Next assignment Mon Feb 22 pick a biofuel

Next assignment Mon Feb 22 pick a biofuel, gene or condition & convince the group in 5 -10 minutes why your choice is best. Suggestions 1. Effects of environment on biofuel production • Drought/humidity/salinity • p. CO 2 • Temperature • Light • Quantity • PFD (photonflux density) = intensity • Photoperiod (daylength) = duration • Light quality = color(s) • Nutrition • Macronutrients = N, P, K, S, Ca, Mg, Fe • Micronutrients, vitamins, adding acetate? 2. Effects of environment on cell walls: Drought? inhibitors?
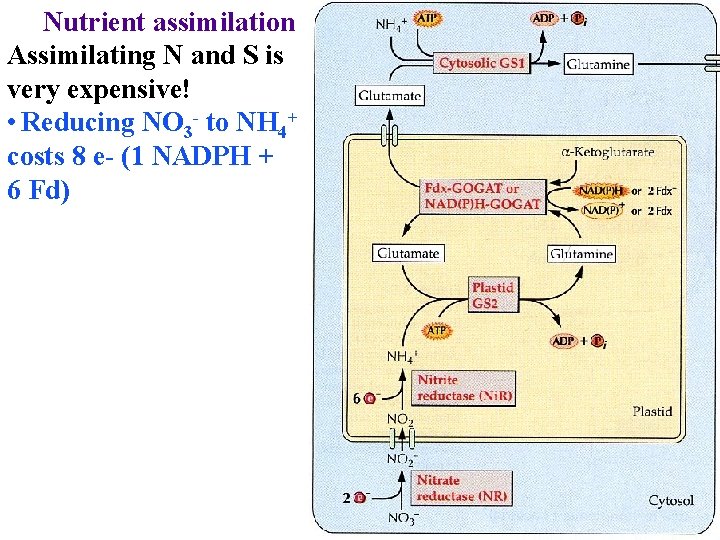
Nutrient assimilation Assimilating N and S is very expensive! • Reducing NO 3 - to NH 4+ costs 8 e- (1 NADPH + 6 Fd)
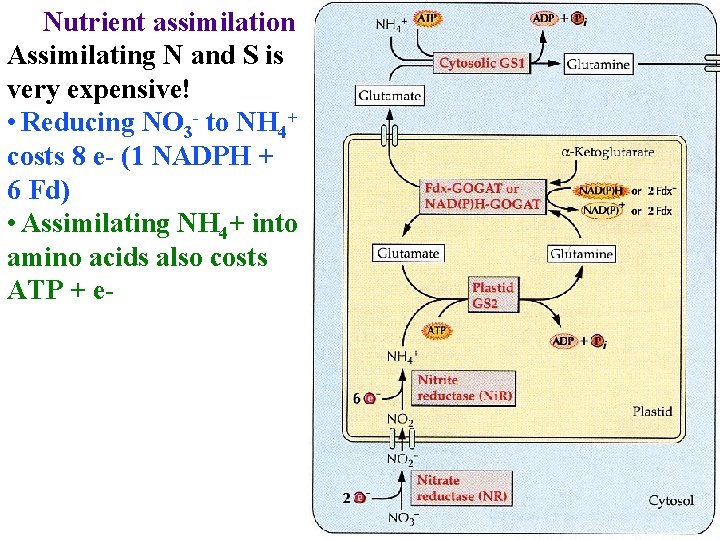
Nutrient assimilation Assimilating N and S is very expensive! • Reducing NO 3 - to NH 4+ costs 8 e- (1 NADPH + 6 Fd) • Assimilating NH 4+ into amino acids also costs ATP + e-
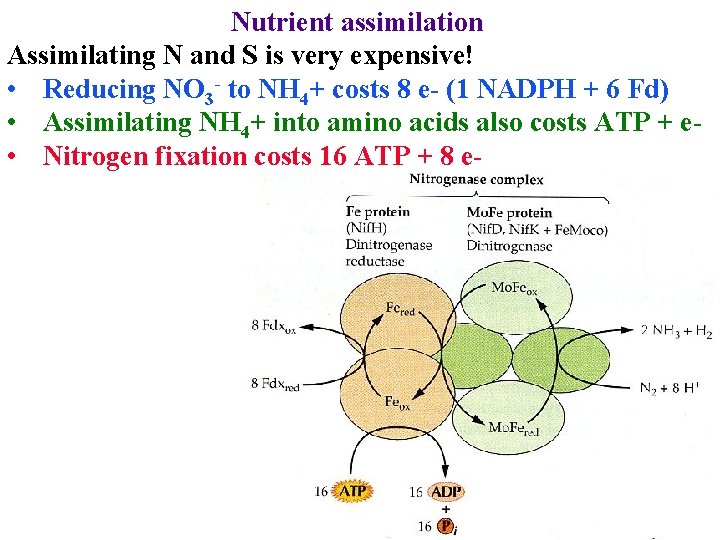
Nutrient assimilation Assimilating N and S is very expensive! • Reducing NO 3 - to NH 4+ costs 8 e- (1 NADPH + 6 Fd) • Assimilating NH 4+ into amino acids also costs ATP + e • Nitrogen fixation costs 16 ATP + 8 e-

Nutrient assimilation Assimilating N and S is very expensive! • Reducing NO 3 - to NH 4+ costs 8 e- (1 NADPH + 6 Fd) • Assimilating NH 4+ into amino acids also costs ATP + e • Nitrogen fixation costs 16 ATP + 8 e • SO 42 - reduction to S 2 - costs 8 e- + 2 ATP

Nutrient assimilation Assimilating N and S is very expensive! • Reducing NO 3 - to NH 4+ costs 8 e- (1 NADPH + 6 Fd) • Assimilating NH 4+ into amino acids also costs ATP + e • Nitrogen fixation costs 16 ATP + 8 e • SO 42 - reduction to S 2 - costs 8 e- + 2 ATP • S 2 - assimilation into Cysteine costs 2 more e • Most explosives are based on N or S!
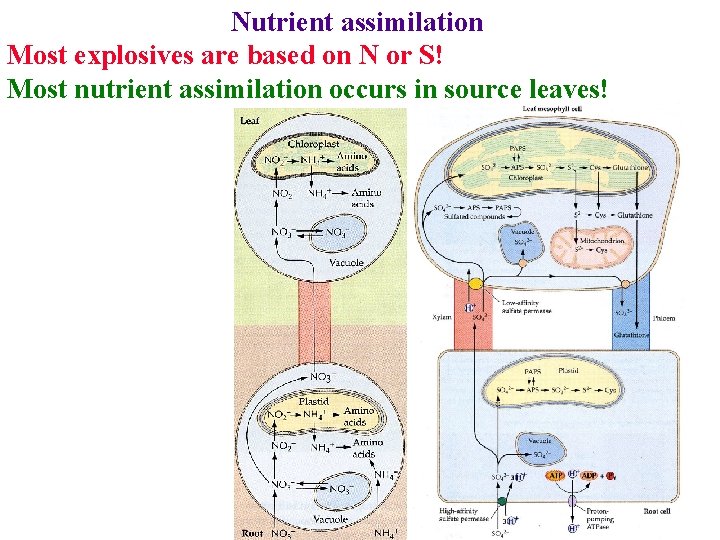
Nutrient assimilation Most explosives are based on N or S! Most nutrient assimilation occurs in source leaves!
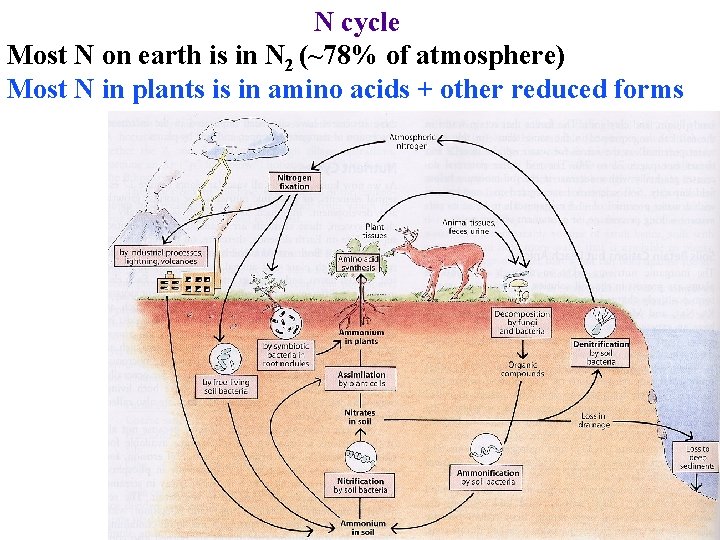
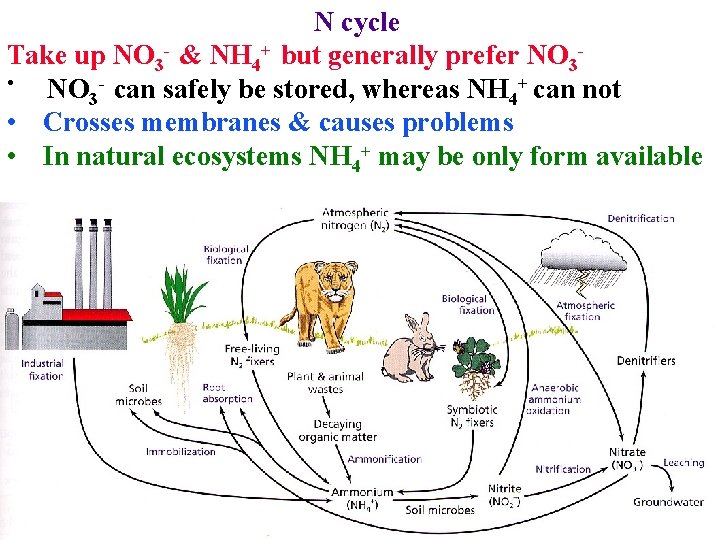
N cycle Most N on earth is in N 2 (~78% of atmosphere) Most N in plants is in amino acids + other reduced forms
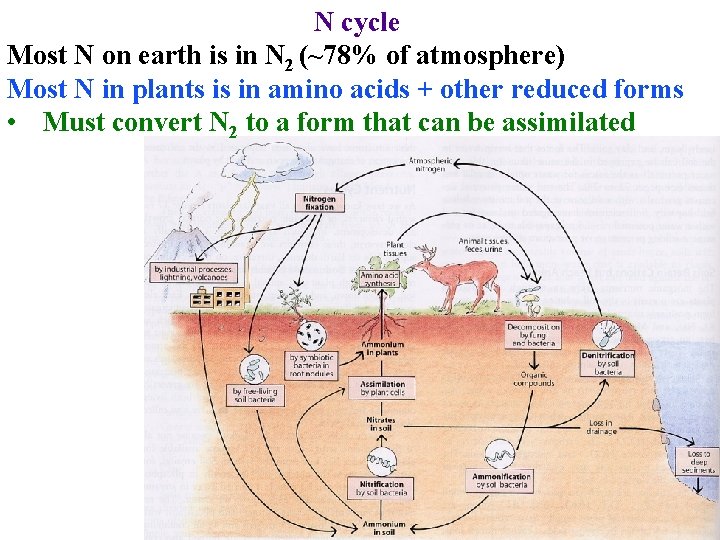
N cycle Most N on earth is in N 2 (~78% of atmosphere) Most N in plants is in amino acids + other reduced forms • Must convert N 2 to a form that can be assimilated
