NSFEU Cyberinfrastructure Meeting Washington DC Annotating Metagenomes Using

NSF/EU Cyberinfrastructure Meeting, Washington, DC. Annotating Metagenomes Using the SEED Rob Edwards Department of Computer Sciences, San Diego State University Mathematics and Computer Sciences Division, Argonne National Laboratory www. nmpdr. org www. theseed. org

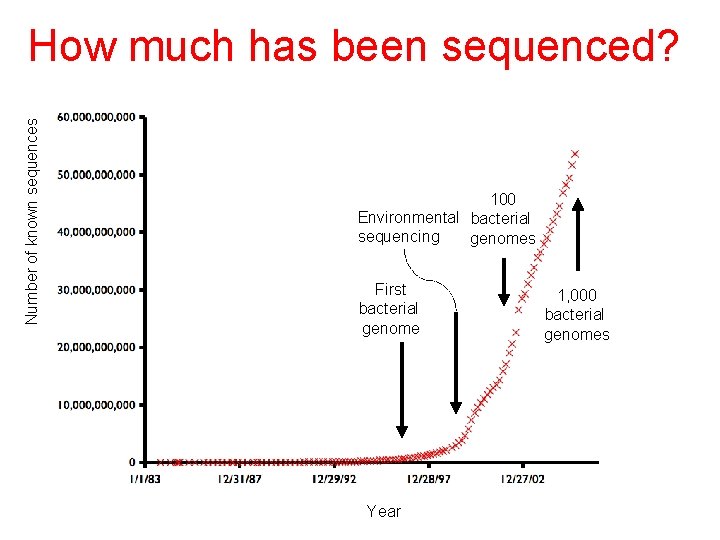
Number of known sequences How much has been sequenced? 100 Environmental bacterial sequencing genomes First bacterial genome Year 1, 000 bacterial genomes
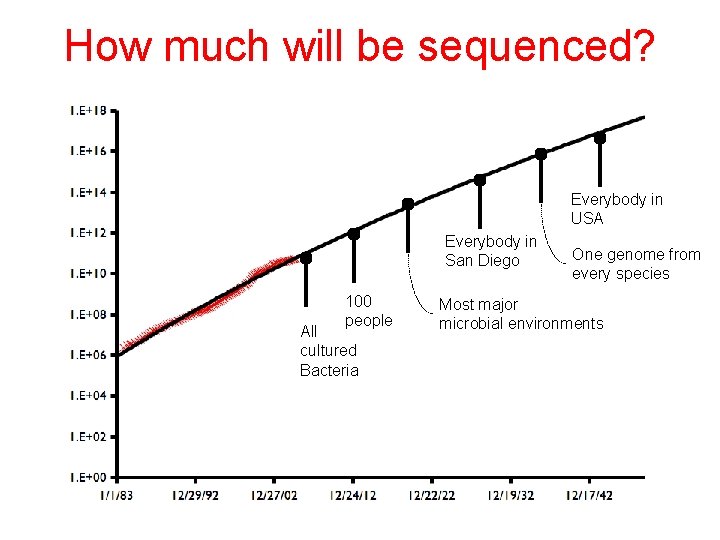
How much will be sequenced? Everybody in USA Everybody in San Diego 100 people All cultured Bacteria One genome from every species Most major microbial environments

What do we want from annotations? Consistent Accurate Available Reliable www. nmpdr. org www. theseed. org

Consistent www. nmpdr. org www. theseed. org

The Importance of Consistency • Consistency: same genes connected to same functional role • Enables communication • Required for most comparative genomics assays www. nmpdr. org www. theseed. org

his. A FIG function: Phosphoribosylformimino-5 -aminoimidazole carboxamide ribotide isomerase (EC 5. 3. 1. 16) Other functions in Ref. Seq: phosphoribosylformimino-5 -aminoimidazole carboxamide ribotide isomerase phosphoribosylformimino-5 -aminoimidazole carboxamide ribotide. . . 1 -(5 -phosphoribosyl)-5 -[(5 - phosphoribosylamino)methylideneamino] imidazole-4 -carboxamide isomerase N-(5 -phospho-L-ribosyl-formimino)-5 -amino-1 -(5 - phosphoribosyl)-4 -imidazolecarboxamide isomerase N-(5'-phospho-L-ribosyl-formimino)-5 -amino-1 -(5'-phosphoribosyl)-4 -imidazolecarboxamide isomerase N-(5'-phospho-L-ribosyl-formimino)-5 -amino-1 - (5'-phosphoribosyl)-4 - imidazolecarboxamide isomerase Phosphoribosyl isomerase A [1 -[5 -phosphoribosyl]-5 -[[5 -phosphoribosylamino]methylideneamino] imidazole-4 -carboxamide isomerase] www. nmpdr. org www. theseed. org

