Nitrogen Metabolism 1 Andy Howard Biochemistry Lectures Spring

Nitrogen Metabolism 1 Andy Howard Biochemistry Lectures, Spring 2019 Thursday 25 April 2019

Nitrogen metabolism n n Amino acid anabolism sometimes involves little more than transamination, but often is more complex Amino acid catabolism feeds the TCA cycle and acetyl Co. A 04/25/2019 Nitrogen Metabolism 1 p. 2 of 56

What we’ll discuss n n n Nitrogenase Nitrogen cycles AA anabolism n n Glutamate Transaminations Simple syntheses Complex syntheses 04/25/2019 n AA catabolism n n Transaminations Glucogenic & Ketogenic aa’s Nitrogen Metabolism 1 p. 3 of 56

The nitrogen pool n n n Nitrogen fixation from air (N 2 NH 3) doesn’t produce a large percentage of circulating biological nitrogen but it’s the ultimate source of most of it Other entries in pool: nitrate (NO 3 -), nitrite (NO 2 -) Most of this difficult biochemistry is bacterial 04/25/2019 Nitrogen Metabolism 1 p. 4 of 56

Nitrogenase n n Enzyme found in Rhizobium, a bacterium that colonizes & lives symbiotically in the root nodules of legumes and a few other plants Also in free-living microorganisms like Azotobacter Energetically expensive but irreversible path to reduction of dinitrogen to ammonia: N 2 + 8 H+ + 8 e- + 16 ATP 2 NH 3 + H 2 + 16 ADP + 16 Pi 04/25/2019 Nitrogen Metabolism 1 p. 5 of 56
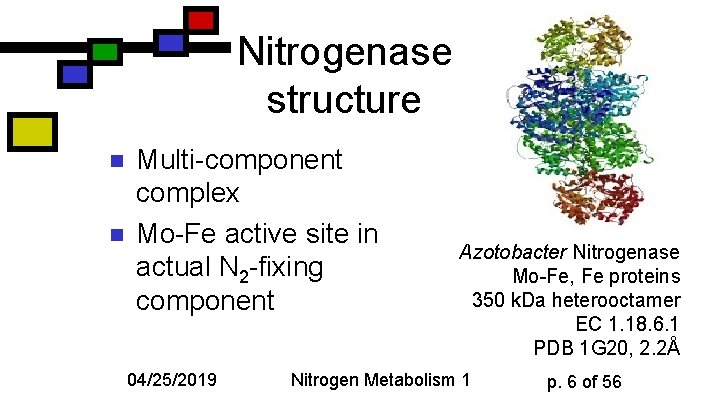
Nitrogenase structure n n Multi-component complex Mo-Fe active site in actual N 2 -fixing component 04/25/2019 Azotobacter Nitrogenase Mo-Fe, Fe proteins 350 k. Da heterooctamer EC 1. 18. 6. 1 PDB 1 G 20, 2. 2Å Nitrogen Metabolism 1 p. 6 of 56
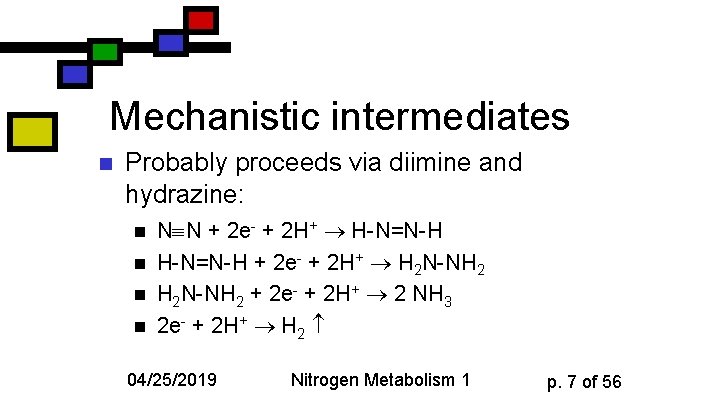
Mechanistic intermediates n Probably proceeds via diimine and hydrazine: n n N N + 2 e- + 2 H+ H-N=N-H + 2 e- + 2 H+ H 2 N-NH 2 + 2 e- + 2 H+ 2 NH 3 2 e- + 2 H+ H 2 04/25/2019 Nitrogen Metabolism 1 p. 7 of 56

