NMR Spectroscopy Judith KleinSeetharaman Department of Structural Biology

NMR Spectroscopy Judith Klein-Seetharaman Department of Structural Biology jks 33@pitt. edu Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture

Objectives of this Lecture and Practicum • • 12/5/2020 Resources Physical principle Sample requirements Parameters that are measured by NMR Dynamics by NMR Limitations Practical aspects Setup of NMR experiments (downstairs) Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 2

Resources Websites • • • • http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf http: //www. bmrb. wisc. edu/ http: //www. biochem. ucl. ac. uk/bsm/nmr/ubq/ http: //nobelprize. org/nobel_prizes/chemistry/laureates/2002/wutrich-lecture. pdf http: //www. cis. rit. edu/htbooks/nmr/ http: //www. ch. ic. ac. uk/local/organic/nmr. html http: //www. spectroscopynow. com/ http: //www. chem. queensu. ca/FACILITIES/NMR/nmr/webcourse/ http: //spincore. com/nmrinfo/ http: //www. chembio. uoguelph. ca/driguana/NMR/TOC. HTM http: //www. embl-heidelberg. de/nmr/sattler/embo/handouts/griesinger_lecture_pof. pdf http: //dupont. molbio. ku. dk/teach/course/intro. NMR. pdf http: //www. infochembio. ethz. ch/links/en/spectrosc_nmr_lehr. html 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 3

Resources Books NMR Books: • Protein NMR Techniques (Methods in Molecular Biology) by A. Kristina Downing (Editor) • Protein NMR Spectroscopy: Principles and Practice by John Cavanagh, Wayne J. Fairbrother, III, Arthur G. Palmer, Nicholas J. Skelton, Mark Rance • Spin Dynamics: Basics of Nuclear Magnetic Resonance by Malcolm H. Levitt • Principles of Nuclear Magnetic Resonance in One and Two Dimensions by Richard R. Ernst, Geoffrey Bodenhausen, Alexander Wokaun • 200 and More NMR Experiments: A Practical Course by Stefan Berger, Siegmar Braun • Basic One- and Two-Dimensional NMR Spectroscopy by Horst Friebolin • NMR Spectroscopy: Basic Principles, Concepts, and Applications in Chemistry by Harald Günther • NMR Data Processing by Hoch • NMR: The Toolkit by P. J. Hore, J. A. Jones, S. Wimperis • Nuclear Magnetic Resonance by P. J. Hore • NMR for Physical and Biological Scientists by Thoma Pochapsky • Understanding NMR Spectroscopy by James Keeler • NMR of Proteins (Topics on Molecular and Structural Biology) by G. M. Clore, A. M. Gronenborn • The Nuclear Overhauser Effect in Structural and Conformational Analysis by David Neuhaus, Michael P. Williamson Biophysics Books with chapters on NMR: • Biophysical Chemistry: Part II: Techniques for the Study of Biological Structure and Function by Charles R Cantor, Paul R Schimmel • Principles of Physical Biochemistry by Kensal E van Holde, Curtis Johnson, Pui Shing Ho 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 4

Objectives of this Lecture and Practicum • • 12/5/2020 Resources Physical principle Sample requirements Parameters that are measured by NMR Dynamics by NMR Limitations Practical aspects Setup of NMR experiments (downstairs) Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 5
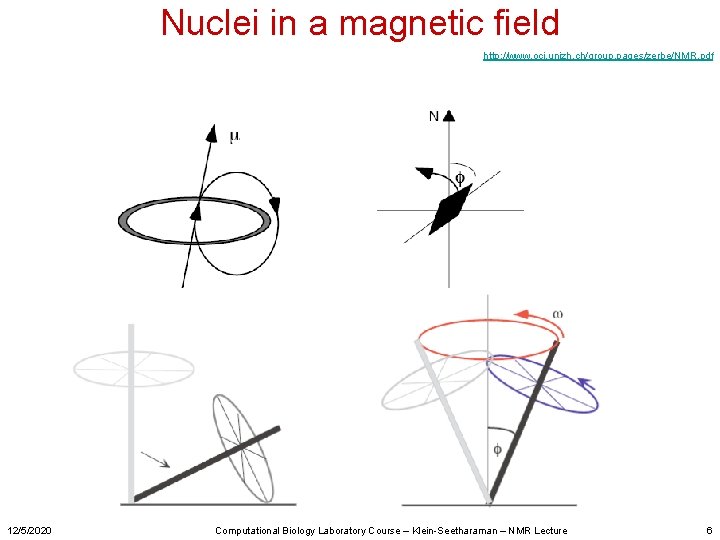
Nuclei in a magnetic field http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 6
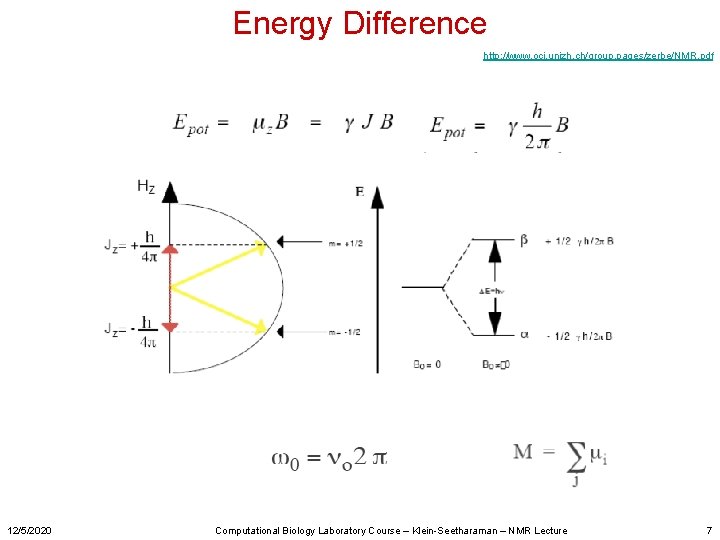
Energy Difference http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 7
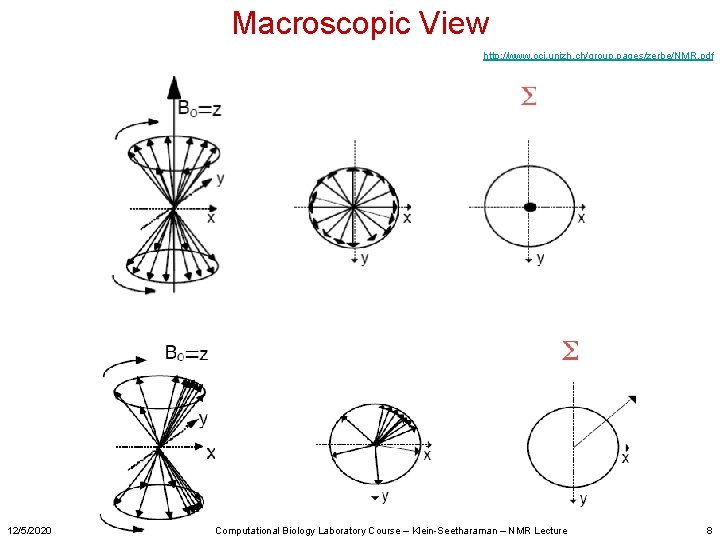
Macroscopic View http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 8
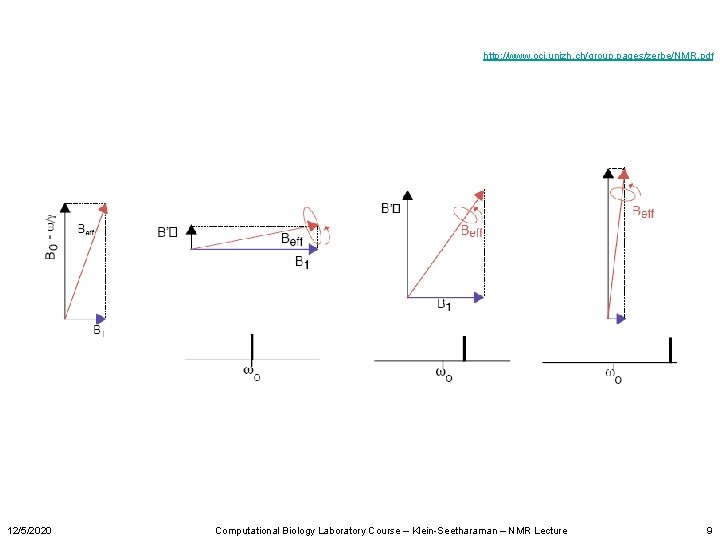
http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 9
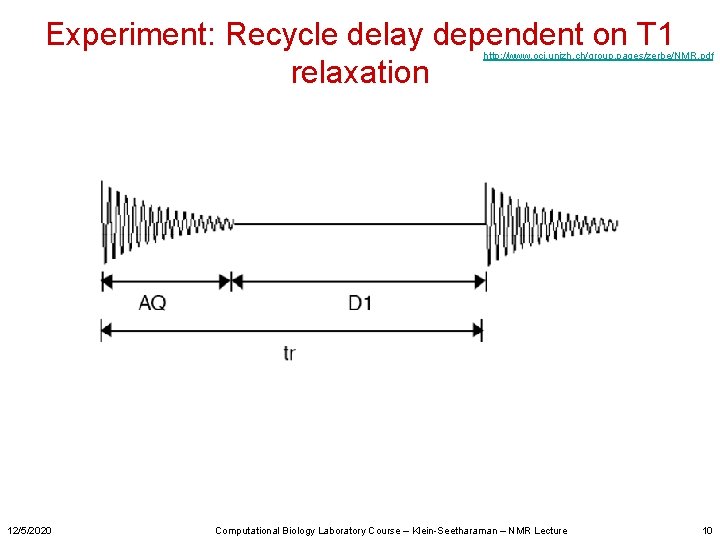
Experiment: Recycle delay dependent on T 1 relaxation http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 10
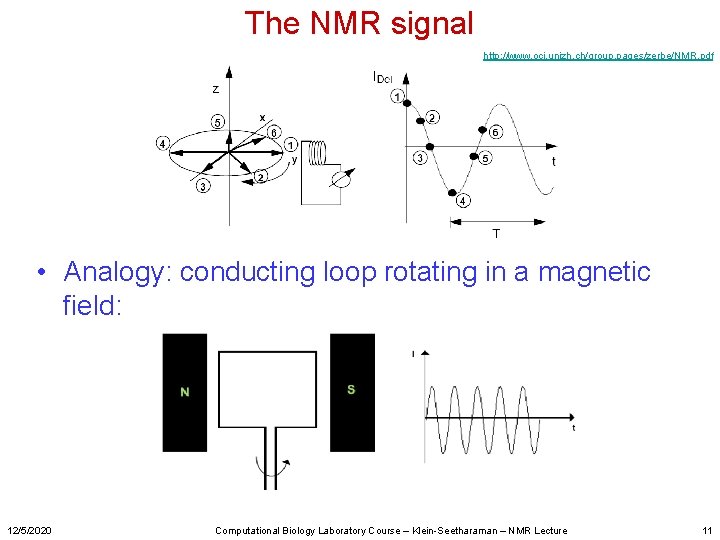
The NMR signal http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf • Analogy: conducting loop rotating in a magnetic field: 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 11
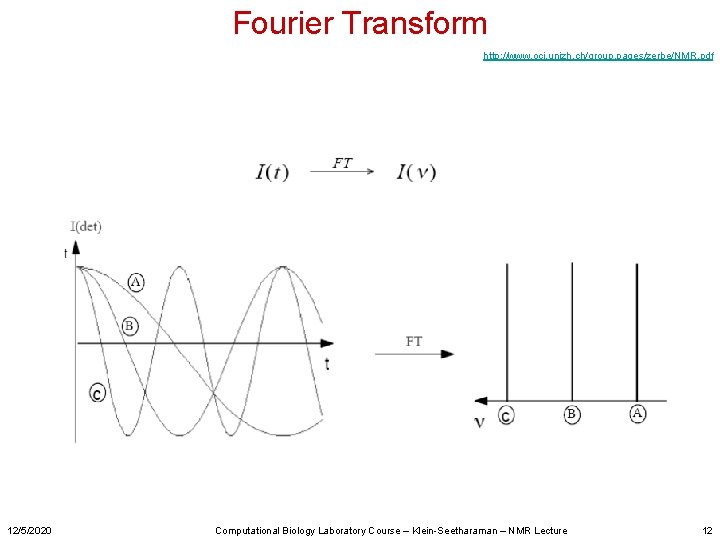
Fourier Transform http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 12
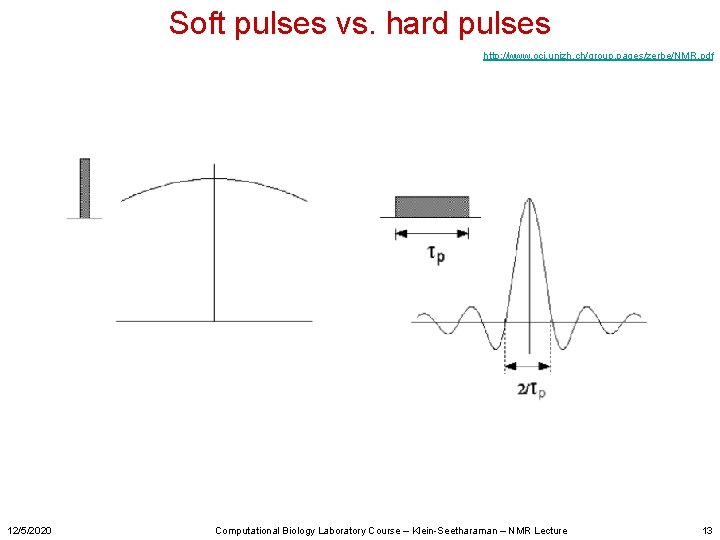
Soft pulses vs. hard pulses http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 13
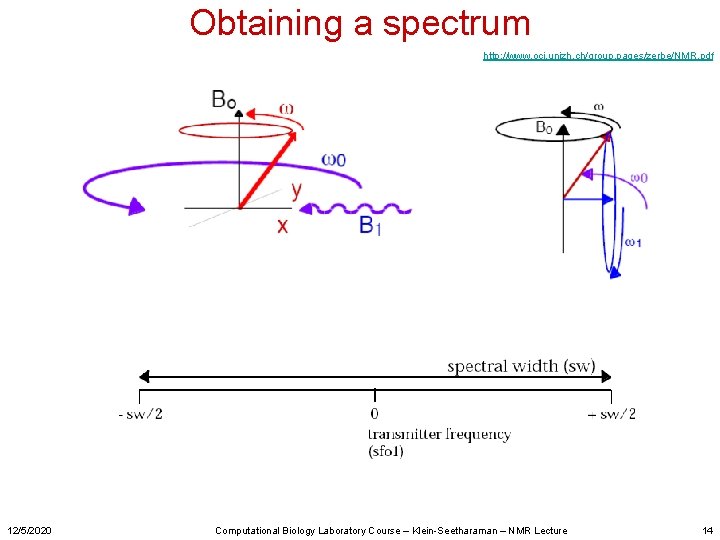
Obtaining a spectrum http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 14