Measuring Consistency • Define a set of protein families such that each family contains genes playing the same function • Attach functional roles to protein families • Measure the consistency of the annotations made to genes within each family 1. "consistency" is the odds that two proteins from the same family have the same function 2. Evaluate both families and functions. www. nmpdr. org www. theseed. org
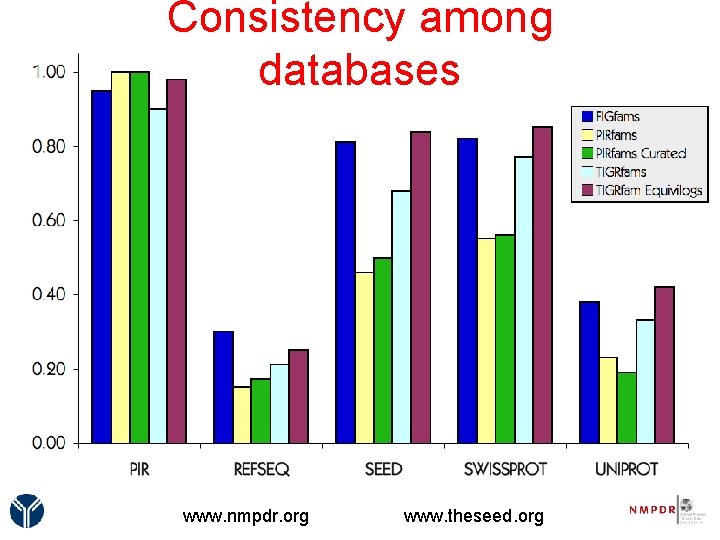
Consistency among databases www. nmpdr. org www. theseed. org

Accurate www. nmpdr. org www. theseed. org

How to measure accuracy • If everything was called “hypothetical protein” the database would be 100% consistent • Need to measure accuracy (specificity) as well as consistency • Sample 100 proteins at random from “curated” set (i. e. that are believed to be correct) • Manually inspect annotations to score correctness www. nmpdr. org www. theseed. org

