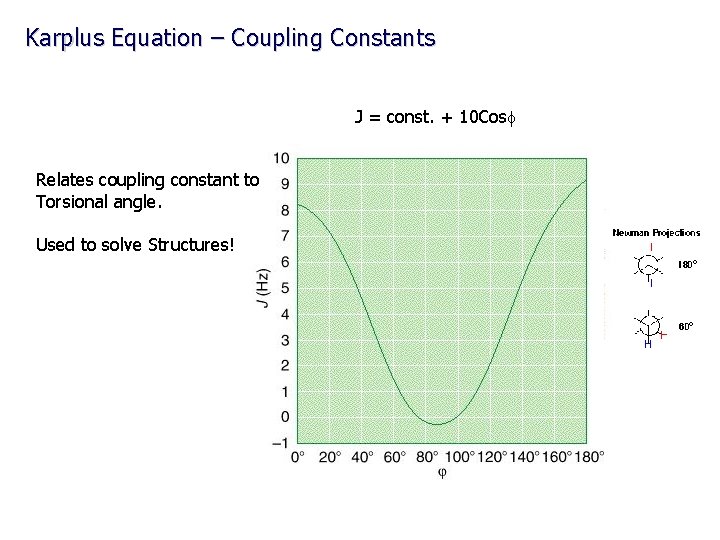
Karplus Equation – Coupling Constants J = const. + 10 Cosf Relates coupling constant to Torsional angle. Used to solve Structures!

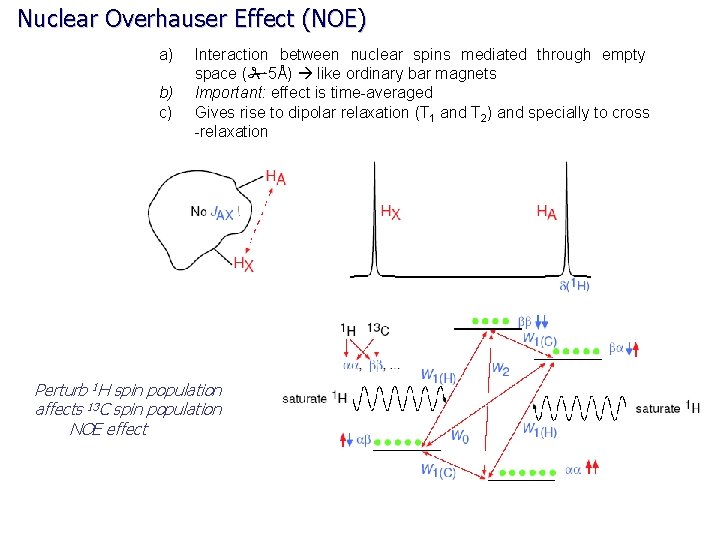
Nuclear Overhauser Effect (NOE) a) b) c) Interaction between nuclear spins mediated through empty space (#5Å) like ordinary bar magnets Important: effect is time-averaged Gives rise to dipolar relaxation (T 1 and T 2) and specially to cross -relaxation Perturb 1 H spin population affects 13 C spin population NOE effect

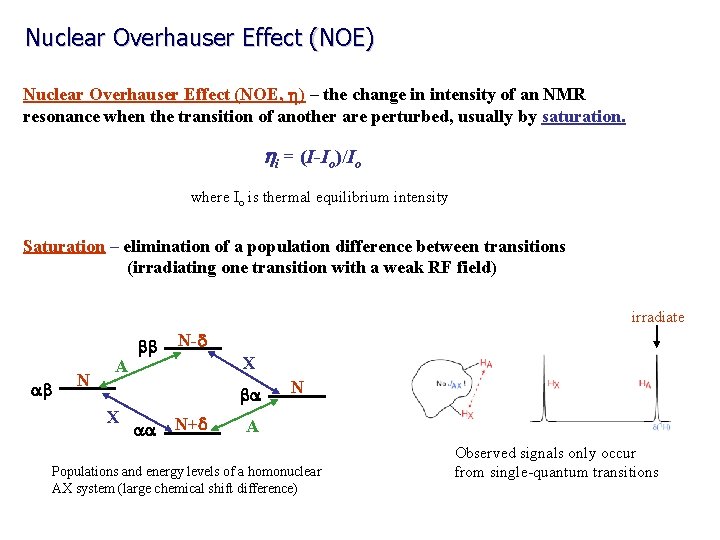
Nuclear Overhauser Effect (NOE) Nuclear Overhauser Effect (NOE, h) – the change in intensity of an NMR resonance when the transition of another are perturbed, usually by saturation. hi = (I-Io)/Io where Io is thermal equilibrium intensity Saturation – elimination of a population difference between transitions (irradiating one transition with a weak RF field) irradiate ab N A bb N-d X ba X aa N+d N A Populations and energy levels of a homonuclear AX system (large chemical shift difference) Observed signals only occur from single-quantum transitions

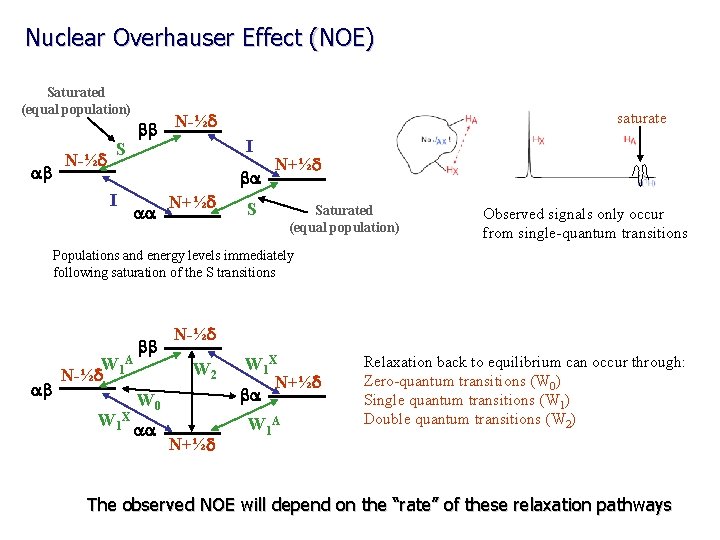
Nuclear Overhauser Effect (NOE) Saturated (equal population) ab N-½d S bb N-½d saturate I ba I aa N+½d S Saturated (equal population) Observed signals only occur from single-quantum transitions Populations and energy levels immediately following saturation of the S transitions ab W N-½d 1 A W 1 X bb N-½d W 2 W 0 aa N+½d W 1 X N+½d ba W 1 A Relaxation back to equilibrium can occur through: Zero-quantum transitions (W 0) Single quantum transitions (W 1) Double quantum transitions (W 2) The observed NOE will depend on the “rate” of these relaxation pathways