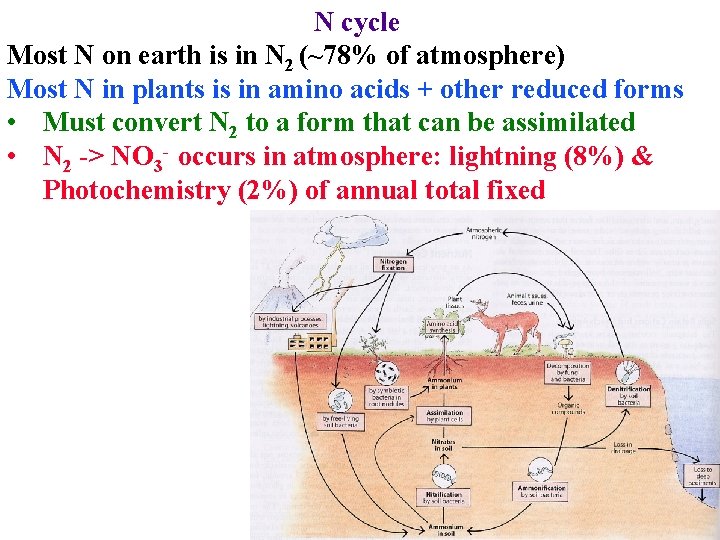
N cycle Most N on earth is in N 2 (~78% of atmosphere) Most N in plants is in amino acids + other reduced forms • Must convert N 2 to a form that can be assimilated • N 2 -> NO 3 - occurs in atmosphere: lightning (8%) & Photochemistry (2%) of annual total fixed
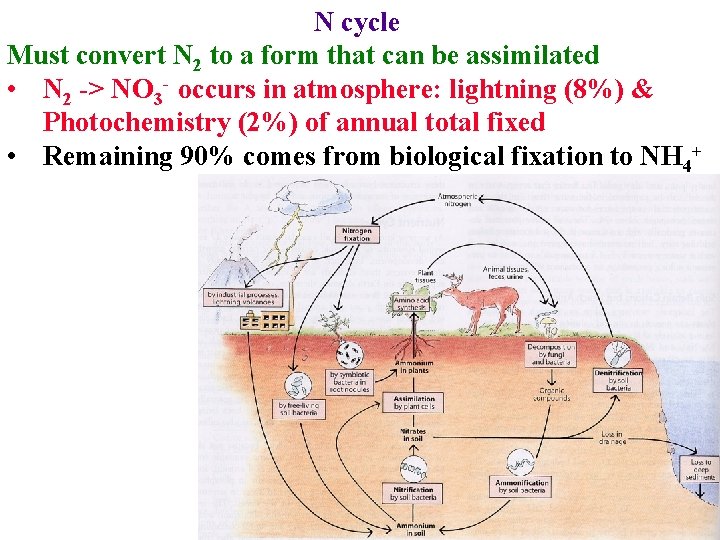
N cycle Must convert N 2 to a form that can be assimilated • N 2 -> NO 3 - occurs in atmosphere: lightning (8%) & Photochemistry (2%) of annual total fixed • Remaining 90% comes from biological fixation to NH 4+
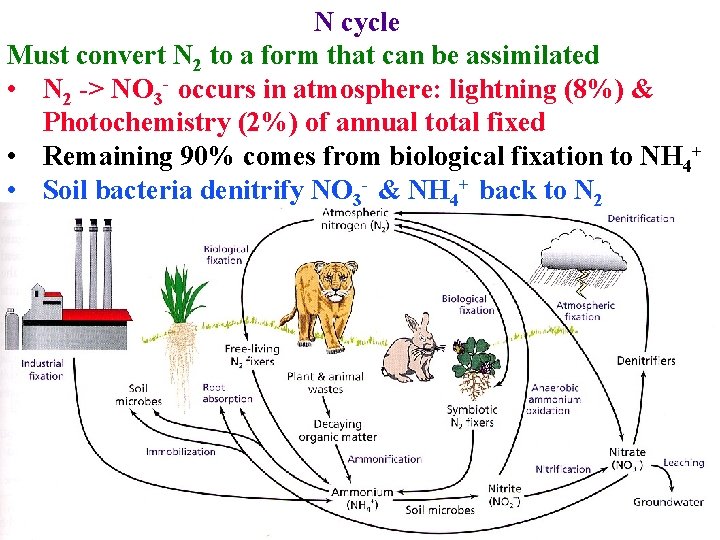
N cycle Must convert N 2 to a form that can be assimilated • N 2 -> NO 3 - occurs in atmosphere: lightning (8%) & Photochemistry (2%) of annual total fixed • Remaining 90% comes from biological fixation to NH 4+ • Soil bacteria denitrify NO 3 - & NH 4+ back to N 2
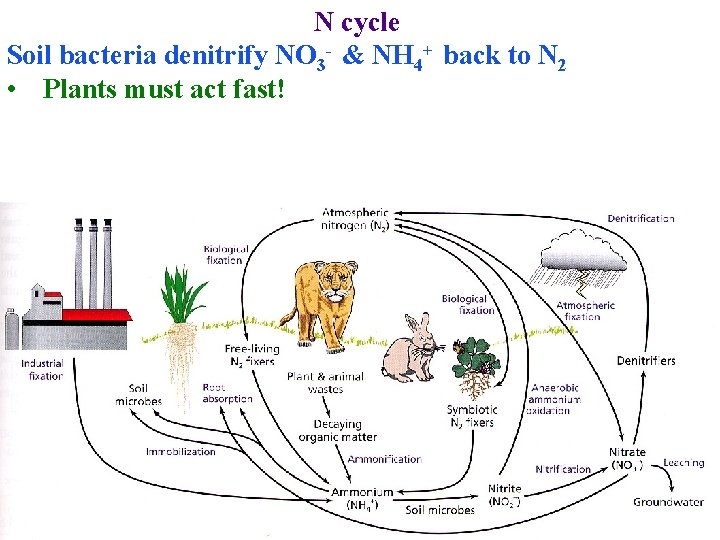
N cycle Soil bacteria denitrify NO 3 - & NH 4+ back to N 2 • Plants must act fast!
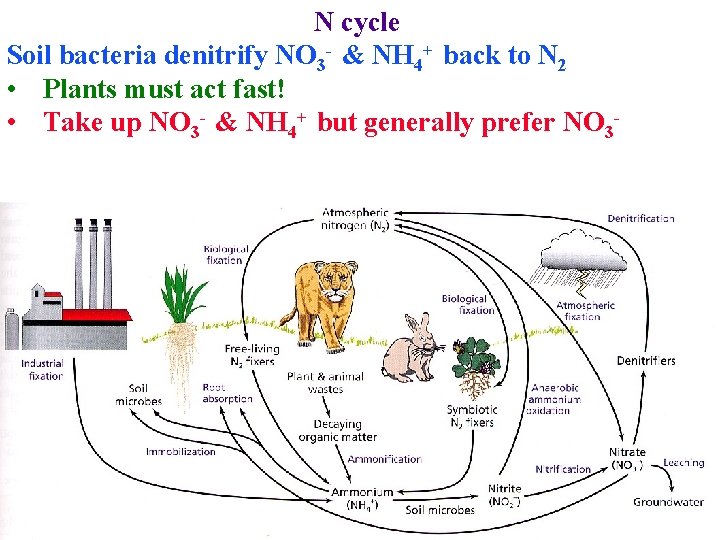
N cycle Soil bacteria denitrify NO 3 - & NH 4+ back to N 2 • Plants must act fast! • Take up NO 3 - & NH 4+ but generally prefer NO 3 -
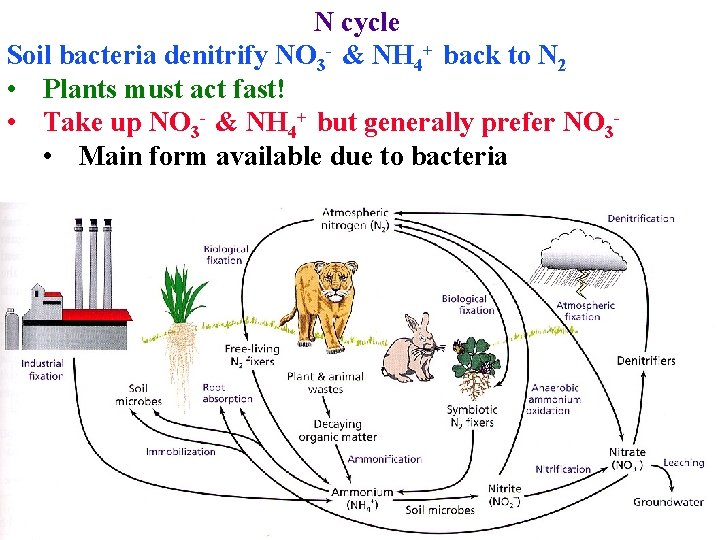
N cycle Soil bacteria denitrify NO 3 - & NH 4+ back to N 2 • Plants must act fast! • Take up NO 3 - & NH 4+ but generally prefer NO 3 • Main form available due to bacteria
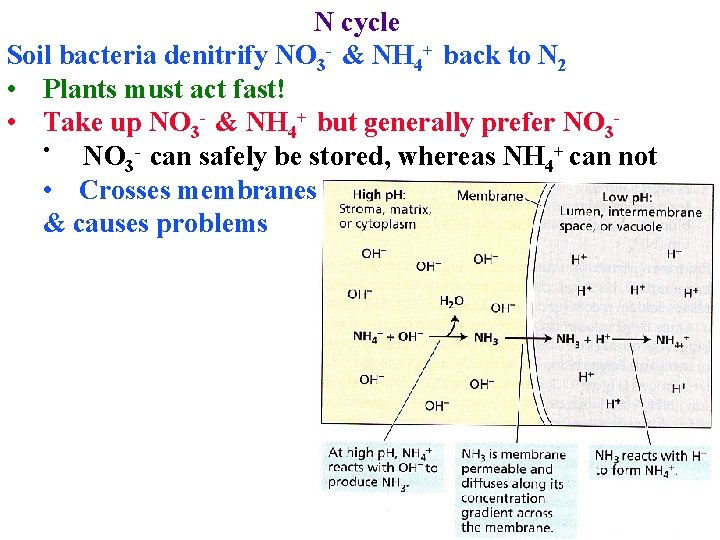
N cycle Soil bacteria denitrify NO 3 - & NH 4+ back to N 2 • Plants must act fast! • Take up NO 3 - & NH 4+ but generally prefer NO 3 • NO 3 - can safely be stored, whereas NH 4+ can not • Crosses membranes & causes problems
N cycle Take up NO 3 - & NH 4+ but generally prefer NO 3 • NO 3 - can safely be stored, whereas NH 4+ can not • Crosses membranes & causes problems • In natural ecosystems NH 4+ may be only form available
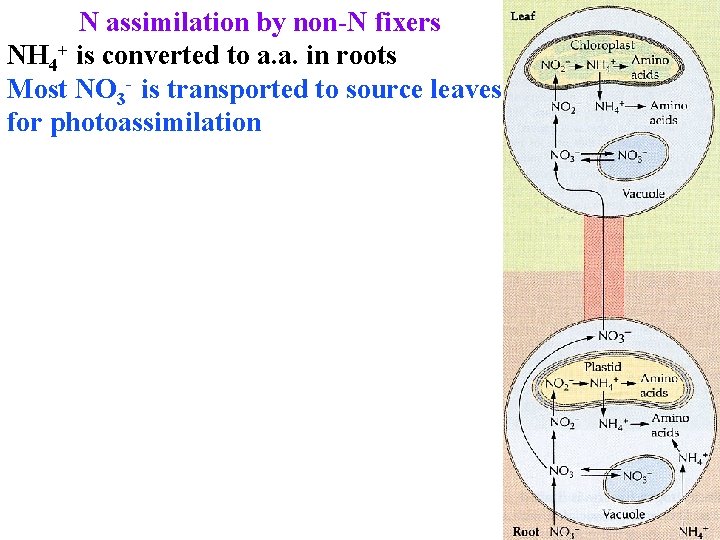
N assimilation by non-N fixers NH 4+ is converted to a. a. in roots Most NO 3 - is transported to source leaves for photoassimilation
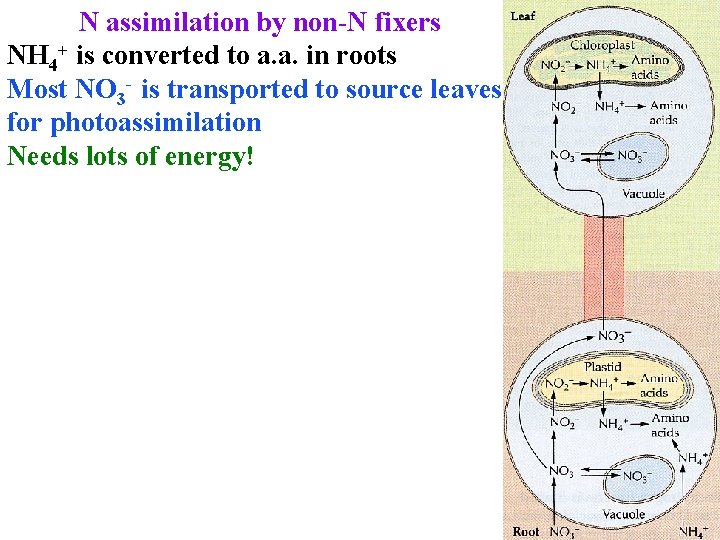
N assimilation by non-N fixers NH 4+ is converted to a. a. in roots Most NO 3 - is transported to source leaves for photoassimilation Needs lots of energy!