Available www. nmpdr. org www. theseed. org

http: //metagenomics. theseed. org Free service User registration/log in Free to upload sequences in several formats Automatically annotates sequences Download in several formats Complete genomes too: http: //www. nmpdr. org/anno-server Soon to come: Plasmids, phages, other short genomes
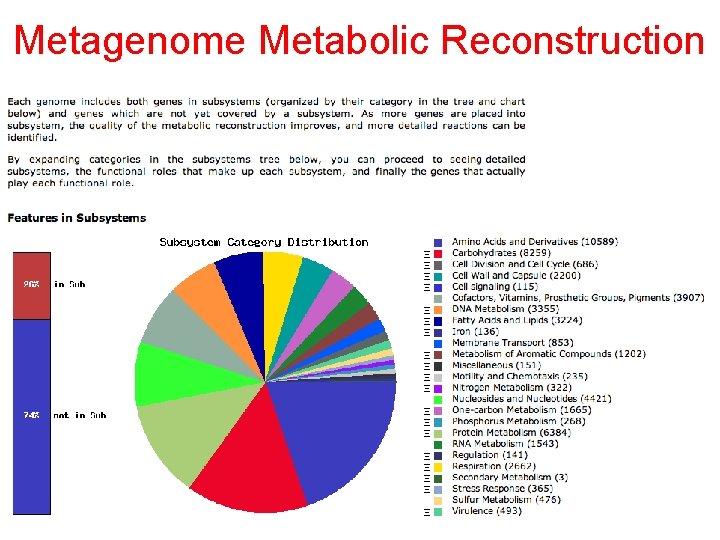
Metagenome Metabolic Reconstruction
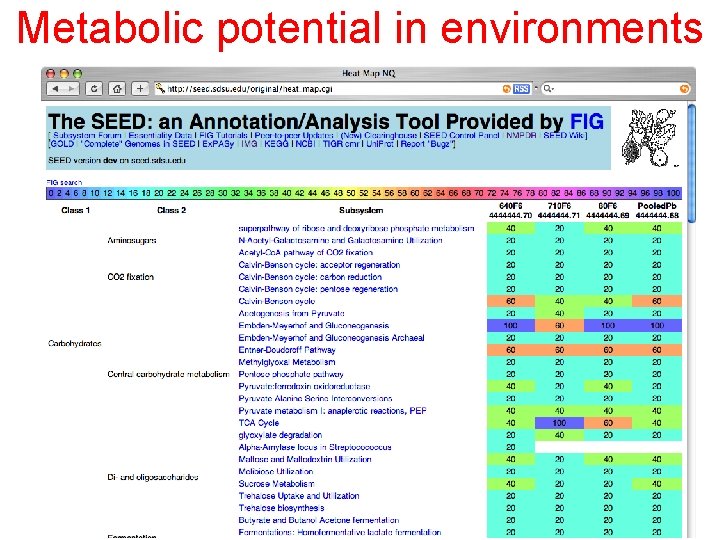
Metabolic potential in environments
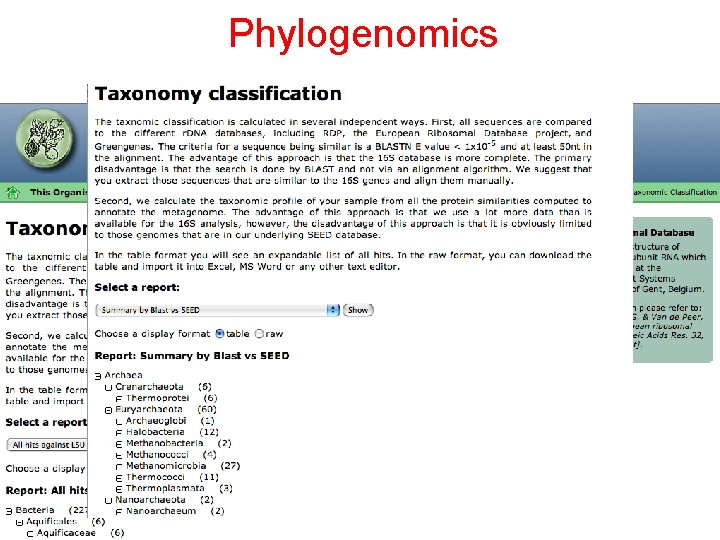
Phylogenomics
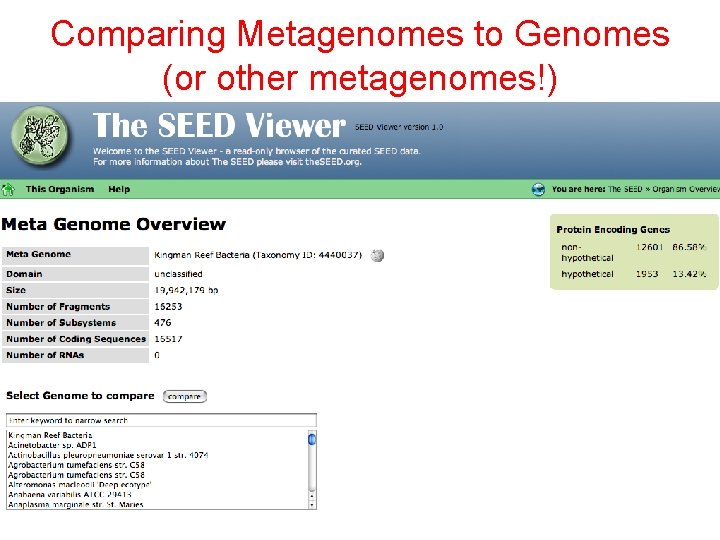
Comparing Metagenomes to Genomes (or other metagenomes!)