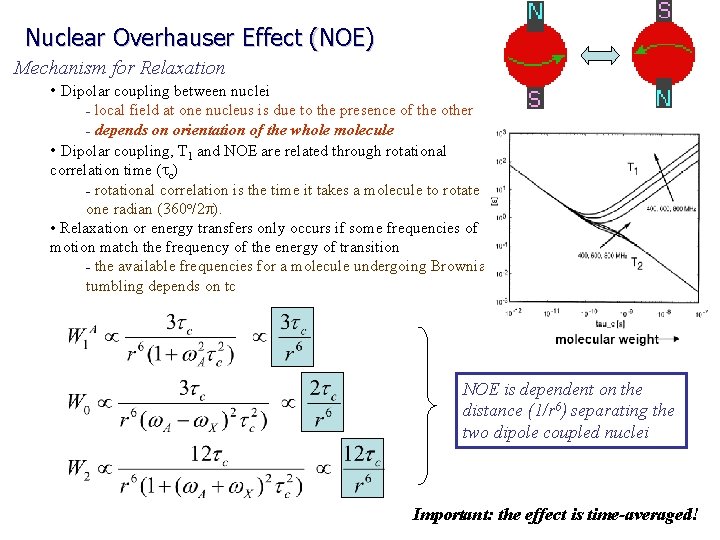
Nuclear Overhauser Effect (NOE) Mechanism for Relaxation • Dipolar coupling between nuclei local field at one nucleus is due to the presence of the other – depends on orientation of the whole molecule • Dipolar coupling, T 1 and NOE are related through rotational correlation time (tc) – rotational correlation is the time it takes a molecule to rotate one radian (360 o/2 p). • Relaxation or energy transfers only occurs if some frequencies of motion match the frequency of the energy of transition – the available frequencies for a molecule undergoing Brownian tumbling depends on tc – NOE is dependent on the distance (1/r 6) separating the two dipole coupled nuclei Important: the effect is time-averaged!

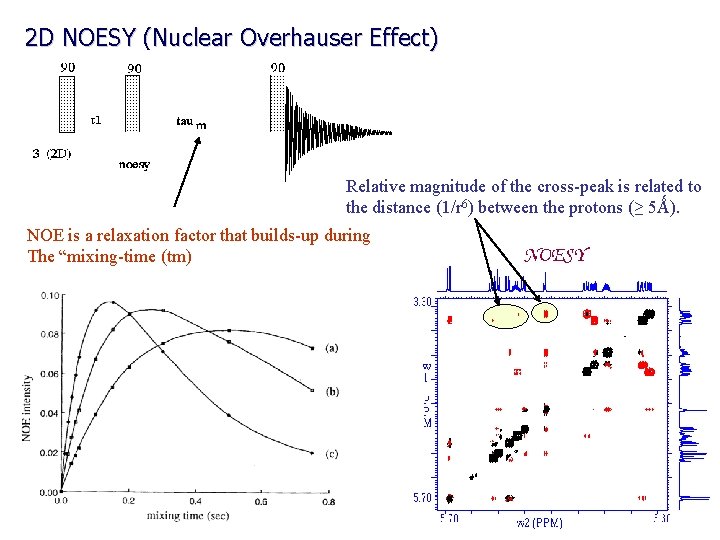
2 D NOESY (Nuclear Overhauser Effect) Relative magnitude of the cross-peak is related to the distance (1/r 6) between the protons (≥ 5Ǻ). NOE is a relaxation factor that builds-up during The “mixing-time (tm)

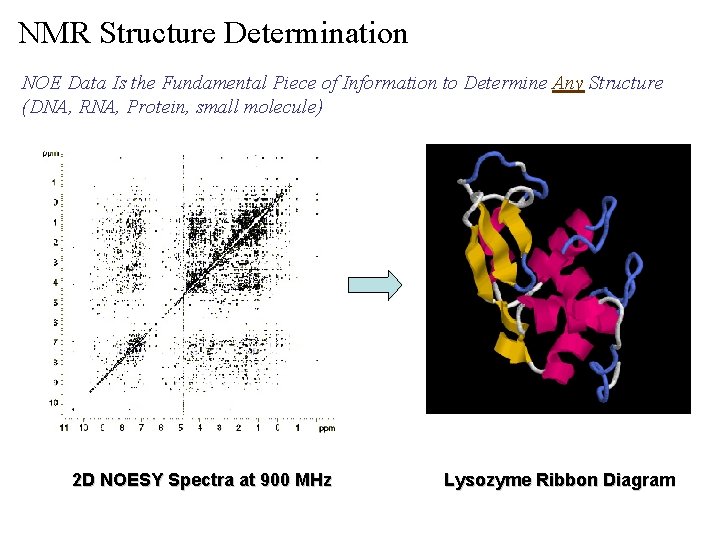
NMR Structure Determination NOE Data Is the Fundamental Piece of Information to Determine Any Structure (DNA, RNA, Protein, small molecule) 2 D NOESY Spectra at 900 MHz Lysozyme Ribbon Diagram

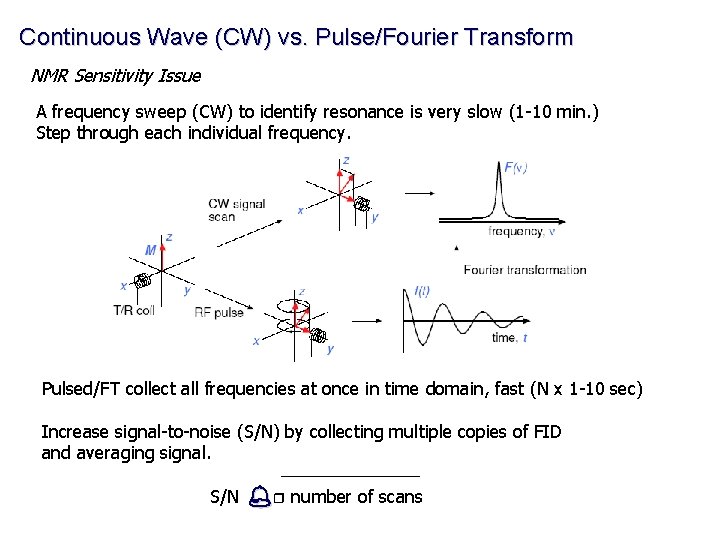
Continuous Wave (CW) vs. Pulse/Fourier Transform NMR Sensitivity Issue A frequency sweep (CW) to identify resonance is very slow (1 -10 min. ) Step through each individual frequency. Pulsed/FT collect all frequencies at once in time domain, fast (N x 1 -10 sec) Increase signal-to-noise (S/N) by collecting multiple copies of FID and averaging signal. S/N % r number of scans