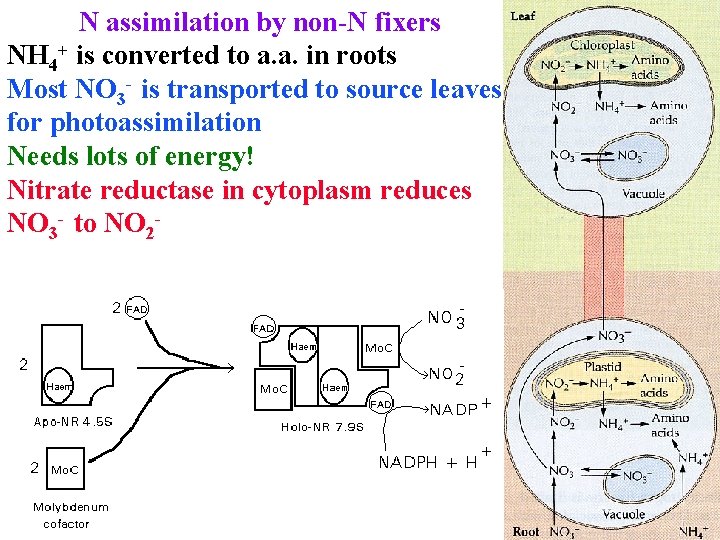
N assimilation by non-N fixers NH 4+ is converted to a. a. in roots Most NO 3 - is transported to source leaves for photoassimilation Needs lots of energy! Nitrate reductase in cytoplasm reduces NO 3 - to NO 2 -
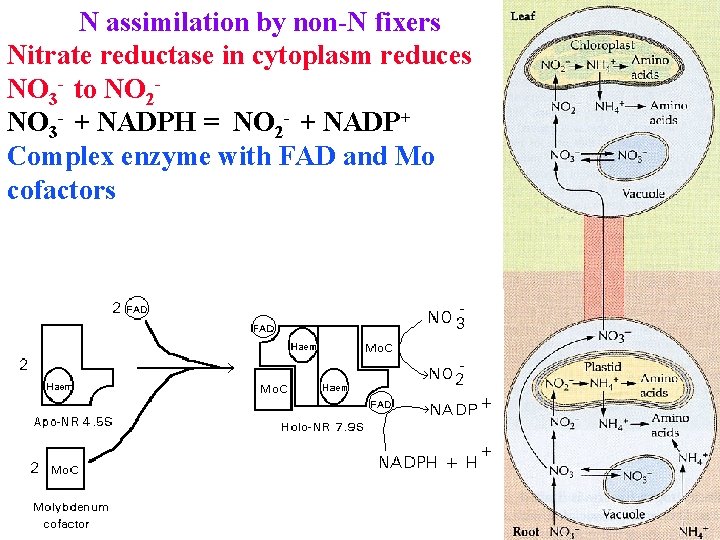
N assimilation by non-N fixers Nitrate reductase in cytoplasm reduces NO 3 - to NO 2 NO 3 - + NADPH = NO 2 - + NADP+ Complex enzyme with FAD and Mo cofactors
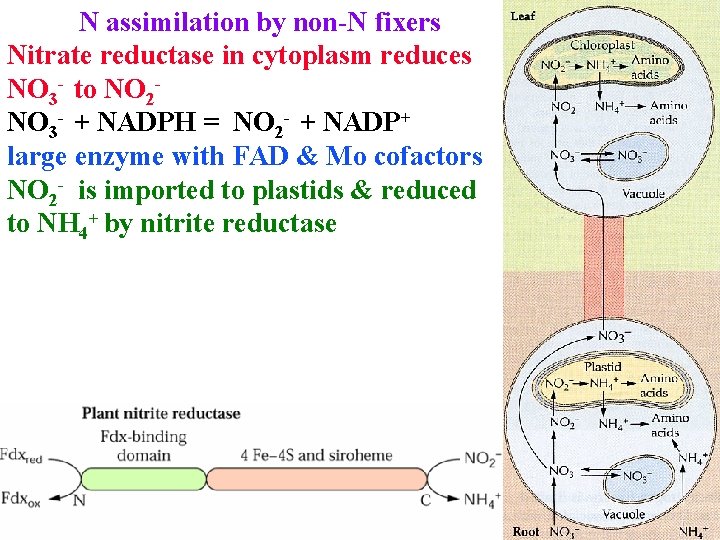
N assimilation by non-N fixers Nitrate reductase in cytoplasm reduces NO 3 - to NO 2 NO 3 - + NADPH = NO 2 - + NADP+ large enzyme with FAD & Mo cofactors NO 2 - is imported to plastids & reduced to NH 4+ by nitrite reductase
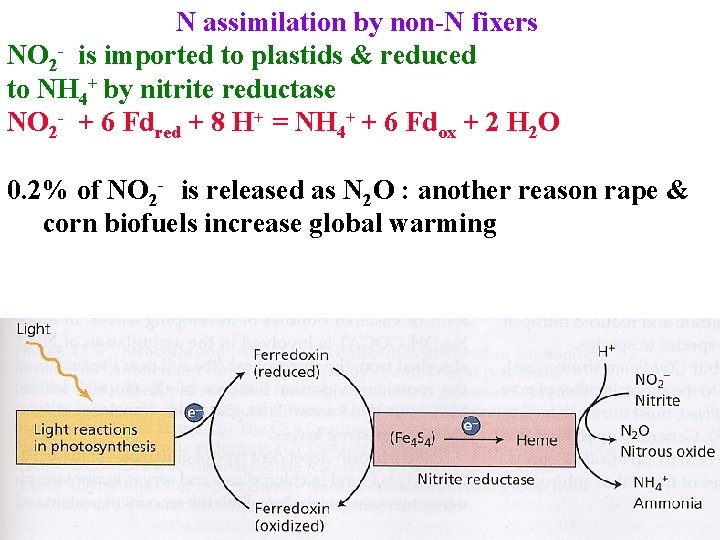
N assimilation by non-N fixers NO 2 - is imported to plastids & reduced to NH 4+ by nitrite reductase NO 2 - + 6 Fdred + 8 H+ = NH 4+ + 6 Fdox + 2 H 2 O 0. 2% of NO 2 - is released as N 2 O : another reason rape & corn biofuels increase global warming

N assimilation by non-N fixers NO 2 - is imported to plastids & reduced to NH 4+ by nitrite reductase NO 2 - + 6 Fdred + 8 H+ = NH 4+ + 6 Fdox + 2 H 2 O 0. 2% of NO 2 - is released as N 2 O : another reason rape & corn biofuels increase global warming Regulated at NO 3 - reductase; always << NO 2 - reductase NO 2 - is toxic!
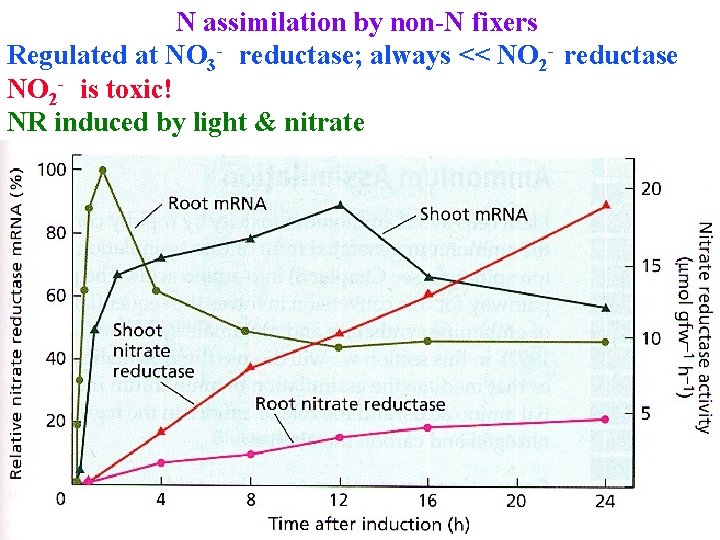
N assimilation by non-N fixers Regulated at NO 3 - reductase; always << NO 2 - reductase NO 2 - is toxic! NR induced by light & nitrate
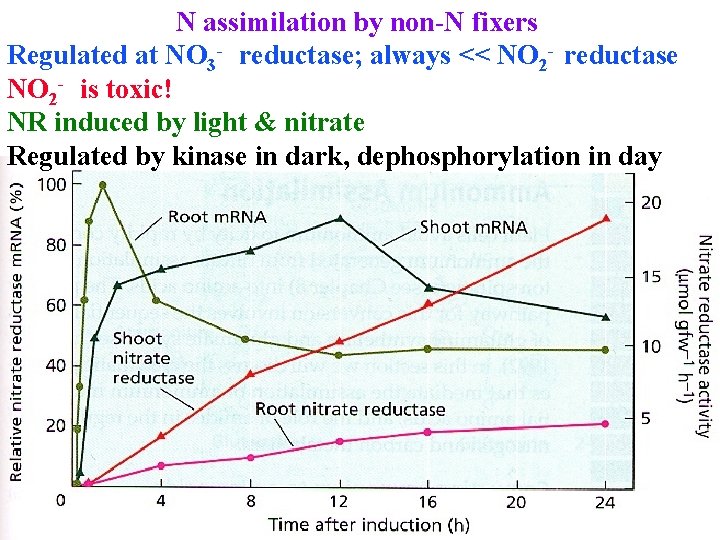
N assimilation by non-N fixers Regulated at NO 3 - reductase; always << NO 2 - reductase NO 2 - is toxic! NR induced by light & nitrate Regulated by kinase in dark, dephosphorylation in day
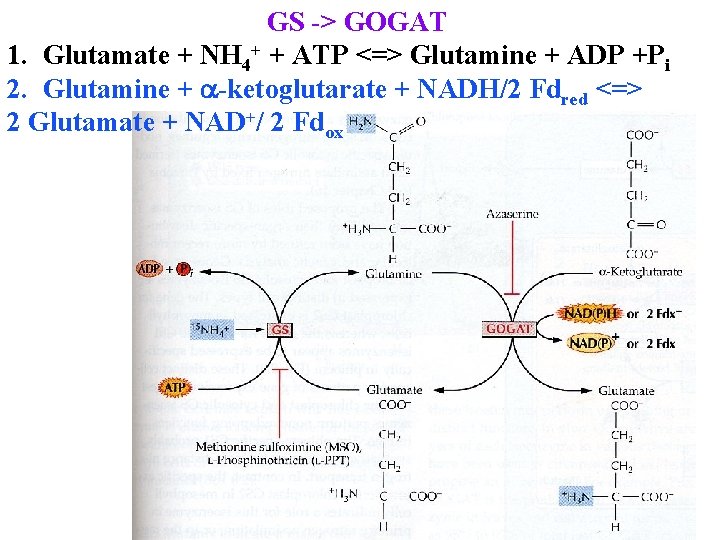
GS -> GOGAT 1. Glutamate + NH 4+ + ATP <=> Glutamine + ADP +Pi 2. Glutamine + a-ketoglutarate + NADH/2 Fdred <=> 2 Glutamate + NAD+/ 2 Fdox
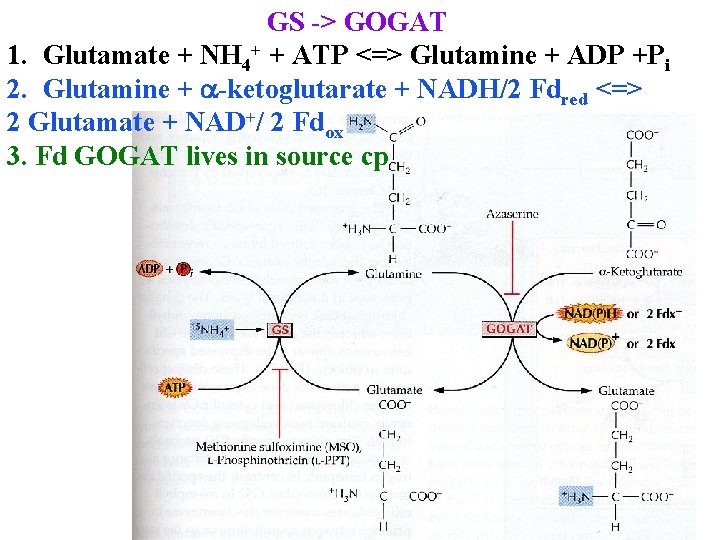
GS -> GOGAT 1. Glutamate + NH 4+ + ATP <=> Glutamine + ADP +Pi 2. Glutamine + a-ketoglutarate + NADH/2 Fdred <=> 2 Glutamate + NAD+/ 2 Fdox 3. Fd GOGAT lives in source cp