Reliable (Believable)
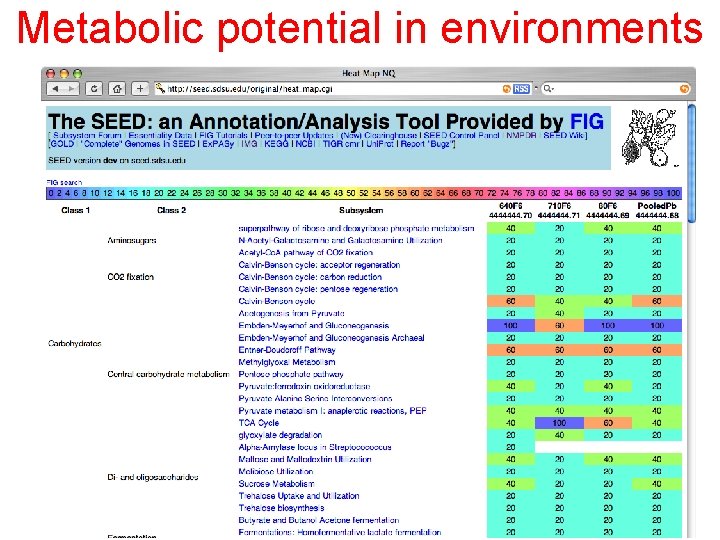
Metabolic potential in environments
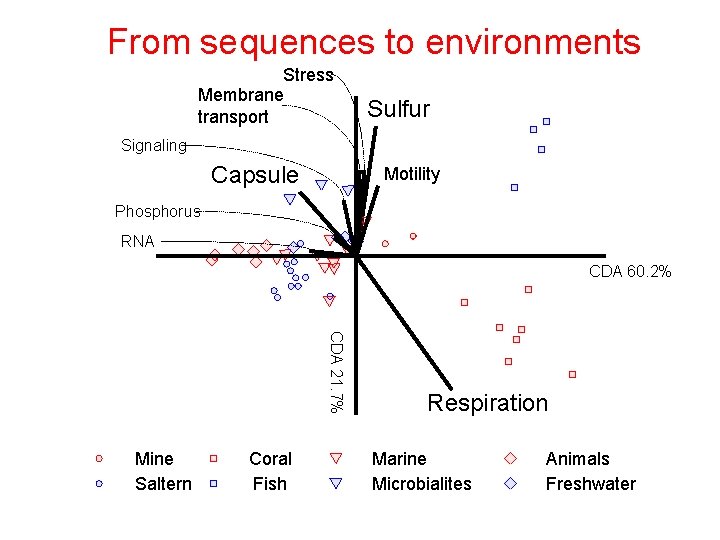
From sequences to environments Stress Membrane transport Sulfur Signaling Capsule Motility Phosphorus RNA CDA 60. 2% CDA 21. 7% Mine Saltern Coral Fish Respiration Marine Microbialites Animals Freshwater

What do we want from annotations? W t Consistent n a w Accurate e w W o Available d NO n e h Reliable ti ?

Acknowledgements Environmental Genomics Statistics Forest Rohwer Liz Dinsdale Rohwer lab members Dana Hall All the labs that Beltran Rodriguez-Brito provided sequence Metagenomics Annotation Server FIG Rick Stevens Daniel Paarman Folker Meyer Bob Olsen Ross Overbeek Veronika Vonstein Annotators

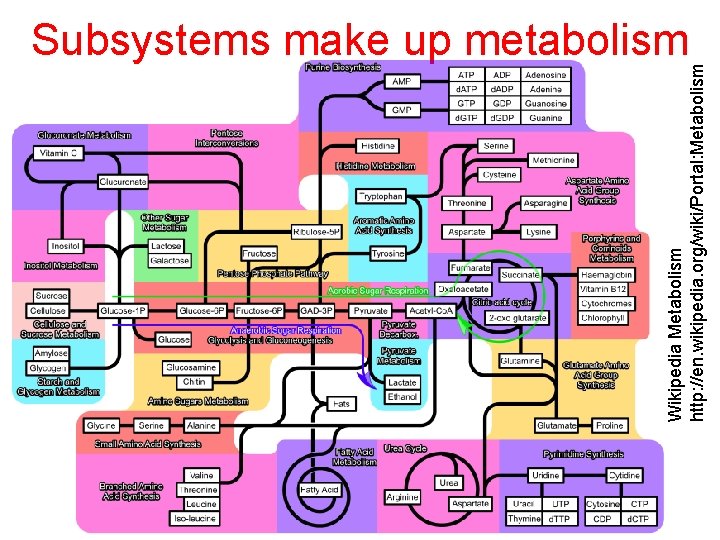
Wikipedia Metabolism http: //en. wikipedia. org/wiki/Portal: Metabolism Subsystems make up metabolism
- Slides: 25