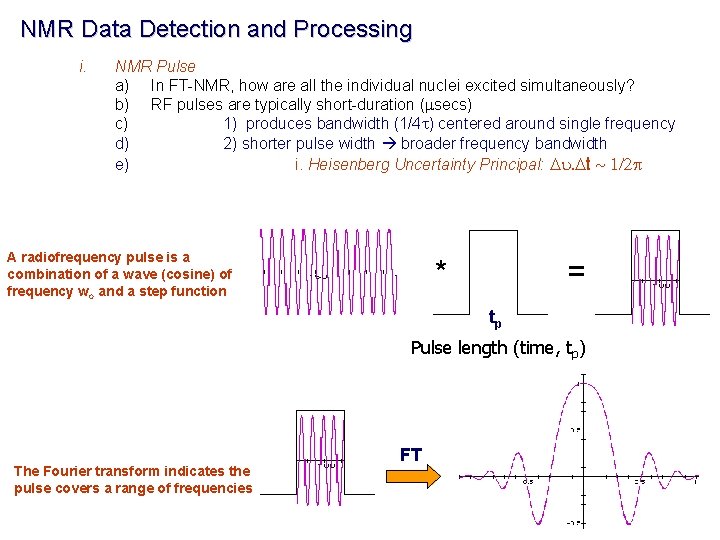
NMR Data Detection and Processing i. NMR Pulse a) In FT-NMR, how are all the individual nuclei excited simultaneously? b) RF pulses are typically short-duration (msecs) c) 1) produces bandwidth (1/4 t) centered around single frequency d) 2) shorter pulse width broader frequency bandwidth e) i. Heisenberg Uncertainty Principal: Du. Dt ~ 1/2 p A radiofrequency pulse is a combination of a wave (cosine) of frequency wo and a step function * = tp Pulse length (time, tp) The Fourier transform indicates the pulse covers a range of frequencies FT

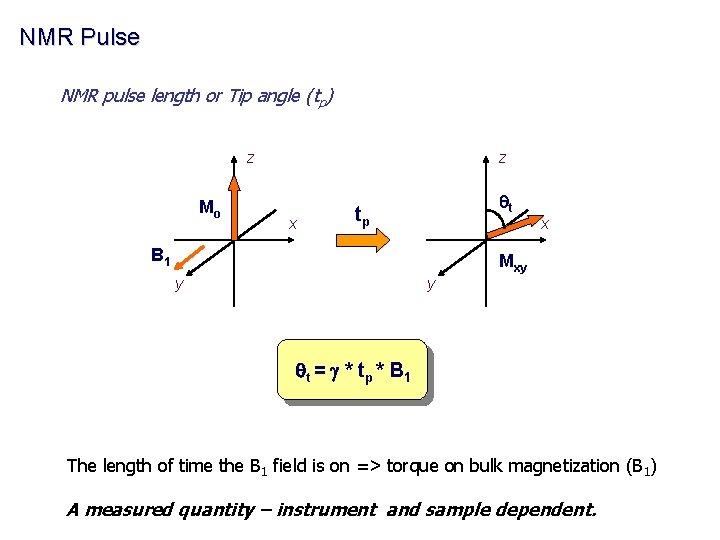
NMR Pulse NMR pulse length or Tip angle (tp) z Mo z x qt tp x B 1 Mxy y y q t = g * tp * B 1 The length of time the B 1 field is on => torque on bulk magnetization (B 1) A measured quantity – instrument and sample dependent.

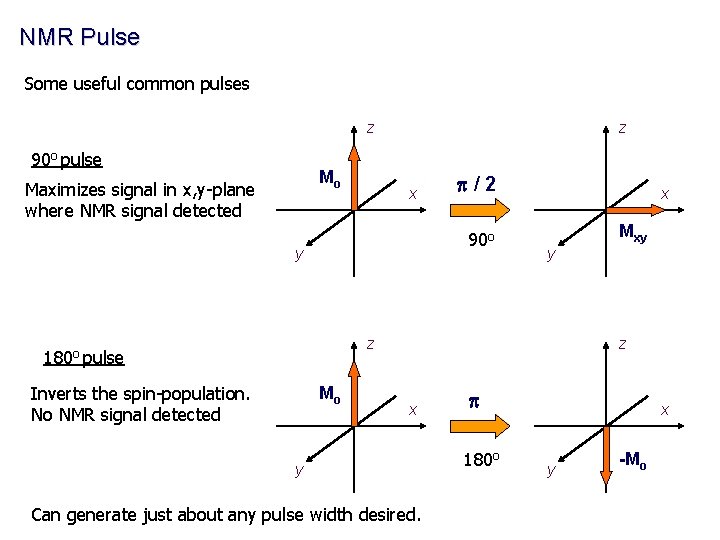
NMR Pulse Some useful common pulses z 90 o pulse Mo Maximizes signal in x, y-plane where NMR signal detected z x p/2 90 o y x Mxy y z 180 o pulse Inverts the spin-population. No NMR signal detected Mo z x y Can generate just about any pulse width desired. p 180 o x y -Mo

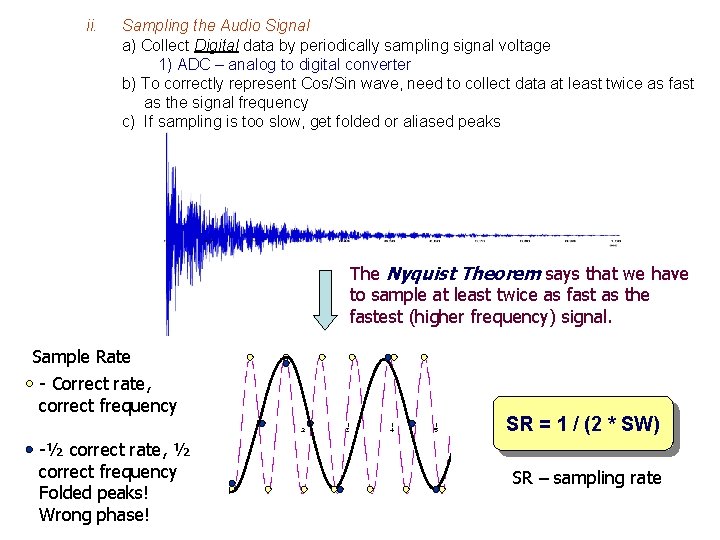
ii. Sampling the Audio Signal a) Collect Digital data by periodically sampling signal voltage 1) ADC – analog to digital converter b) To correctly represent Cos/Sin wave, need to collect data at least twice as fast as the signal frequency c) If sampling is too slow, get folded or aliased peaks The Nyquist Theorem says that we have to sample at least twice as fast as the fastest (higher frequency) signal. Sample Rate - Correct rate, correct frequency -½ correct rate, ½ correct frequency Folded peaks! Wrong phase! SR = 1 / (2 * SW) SR – sampling rate