n n Ammonia, nitrate, nitrite Ammonia comes from decayed organisms and is oxidized in soil bacteria to nitrate (nitrification) Nitrate reductase and nitrite reductase found in plants and microorganisms 04/25/2019 Nitrogen Metabolism 1 Alcaligines Nitrite Reductase 111 k. Da trimer monomer shown PDB 2 BO 0, 1. 35Å p. 8 of 56
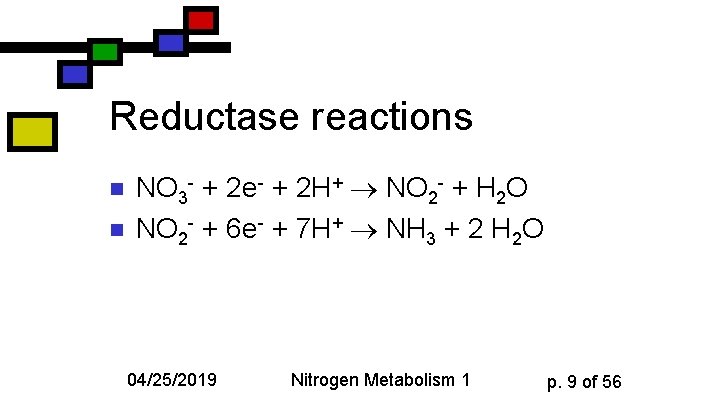
Reductase reactions n n NO 3 - + 2 e- + 2 H+ NO 2 - + H 2 O NO 2 - + 6 e- + 7 H+ NH 3 + 2 H 2 O 04/25/2019 Nitrogen Metabolism 1 p. 9 of 56

Essential and non-essential amino acids n n n An amino acid is defined as essential if it must be obtained within the diet In general the essential amino acids are the ones that have complicated and highly ATPdependent biosynthetic pathways Of course, it depends on the organism 04/25/2019 Nitrogen Metabolism 1 p. 10 of 56
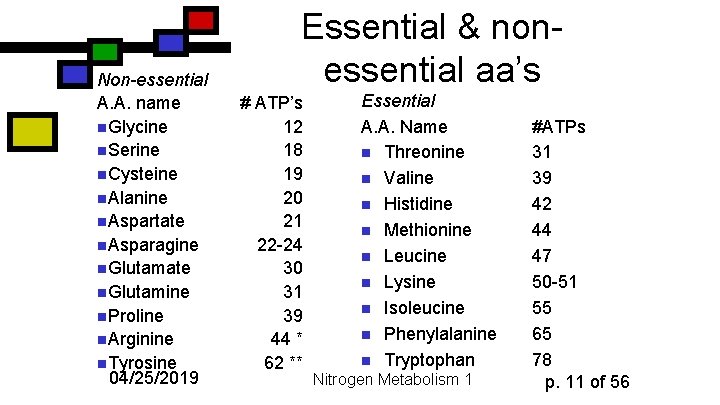
Non-essential A. A. name n. Glycine n. Serine n. Cysteine n. Alanine n. Aspartate n. Asparagine n. Glutamate n. Glutamine n. Proline n. Arginine n. Tyrosine 04/25/2019 Essential & nonessential aa’s # ATP’s 12 18 19 20 21 22 -24 30 31 39 44 * 62 ** Essential A. A. Name n Threonine n Valine n Histidine n Methionine n Leucine n Lysine n Isoleucine n Phenylalanine n Tryptophan Nitrogen Metabolism 1 #ATPs 31 39 42 44 47 50 -51 55 65 78 p. 11 of 56
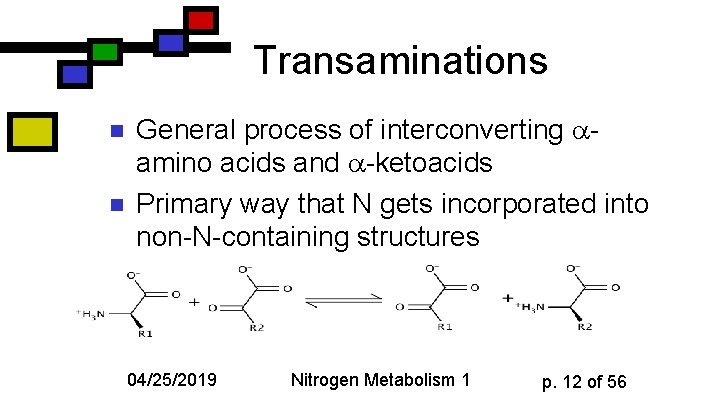
Transaminations n n General process of interconverting amino acids and -ketoacids Primary way that N gets incorporated into non-N-containing structures 04/25/2019 Nitrogen Metabolism 1 p. 12 of 56

Reaction dynamics n n n All (? ) transaminations involve PLP as a cofactor: see mechanism in textbook E. coli Aspartate These are actually oxidation-reduction aminotransferase reactions, since we’re swapping an EC 2. 6. 1. 1 amine (carbon oxidation state +2) for a 87 k. Da dimer; carbonyl (carbon oxidation state 0) Monomer shown But there is no external oxidizing agent PDB 2 Q 7 W, 1. 4Å 04/25/2019 Nitrogen Metabolism 1 p. 13 of 56
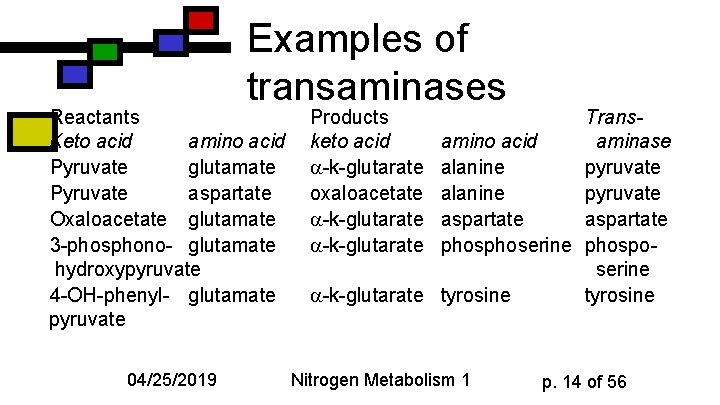
Examples of transaminases Reactants Keto acid amino acid Pyruvate glutamate Pyruvate aspartate Oxaloacetate glutamate 3 -phosphono- glutamate hydroxypyruvate 4 -OH-phenyl- glutamate pyruvate 04/25/2019 Products keto acid -k-glutarate oxaloacetate -k-glutarate Transamino acid aminase alanine pyruvate aspartate phoserine phosposerine -k-glutarate tyrosine Nitrogen Metabolism 1 p. 14 of 56