Product Operator Formalism http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 15
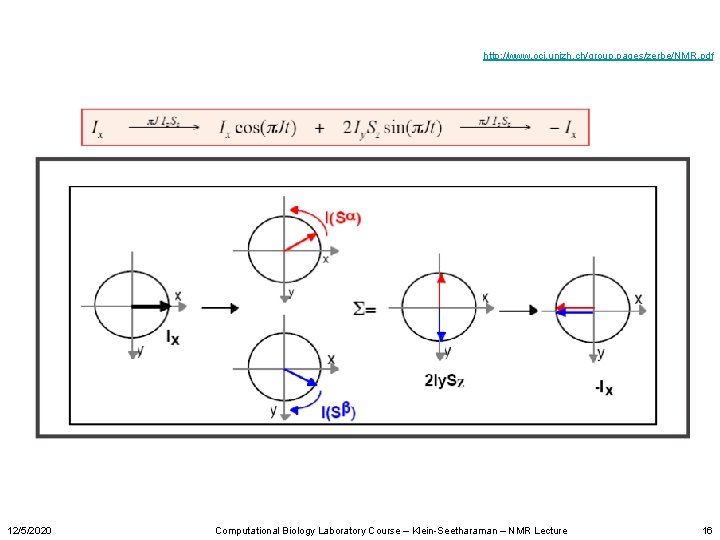
http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 16

http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 17
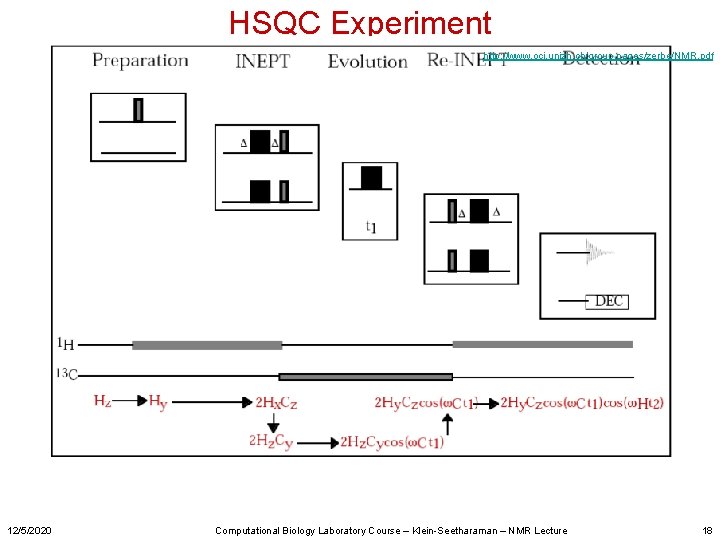
HSQC Experiment http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 18
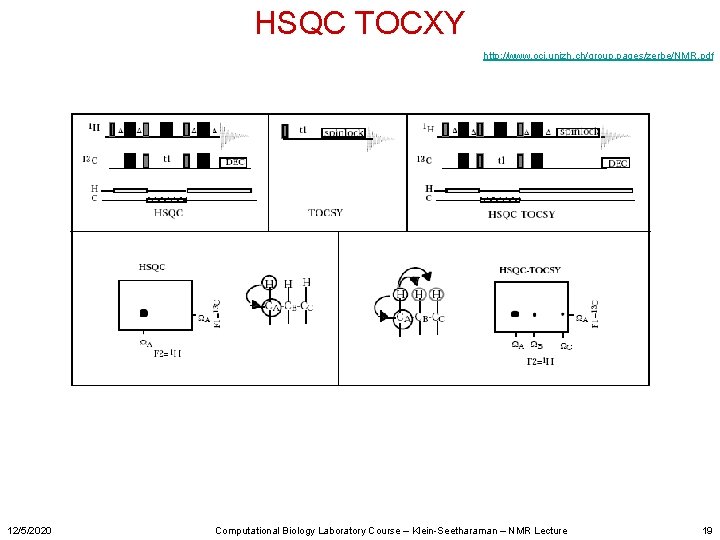
HSQC TOCXY http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 19
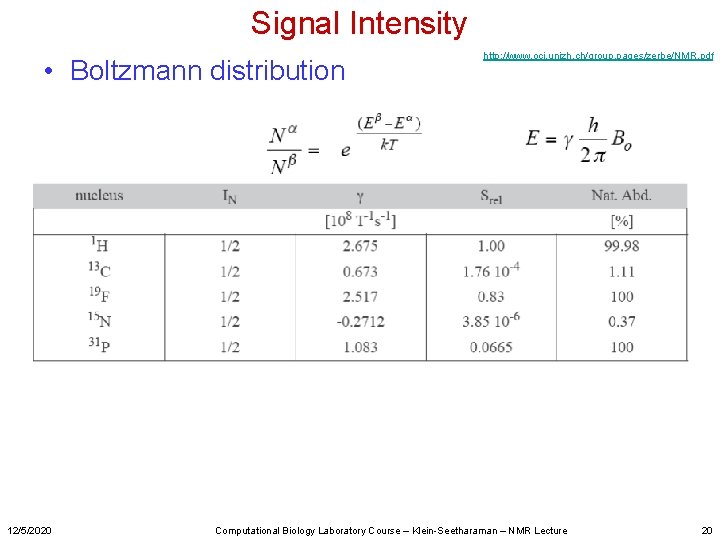
Signal Intensity • Boltzmann distribution 12/5/2020 http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 20

Objectives of this Lecture and Practicum • • 12/5/2020 Resources Physical principle Sample requirements Parameters that are measured by NMR Dynamics by NMR Limitations Practical aspects Setup of NMR experiments (downstairs) Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 21
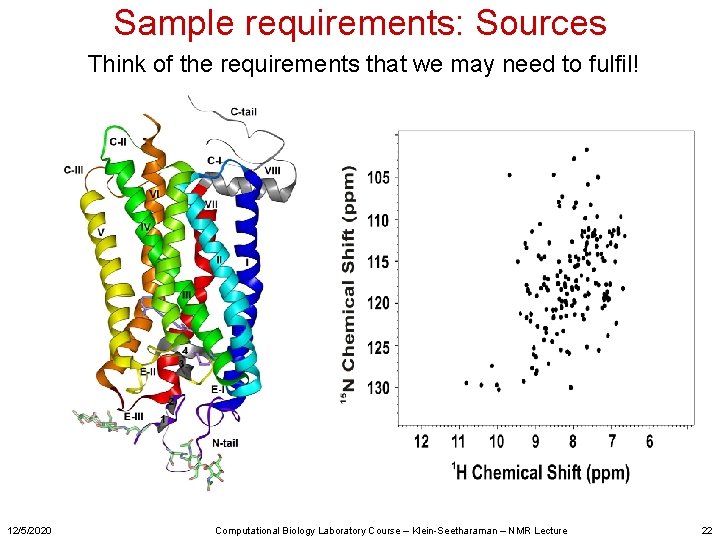
Sample requirements: Sources Think of the requirements that we may need to fulfil! 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 22
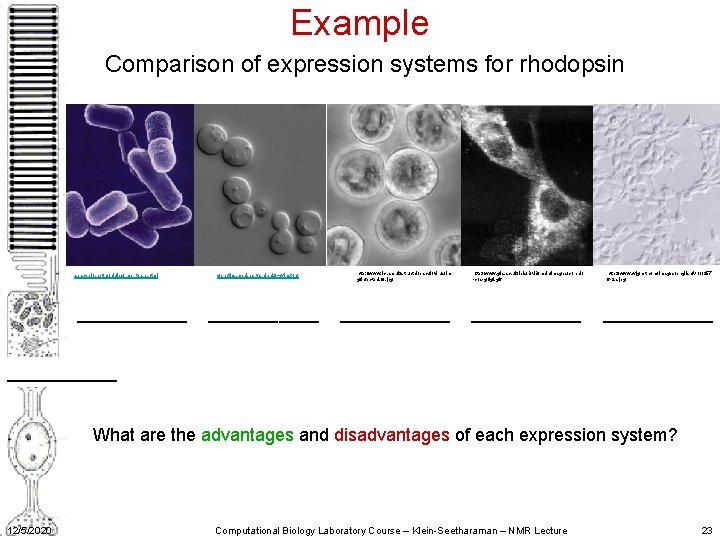
Example Comparison of expression systems for rhodopsin spacebio. net/modules/ mb_teare. html ______ genetics. med. harvard. edu/ ~winston/ ______ http: //www. icr. ac. uk/structbi/baculovirus/im g/infectedsf 9. jpg ______ http: //www. gla. ac. uk/ibls/BMB/mdh/images/conrd 1 -cos-golgi. gif ______ http: //www. wjgnet. com/images/english/V 11/257 6 -2 a. jpg ___________ What are the advantages and disadvantages of each expression system? 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 23
Where can we get these molecules from? 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 24

Sources of biomolecules • Native sources Summary – Best quality (correct fold, posttranslational modifications etc. ) – Not always best quantity – Limitations in labeling – No mutants • Chemical synthesis – Good for small molecules – Not good for large proteins • Biosynthesis – A variety of expression systems exist, all with their advantages and disadvantages. – Required for isotope labeling for NMR 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 25

