NGS Data Consortium October 8 2012 HLA Genotyping

NGS Data Consortium October 8, 2012 HLA Genotyping Data Generated by 454 Sequencing Cherie Holcomb, Ph. D. Roche Molecular Systems picture placeholder

Medium, High, and Very High Resolution HLA Genotyping Systems Targeted Amplicon sequencing – Primers target up to 9 loci (format as fusion primers or “ 4 primer system” w/use of Fluidigm Access Array™) • NOTE: After amplification of g. DNA, amplicons contain all adapters & MIDs etc. for NGS sequencing – Workflow: Amplicons processed either individually or pooled (fusion primers) or pooled (“ 4 primer system”) – 454 Life Sciences GS FLX or GS Junior for NGS – Conexio Assign ATF 454 software (commercially available will perform MR and HR; for VHR limited early access to Conexio Assign MPS v 1. 0)

Amplicon Sequencing using 454 GS FLX* *GS Junior can also be used, ~8 x fewer samples per run ∆Abstract #1025 -LB ǂAbstract #ORO 1 -02 NOTE: All current products are for Research Use Only
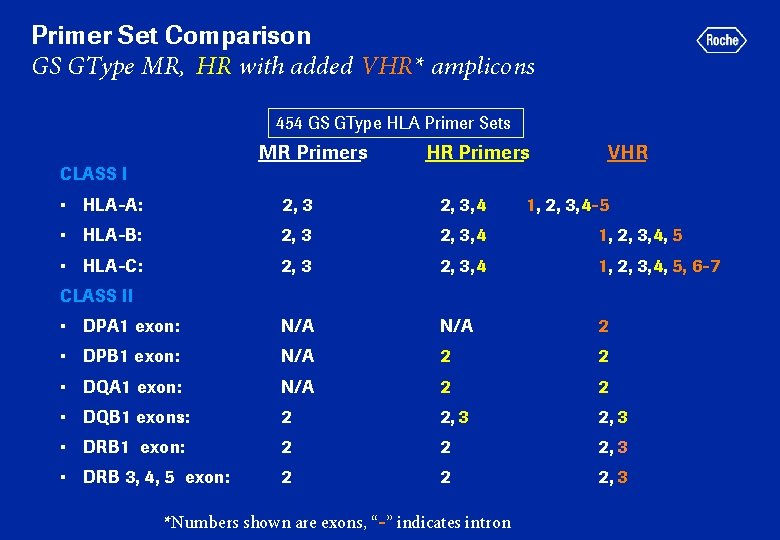