Catabolic or anabolic? n n From the point of view of available pools of amino acids, these are amphibolic: They involve synthesis of one amino acid at the expense of another 04/25/2019 Nitrogen Metabolism 1 p. 15 of 56

i. Clicker question #1 1. The distinction n between essential and nonn -essential amino acids relates to n n 04/25/2019 (a) whether an organism can synthesize the amino acid (b) how we degrade the amino acid (c) how many k. J/mol can be derived from its catabolism (d) how many sulfurs are present Nitrogen Metabolism 1 p. 16 of 56

Biosynthetic pathways to specific amino acids n n Some are complex and energy-requiring Can be logically divided according to chemical properties of the target amino acids: 04/25/2019 n n n n Small Branched-chain aliphatic Neutral polar Acidic Basic Aromatic Sulfur-containing Nitrogen Metabolism 1 p. 17 of 56
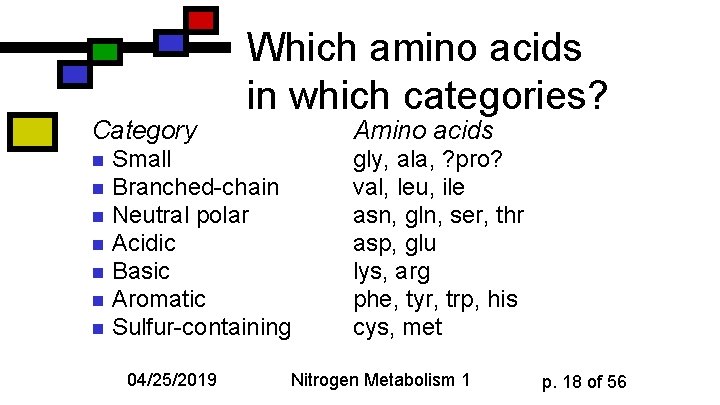
Category Which amino acids in which categories? Amino acids Small n Branched-chain n Neutral polar n Acidic n Basic n Aromatic n Sulfur-containing n 04/25/2019 gly, ala, ? pro? val, leu, ile asn, gln, ser, thr asp, glu lys, arg phe, tyr, trp, his cys, met Nitrogen Metabolism 1 p. 18 of 56

Glutamate n n Glutamate is a critical metabolite because so many of the transaminations start with it as the amine donor It is produced in E. coli, etc. via glutamate dehydrogenase using ammonium ion as nitrogen donor: 04/25/2019 Nitrogen Metabolism 1 Glu dehydrogenase PDB 1 BGV 296 k. Da hexamer monomer shown EC 1. 4. 1. 2, 1. 9Å Clostridium p. 19 of 56

Glutamate dehydrogenase reaction n -ketoglutarate + NH 4+ + NAD(P)H + H+ NAD(P)+ + H 2 O + glutamate 04/25/2019 Nitrogen Metabolism 1 p. 20 of 56

Glutamine n n Glutamate can be aminated with expenditure of ATP to form glutamine: glutamate + NH 4+ + ATP glutamine + ADP + Pi Note that glutamine synthetase is a ligase: the ATP is an energyprovider, not a phosphate donor 04/25/2019 Nitrogen Metabolism 1 Human Glutamine synthetase EC 6. 3. 1. 2 211 k. Da pentamer PDB 2 OJW, 2. 05Å p. 21 of 56

Aspartate and asparagine n n Asp is simple: transamination of oxaloacetate E. coli Asn is straightforward too Asparagine asparagine synthetase moves amine from synthetase B EC 6. 3. 5. 4 gln to asp, leaving glu (another ligase) 243 k. Da Gln + asp + ATP AMP + PPi + glu + asn tetramer PDB 1 CT 9, 2Å 04/25/2019 Nitrogen Metabolism 1 p. 22 of 56

Simple: ala, gly, ser n n n Alanine by transamination from pyruvate Glycine from serine by SHMT (q. v. ) Serine from 3 -phosphoglycerate: n n n Human 3 -phosphoglycerate + NAD+ Phosphoserine + NADH + H phosphatase + 3 -phosphohydroxypyruvate 49 k. Da dimer 3 -phosphohydroxypyruvate + glutamate EC 3. 1. 3. 3 3 -phoserine + -ketoglutarate PDB 1 NNL, 1. 53Å 3 -phoserine + H 2 O serine + Pi 04/25/2019 Nitrogen Metabolism 1 p. 23 of 56