Objectives of this Lecture and Practicum • • 12/5/2020 Resources Physical principle Sample requirements Parameters that are measured by NMR Dynamics by NMR Limitations Practical aspects Setup of NMR experiments (downstairs) Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 26
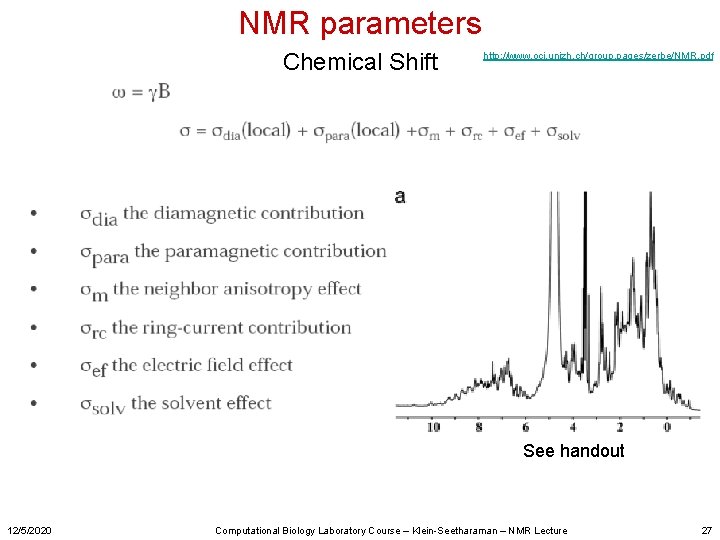
NMR parameters Chemical Shift http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf See handout 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 27
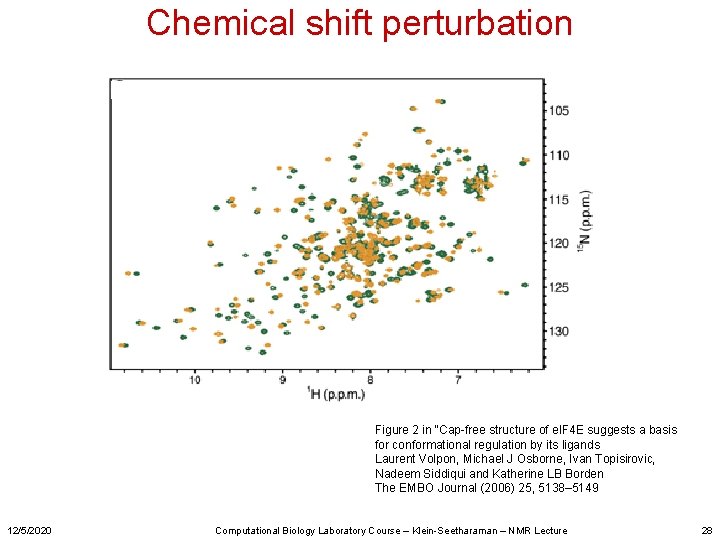
Chemical shift perturbation Figure 2 in “Cap-free structure of e. IF 4 E suggests a basis for conformational regulation by its ligands Laurent Volpon, Michael J Osborne, Ivan Topisirovic, Nadeem Siddiqui and Katherine LB Borden The EMBO Journal (2006) 25, 5138– 5149 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 28
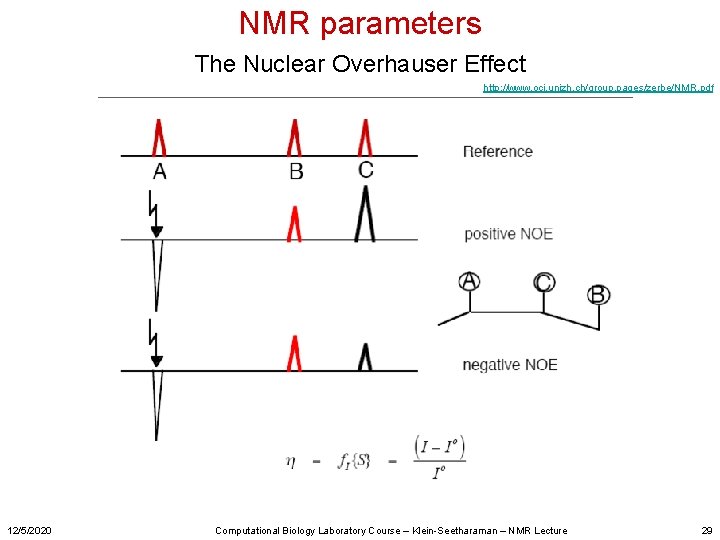
NMR parameters The Nuclear Overhauser Effect http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 29
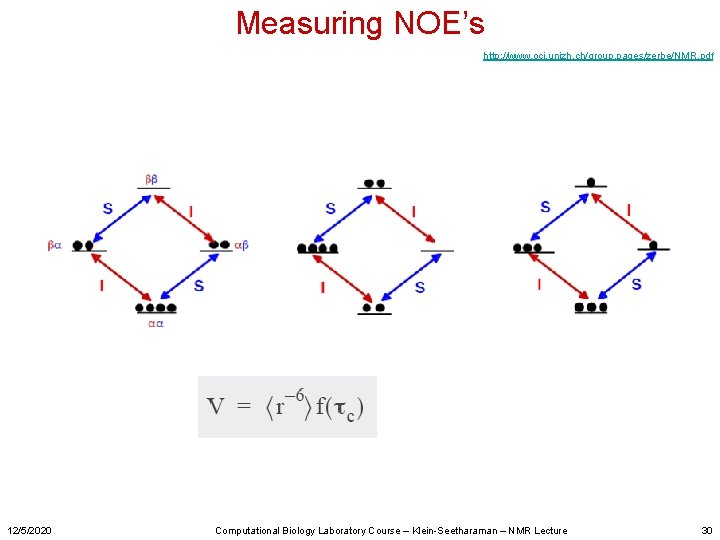
Measuring NOE’s http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 30
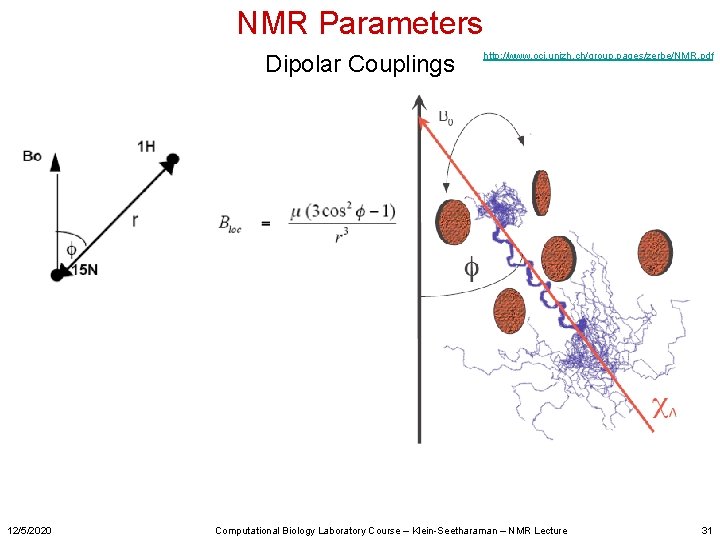
NMR Parameters Dipolar Couplings 12/5/2020 http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 31
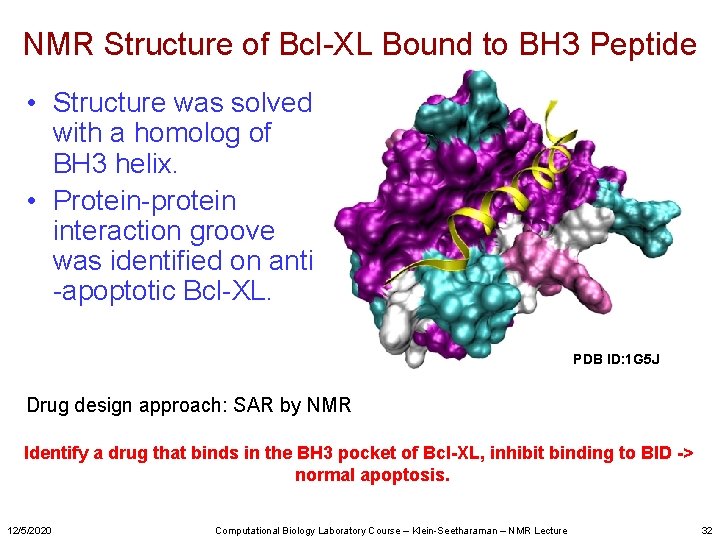
NMR Structure of Bcl-XL Bound to BH 3 Peptide • Structure was solved with a homolog of BH 3 helix. • Protein-protein interaction groove was identified on anti -apoptotic Bcl-XL. PDB ID: 1 G 5 J Drug design approach: SAR by NMR Identify a drug that binds in the BH 3 pocket of Bcl-XL, inhibit binding to BID -> normal apoptosis. 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 32
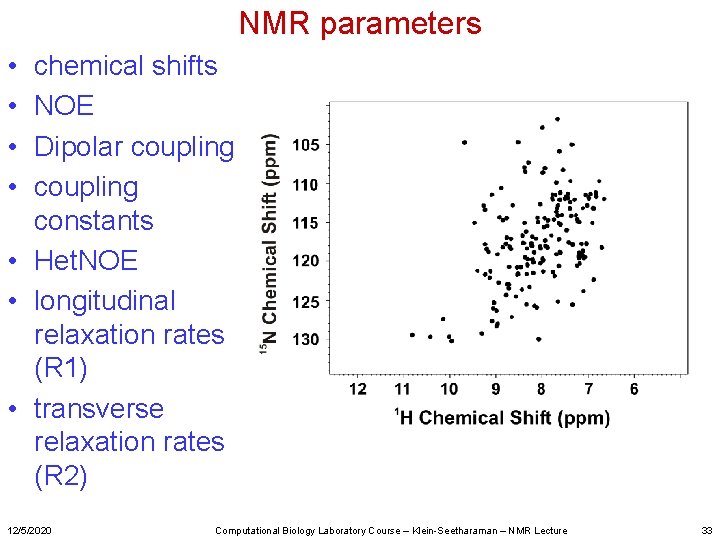
NMR parameters • • chemical shifts NOE Dipolar coupling constants • Het. NOE • longitudinal relaxation rates (R 1) • transverse relaxation rates (R 2) 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 33

Objectives of this Lecture and Practicum • • 12/5/2020 Resources Physical principle Sample requirements Parameters that are measured by NMR Dynamics by NMR Limitations Practical aspects Setup of NMR experiments (downstairs) Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 34
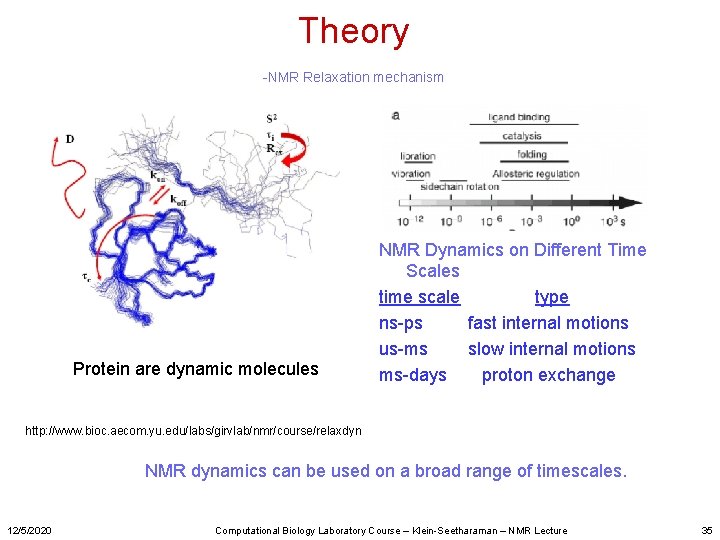
Theory -NMR Relaxation mechanism Protein are dynamic molecules NMR Dynamics on Different Time Scales time scale type ns-ps fast internal motions us-ms slow internal motions ms-days proton exchange http: //www. bioc. aecom. yu. edu/labs/girvlab/nmr/course/relaxdyn NMR dynamics can be used on a broad range of timescales. 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 35

Relaxation • Longitudinal relaxation (T 1): return of longitudinal (z-component) to its equilibrium value • Transverse relaxation (T 2): decay of transverse (x, y-component) 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 36