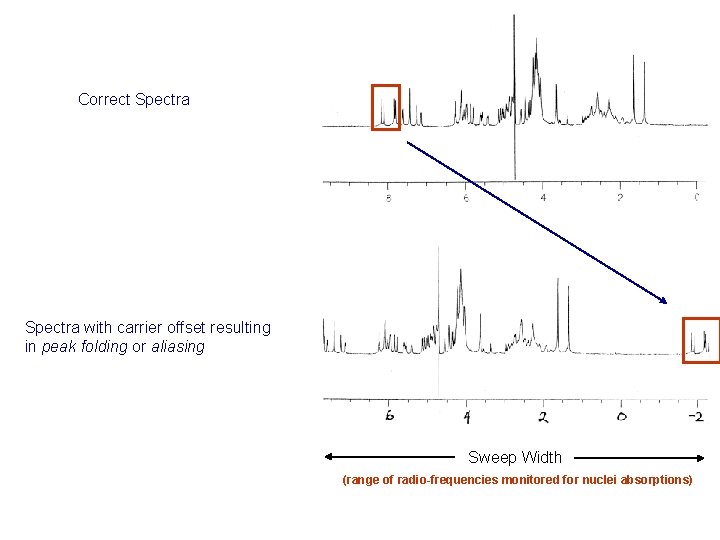
Correct Spectra with carrier offset resulting in peak folding or aliasing Sweep Width (range of radio-frequencies monitored for nuclei absorptions)

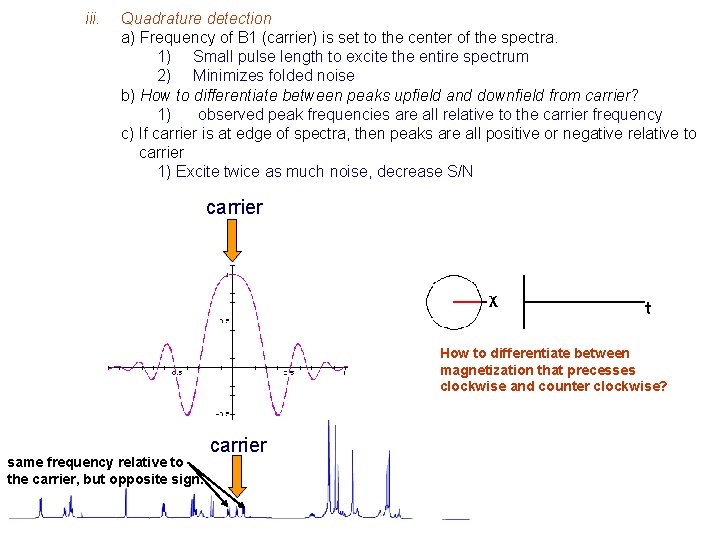
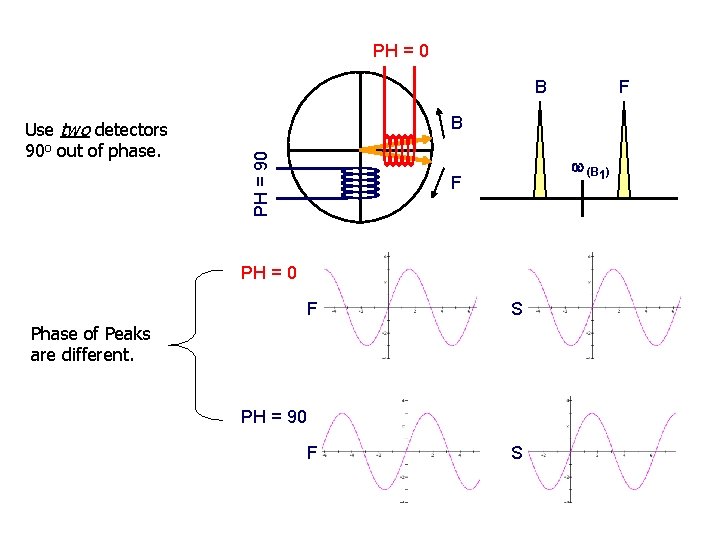
iii. Quadrature detection a) Frequency of B 1 (carrier) is set to the center of the spectra. 1) Small pulse length to excite the entire spectrum 2) Minimizes folded noise b) How to differentiate between peaks upfield and downfield from carrier? 1) observed peak frequencies are all relative to the carrier frequency c) If carrier is at edge of spectra, then peaks are all positive or negative relative to carrier 1) Excite twice as much noise, decrease S/N carrier How to differentiate between magnetization that precesses clockwise and counter clockwise? same frequency relative to the carrier, but opposite sign. carrier

PH = 0 B B PH = 90 Use two detectors 90 o out of phase. F w (B 1) F PH = 0 F S Phase of Peaks are different. PH = 90

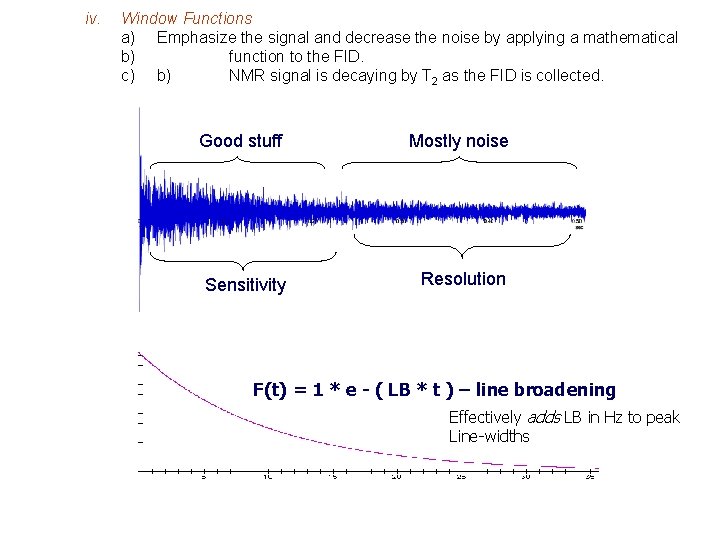
iv. Window Functions a) Emphasize the signal and decrease the noise by applying a mathematical b) function to the FID. c) b) NMR signal is decaying by T 2 as the FID is collected. Good stuff Mostly noise Sensitivity Resolution F(t) = 1 * e - ( LB * t ) – line broadening Effectively adds LB in Hz to peak Line-widths