Serine hydroxymethyltransferase n n n Serine + tetrahydrofolate H 2 O + glycine + 5, 10 -methylenetetrahydrofolate This can be viewed as a source of methylene units for other biosyntheses PLP-dependent reaction 04/25/2019 Nitrogen Metabolism 1 Thermus thermophilus SHMT 90 k. Da dimer EC 2. 1 PDB 2 DKJ, 1. 15Å p. 24 of 56

Arginine & proline n n Glutamate semialdehyde Two routes: Glutamate to glutamate semialdehyde n n n that cyclizes to 1 -pyrroline 5 -carboxylate and thence to proline Glutamate semialdehyde can also be converted to ornithine and thence to arg Alternative: glutamate acetylated to N-acetyl-glutamate-5 -semialdehyde and thence to ornithine 04/25/2019 Nitrogen Metabolism 1 ornithine p. 25 of 56

Glutamate to P 5 C n n Single enzyme can interconvert glutamate and 1 -pyrroline carboxylate: 1 -pyrroline-5 -carboxylate dehydrogenase 3 -layer sandwich protein 04/25/2019 Nitrogen Metabolism 1 Thermus thermophilus PCD 300 k. Da hexamer dimer shown EC 1. 5. 1. 12 PDB 2 BJA, 1. 9Å p. 26 of 56
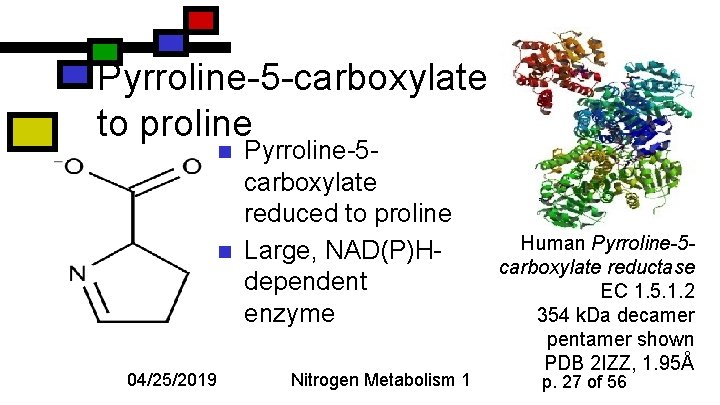
Pyrroline-5 -carboxylate to proline n n 04/25/2019 Pyrroline-5 carboxylate reduced to proline Large, NAD(P)Hdependent enzyme Nitrogen Metabolism 1 Human Pyrroline-5 carboxylate reductase EC 1. 5. 1. 2 354 k. Da decamer pentamer shown PDB 2 IZZ, 1. 95Å p. 27 of 56

Glutamate to Glu semialdehyde Glu is -phosphorylated: glu + ATP glu-5 -P +ADP (2. 7. 2. 11) n Glu-5 -P is reduced and Thermatoga maritima dephosphorylated: -glutamyl phosphate + glu-5 -P + NADPH + H reductase 47 k. Da monomer glu-5 -semialdehyde + NADP+ + Pi EC 1. 2. 1. 41 Glu-5 -P n PDB 1 O 20, 2Å 04/25/2019 Nitrogen Metabolism 1 p. 28 of 56

n n Glu semialdehyde to ornithine This is just another transamination, catalyzed by ornithine aminotransferase: glu-5 -semialdehyde + glu/asp ornithine + -ketoglutarate / oxaloacetate Typical PLP-dependent reaction ornithine 04/25/2019 Nitrogen Metabolism 1 Human OAT 193 k. Da tetramer EC 2. 6. 1. 13 PDB 2 OAT, 1. 95Å p. 29 of 56
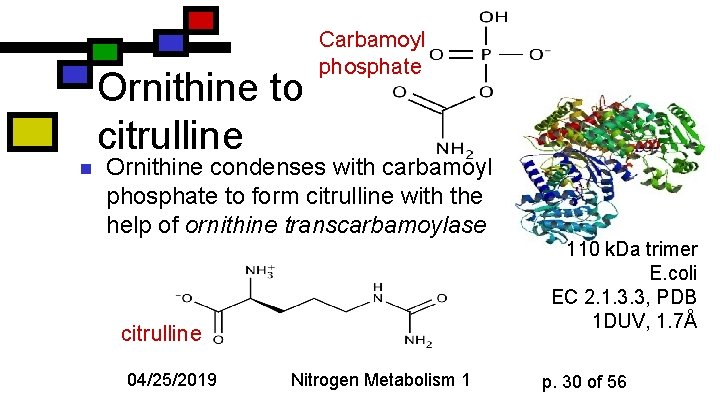
Ornithine to citrulline n Carbamoyl phosphate Ornithine condenses with carbamoyl phosphate to form citrulline with the help of ornithine transcarbamoylase citrulline 04/25/2019 Nitrogen Metabolism 1 110 k. Da trimer E. coli EC 2. 1. 3. 3, PDB 1 DUV, 1. 7Å p. 30 of 56
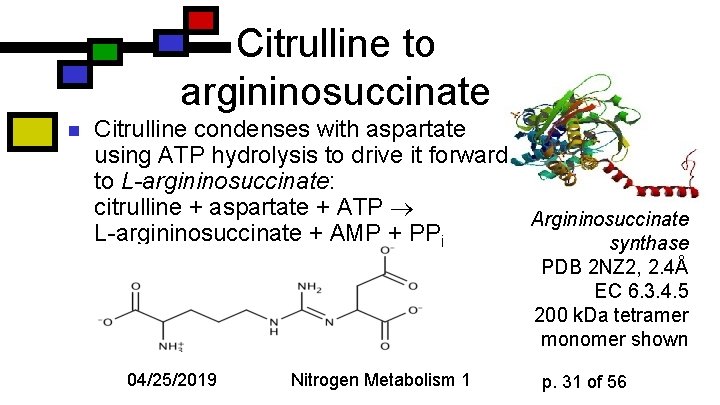
Citrulline to argininosuccinate n Citrulline condenses with aspartate using ATP hydrolysis to drive it forward to L-argininosuccinate: citrulline + aspartate + ATP Argininosuccinate L-argininosuccinate + AMP + PPi synthase PDB 2 NZ 2, 2. 4Å EC 6. 3. 4. 5 200 k. Da tetramer monomer shown 04/25/2019 Nitrogen Metabolism 1 p. 31 of 56