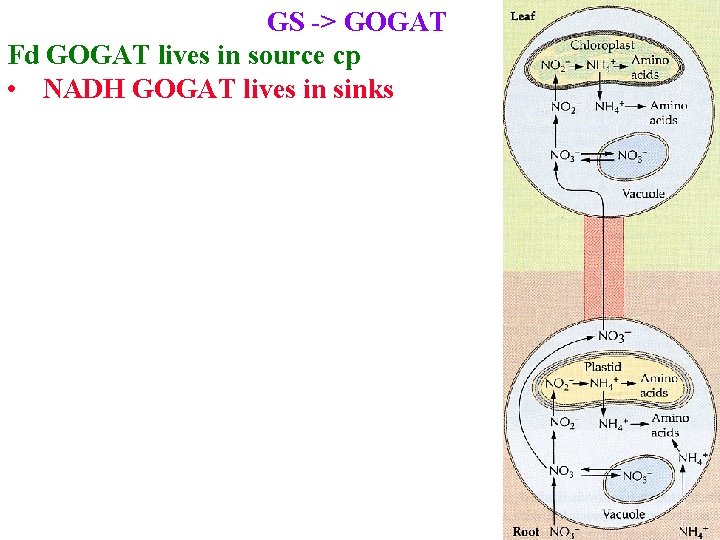
GS -> GOGAT Fd GOGAT lives in source cp • NADH GOGAT lives in sinks
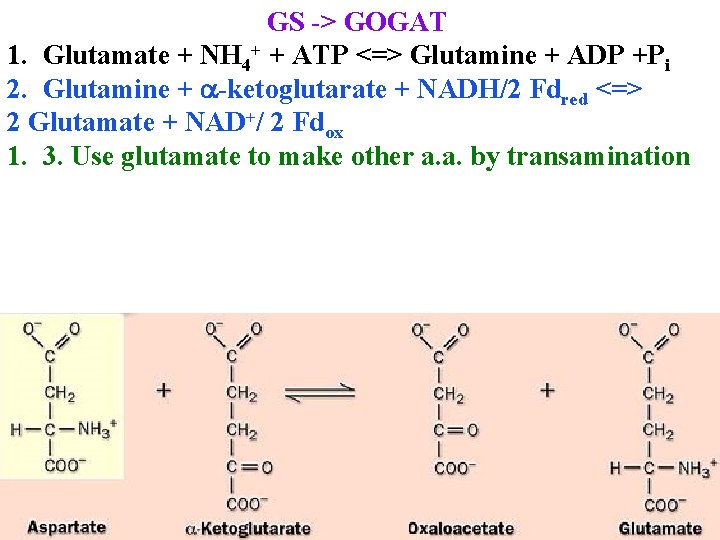
GS -> GOGAT 1. Glutamate + NH 4+ + ATP <=> Glutamine + ADP +Pi 2. Glutamine + a-ketoglutarate + NADH/2 Fdred <=> 2 Glutamate + NAD+/ 2 Fdox 1. 3. Use glutamate to make other a. a. by transamination
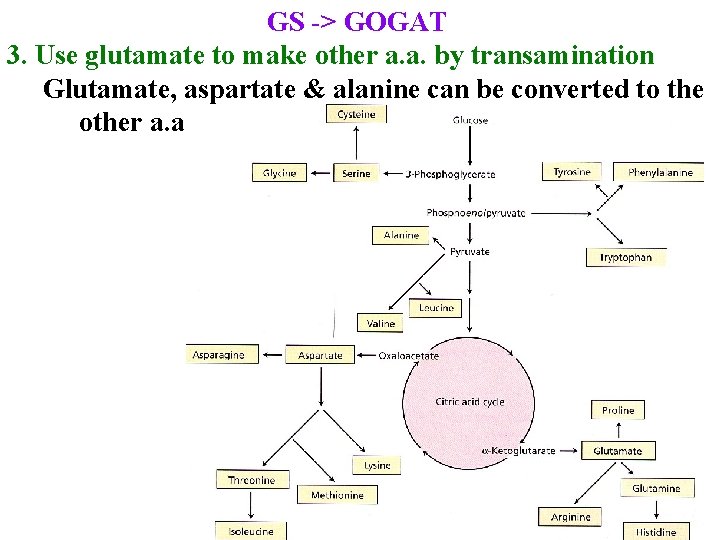
GS -> GOGAT 3. Use glutamate to make other a. a. by transamination Glutamate, aspartate & alanine can be converted to the other a. a.

N assimilation by N fixers Exclusively performed by prokaryotes Dramatically improve the growth of many plants
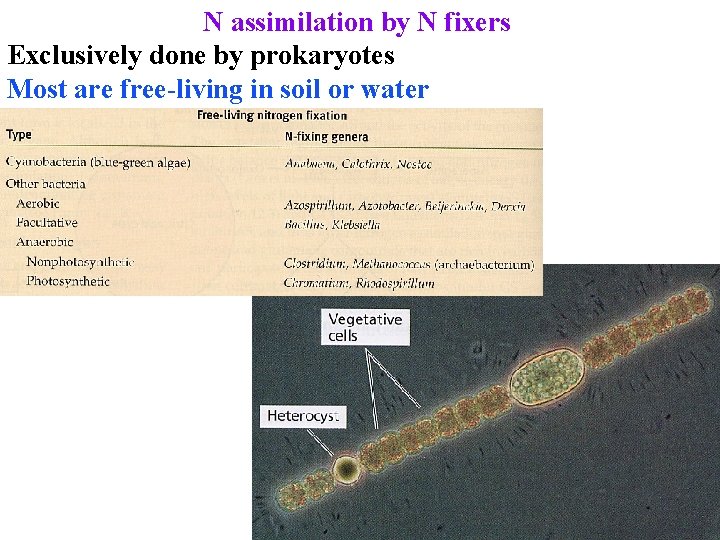
N assimilation by N fixers Exclusively done by prokaryotes Most are free-living in soil or water

N assimilation by N fixers Exclusively done by prokaryotes Most are free-living in soil or water Some form symbioses with plants

N assimilation by N fixers Exclusively done by prokaryotes Most are free-living in soil or water Some form symbioses with plants Legumes are best-known, but many others including mosses, ferns, lichens

N assimilation by N fixers Exclusively done by prokaryotes Most are free-living in soil or water Some form symbioses with plants Legumes are best-known, but many others including mosses, ferns, lichens Also have associations where N-fixers form films on leaves or roots and are fed by plant

N assimilation by N fixers Exclusively done by prokaryotes Also have associations where N-fixers form films on leaves or roots and are fed by plant All must form O 2 -free environment for nitrogenase
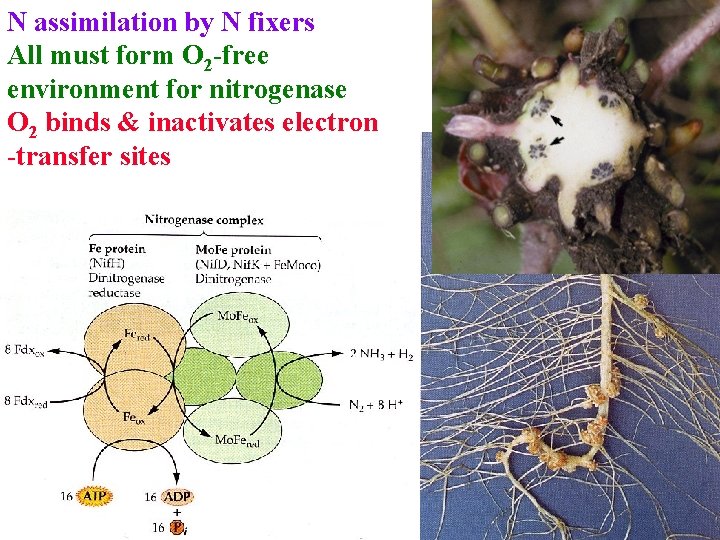
N assimilation by N fixers All must form O 2 -free environment for nitrogenase O 2 binds & inactivates electron -transfer sites
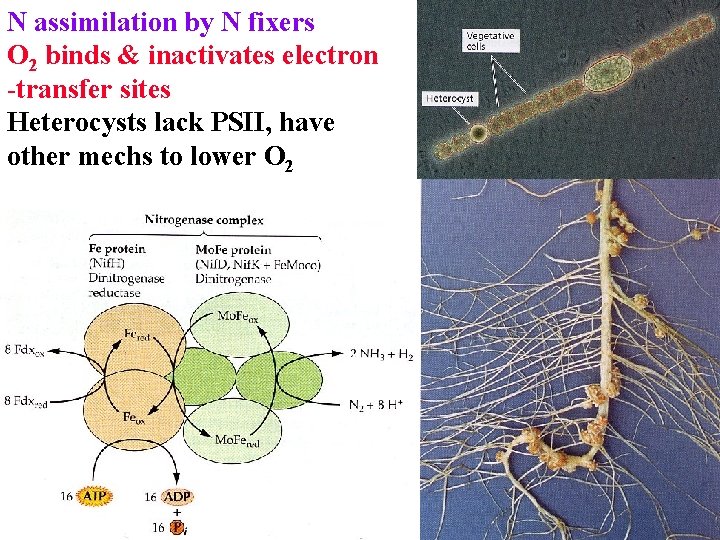
N assimilation by N fixers O 2 binds & inactivates electron -transfer sites Heterocysts lack PSII, have other mechs to lower O 2
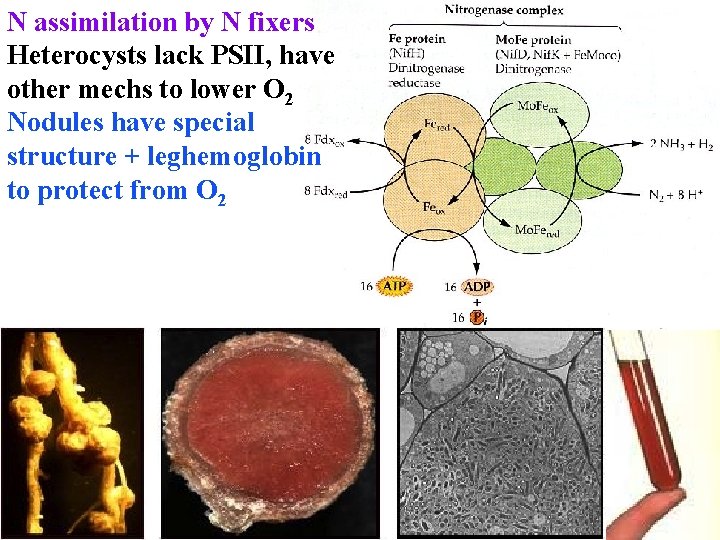