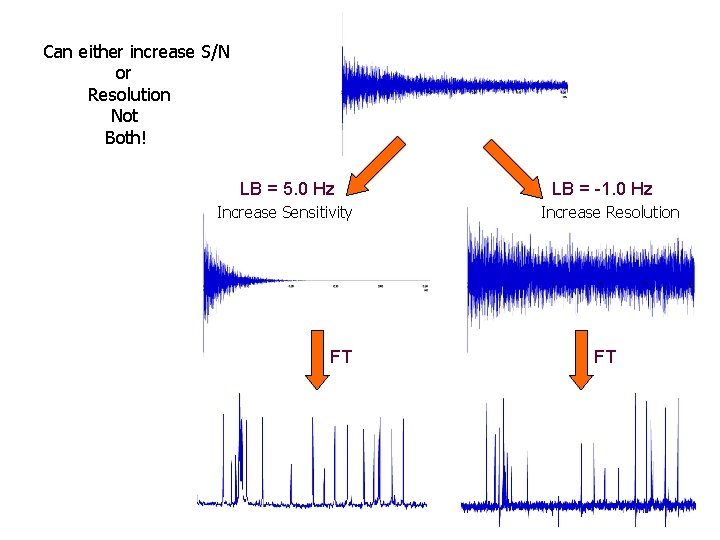
Can either increase S/N or Resolution Not Both! LB = 5. 0 Hz Increase Sensitivity FT LB = -1. 0 Hz Increase Resolution FT

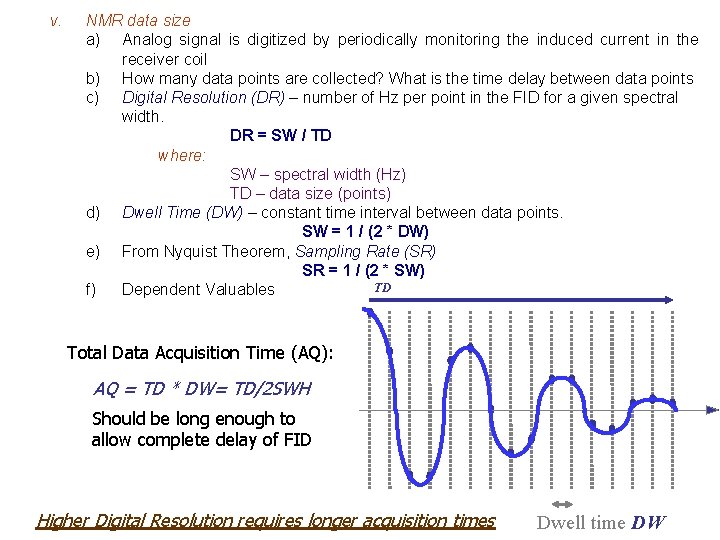
v. NMR data size a) Analog signal is digitized by periodically monitoring the induced current in the receiver coil b) How many data points are collected? What is the time delay between data points c) Digital Resolution (DR) – number of Hz per point in the FID for a given spectral width. DR = SW / TD where: SW – spectral width (Hz) TD – data size (points) d) Dwell Time (DW) – constant time interval between data points. SW = 1 / (2 * DW) e) From Nyquist Theorem, Sampling Rate (SR) SR = 1 / (2 * SW) TD f) Dependent Valuables Total Data Acquisition Time (AQ): AQ = TD * DW= TD/2 SWH Should be long enough to allow complete delay of FID Higher Digital Resolution requires longer acquisition times Dwell time DW

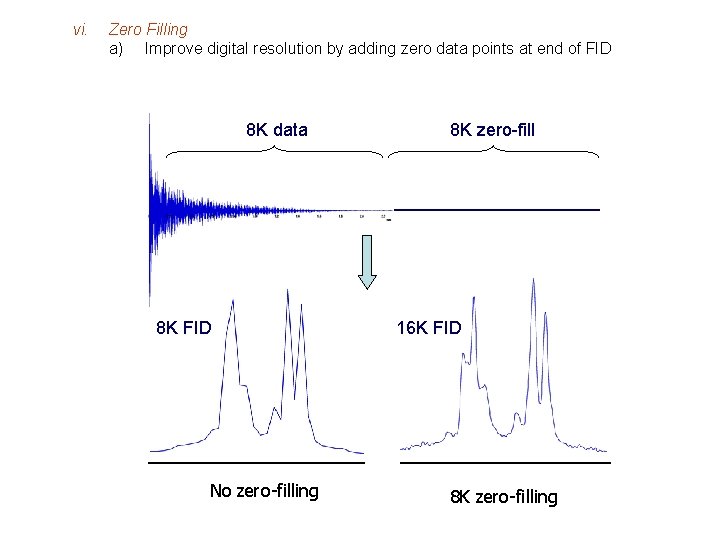
vi. Zero Filling a) Improve digital resolution by adding zero data points at end of FID 8 K data 8 K FID No zero-filling 8 K zero-fill 16 K FID 8 K zero-filling

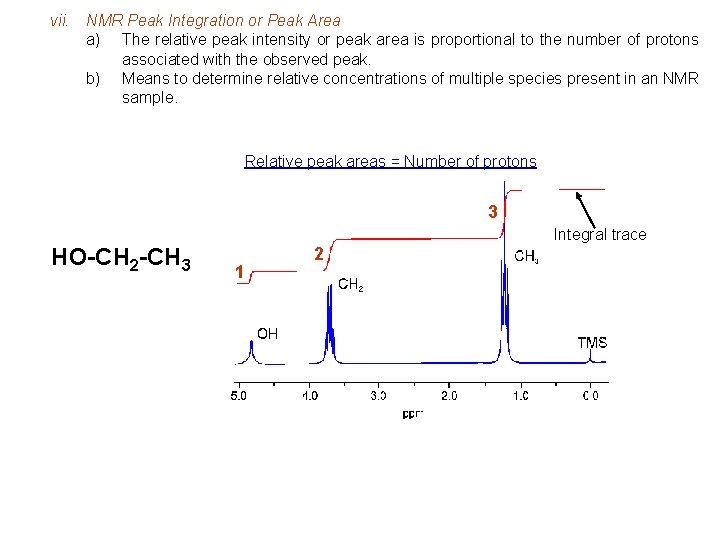
vii. NMR Peak Integration or Peak Area a) The relative peak intensity or peak area is proportional to the number of protons associated with the observed peak. b) Means to determine relative concentrations of multiple species present in an NMR sample. Relative peak areas = Number of protons 3 Integral trace HO-CH 2 -CH 3 1 2