Primer Set Comparison GS GType MR, HR with added VHR* amplicons 454 GS GType HLA Primer Sets MR Primers CLASS I HR Primers VHR • HLA-A: 2, 3, 4 • HLA-B: 2, 3, 4 1, 2, 3, 4, 5 • HLA-C: 2, 3, 4 1, 2, 3, 4, 5, 6 -7 • DPA 1 exon: N/A 2 • DPB 1 exon: N/A 2 2 • DQA 1 exon: N/A 2 2 • DQB 1 exons: 2 2, 3 • DRB 1 exon: 2 2 2, 3 • DRB 3, 4, 5 exon: 2 2 2, 3 1, 2, 3, 4 -5 CLASS II *Numbers shown are exons, “-” indicates intron
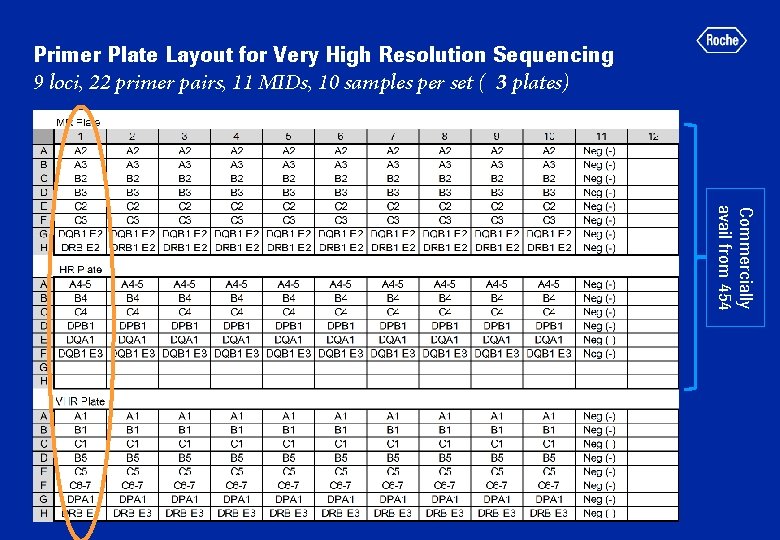
Primer Plate Layout for Very High Resolution Sequencing 9 loci, 22 primer pairs, 11 MIDs, 10 samples per set ( 3 plates) Commercially avail from 454
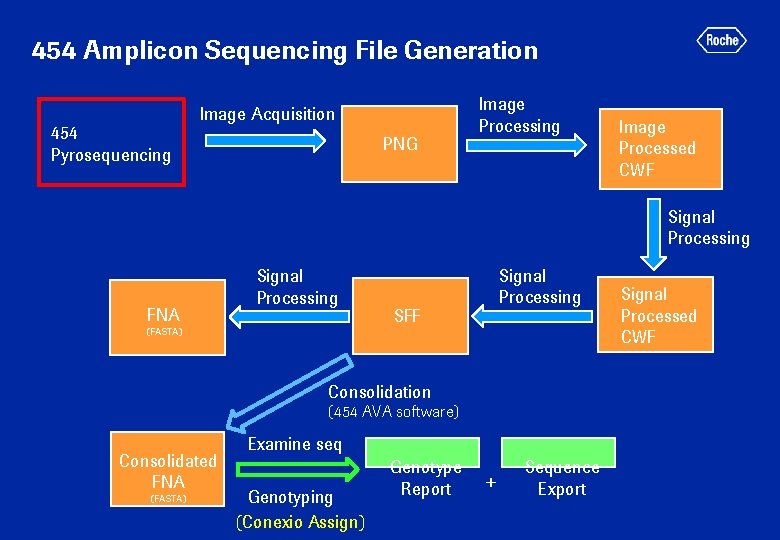
454 Amplicon Sequencing File Generation 454 Pyrosequencing Image Acquisition PNG Image Processing Image Processed CWF Signal Processing FNA Signal Processing (FASTA) Signal Processing SFF Consolidation (454 AVA software) Consolidated FNA (FASTA) Examine seq Genotyping (Conexio Assign) Genotype Report + Sequence Export Signal Processed CWF
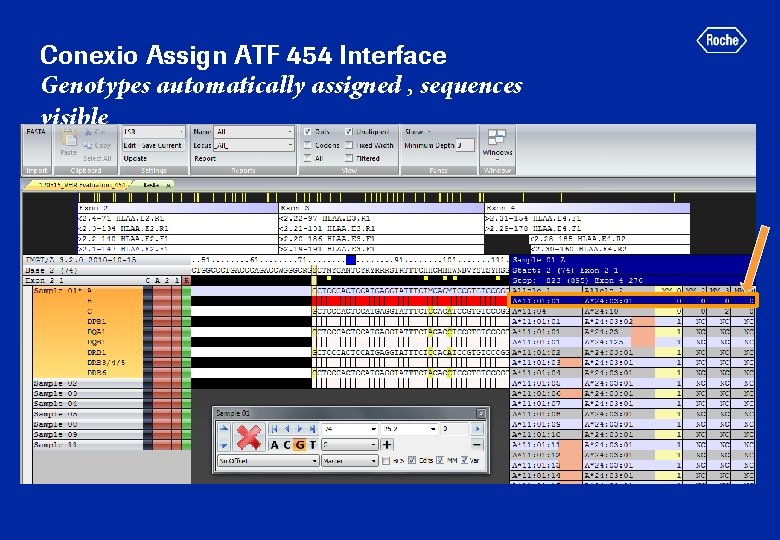
Conexio Assign ATF 454 Interface Genotypes automatically assigned , sequences visible
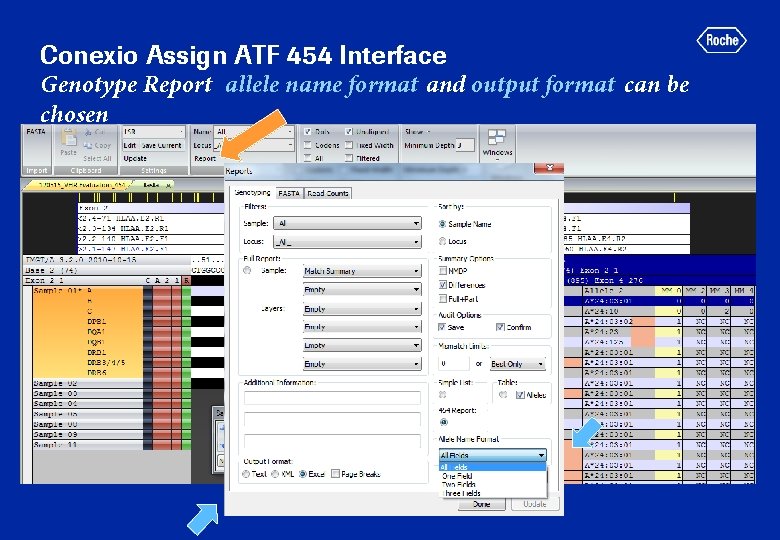
Conexio Assign ATF 454 Interface Genotype Report allele name format and output format can be chosen