N assimilation by N fixers Heterocysts lack PSII, have other mechs to lower O 2 Nodules have special structure + leghemoglobin to protect from O 2
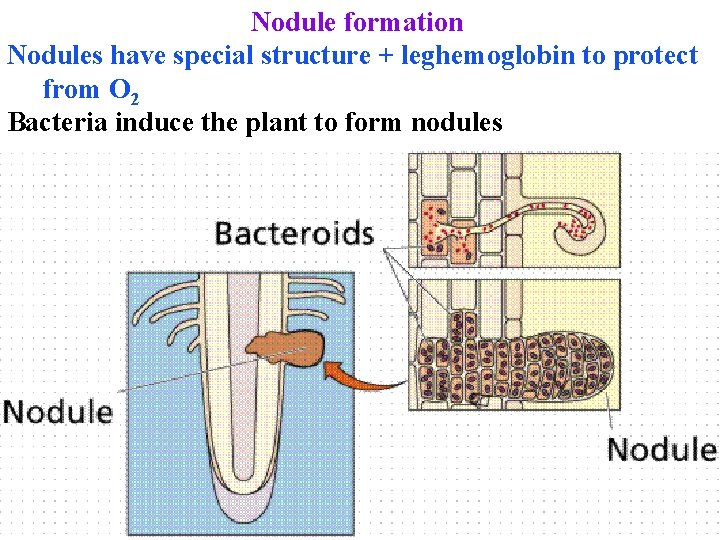
Nodule formation Nodules have special structure + leghemoglobin to protect from O 2 Bacteria induce the plant to form nodules
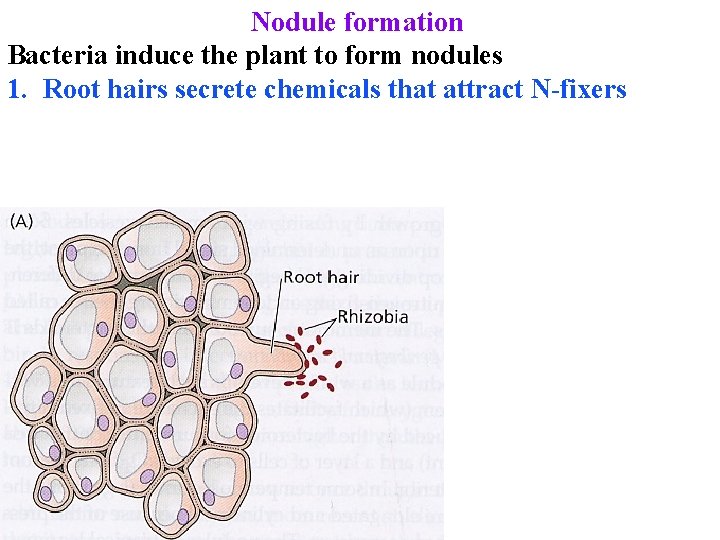
Nodule formation Bacteria induce the plant to form nodules 1. Root hairs secrete chemicals that attract N-fixers
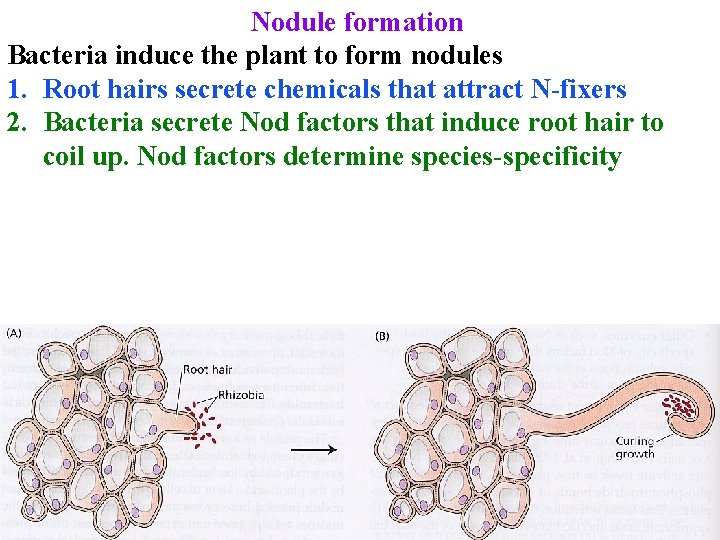
Nodule formation Bacteria induce the plant to form nodules 1. Root hairs secrete chemicals that attract N-fixers 2. Bacteria secrete Nod factors that induce root hair to coil up. Nod factors determine species-specificity
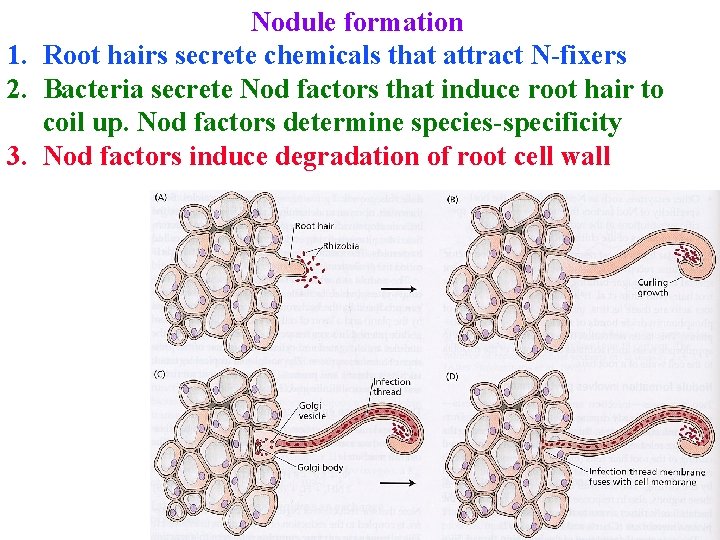
Nodule formation 1. Root hairs secrete chemicals that attract N-fixers 2. Bacteria secrete Nod factors that induce root hair to coil up. Nod factors determine species-specificity 3. Nod factors induce degradation of root cell wall
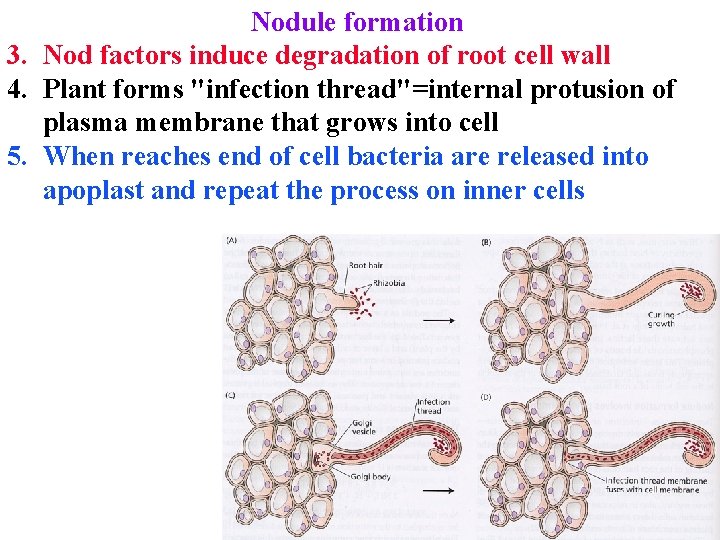
Nodule formation 3. Nod factors induce degradation of root cell wall 4. Plant forms "infection thread"=internal protusion of plasma membrane that grows into cell 5. When reaches end of cell bacteria are released into apoplast and repeat the process on inner cells
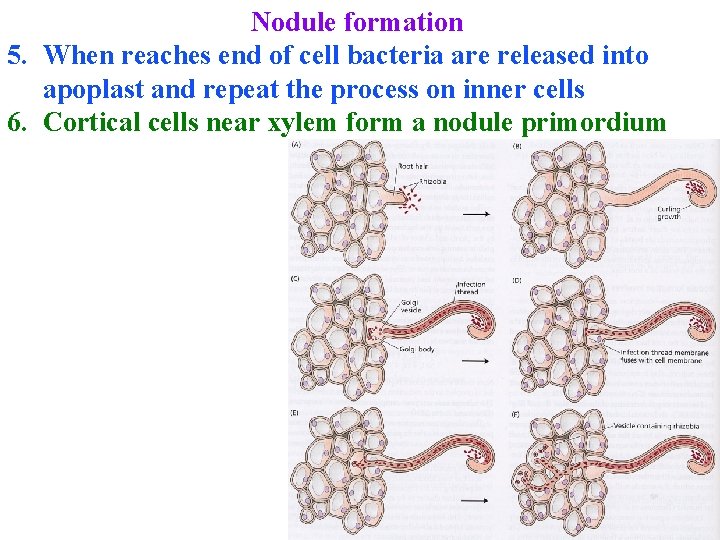
Nodule formation 5. When reaches end of cell bacteria are released into apoplast and repeat the process on inner cells 6. Cortical cells near xylem form a nodule primordium
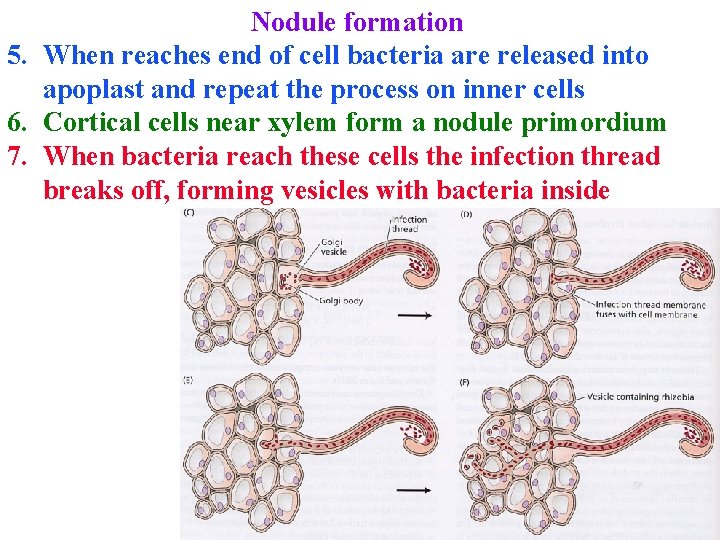
Nodule formation 5. When reaches end of cell bacteria are released into apoplast and repeat the process on inner cells 6. Cortical cells near xylem form a nodule primordium 7. When bacteria reach these cells the infection thread breaks off, forming vesicles with bacteria inside
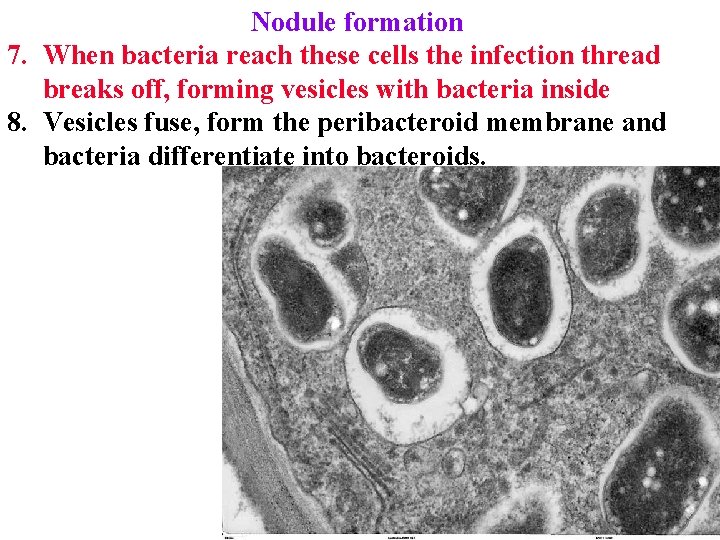
Nodule formation 7. When bacteria reach these cells the infection thread breaks off, forming vesicles with bacteria inside 8. Vesicles fuse, form the peribacteroid membrane and bacteria differentiate into bacteroids.
