Normalization and Statistical Analysis DNA Microarray Bioinformatics 27612

Normalization and Statistical Analysis DNA Microarray Bioinformatics - #27612
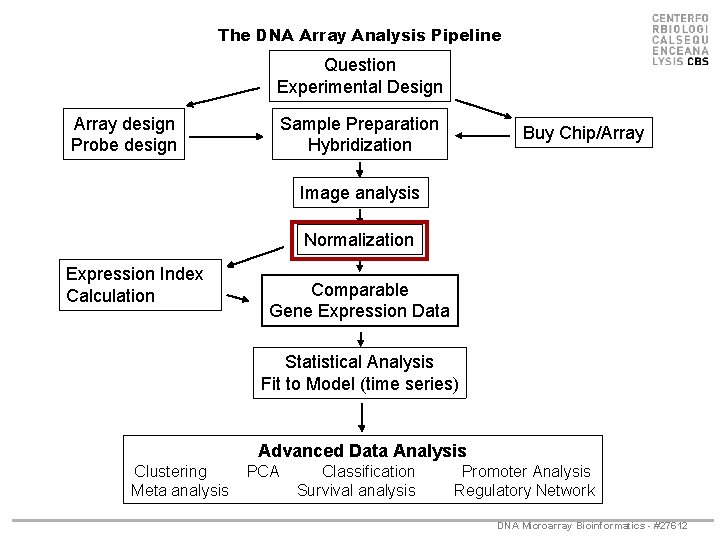
The DNA Array Analysis Pipeline Question Experimental Design Array design Probe design Sample Preparation Hybridization Buy Chip/Array Image analysis Normalization Expression Index Calculation Comparable Gene Expression Data Statistical Analysis Fit to Model (time series) Advanced Data Analysis Clustering Meta analysis PCA Classification Survival analysis Promoter Analysis Regulatory Network DNA Microarray Bioinformatics - #27612
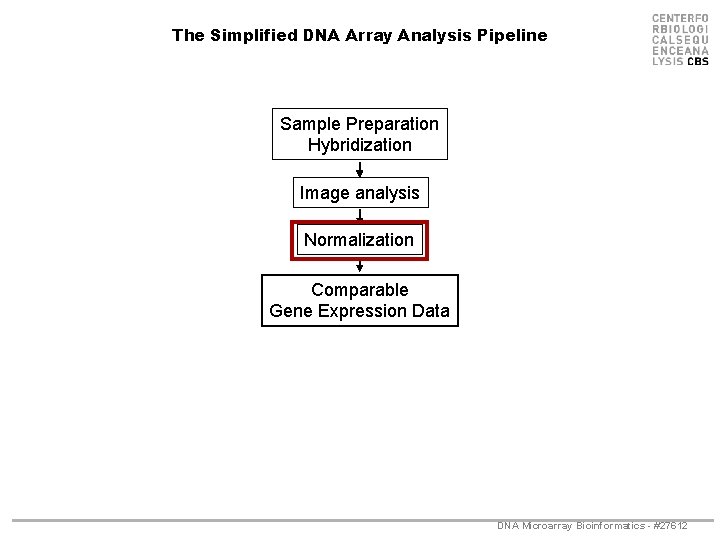
The Simplified DNA Array Analysis Pipeline Sample Preparation Hybridization Image analysis Normalization Comparable Gene Expression Data DNA Microarray Bioinformatics - #27612
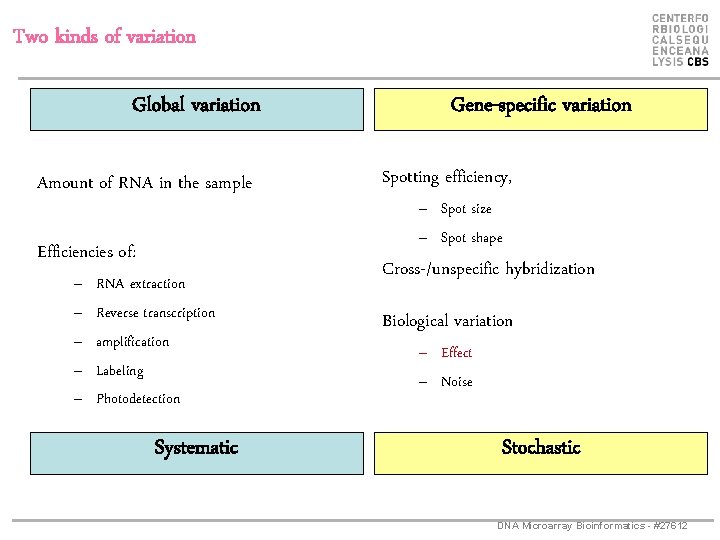
Two kinds of variation Global variation Amount of RNA in the sample Efficiencies of: – – – RNA extraction Reverse transcription amplification Labeling Photodetection Systematic Gene-specific variation Spotting efficiency, – Spot size – Spot shape Cross-/unspecific hybridization Biological variation – Effect – Noise Stochastic DNA Microarray Bioinformatics - #27612