Conexio Assign. MPS v 1. 0 Genotyping Report All fields, MS Excel format A CWD filter OR “highlighting” of CWD alleles in report (preferred) has been requested 454 GS GType HLA +VHR primers—only part of report shown; assay includes DQA 1, DQB 1, DRB 1 and DRB 3/4/5
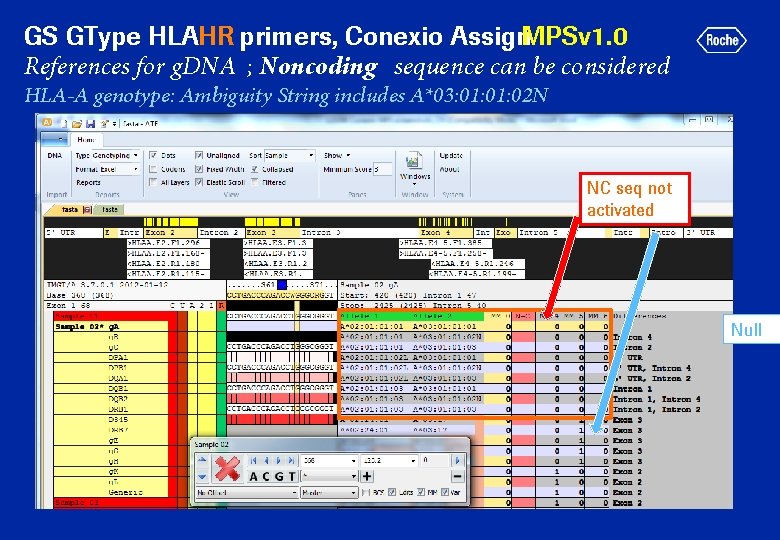
GS GType HLAHR primers, Conexio Assign. MPSv 1. 0 References for g. DNA ; Noncoding sequence can be considered HLA-A genotype: Ambiguity String includes A*03: 01: 02 N NC seq not activated Null
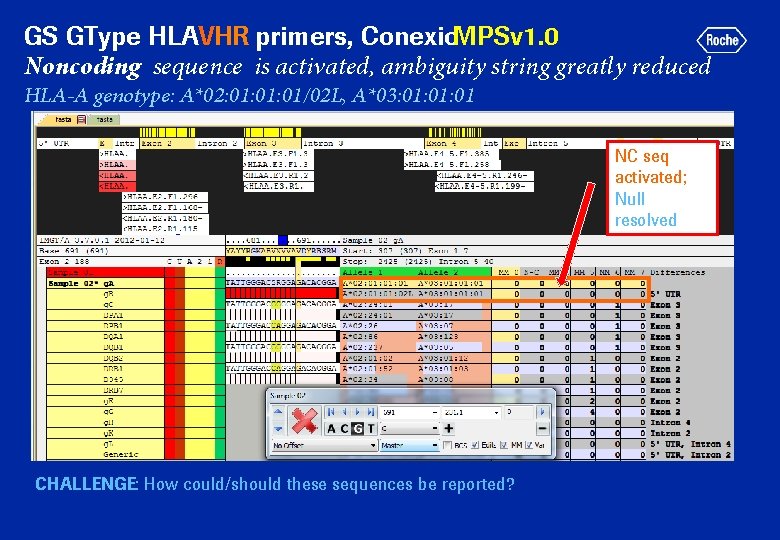
GS GType HLAVHR primers, Conexio. MPSv 1. 0 Noncoding sequence is activated, ambiguity string greatly reduced HLA-A genotype: A*02: 01: 01/02 L, A*03: 01: 01 NC seq activated; Null resolved CHALLENGE: How could/should these sequences be reported?