Exchange Rates and NMR Time Scale i. Time Scale Slow Intermediate Fast Range (Sec-1) NMR time scale refers to the chemical shift time scale a) remember – frequency units are in Hz (sec-1) time scale b) exchange rate (k) c) differences in chemical shifts between species in exchange indicate the exchange rate. Chem. Shift (d) k << d. A- d. B k = d. A - d. B k >> d. A - d. B 0 – 1000 Coupling Const. (J) k << JA- JB k = J A - JB k >> JA- JB 0 – 12 T 2 relaxation k << 1/ T 2, A- 1/ T 2, B k = 1/ T 2, A- 1/ T 2, B k >> 1/ T 2, A- 1/ T 2, B 1 - 20 d) For systems in fast exchange, the observed chemical shift is the average of the individual species chemical shifts. dobs = f 1 d 1 + f 2 d 2 f 1 +f 2 =1 where: f 1, f 2 – mole fraction of each species d 1, d 2 – chemical shift of each species

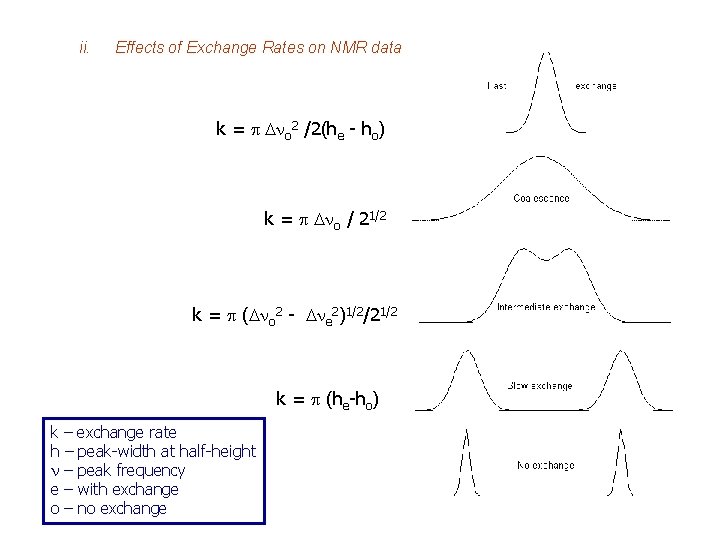
ii. Effects of Exchange Rates on NMR data k = p Dno 2 /2(he - ho) k = p Dno / 21/2 k = p (Dno 2 - Dne 2)1/2/21/2 k = p (he-ho) k – exchange rate h – peak-width at half-height n – peak frequency e – with exchange o – no exchange

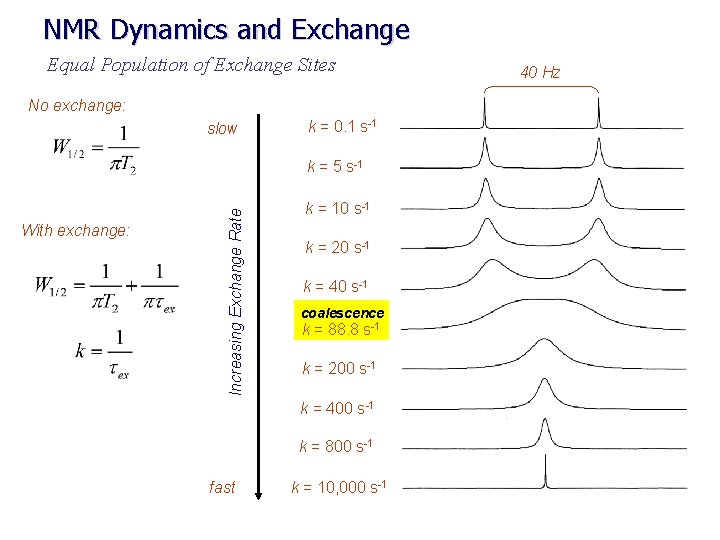
NMR Dynamics and Exchange Equal Population of Exchange Sites No exchange: slow k = 0. 1 s-1 With exchange: Increasing Exchange Rate k = 5 s-1 k = 10 s-1 k = 20 s-1 k = 40 s-1 coalescence k = 88. 8 s-1 k = 200 s-1 k = 400 s-1 k = 800 s-1 fast k = 10, 000 s-1 40 Hz

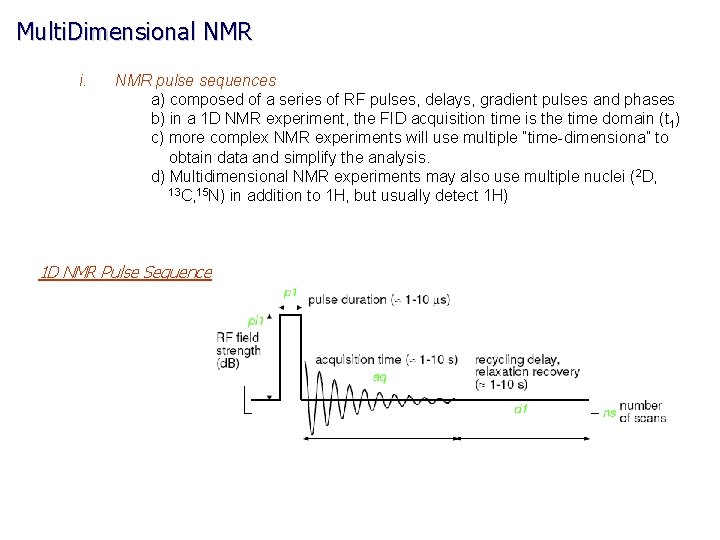
Multi. Dimensional NMR i. NMR pulse sequences a) composed of a series of RF pulses, delays, gradient pulses and phases b) in a 1 D NMR experiment, the FID acquisition time is the time domain (t 1) c) more complex NMR experiments will use multiple “time-dimensiona” to obtain data and simplify the analysis. d) Multidimensional NMR experiments may also use multiple nuclei (2 D, 13 C, 15 N) in addition to 1 H, but usually detect 1 H) 1 D NMR Pulse Sequence