Sources of variation Global variation: Gene-specific variation: Systematic Stochastic • Similar effect on many measurements • Corrections can be estimated from data Normalization • Too random to be explicitly accounted for • “noise” Statistical testing DNA Microarray Bioinformatics - #27612

Intensities are not just m. RNA concentrations • • • Tissue contamination RNA degradation RNA purification Reverse transcription Amplification efficiency Dye effect (cy 3/cy 5) • Spotting • DNA-support binding • Other issues related to array manufacturing • ‘Background’ correction • Image segmentation • Hybridization efficiency and specificity • Spatial effects DNA Microarray Bioinformatics - #27612

Calibration = Normalization = Scaling DNA Microarray Bioinformatics - #27612
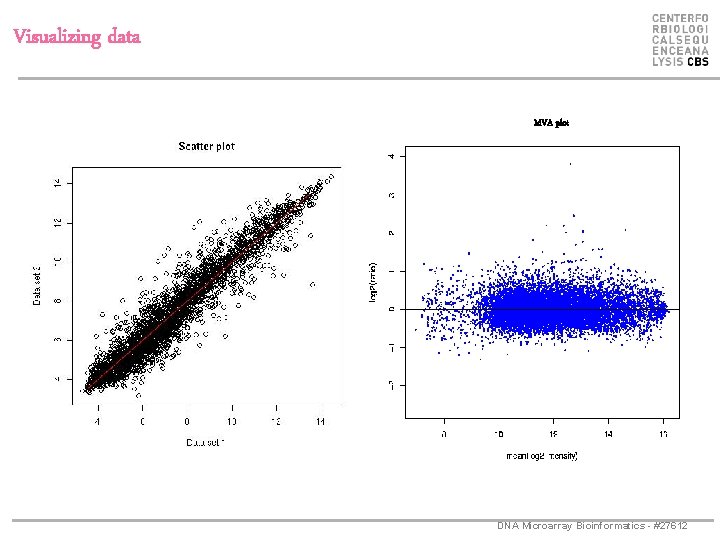
Visualizing data MVA plot DNA Microarray Bioinformatics - #27612

Linear normalization DNA Microarray Bioinformatics - #27612
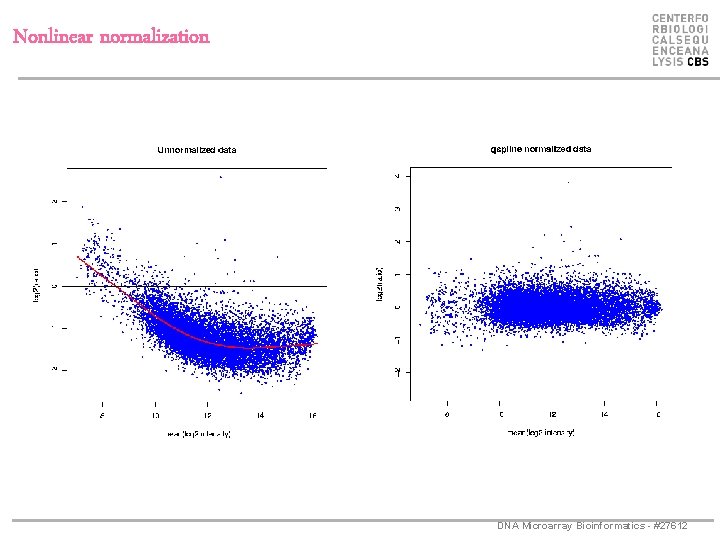
Nonlinear normalization DNA Microarray Bioinformatics - #27612