T 1 Relaxation http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 37
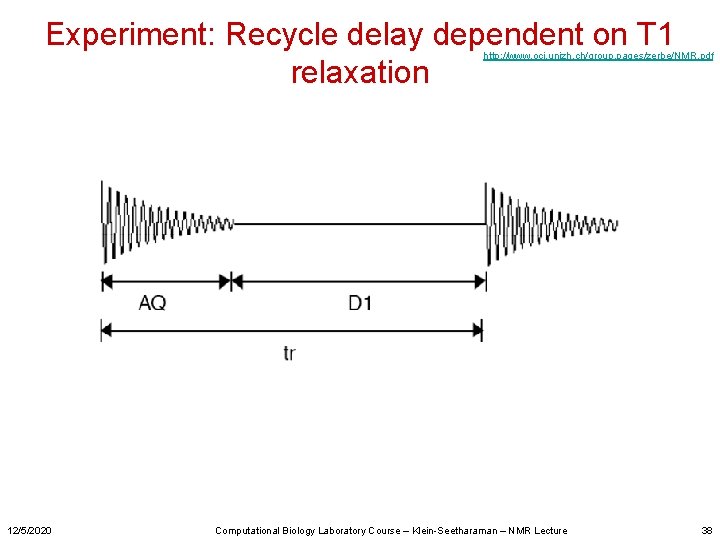
Experiment: Recycle delay dependent on T 1 relaxation http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 38
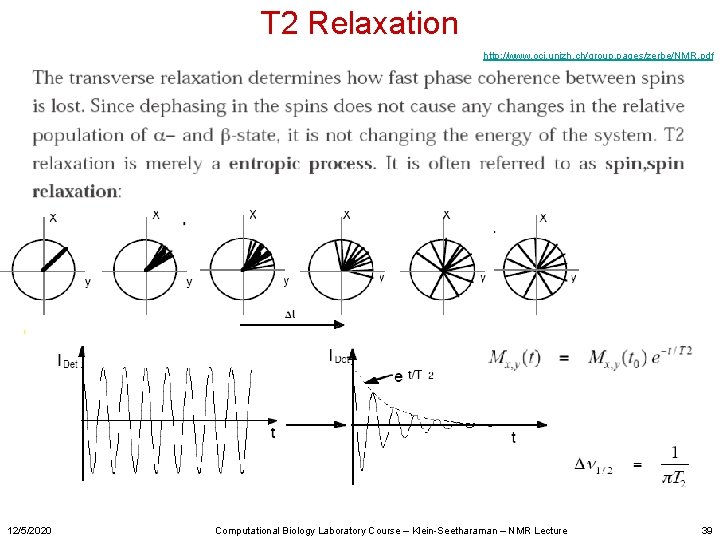
T 2 Relaxation http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 39
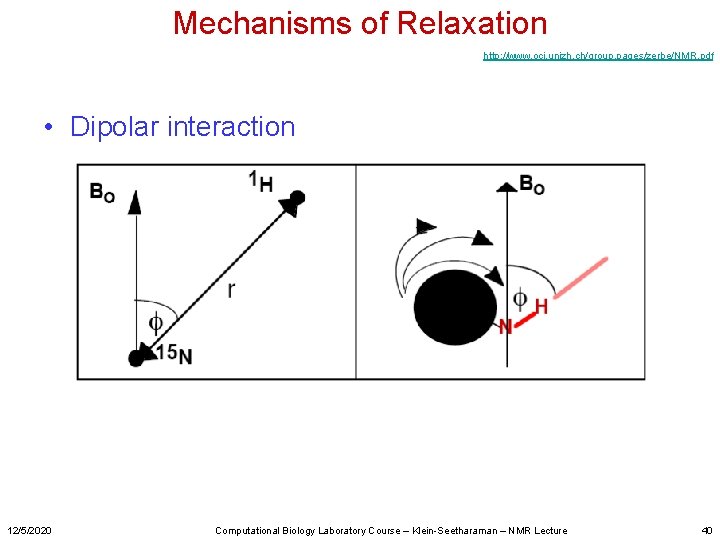
Mechanisms of Relaxation http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf • Dipolar interaction 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 40
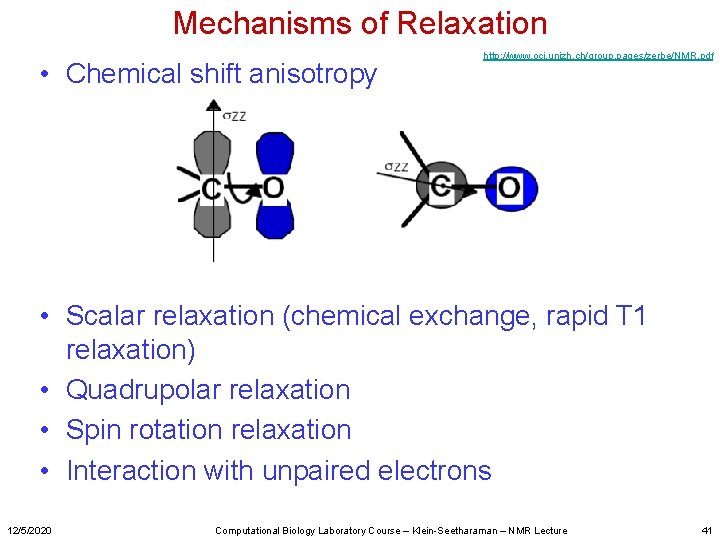
Mechanisms of Relaxation • Chemical shift anisotropy http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf • Scalar relaxation (chemical exchange, rapid T 1 relaxation) • Quadrupolar relaxation • Spin rotation relaxation • Interaction with unpaired electrons 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 41

Quantification of motion - strategies to obtain dynamic information from NMR relaxation experiment Measure R 1, R 2, heteronuclear NOE “model free” approach Get order parameter S 2 , τe, τm 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 42

Lipari-Szabo Model Free Approach http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 43
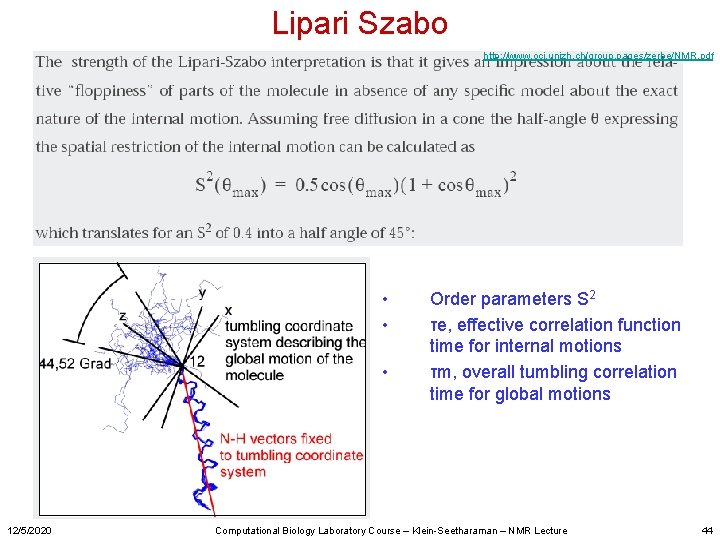
Lipari Szabo http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf • • • 12/5/2020 Order parameters S 2 τe, effective correlation function time for internal motions τm, overall tumbling correlation time for global motions Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 44

Lipari–Szabo “model-free” approach - don’t depend on a specific physical model • • • Estimate (τm) from R 2/R 1 for a selected subset of the residues fits to the observed relaxation data using various regression variables model-selection criteria are used to decide which choice is appropriate for each residue Reoptimize using the selected models. Uncertainties in the optimized parameters were obtained by Monte Carlo simulation. Michael Andrec, Gaetano T. Montelione, Ronald M. Levy Journal of Magnetic Resonance 139, 408– 421 (1999) “Model-free” is the most popular method to calculate dynamic parameters 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 45
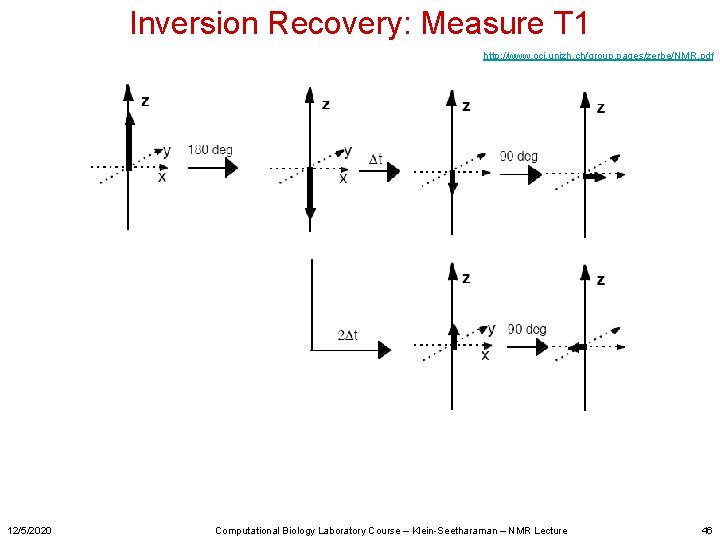
Inversion Recovery: Measure T 1 http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 46
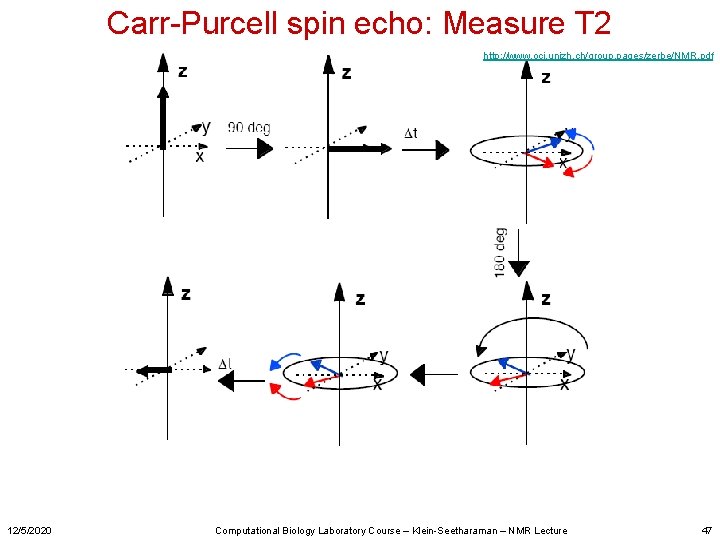
Carr-Purcell spin echo: Measure T 2 http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 47
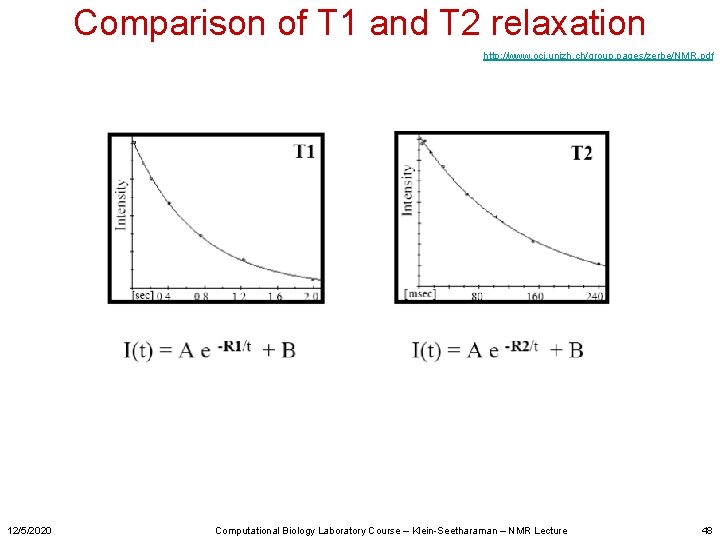
Comparison of T 1 and T 2 relaxation http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 48
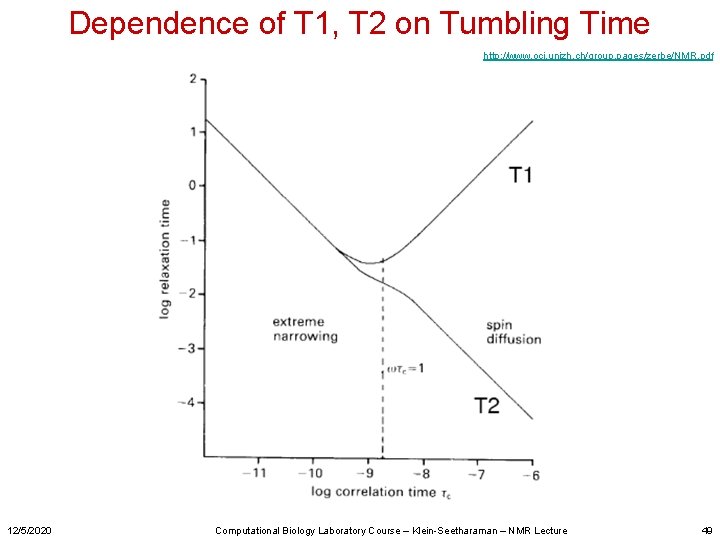
Dependence of T 1, T 2 on Tumbling Time http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 49
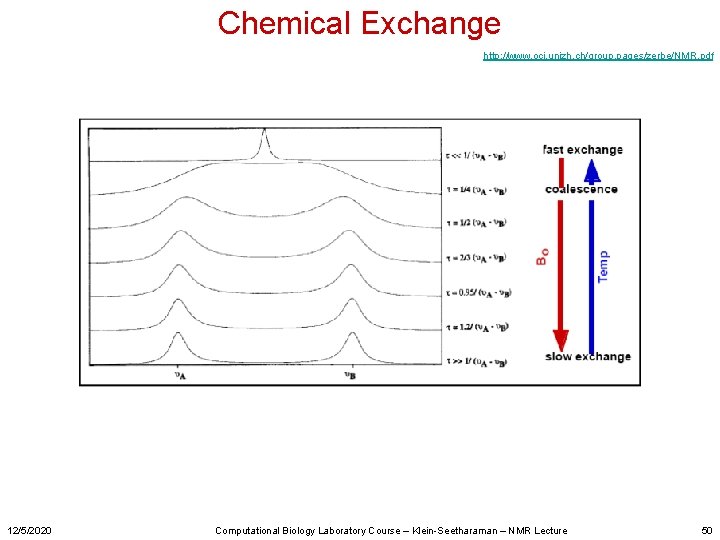
Chemical Exchange http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 50
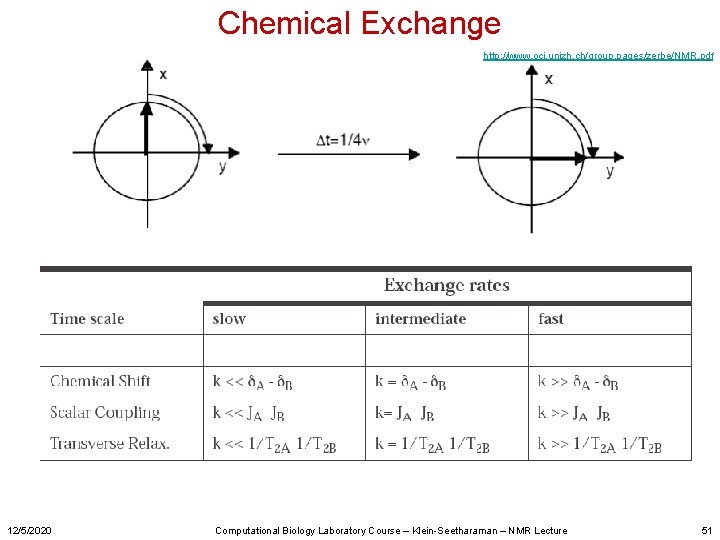
Chemical Exchange http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 51