Reporting of Sequence Information Currently • Can report out (combined) consensusexon sequence that has given rise to list of possible genotypes for a given sample/locus. Can do this easily for all samples. (Q: If community decides sequence is necessary for publications, is this sufficient? ) – Cannot report component(consensus) sequences (with exons matched) to give individual allele(s) – Doesn’t include intronic sequence (but can report consensus of each intron individually—too laborious to be practical) – FASTA format in notepad (Q: Sufficient for publication? )

Reporting of Sequence Information Preferred • Option to report component sequences(with exons and introns matched) to give allele calls—imp for reporting new alleles • For (combined or individual alleles) consensus sequence – Option to report Coding only OR Coding plus Noncoding – Format options including XML (accepted by IMGT)—imp for reporting new alleles Works in Progress
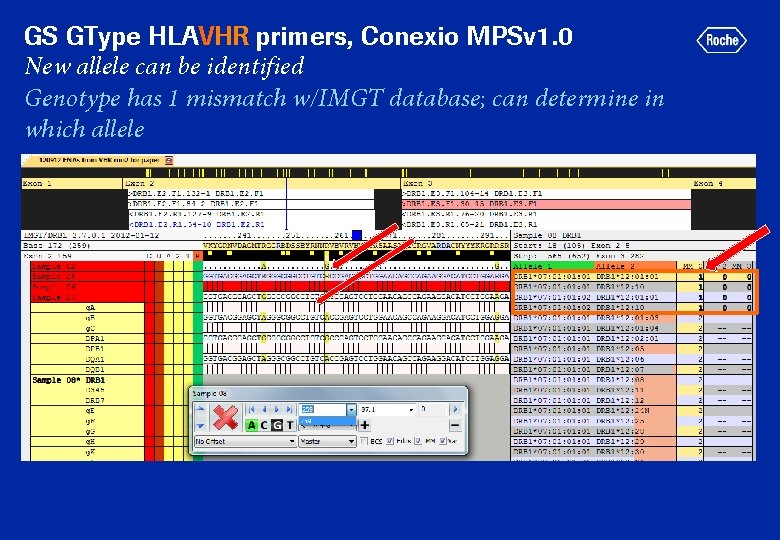
GS GType HLAVHR primers, Conexio MPSv 1. 0 New allele can be identified Genotype has 1 mismatch w/IMGT database; can determine in which allele
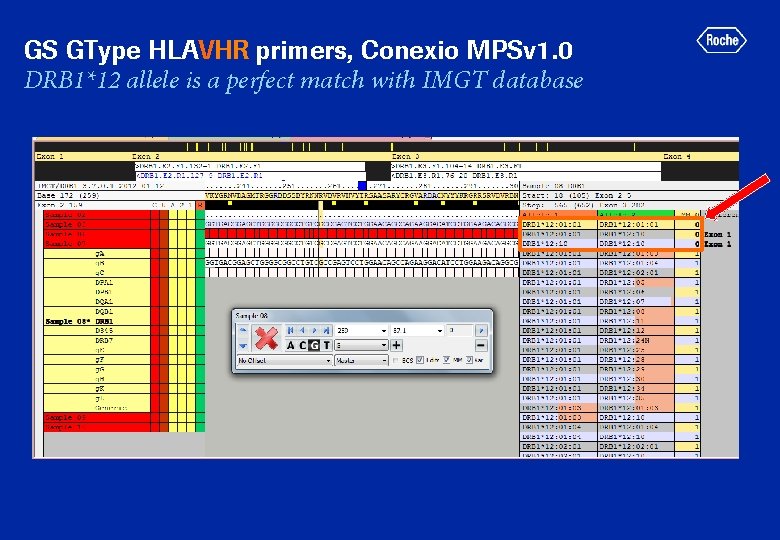
GS GType HLAVHR primers, Conexio MPSv 1. 0 DRB 1*12 allele is a perfect match with IMGT database
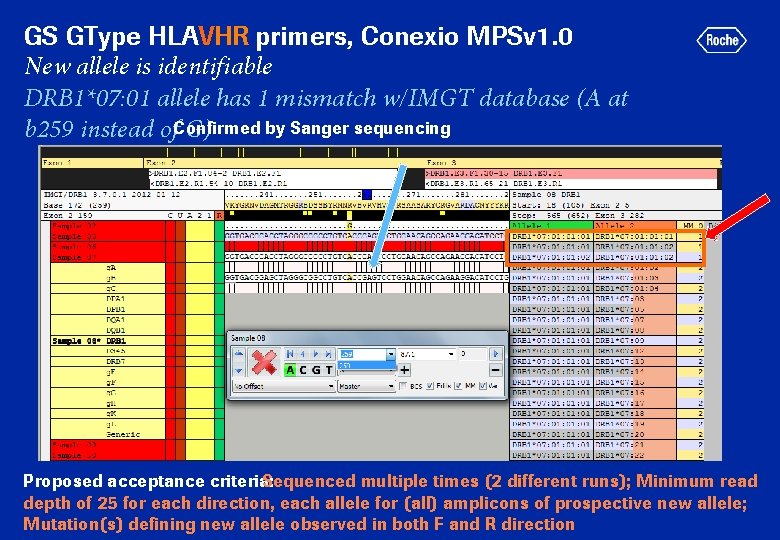
GS GType HLAVHR primers, Conexio MPSv 1. 0 New allele is identifiable DRB 1*07: 01 allele has 1 mismatch w/IMGT database (A at by Sanger sequencing b 259 instead of. Confirmed G) Proposed acceptance criteria: Sequenced multiple times (2 different runs); Minimum read depth of 25 for each direction, each allele for (all) amplicons of prospective new allele; Mutation(s) defining new allele observed in both F and R direction
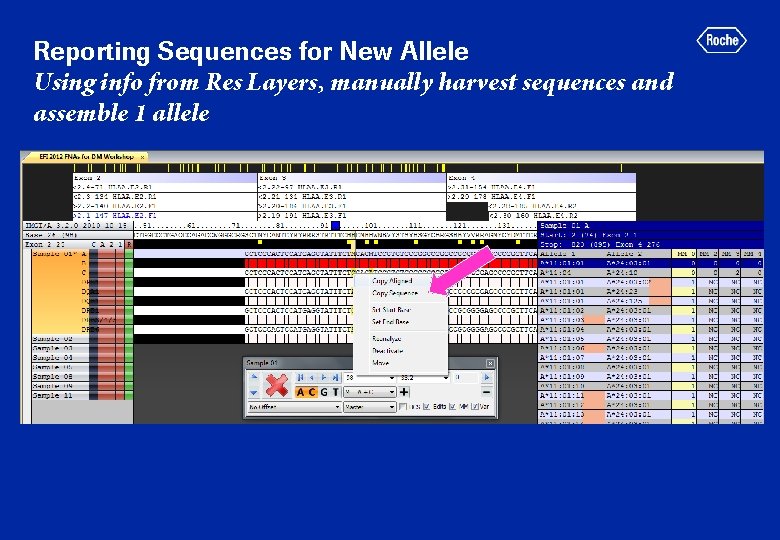
Reporting Sequences for New Allele Using info from Res Layers, manually harvest sequences and assemble 1 allele