MAS 5. 0 Normalization (Affymetrix) • Background: Weighted average of the lowest 2% • Uses MM to calculate the ideal mismatch, then adjust the PM intensity • Uses the Tukey’s biweight estimator to provide a robust mean • Scale to make the means equale for all chips using trimmed mean DNA Microarray Bioinformatics - #27612
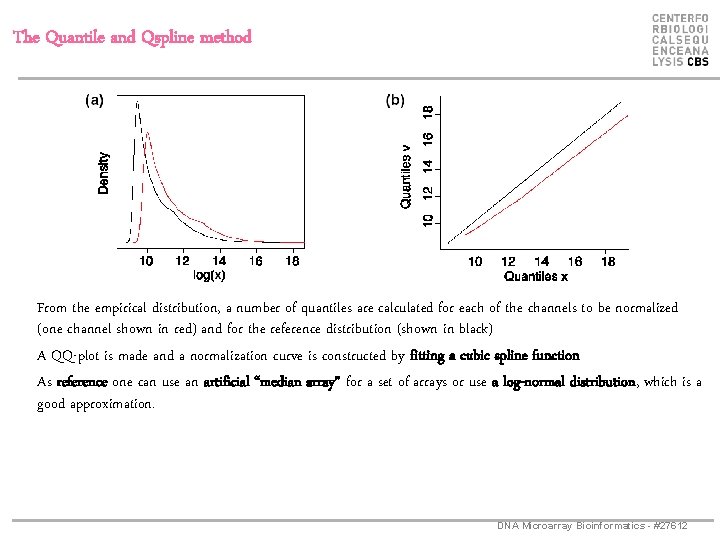
The Quantile and Qspline method From the empirical distribution, a number of quantiles are calculated for each of the channels to be normalized (one channel shown in red) and for the reference distribution (shown in black) A QQ-plot is made and a normalization curve is constructed by fitting a cubic spline function As reference one can use an artificial “median array” for a set of arrays or use a log-normal distribution, which is a good approximation. DNA Microarray Bioinformatics - #27612
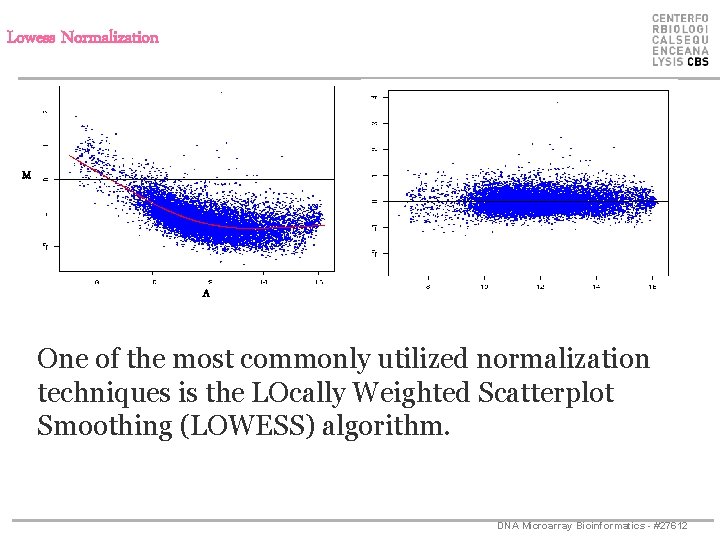
Lowess Normalization * M * * * A One of the most commonly utilized normalization techniques is the LOcally Weighted Scatterplot Smoothing (LOWESS) algorithm. DNA Microarray Bioinformatics - #27612

Invariant set normalization (Li and Wong) A invariant set of probes is used -Probes that does not change intensity rank between arrays -A piecewise linear median line is calculated -This curve is used for normalization DNA Microarray Bioinformatics - #27612

Spatial normalization Raw data After intensity normalization Spatial bias estimate After spatial normalization DNA Microarray Bioinformatics - #27612
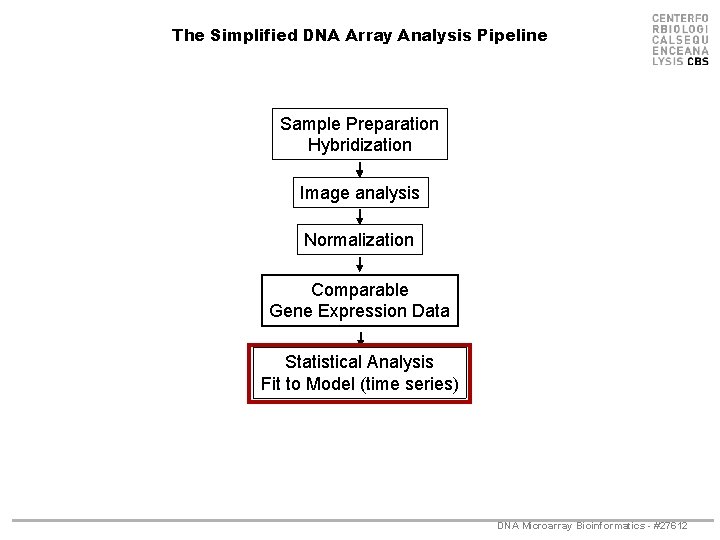
The Simplified DNA Array Analysis Pipeline Sample Preparation Hybridization Image analysis Normalization Comparable Gene Expression Data Statistical Analysis Fit to Model (time series) DNA Microarray Bioinformatics - #27612

The t-test A test for whether two distrubutions has the same mean. Gives a p-value for each gene (need replications). You need to adjust your p-value for multiple testing. Bonferroni correction: P = 0. 05/N DNA Microarray Bioinformatics - #27612
- Slides: 17