Example: GCN 4 -58 complexed with DNA -dynamics of complex formation dynamics of the basic leucine zipper domain of the dimeric yeast transcription factor GCN 4 (GCN 4 -58) as it relates to DNA binding low S 2 indicate high flexibility. S 2 can be used to estimate energetics. 12/5/2020 John Cavanagh and Mikael Akke nature structural biology • volume 7 number 1 • january 2000 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 52
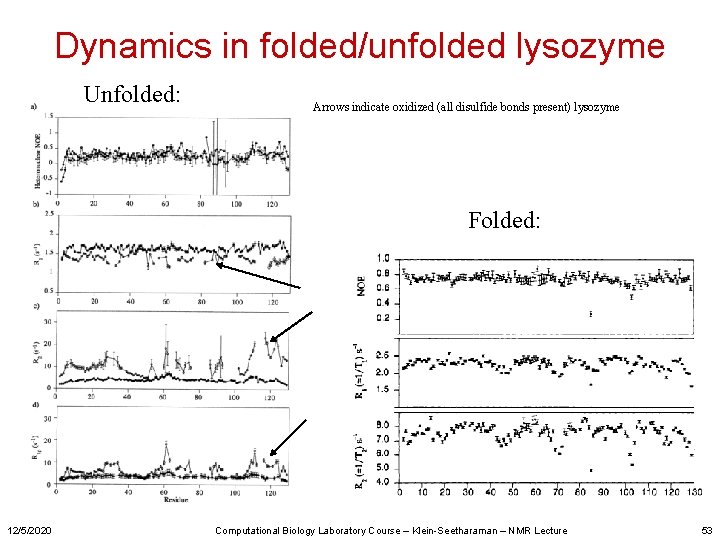
Dynamics in folded/unfolded lysozyme Unfolded: Arrows indicate oxidized (all disulfide bonds present) lysozyme Folded: 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 53

Using NMR to identify residual structure • Can in principle use all parameters: – chemical shifts – coupling constants – Het. NOE – longitudinal relaxation rates (R 1) – transverse relaxation rates (R 2) 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 54
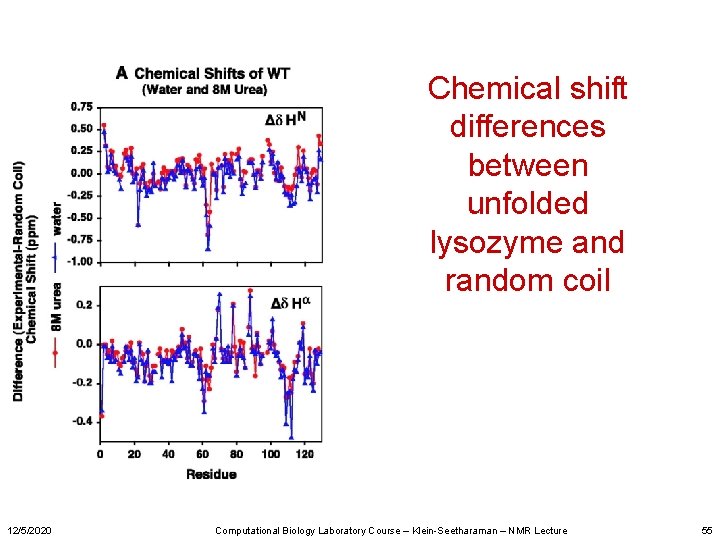
Chemical shift differences between unfolded lysozyme and random coil 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 55
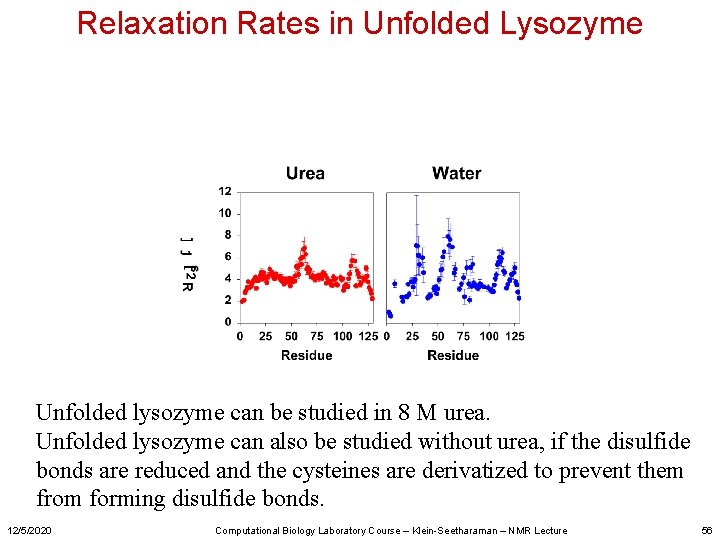
Relaxation Rates in Unfolded Lysozyme Unfolded lysozyme can be studied in 8 M urea. Unfolded lysozyme can also be studied without urea, if the disulfide bonds are reduced and the cysteines are derivatized to prevent them from forming disulfide bonds. 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 56
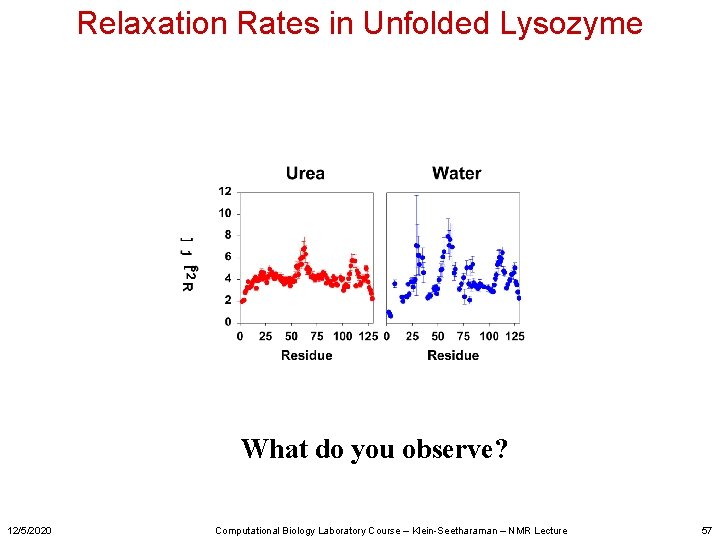
Relaxation Rates in Unfolded Lysozyme What do you observe? 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 57
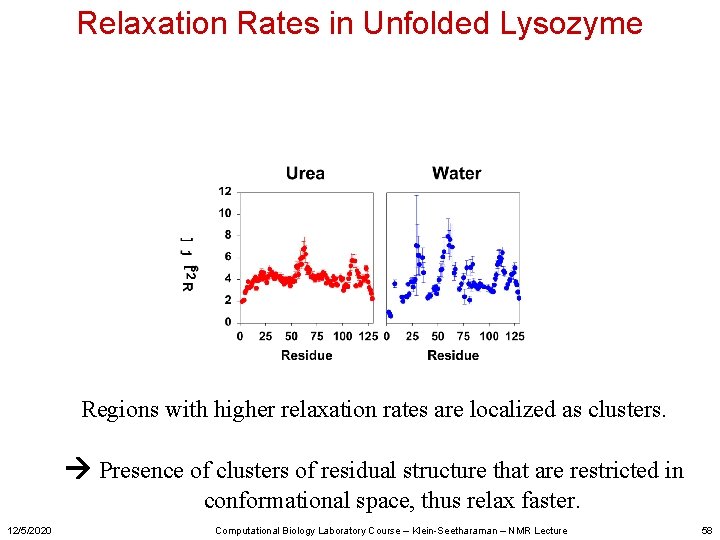
Relaxation Rates in Unfolded Lysozyme Regions with higher relaxation rates are localized as clusters. Presence of clusters of residual structure that are restricted in conformational space, thus relax faster. 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 58