Argininosuccinate to arginine n n Fumarate extracted, leaving arginine Argininosuccinate lyase is also -crystallin, one of the moonlighting proteins: it’s a component of eye lenses E. coli ASL 100 k. Da dimer EC 4. 3. 2. 1, 2. 44Å PDB 1 TJ 7, 2. 44 fumarate 04/25/2019 Nitrogen Metabolism 1 p. 32 of 56

Why all that detail? n n These reactions form 75% of the urea cycle, which is an important path for amino acid and nucleic acid degradation. So we’ll need this later. 04/25/2019 Nitrogen Metabolism 1 p. 33 of 56
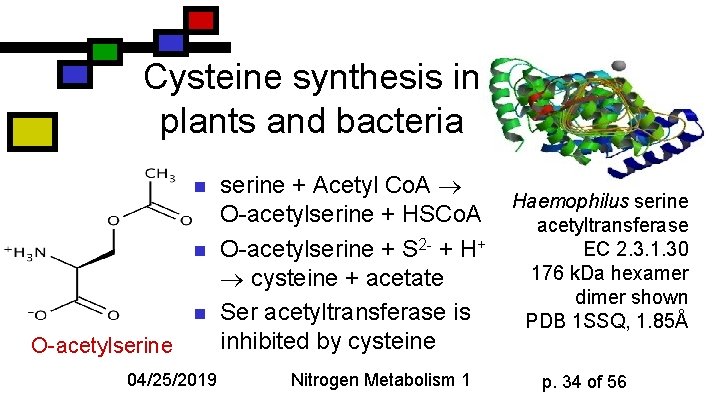
Cysteine synthesis in plants and bacteria n n n O-acetylserine 04/25/2019 serine + Acetyl Co. A O-acetylserine + HSCo. A O-acetylserine + S 2 - + H+ cysteine + acetate Ser acetyltransferase is inhibited by cysteine Nitrogen Metabolism 1 Haemophilus serine acetyltransferase EC 2. 3. 1. 30 176 k. Da hexamer dimer shown PDB 1 SSQ, 1. 85Å p. 34 of 56
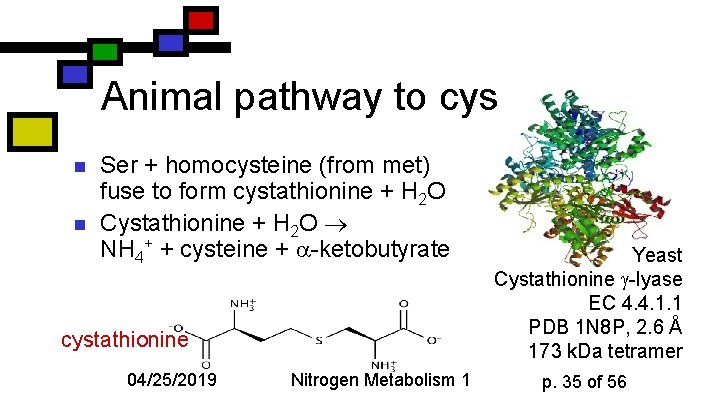
Animal pathway to cys n n Ser + homocysteine (from met) fuse to form cystathionine + H 2 O Cystathionine + H 2 O NH 4+ + cysteine + -ketobutyrate cystathionine 04/25/2019 Nitrogen Metabolism 1 Yeast Cystathionine -lyase EC 4. 4. 1. 1 PDB 1 N 8 P, 2. 6 Å 173 k. Da tetramer p. 35 of 56

n Marching through the list of twenty amino acids Amino acids we’ve already covered n n Essential but straightforward Acids and amides: n met, thr glu, gln, asp, asn n val, leu, ile Simple: n Essential & Ugly: ala, ser, gly lys, phe, tyr, trp, his Others: arg, pro, cys 04/25/2019 Nitrogen Metabolism 1 p. 36 of 56