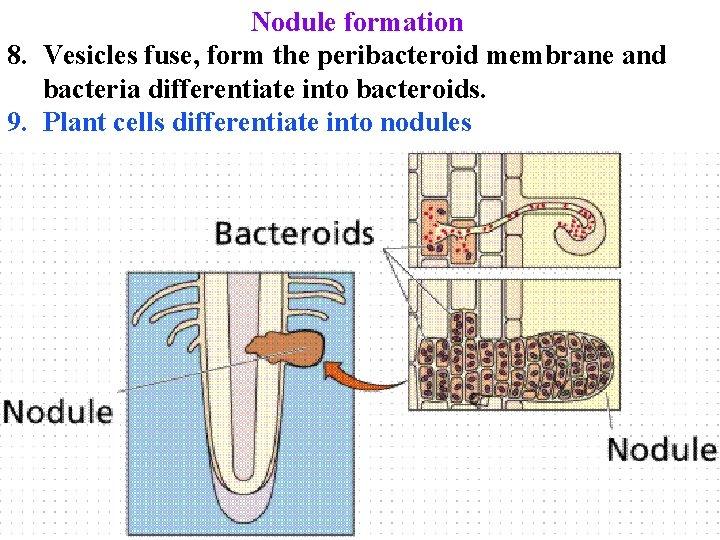
Nodule formation 8. Vesicles fuse, form the peribacteroid membrane and bacteria differentiate into bacteroids. 9. Plant cells differentiate into nodules
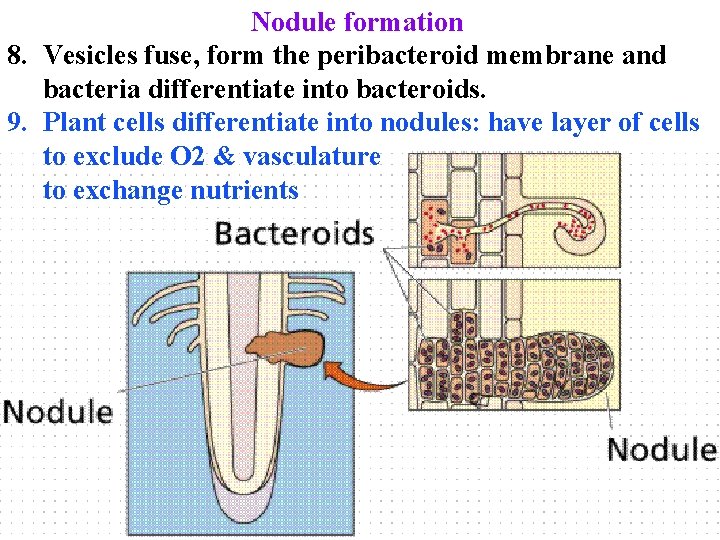
Nodule formation 8. Vesicles fuse, form the peribacteroid membrane and bacteria differentiate into bacteroids. 9. Plant cells differentiate into nodules: have layer of cells to exclude O 2 & vasculature to exchange nutrients
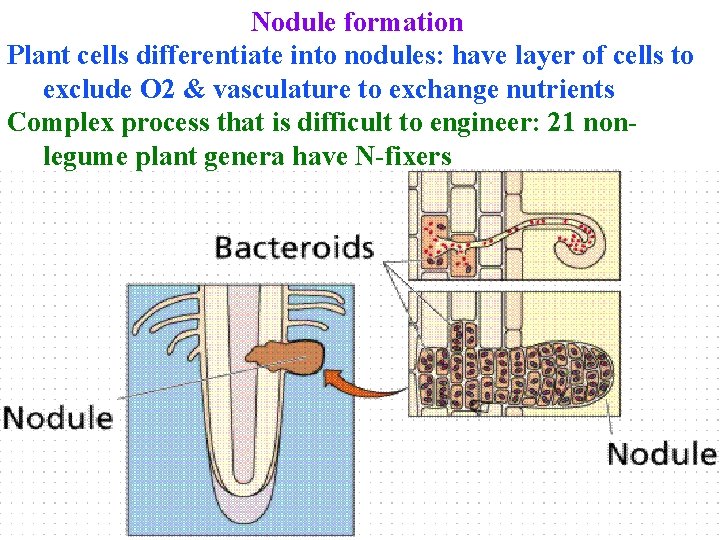
Nodule formation Plant cells differentiate into nodules: have layer of cells to exclude O 2 & vasculature to exchange nutrients Complex process that is difficult to engineer: 21 nonlegume plant genera have N-fixers
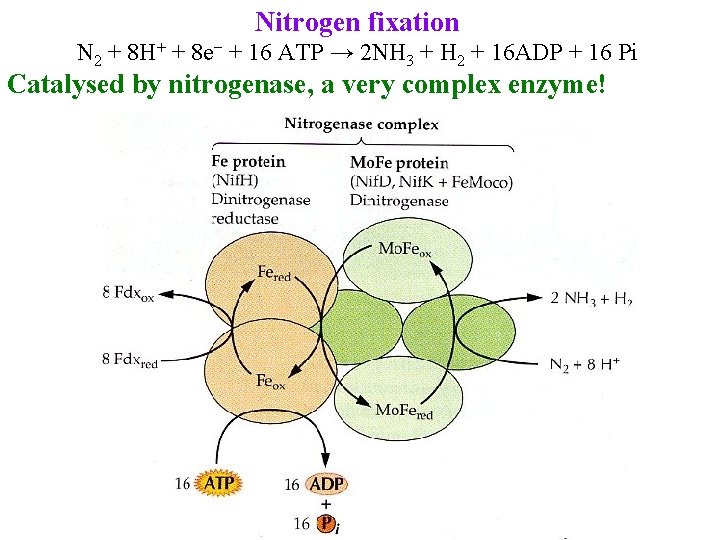
Nitrogen fixation N 2 + 8 H+ + 8 e− + 16 ATP → 2 NH 3 + H 2 + 16 ADP + 16 Pi Catalysed by nitrogenase, a very complex enzyme!

Nitrogen fixation N 2 + 8 H+ + 8 e− + 16 ATP → 2 NH 3 + H 2 + 16 ADP + 16 Pi Catalysed by nitrogenase, a very complex enzyme! Also catalyzes many other reactions Usually assayed by acetylene reduction
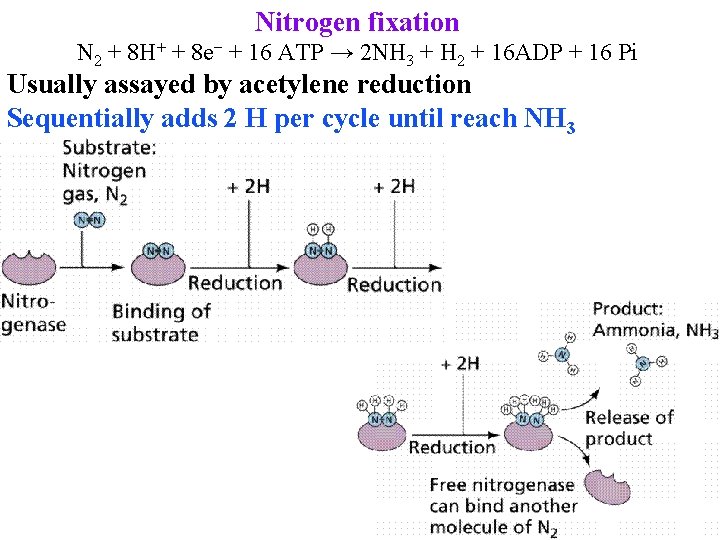
Nitrogen fixation N 2 + 8 H+ + 8 e− + 16 ATP → 2 NH 3 + H 2 + 16 ADP + 16 Pi Usually assayed by acetylene reduction Sequentially adds 2 H per cycle until reach NH 3
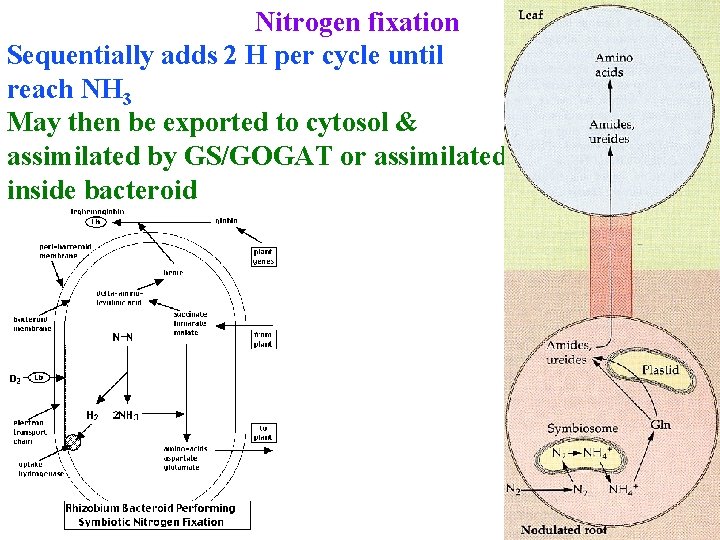
Nitrogen fixation Sequentially adds 2 H per cycle until reach NH 3 May then be exported to cytosol & assimilated by GS/GOGAT or assimilated inside bacteroid
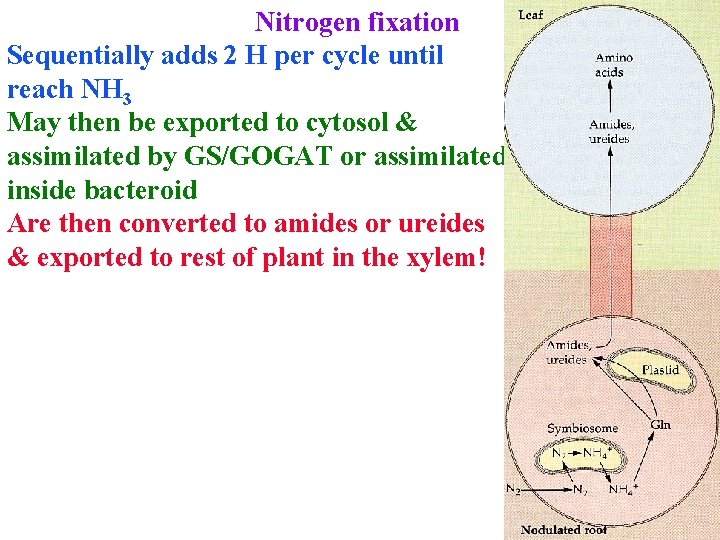
Nitrogen fixation Sequentially adds 2 H per cycle until reach NH 3 May then be exported to cytosol & assimilated by GS/GOGAT or assimilated inside bacteroid Are then converted to amides or ureides & exported to rest of plant in the xylem!
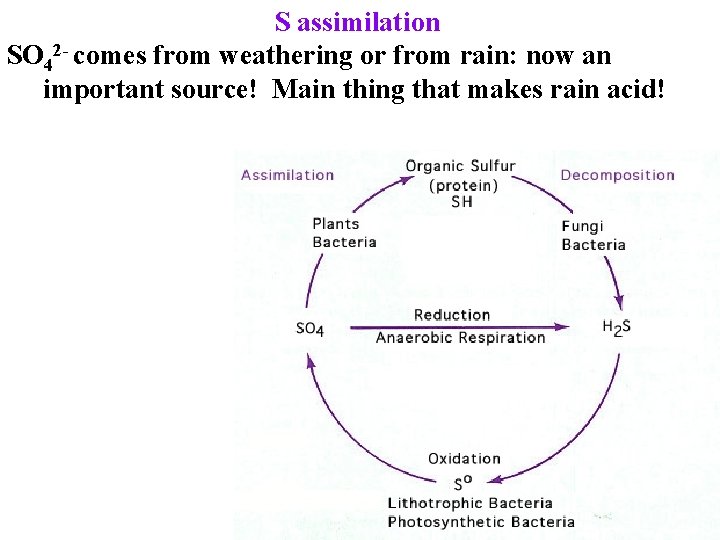
S assimilation SO 42 - comes from weathering or from rain: now an important source! Main thing that makes rain acid!