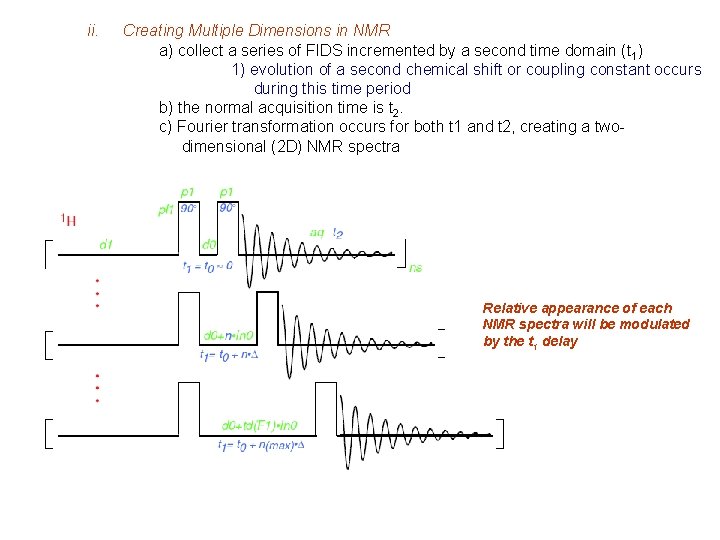
ii. Creating Multiple Dimensions in NMR a) collect a series of FIDS incremented by a second time domain (t 1) 1) evolution of a second chemical shift or coupling constant occurs during this time period b) the normal acquisition time is t 2. c) Fourier transformation occurs for both t 1 and t 2, creating a twodimensional (2 D) NMR spectra Relative appearance of each NMR spectra will be modulated by the t 1 delay

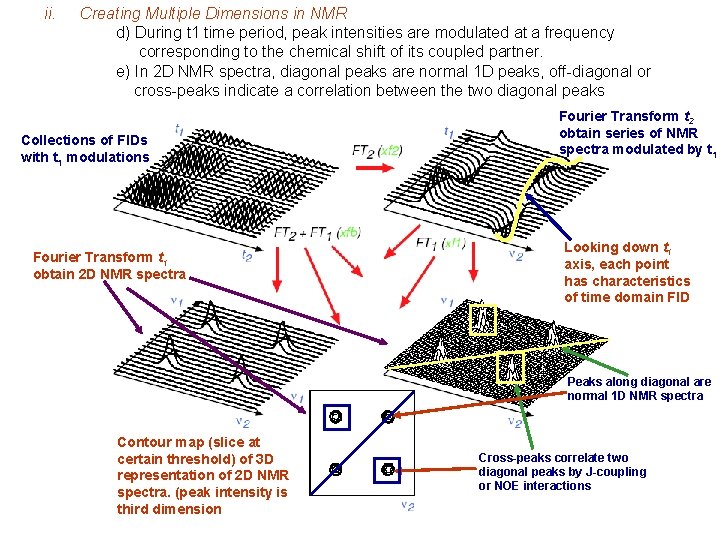
ii. Creating Multiple Dimensions in NMR d) During t 1 time period, peak intensities are modulated at a frequency corresponding to the chemical shift of its coupled partner. e) In 2 D NMR spectra, diagonal peaks are normal 1 D peaks, off-diagonal or cross-peaks indicate a correlation between the two diagonal peaks Collections of FIDs with t 1 modulations Fourier Transform t 1 obtain 2 D NMR spectra Fourier Transform t 2 obtain series of NMR spectra modulated by t 1 Looking down t 1 axis, each point has characteristics of time domain FID Peaks along diagonal are normal 1 D NMR spectra Contour map (slice at certain threshold) of 3 D representation of 2 D NMR spectra. (peak intensity is third dimension Cross-peaks correlate two diagonal peaks by J-coupling or NOE interactions

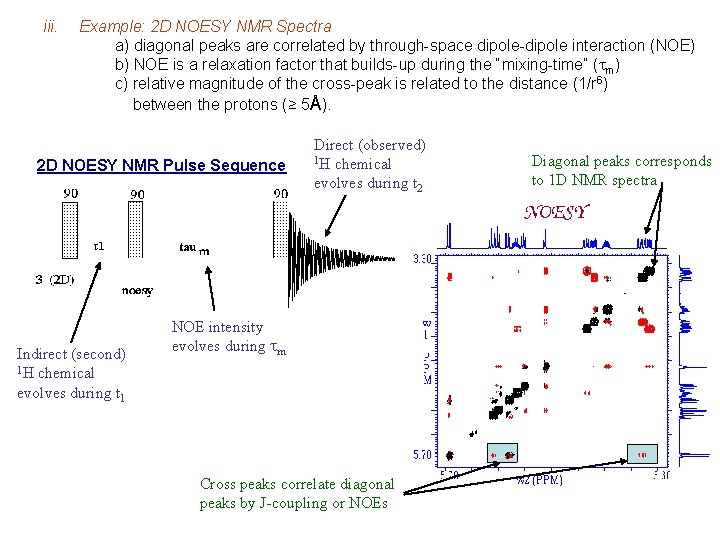
iii. Example: 2 D NOESY NMR Spectra a) diagonal peaks are correlated by through-space dipole-dipole interaction (NOE) b) NOE is a relaxation factor that builds-up during the “mixing-time” (tm) c) relative magnitude of the cross-peak is related to the distance (1/r 6) between the protons (≥ 5Å). 2 D NOESY NMR Pulse Sequence Indirect (second) 1 H chemical evolves during t 1 Direct (observed) 1 H chemical evolves during t 2 NOE intensity evolves during tm Cross peaks correlate diagonal peaks by J-coupling or NOEs Diagonal peaks corresponds to 1 D NMR spectra

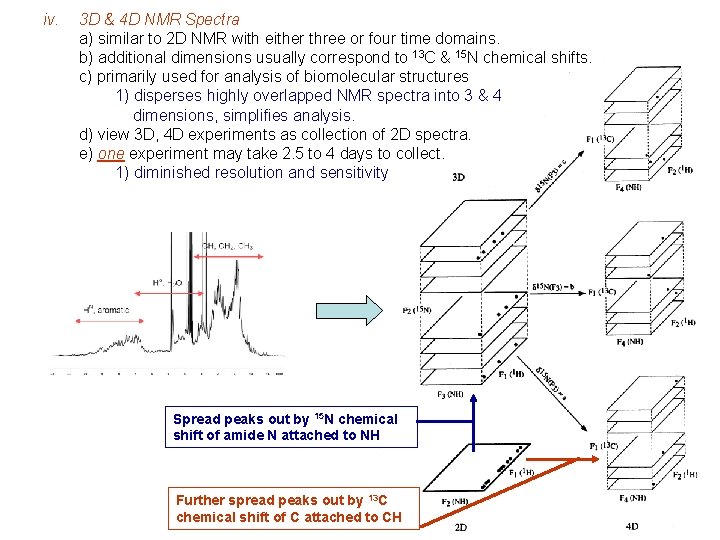
iv. 3 D & 4 D NMR Spectra a) similar to 2 D NMR with either three or four time domains. b) additional dimensions usually correspond to 13 C & 15 N chemical shifts. c) primarily used for analysis of biomolecular structures 1) disperses highly overlapped NMR spectra into 3 & 4 dimensions, simplifies analysis. d) view 3 D, 4 D experiments as collection of 2 D spectra. e) one experiment may take 2. 5 to 4 days to collect. 1) diminished resolution and sensitivity Spread peaks out by 15 N chemical shift of amide N attached to NH Further spread peaks out by 13 C chemical shift of C attached to CH