“Copy Sequence” Output Simple Text file: Copy into Word Pad, Excel, Bioedit Not in FASTA or XML format (currently no way to convert to latter) Assume XML is most appropriate for submission to IMGT database In discussion with Conexio
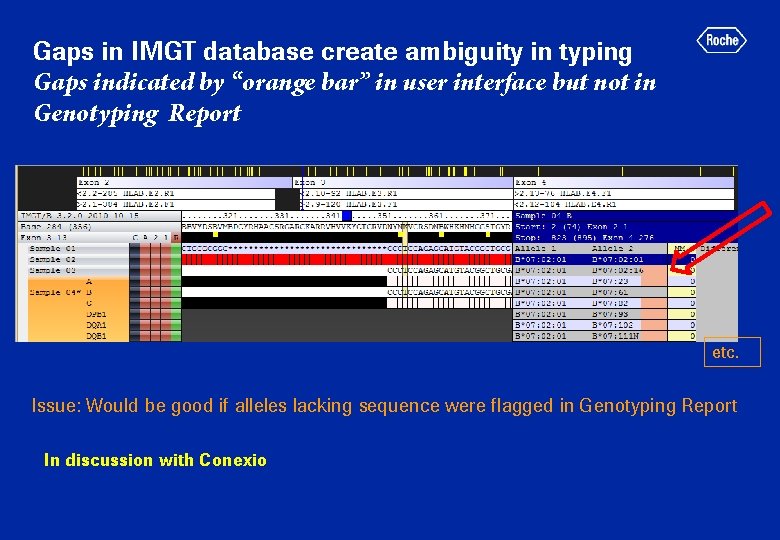
Gaps in IMGT database create ambiguity in typing Gaps indicated by “orange bar” in user interface but not in Genotyping Report etc. Issue: Would be good if alleles lacking sequence were flagged in Genotyping Report In discussion with Conexio