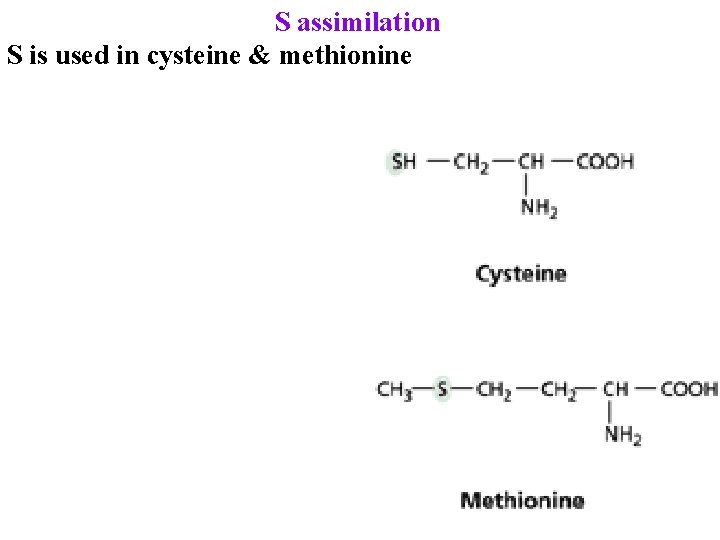
S assimilation S is used in cysteine & methionine
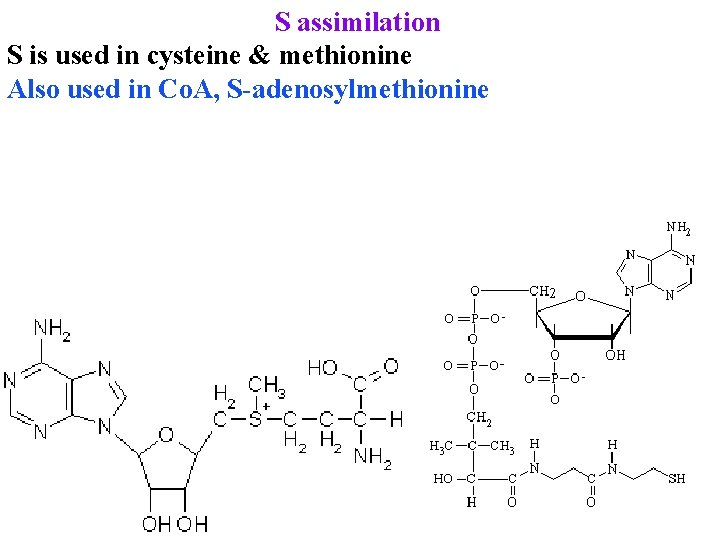
S assimilation S is used in cysteine & methionine Also used in Co. A, S-adenosylmethionine
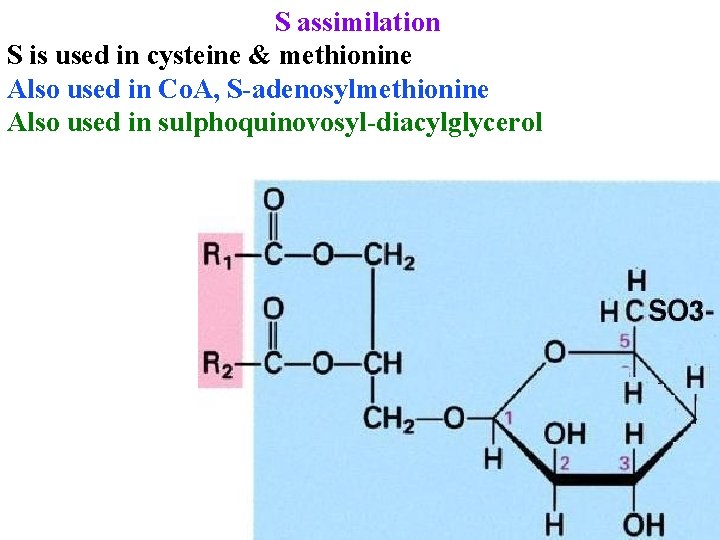
S assimilation S is used in cysteine & methionine Also used in Co. A, S-adenosylmethionine Also used in sulphoquinovosyl-diacylglycerol
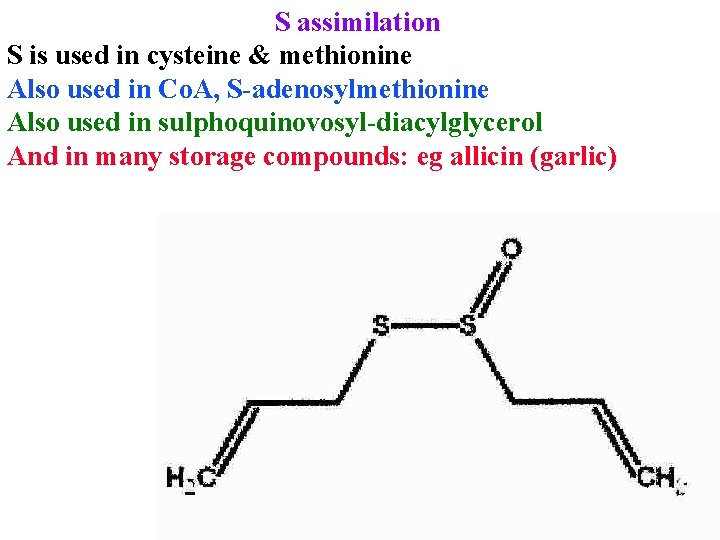
S assimilation S is used in cysteine & methionine Also used in Co. A, S-adenosylmethionine Also used in sulphoquinovosyl-diacylglycerol And in many storage compounds: eg allicin (garlic)
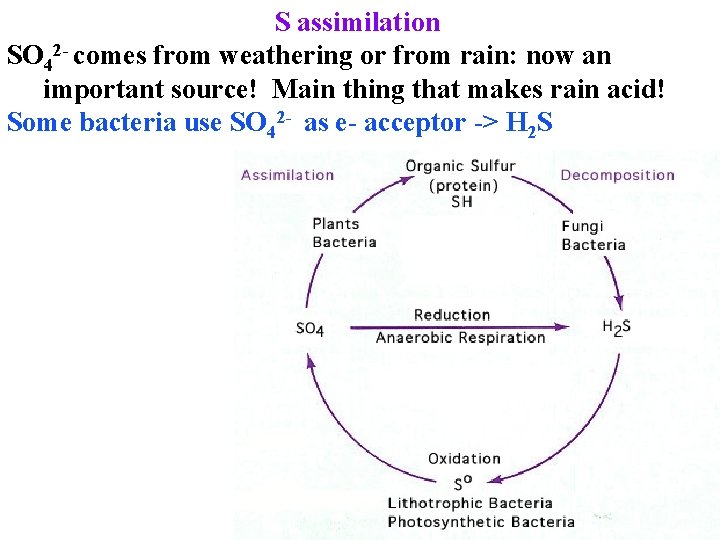
S assimilation SO 42 - comes from weathering or from rain: now an important source! Main thing that makes rain acid! Some bacteria use SO 42 - as e- acceptor -> H 2 S
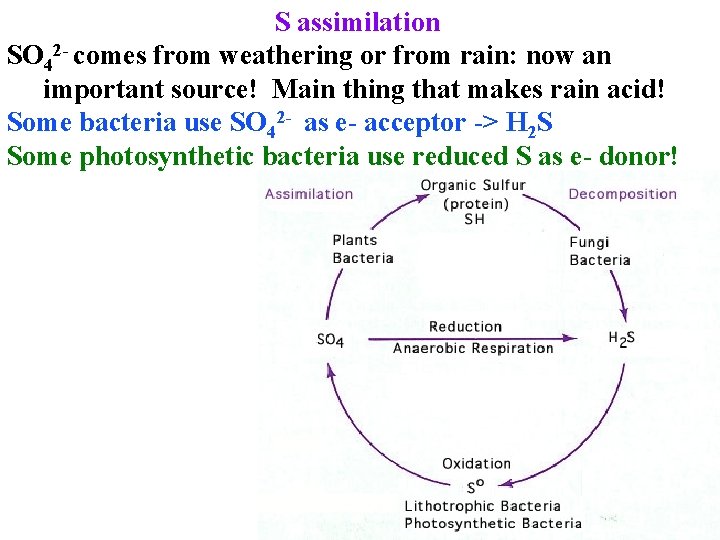
S assimilation SO 42 - comes from weathering or from rain: now an important source! Main thing that makes rain acid! Some bacteria use SO 42 - as e- acceptor -> H 2 S Some photosynthetic bacteria use reduced S as e- donor!

S assimilation SO 42 - comes from weathering or from rain: now an important source! Main thing that makes rain acid! Some bacteria use SO 42 - as e- acceptor -> H 2 S Some photosynthetic bacteria use reduced S as e- donor! Now that acid rain has declined in N. Europe Brassica & wheat need S in many places
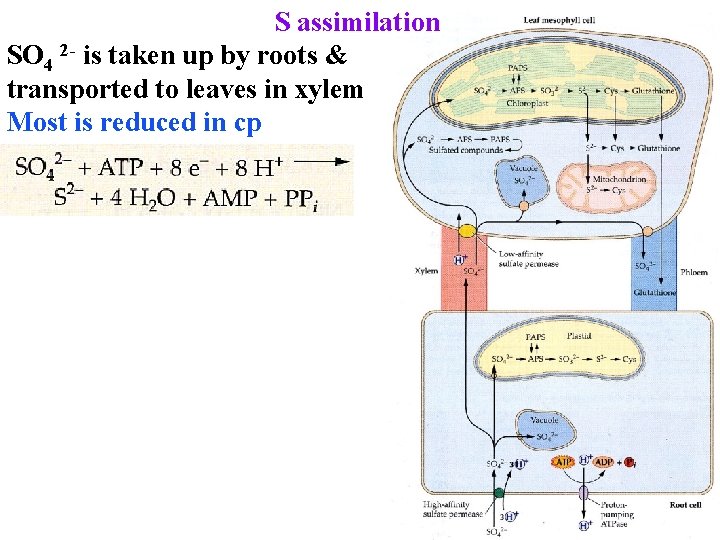
S assimilation SO 4 2 - is taken up by roots & transported to leaves in xylem Most is reduced in cp
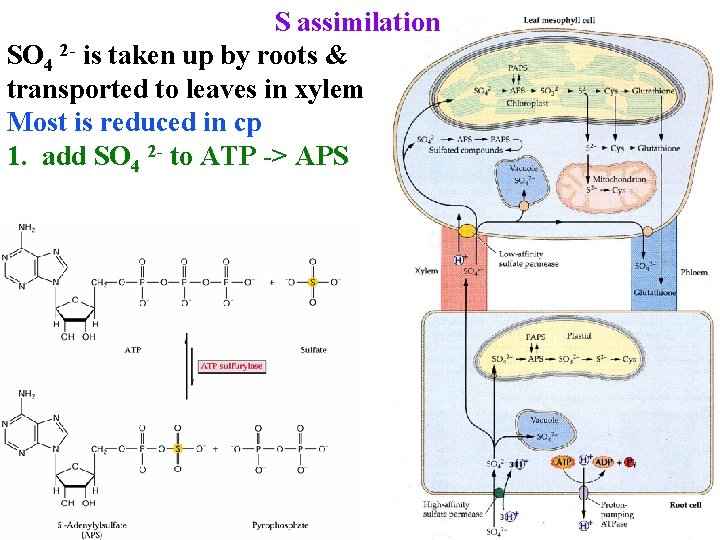
S assimilation SO 4 2 - is taken up by roots & transported to leaves in xylem Most is reduced in cp 1. add SO 4 2 - to ATP -> APS
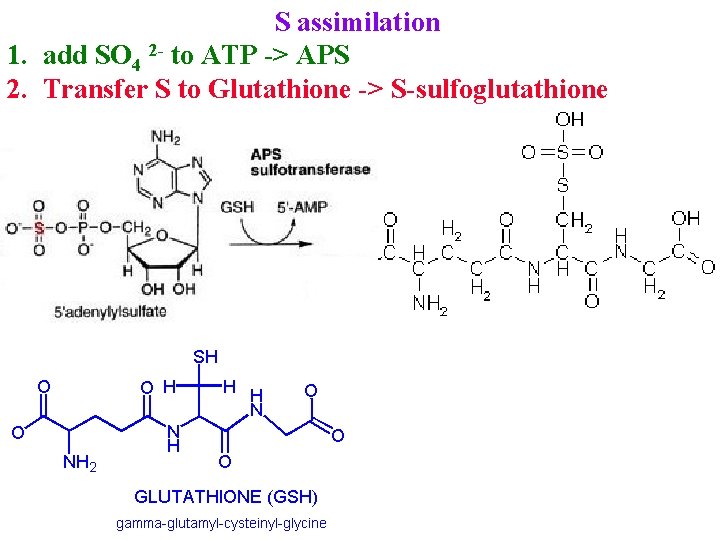