Lys, met, thr n asp gets phosphorylated and becomes a source for all of these: n asp + ATP -aspartyl phosphate + ADP n n n via aspartate kinase -asp P + NADPH + H+ Pi + aspartate -semialdehyde +NADP+ This heads to lys or to homoserine Homoserine converts in a few steps to met or thr 04/25/2019 Nitrogen Metabolism 1 Arabidopsis Aspartate kinase 112 k. Da dimer EC 2. 7. 2. 4 PDB 2 CDQ, 2. 85Å p. 37 of 56

Asp -semialdehyde to homoserine n n -aldehyde reduced to sec-alcohol, which is homoserine Homo is generally a prefix meaning containing an extra methylene group This is precursor to homocysteine homoserine methionine It also leads to threonine 04/25/2019 Nitrogen Metabolism 1 p. 38 of 56
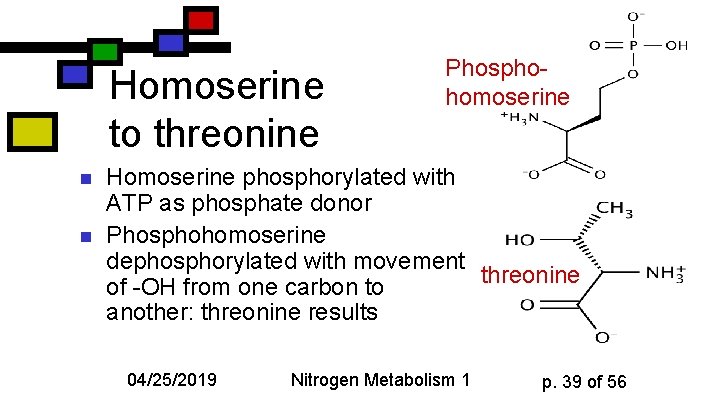
Homoserine to threonine n n Phosphohomoserine Homoserine phosphorylated with ATP as phosphate donor Phosphohomoserine dephosphorylated with movement threonine of -OH from one carbon to another: threonine results 04/25/2019 Nitrogen Metabolism 1 p. 39 of 56

homocysteine Homoserine to met n n Three reactions convert homoserine to homocysteine 5 -methyltetrahydrofolate serves as a methyl donor to convert homocysteine to methionine via methionine synthase This enzyme exists in humans but its activity is low and [homocysteine] is low; So methionine is essential in humans 04/25/2019 Nitrogen Metabolism 1 p. 40 of 56
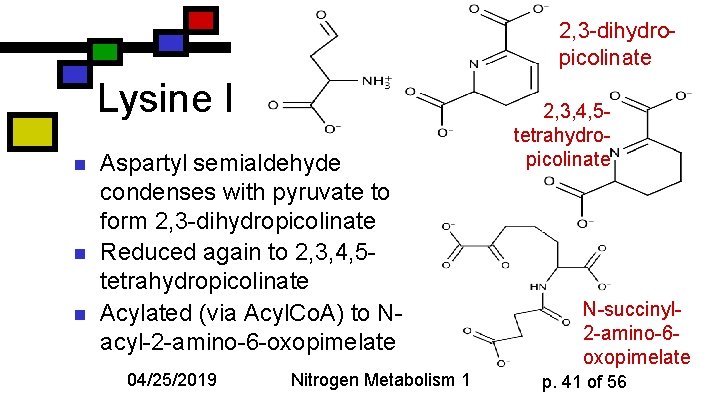
2, 3 -dihydropicolinate Lysine I n n n Aspartyl semialdehyde condenses with pyruvate to form 2, 3 -dihydropicolinate Reduced again to 2, 3, 4, 5 tetrahydropicolinate Acylated (via Acyl. Co. A) to Nacyl-2 -amino-6 -oxopimelate 04/25/2019 Nitrogen Metabolism 1 2, 3, 4, 5 tetrahydropicolinate N-succinyl 2 -amino-6 oxopimelate p. 41 of 56
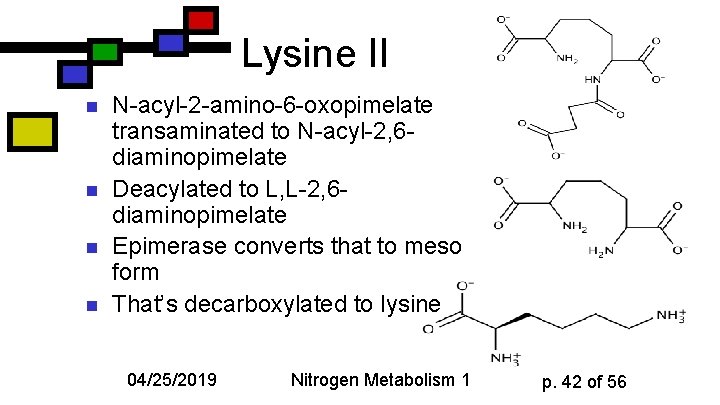
Lysine II n n N-acyl-2 -amino-6 -oxopimelate transaminated to N-acyl-2, 6 diaminopimelate Deacylated to L, L-2, 6 diaminopimelate Epimerase converts that to meso form That’s decarboxylated to lysine 04/25/2019 Nitrogen Metabolism 1 p. 42 of 56