Additional info & Summary • Using HR or VHR 454 sequencing HLA genotyping system including Conexio Assign ATF 454 or MPS v 1. 0 software, respectively: – Ambiguity string lengths are reduced to a practically reportable size – Genotype/ allele ambiguity strings (in various formats using combinations of delineation in columns, “+”, “or”, “, ”) can be reported in Excel, text and XML(? ? ) format at 1, 2, or all field level. – NMDP codes supported – Most recent IMGT nomenclature and references supported (updated with periodicity, 6 months); version of references used is reported – Export of consensussequence used to make genotype calls for all loci/all samples is easily accomplished in FASTA format—currently doesn’t include NC sequence. – New alleles readily identifiable, however, reporting of amplicon sequences currently only possible by manual “harvesting” into text file.

Acknowledgements • Roche Molecular Systems – Henry Erlich • Conexio Genomics – Damian Goodridge

We Innovate Healthcare 22

Back-up slides 23
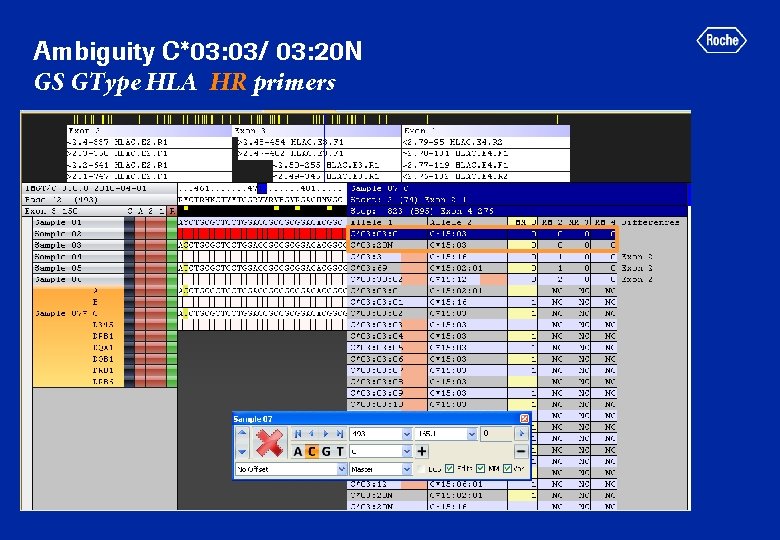
Ambiguity C*03: 03/ 03: 20 N GS GType HLA HR primers
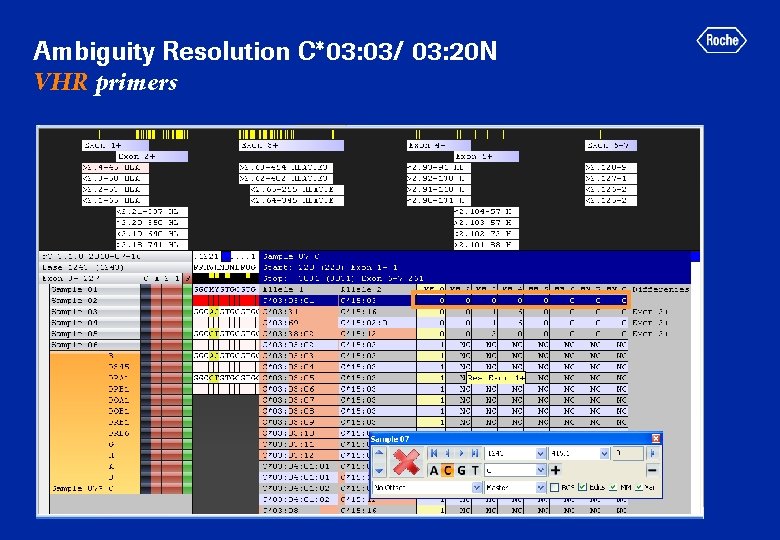
Ambiguity Resolution C*03: 03/ 03: 20 N VHR primers
- Slides: 25