Analysis of the relaxation data Three means of analysis have been proposed: a. Model-free approach b. Cole-Cole distributions c. Gaussian clusters However: What gives rise to these clusters is not known. 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 59
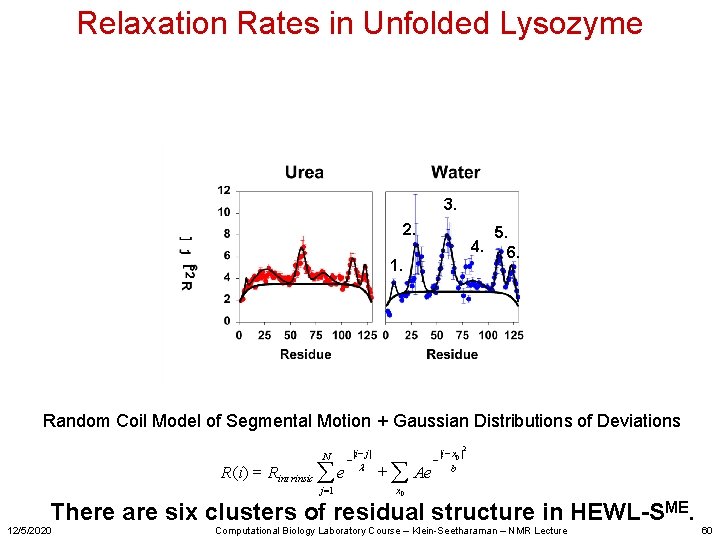
Relaxation Rates in Unfolded Lysozyme 3. 2. 4. 1. 5. 6. Random Coil Model of Segmental Motion + Gaussian Distributions of Deviations N R (i ) = Rint rinsic å e j =1 - |i - j | l + å Ae - |i - x 0 | b 2 x 0 There are six clusters of residual structure in HEWL-SME. 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 60
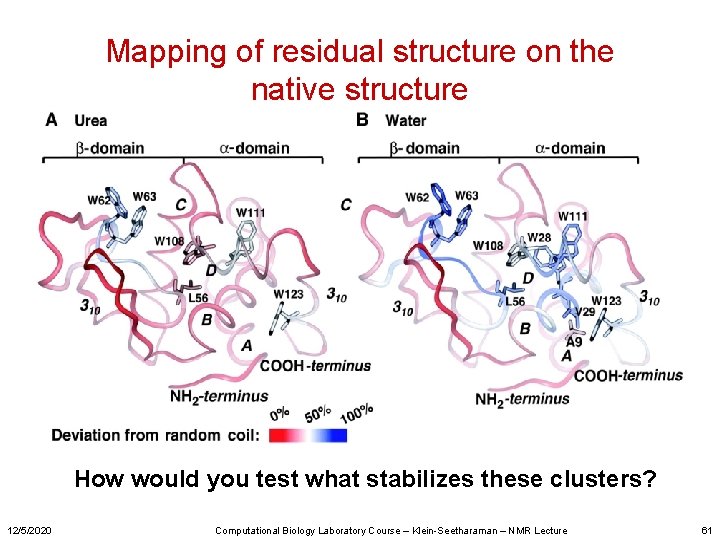
Mapping of residual structure on the native structure How would you test what stabilizes these clusters? 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 61
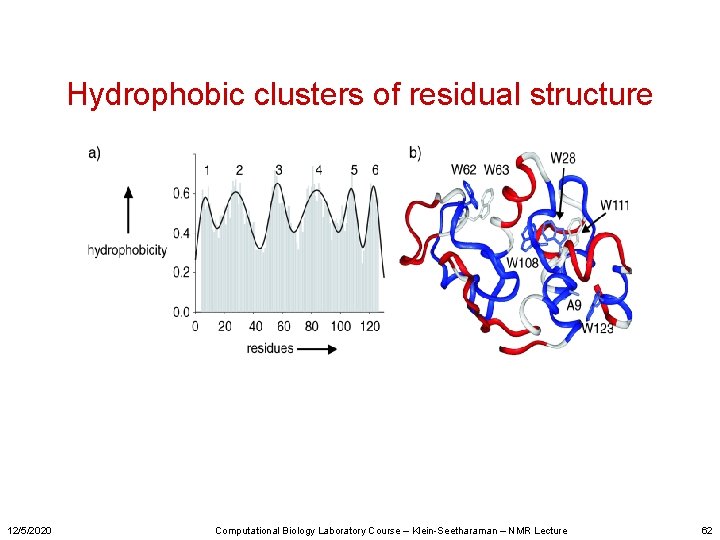
Hydrophobic clusters of residual structure 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 62
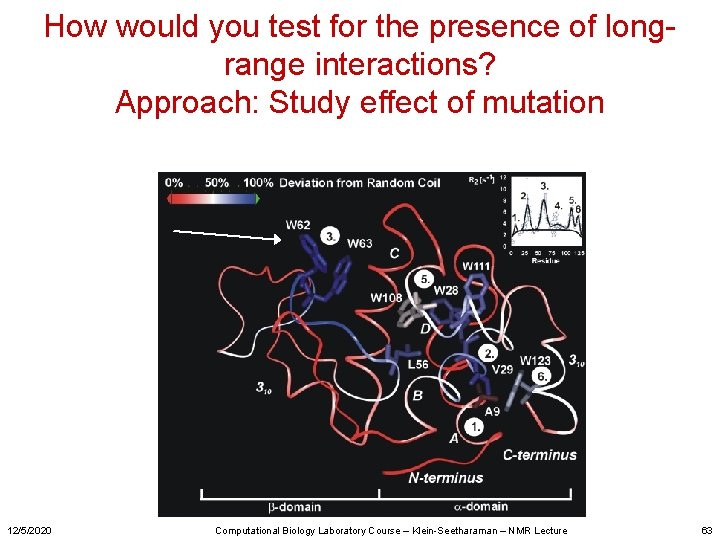
How would you test for the presence of longrange interactions? Approach: Study effect of mutation 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 63
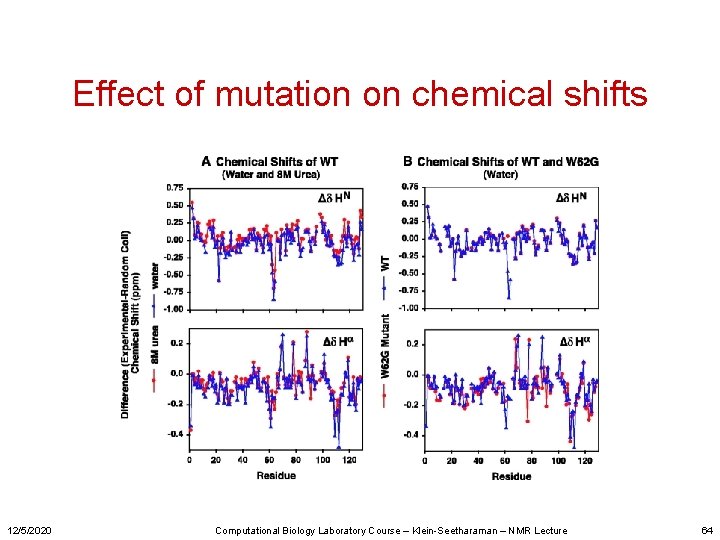
Effect of mutation on chemical shifts 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 64
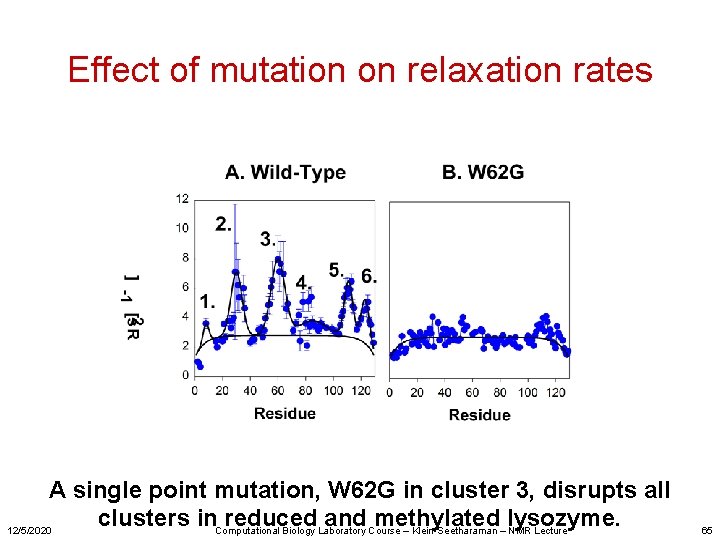
Effect of mutation on relaxation rates A single point mutation, W 62 G in cluster 3, disrupts all clusters in. Computational reduced and methylated lysozyme. 12/5/2020 Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 65
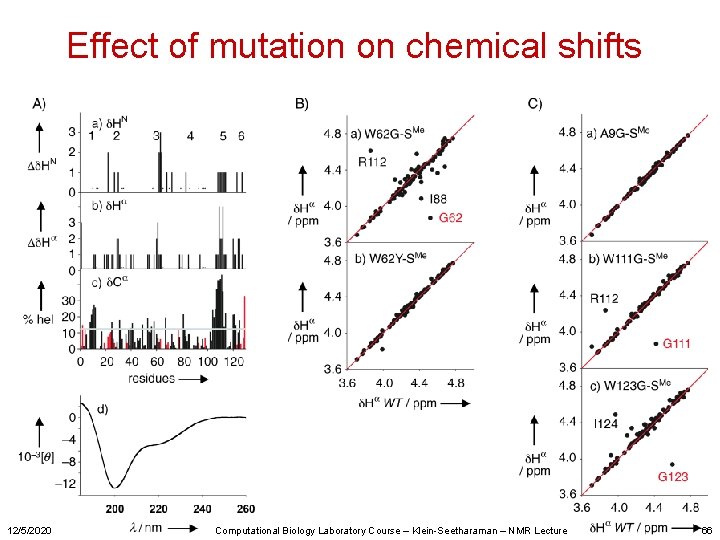
Effect of mutation on chemical shifts 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 66
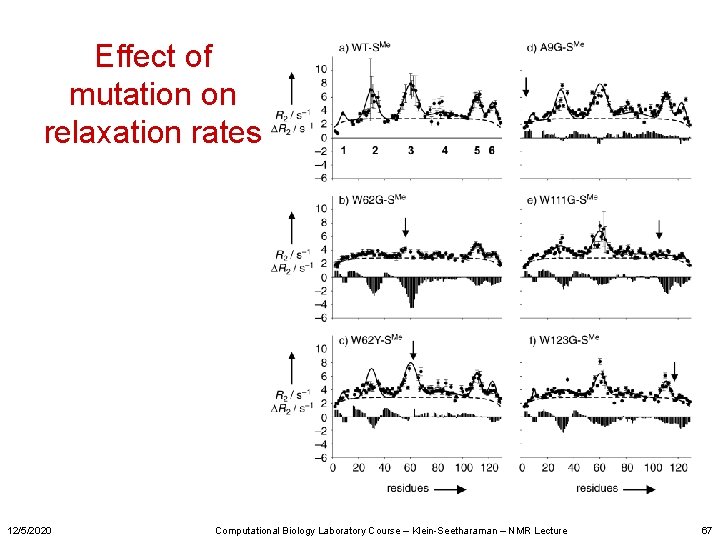
Effect of mutation on relaxation rates 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 67
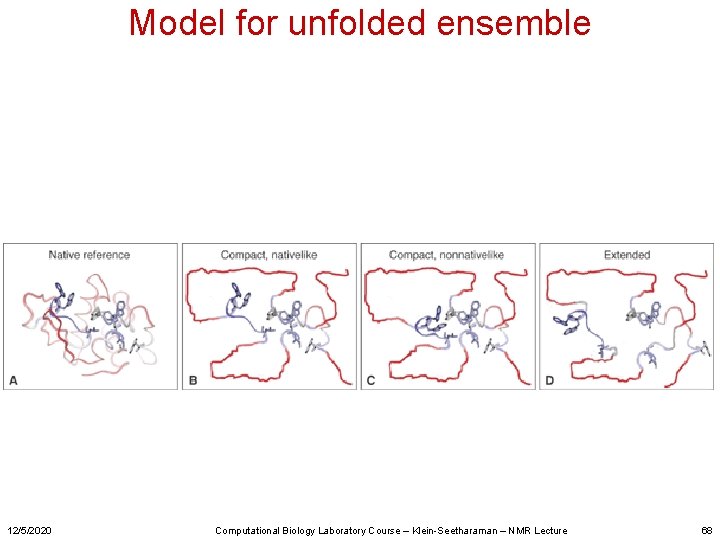
Model for unfolded ensemble 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 68
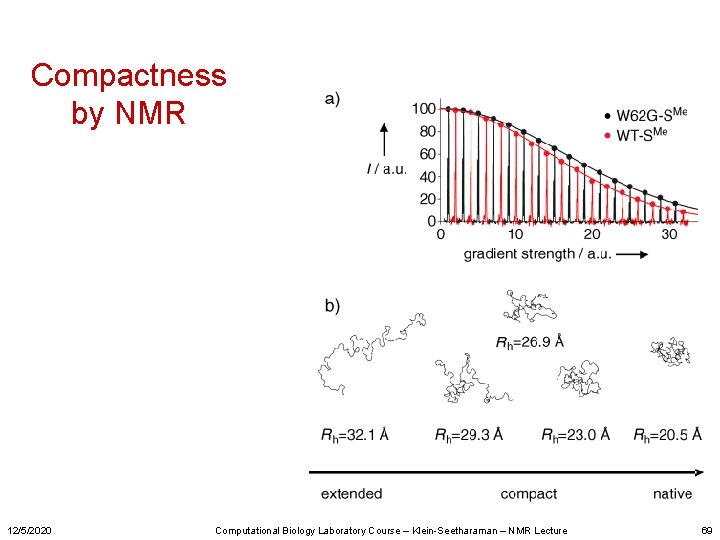
Compactness by NMR 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 69

Objectives of this Lecture and Practicum • • 12/5/2020 Resources Physical principle Sample requirements Parameters that are measured by NMR Dynamics by NMR Limitations Practical aspects Setup of NMR experiments (downstairs) Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 70