Non-essential A. A. name n. Glycine 12 n. Serine n. Cysteine n. Alanine 20 n. Aspartate n. Asparagine n. Glutamate n. Glutamine n. Proline n. Arginine 04/25/2019 n. Tyrosine What’s left? ( = “done”) # ATP’s 18 19 Essential A. A. Name n Threonine n Valine n Histidine n Methionine n Leucine n Lysine n Isoleucine n Phenylalanine n Tryptophan 21 22 -24 30 31 39 44 * Nitrogen Metabolism 1 62 ** #ATPs 31 39 42 44 47 50 -51 55 65 78 p. 43 of 56
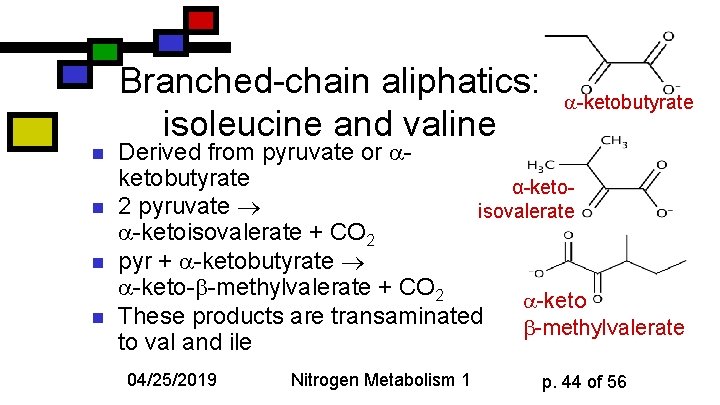
n n Branched-chain aliphatics: isoleucine and valine -ketobutyrate Derived from pyruvate or ketobutyrate α-keto 2 pyruvate isovalerate -ketoisovalerate + CO 2 pyr + -ketobutyrate -keto- -methylvalerate + CO 2 -keto These products are transaminated -methylvalerate to val and ile 04/25/2019 Nitrogen Metabolism 1 p. 44 of 56
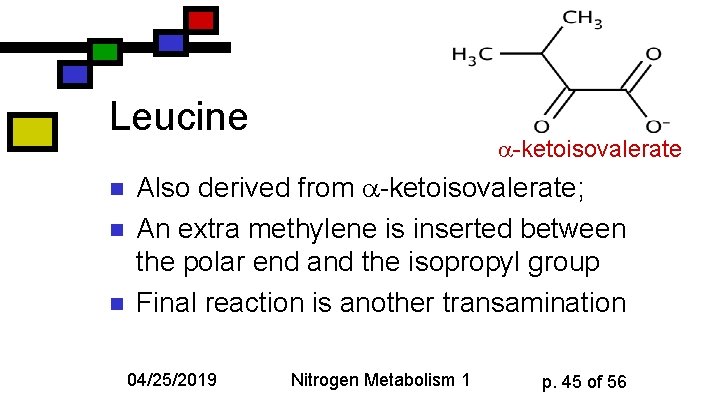
Leucine n n n -ketoisovalerate Also derived from -ketoisovalerate; An extra methylene is inserted between the polar end and the isopropyl group Final reaction is another transamination 04/25/2019 Nitrogen Metabolism 1 p. 45 of 56
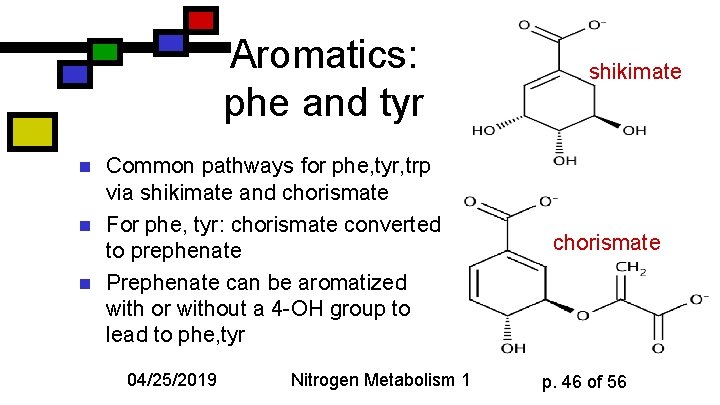
Aromatics: phe and tyr n n n Common pathways for phe, tyr, trp via shikimate and chorismate For phe, tyr: chorismate converted to prephenate Prephenate can be aromatized with or without a 4 -OH group to lead to phe, tyr 04/25/2019 Nitrogen Metabolism 1 shikimate chorismate p. 46 of 56
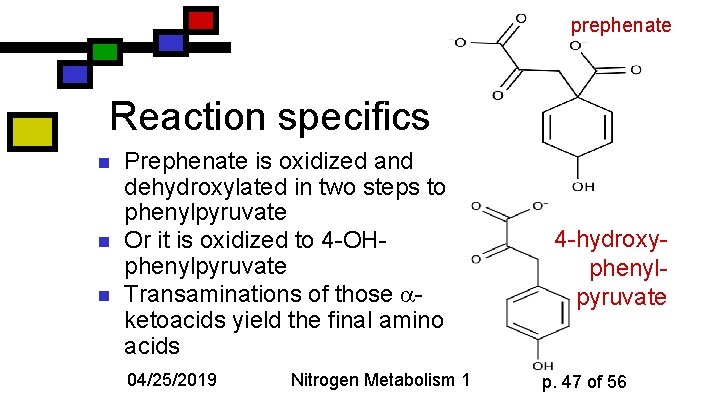
prephenate Reaction specifics n n n Prephenate is oxidized and dehydroxylated in two steps to phenylpyruvate Or it is oxidized to 4 -OHphenylpyruvate Transaminations of those ketoacids yield the final amino acids 04/25/2019 Nitrogen Metabolism 1 4 -hydroxyphenylpyruvate p. 47 of 56