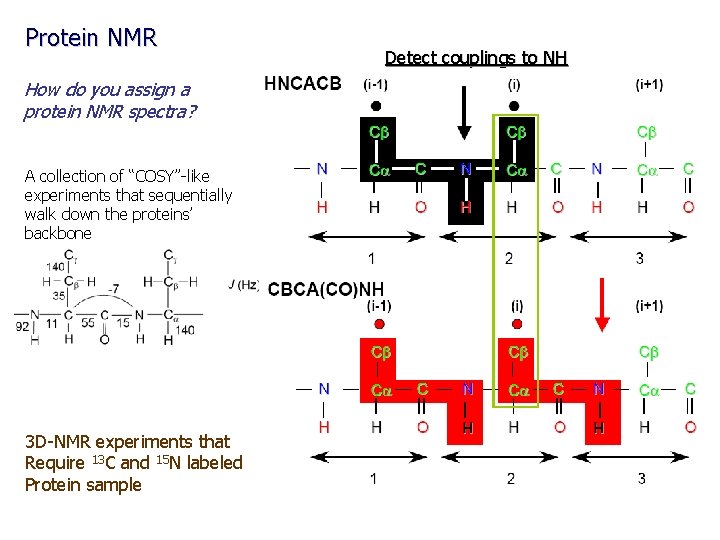
Protein NMR How do you assign a protein NMR spectra? A collection of “COSY”-like experiments that sequentially walk down the proteins’ backbone 3 D-NMR experiments that Require 13 C and 15 N labeled Protein sample Detect couplings to NH

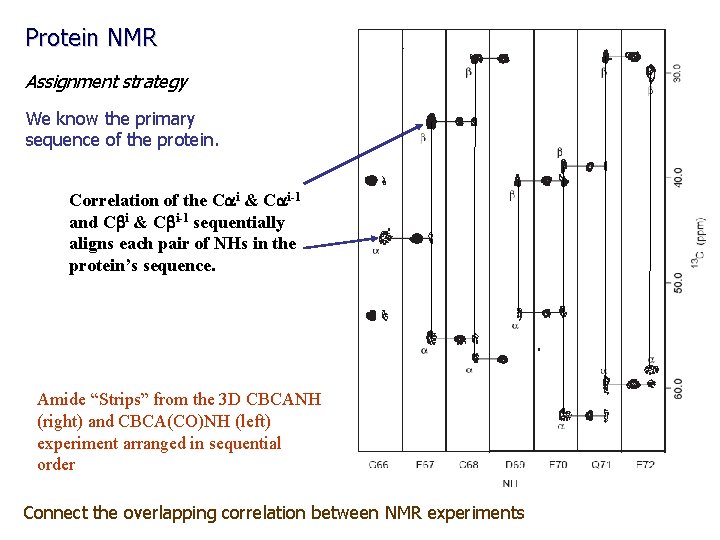
Protein NMR Assignment strategy We know the primary sequence of the protein. Correlation of the Cai & Cai-1 and Cbi & Cbi-1 sequentially aligns each pair of NHs in the protein’s sequence. Amide “Strips” from the 3 D CBCANH (right) and CBCA(CO)NH (left) experiment arranged in sequential order Connect the overlapping correlation between NMR experiments


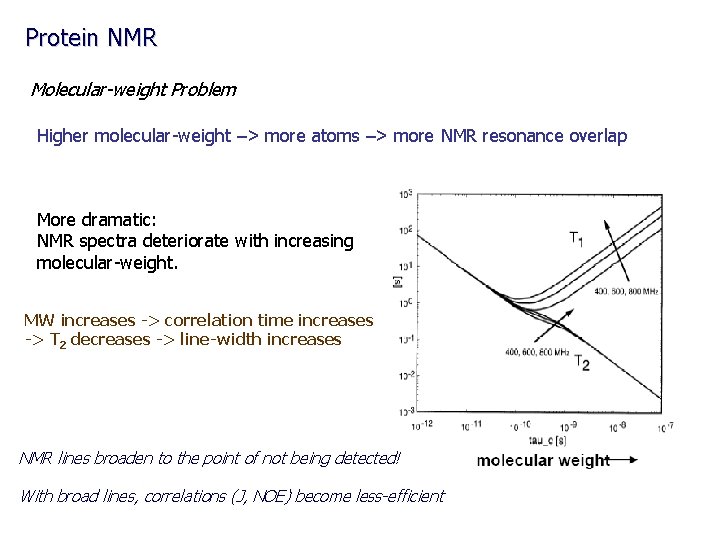
Protein NMR Molecular-weight Problem Higher molecular-weight –> more atoms –> more NMR resonance overlap More dramatic: NMR spectra deteriorate with increasing molecular-weight. MW increases -> correlation time increases -> T 2 decreases -> line-width increases NMR lines broaden to the point of not being detected! With broad lines, correlations (J, NOE) become less-efficient

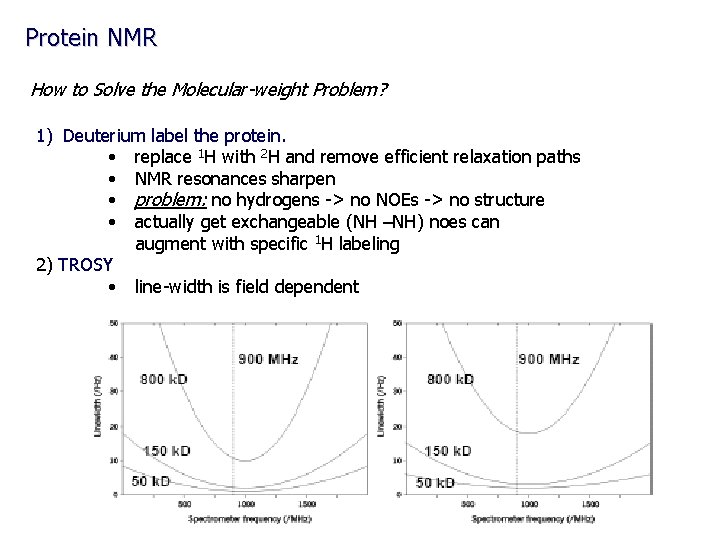
Protein NMR How to Solve the Molecular-weight Problem? 1) Deuterium label the protein. • replace 1 H with 2 H and remove efficient relaxation paths • NMR resonances sharpen • problem: no hydrogens -> no NOEs -> no structure • actually get exchangeable (NH –NH) noes can augment with specific 1 H labeling 2) TROSY • line-width is field dependent