NMR spectroscopy General limitations • • • 12/5/2020 Size Stability Sample homogeneity Need for labeling Quantities and source of biomolecules Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 71
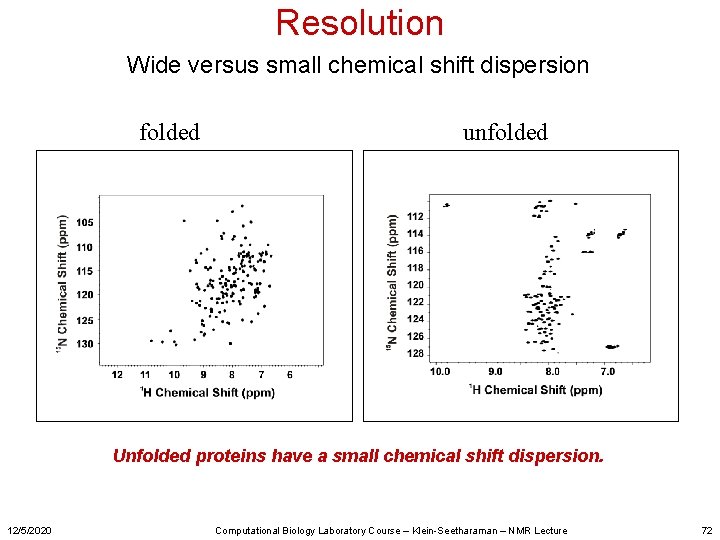
Resolution Wide versus small chemical shift dispersion folded unfolded Unfolded proteins have a small chemical shift dispersion. 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 72
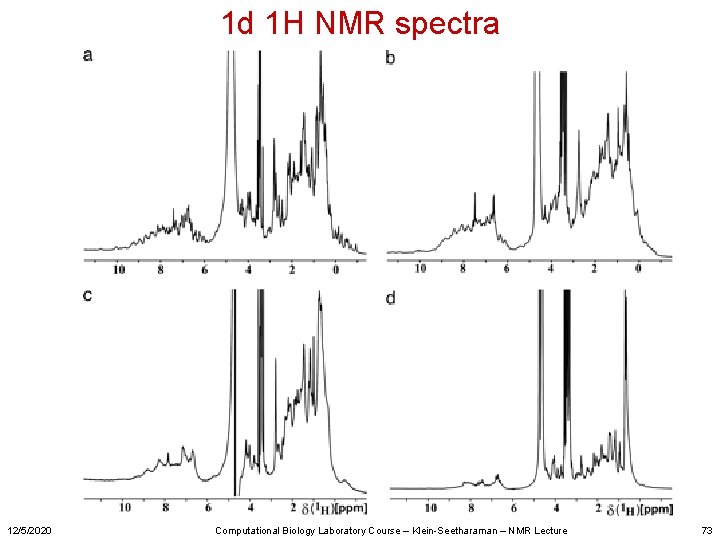
1 d 1 H NMR spectra 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 73
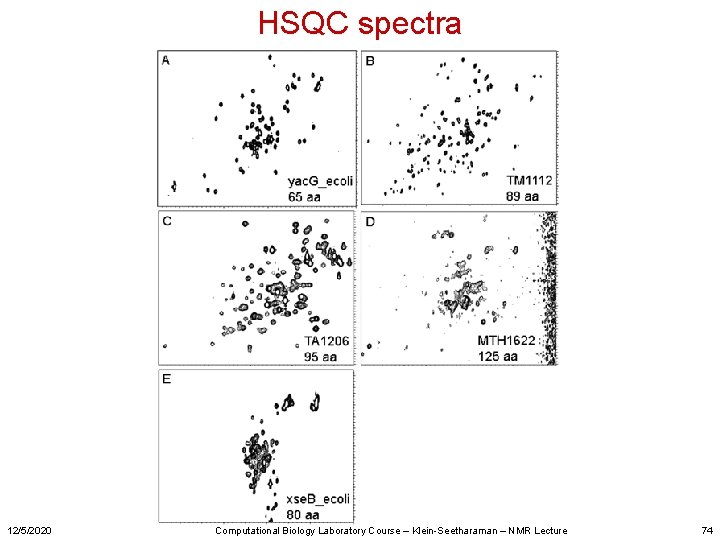
HSQC spectra 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 74
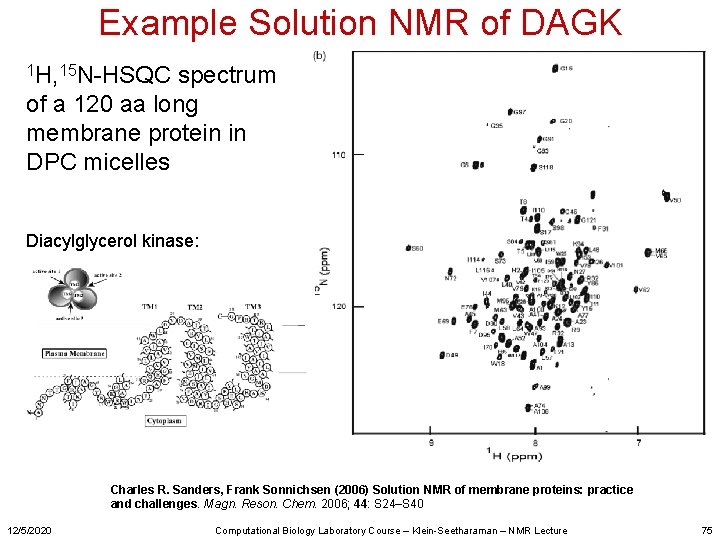
Example Solution NMR of DAGK 1 H, 15 N-HSQC spectrum of a 120 aa long membrane protein in DPC micelles Diacylglycerol kinase: Charles R. Sanders, Frank Sonnichsen (2006) Solution NMR of membrane proteins: practice and challenges. Magn. Reson. Chem. 2006; 44: S 24–S 40 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 75
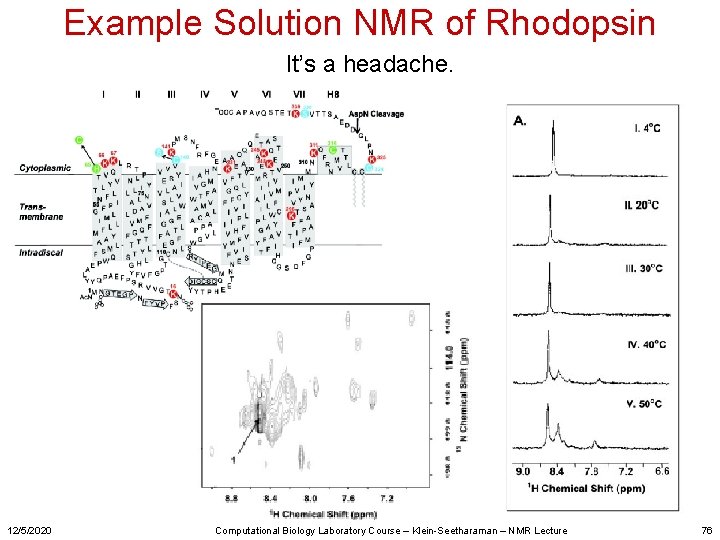
Example Solution NMR of Rhodopsin It’s a headache. 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 76
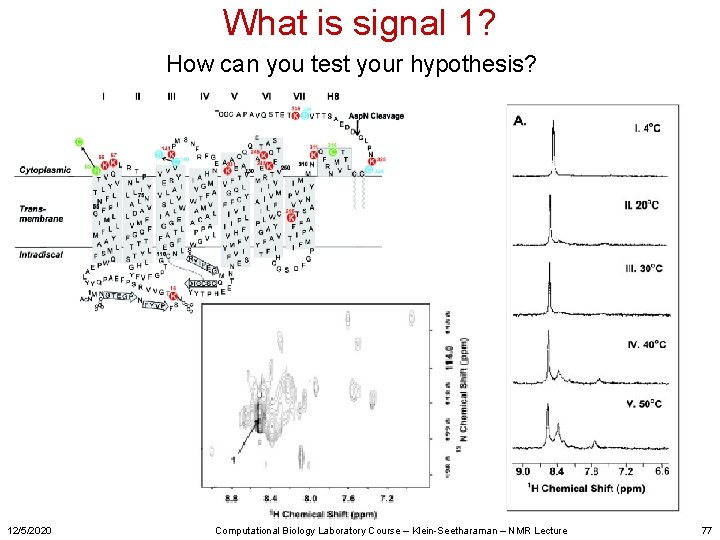
What is signal 1? How can you test your hypothesis? 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 77

Assignment of Signal 1 NMR Spectroscopy An enzyme was used to cleave off the C-terminus at the site indicated below: 12/5/2020 Black: original spectrum, red: C-terminus, green: Nterminus (after Asp. N cleavage) Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 78
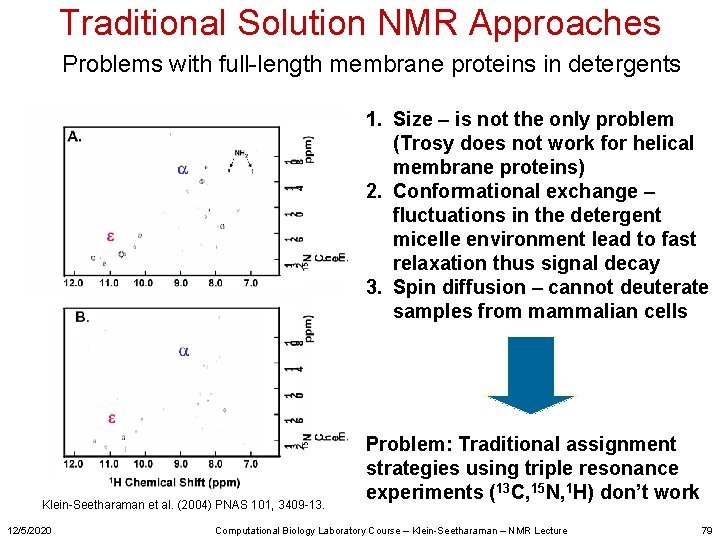
Traditional Solution NMR Approaches Problems with full-length membrane proteins in detergents 1. Size – is not the only problem (Trosy does not work for helical membrane proteins) 2. Conformational exchange – fluctuations in the detergent micelle environment lead to fast relaxation thus signal decay 3. Spin diffusion – cannot deuterate samples from mammalian cells Klein-Seetharaman et al. (2004) PNAS 101, 3409 -13. 12/5/2020 Problem: Traditional assignment strategies using triple resonance experiments (13 C, 15 N, 1 H) don’t work Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 79
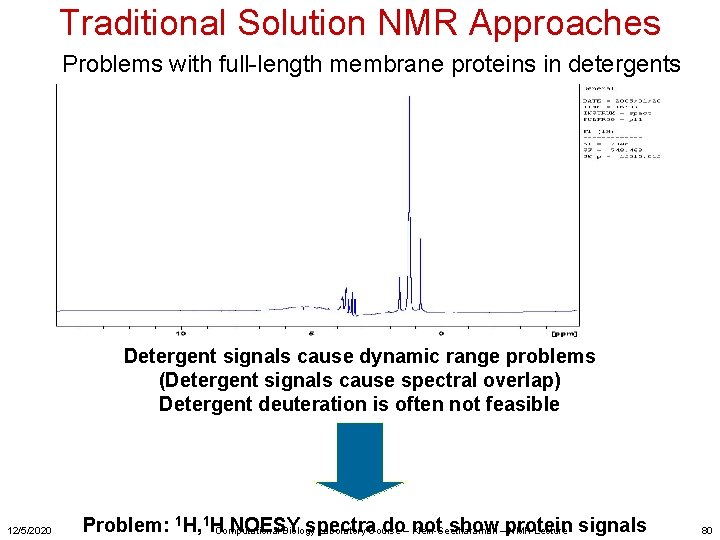
Traditional Solution NMR Approaches Problems with full-length membrane proteins in detergents Detergent signals cause dynamic range problems (Detergent signals cause spectral overlap) Detergent deuteration is often not feasible 12/5/2020 Problem: 1 H, 1 HComputational NOESY spectra do– Klein-Seetharaman not show–protein Biology Laboratory Course NMR Lecture signals 80
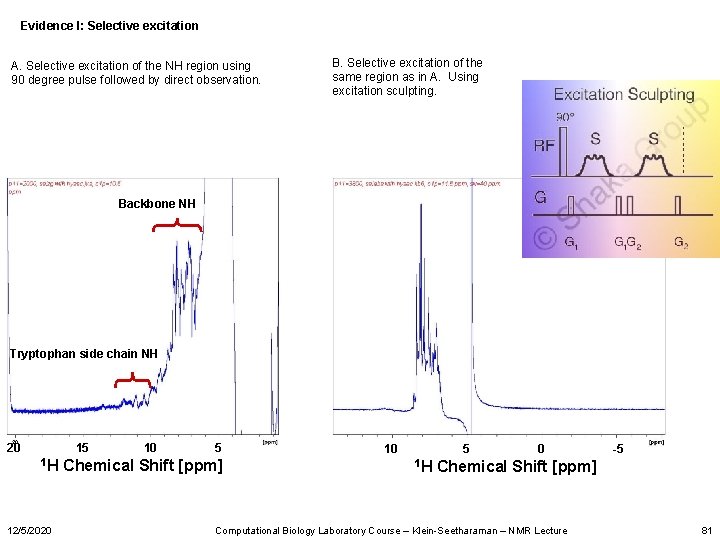
Evidence I: Selective excitation A. Selective excitation of the NH region using 90 degree pulse followed by direct observation. B. Selective excitation of the same region as in A. Using excitation sculpting. Backbone NH Tryptophan side chain NH 20 15 1 H 12/5/2020 10 5 Chemical Shift [ppm] 10 5 1 H 0 -5 Chemical Shift [ppm] Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 81