Chorismate mutase n n Isomerase, converts chorismate to prephenate In E. coli: 2 versions depending on which path the product is heading to Active sites are similar in all organisms but architecture is very different Catalytic triad similar to serine proteases 04/25/2019 Nitrogen Metabolism 1 B. subtilis chorismate mutase 42 k. Da trimer EC 5. 4. 99. 5 PDB 1 DBF, 1. 3Å p. 48 of 56
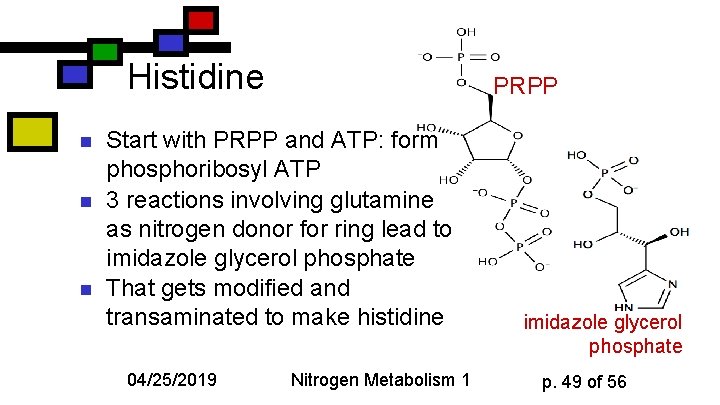
Histidine n n n PRPP Start with PRPP and ATP: form phosphoribosyl ATP 3 reactions involving glutamine as nitrogen donor for ring lead to imidazole glycerol phosphate That gets modified and transaminated to make histidine 04/25/2019 Nitrogen Metabolism 1 imidazole glycerol phosphate p. 49 of 56

Tryptophan n n anthranilate Also via chorismate Sidechain amide transferred to chorismate Conversion to anthranilate PRPP provides phosphoribosyl moiety; this sets up indole glycerol phosphate Tryptophan synthase eliminates glyc-P, adds ser 04/25/2019 Nitrogen Metabolism 1 p. 50 of 56

i. Clicker question #2 2. Which of these amino acids is not synthesized through a pathway involving chorismate? n n n 04/25/2019 (a) phe (b) his (c) trp (d) tyr (e) all of those involve chorismate Nitrogen Metabolism 1 p. 51 of 56

What do we do with amino acids? n n Obviously a lot of them serve as building-blocks for protein and peptide synthesis via ribosomal mechanisms Also serve as metabolites, getting converted to other compounds (including nucleic acids) or getting oxidized as fuel 04/25/2019 Nitrogen Metabolism 1 p. 52 of 56
Transaminations n Generally two stages: n n n amino acid + -ketoglutarate -keto acid + glutamate Glutamate + NAD+ + H 2 O -ketoglutarate + NADH + H+ + NH 4+ Net reaction is amino acid + NAD+ + H 2 O -keto acid- + NADH + H+ + NH 4+ 04/25/2019 Nitrogen Metabolism 1 p. 53 of 56

Glucogenic amino acids n Degradation of many amino acids lead to TCA cycle intermediates or pyruvate n n These can be built back up to glucose; So these are called glucogenic 04/25/2019 Nitrogen Metabolism 1 p. 54 of 56

Ketogenic amino acids n Degradation of others leads to acetyl Co. A and related compounds n n these cannot be built back up to glucose except via the glyoxalate shuttle these are called ketogenic 04/25/2019 Nitrogen Metabolism 1 p. 55 of 56

Glucogenic amino acids n n n Amino acids that can be catabolized to produce building blocks that lead to glucose without help of glyoxalate pathway Most produce pyruvate, -ketoglutarate, succinyl Co. A, succinate, fumarate, or oxaloacetate By convention, amino acids that produce both TCA-cycle intermediates and acetyl Co. A are labeled as glucogenic 04/25/2019 Nitrogen Metabolism 1 p. 56 of 56
- Slides: 56