You might also want to develop your own biophysical approaches… 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 82
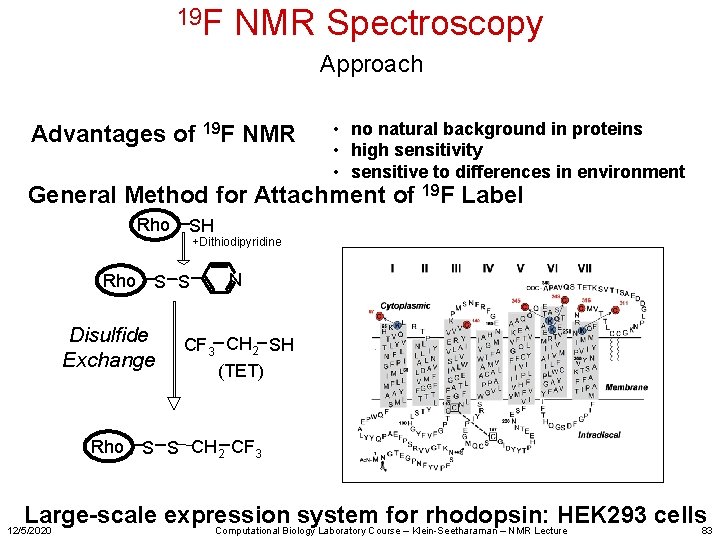
19 F NMR Spectroscopy Approach Advantages of 19 F NMR • no natural background in proteins • high sensitivity • sensitive to differences in environment General Method for Attachment of 19 F Label Rho +Dithiodipyridine S S Disulfide Exchange Rho SH N CF 3 CH 2 SH (TET) S S CH 2 CF 3 Large-scale expression system for rhodopsin: HEK 293 cells 83 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 12/5/2020
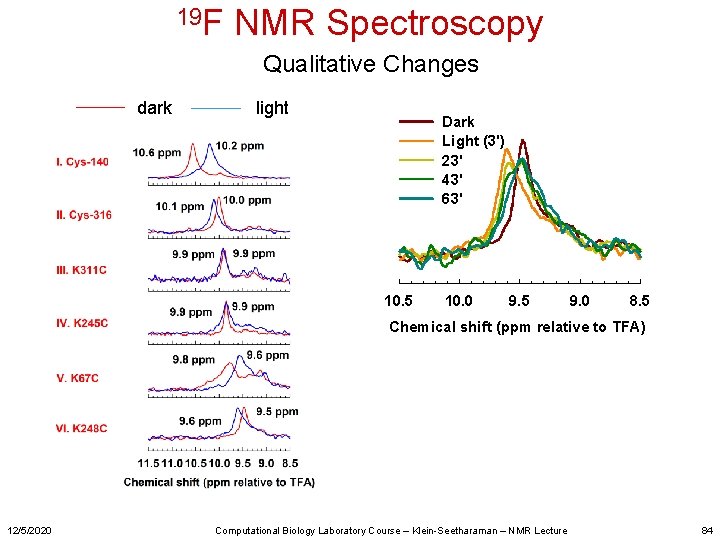
19 F NMR Spectroscopy Qualitative Changes dark light Dark Light (3') 23' 43' 63' 10. 5 10. 0 9. 5 9. 0 8. 5 Chemical shift (ppm relative to TFA) 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 84
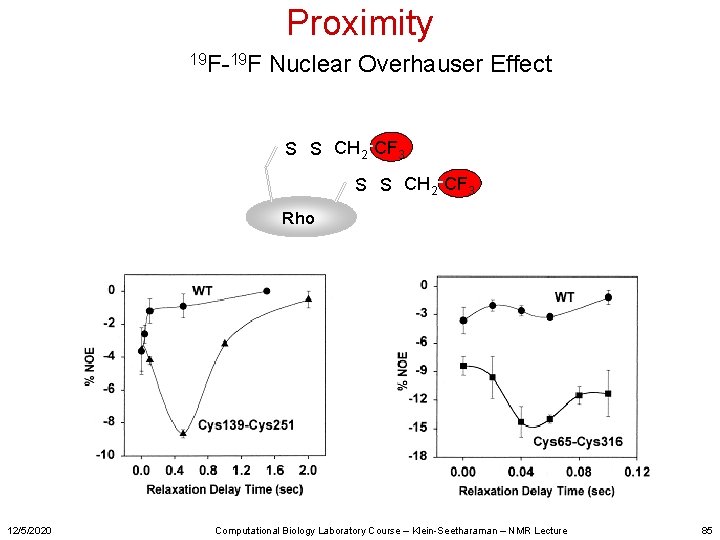
Proximity 19 F-19 F Nuclear Overhauser Effect S S CH 2 CF 3 Rho 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 85
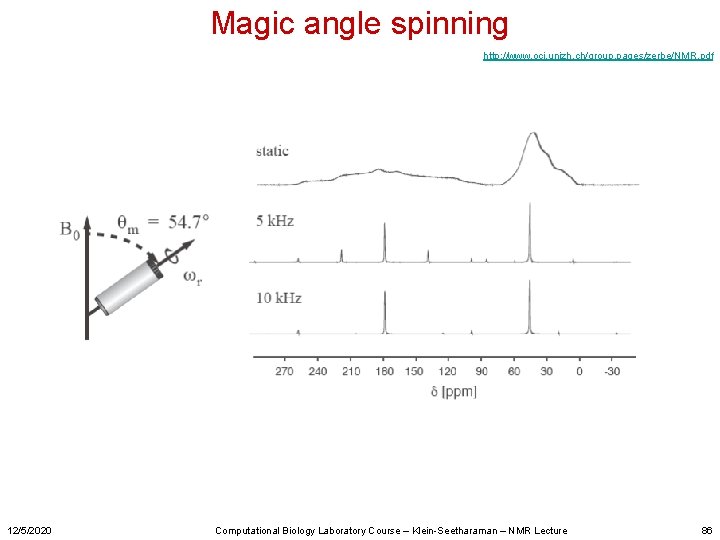
Magic angle spinning http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 86
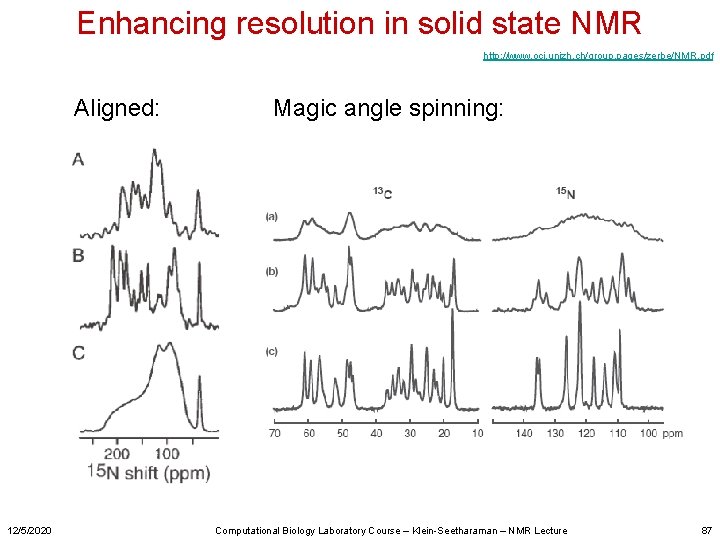
Enhancing resolution in solid state NMR http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf Aligned: 12/5/2020 Magic angle spinning: Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 87
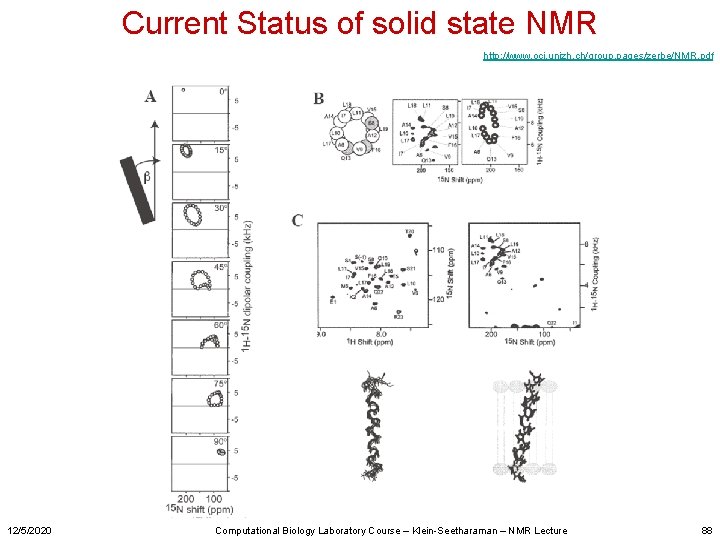
Current Status of solid state NMR http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 88
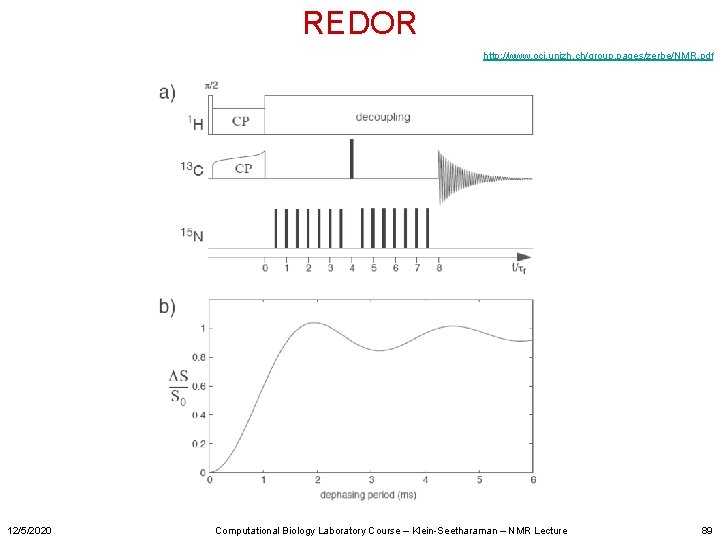
REDOR http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 89

Objectives of this Lecture and Practicum • • 12/5/2020 Resources Physical principle Sample requirements Parameters that are measured by NMR Dynamics by NMR Limitations Practical aspects Setup of NMR experiments (downstairs) Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 90

The components of an NMR spectrometer • • 12/5/2020 A magnet Probehead(s) Radiofrequency sources Amplifiers Analog to digital converters The lock system The shim system A computer Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 91
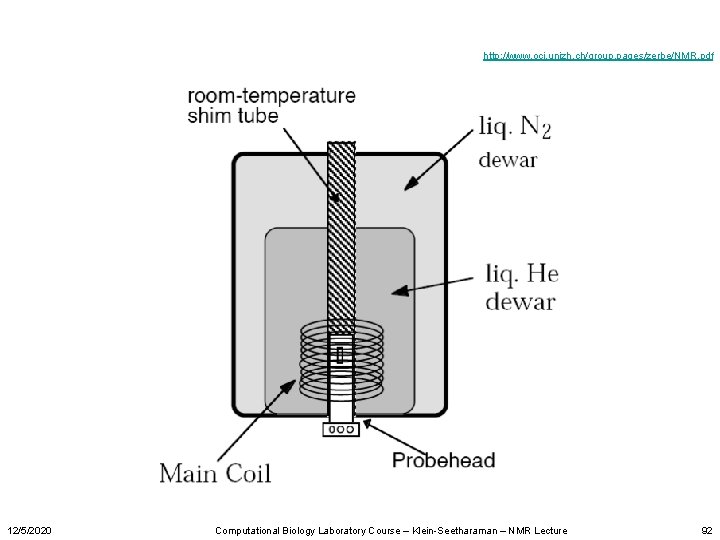
http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 92
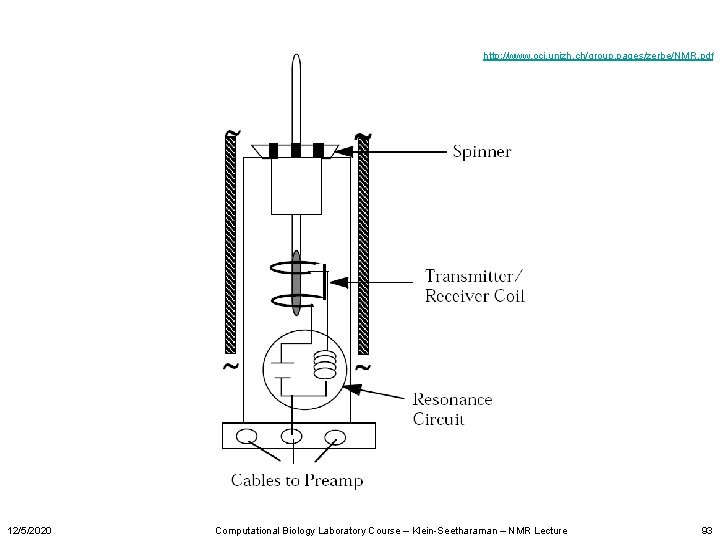
http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 93

The Shim System http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 94
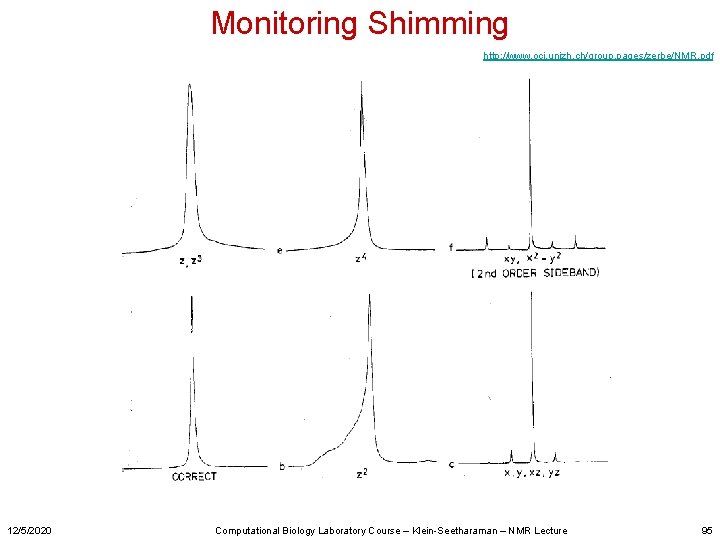
Monitoring Shimming http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 95

Monitoring shimming http: //www. oci. unizh. ch/group. pages/zerbe/NMR. pdf 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 96

Objectives of this Lecture and Practicum • • • 12/5/2020 Resources Physical principle Sample requirements Parameters that are measured by NMR Dynamics by NMR Limitations Practical aspects Setup of NMR experiments (downstairs) Bring your coats!! Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 97

Outlook for next week • Lecture: The structure determination pipeline • Practical analysis of NMR data in computer lab: – Topspin – NMRpipe – NMRview. J 12/5/2020 Computational Biology Laboratory Course – Klein-Seetharaman – NMR Lecture 98
- Slides: 98