S assimilation 1. add SO 4 2 - to ATP -> APS 2. Transfer S to Glutathione -> S-sulfoglutathione
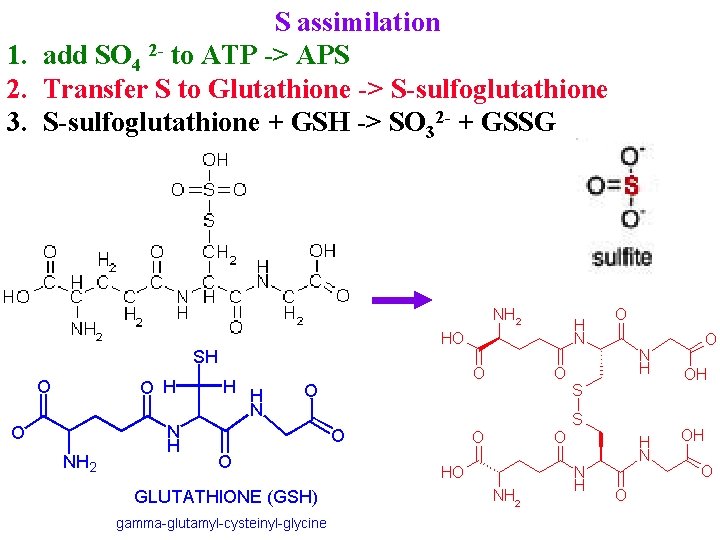
S assimilation 1. add SO 4 2 - to ATP -> APS 2. Transfer S to Glutathione -> S-sulfoglutathione 3. S-sulfoglutathione + GSH -> SO 32 - + GSSG
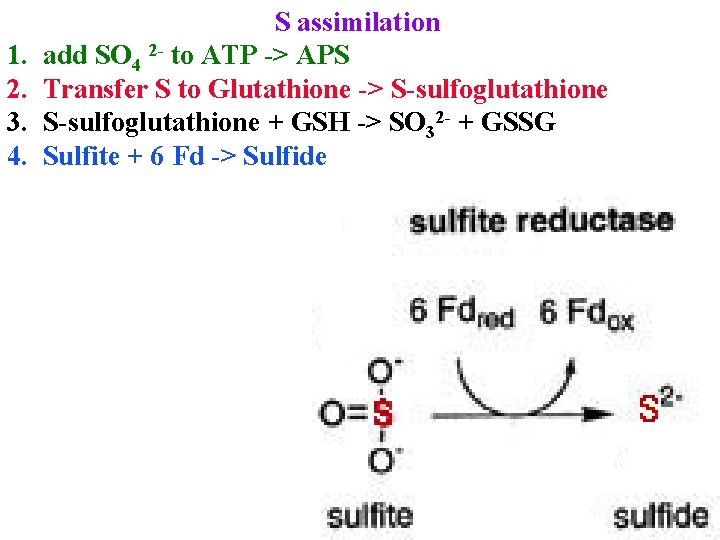
1. 2. 3. 4. S assimilation add SO 4 2 - to ATP -> APS Transfer S to Glutathione -> S-sulfoglutathione + GSH -> SO 32 - + GSSG Sulfite + 6 Fd -> Sulfide
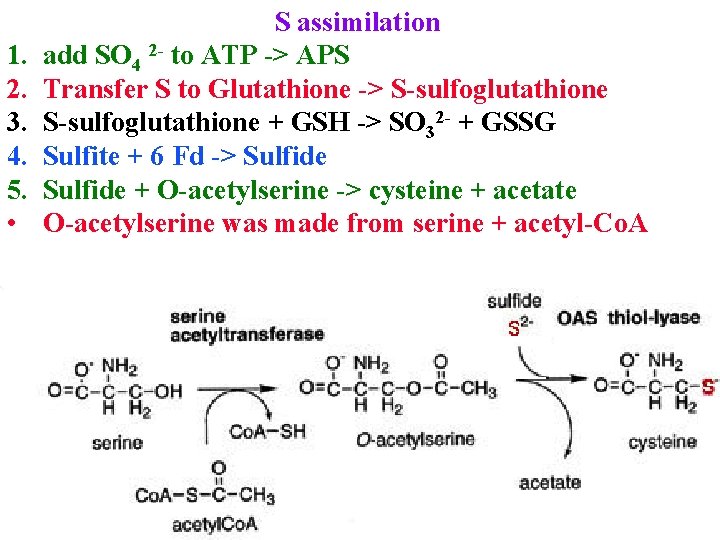
1. 2. 3. 4. 5. • S assimilation add SO 4 2 - to ATP -> APS Transfer S to Glutathione -> S-sulfoglutathione + GSH -> SO 32 - + GSSG Sulfite + 6 Fd -> Sulfide + O-acetylserine -> cysteine + acetate O-acetylserine was made from serine + acetyl-Co. A
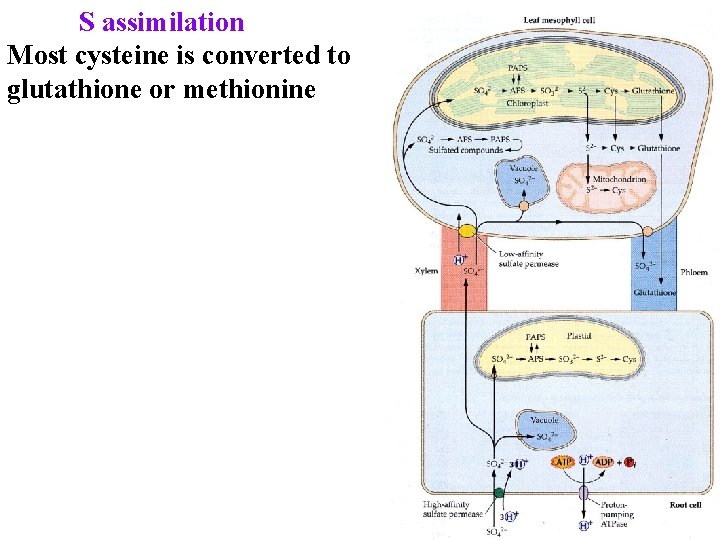
S assimilation Most cysteine is converted to glutathione or methionine
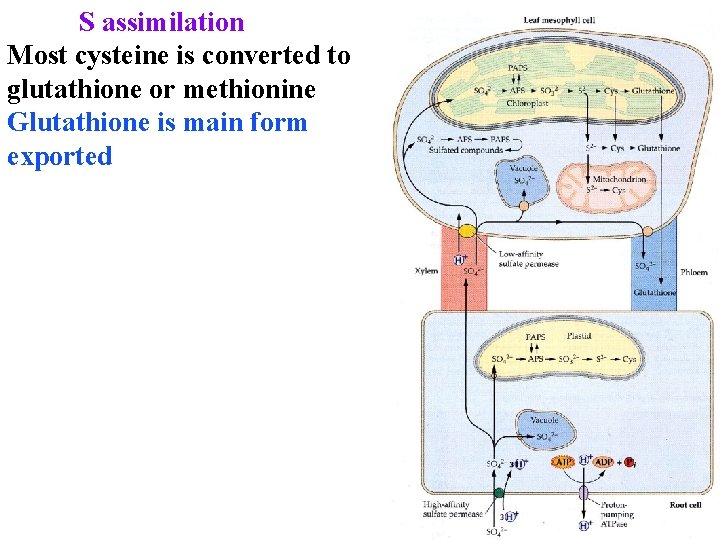
S assimilation Most cysteine is converted to glutathione or methionine Glutathione is main form exported
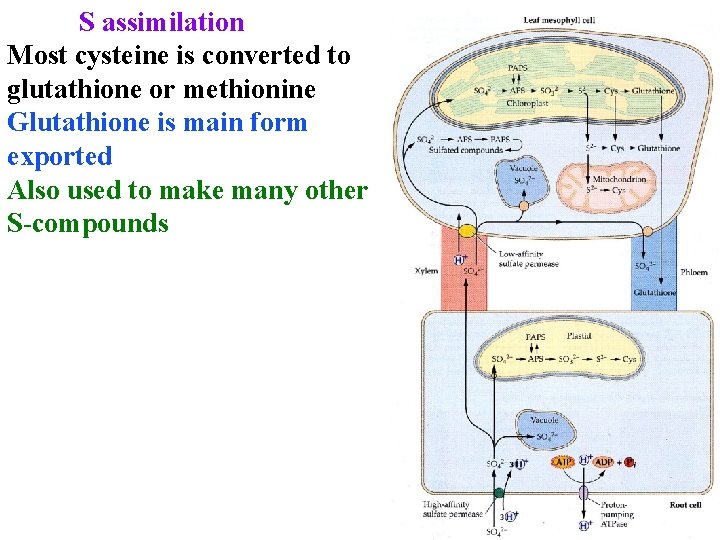
S assimilation Most cysteine is converted to glutathione or methionine Glutathione is main form exported Also used to make many other S-compounds
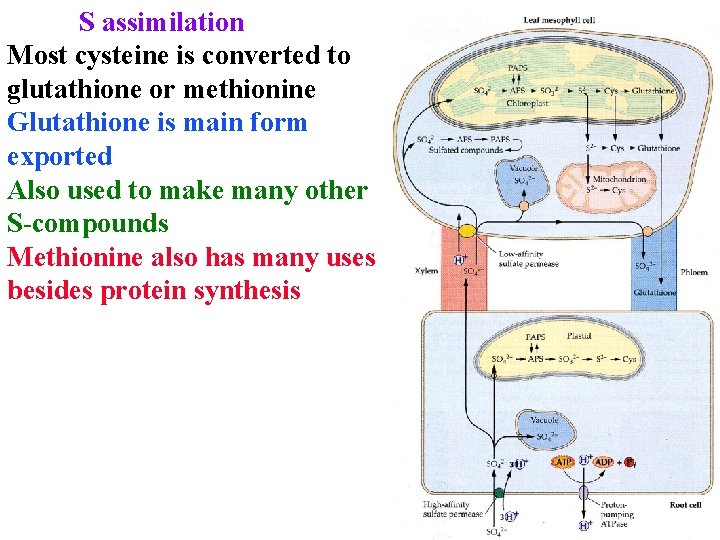
S assimilation Most cysteine is converted to glutathione or methionine Glutathione is main form exported Also used to make many other S-compounds Methionine also has many uses besides protein synthesis
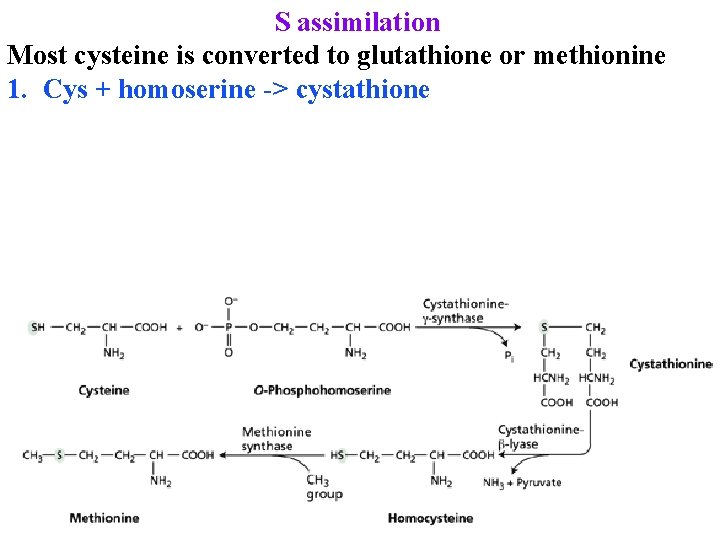
S assimilation Most cysteine is converted to glutathione or methionine 1. Cys + homoserine -> cystathione
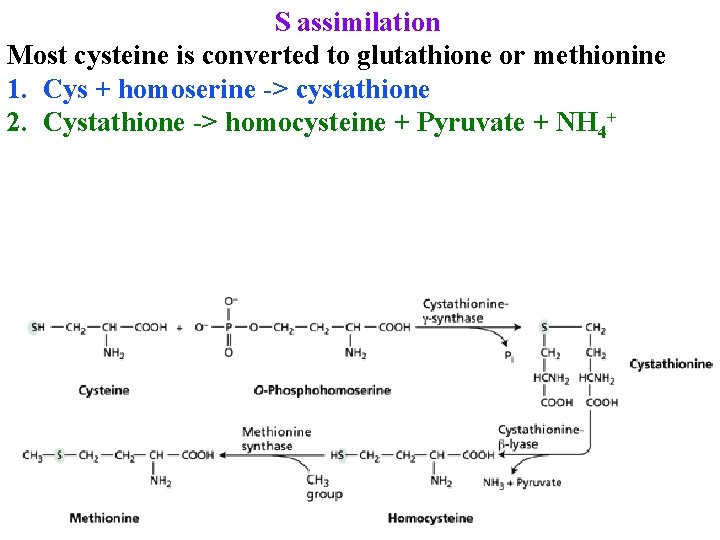
S assimilation Most cysteine is converted to glutathione or methionine 1. Cys + homoserine -> cystathione 2. Cystathione -> homocysteine + Pyruvate + NH 4+
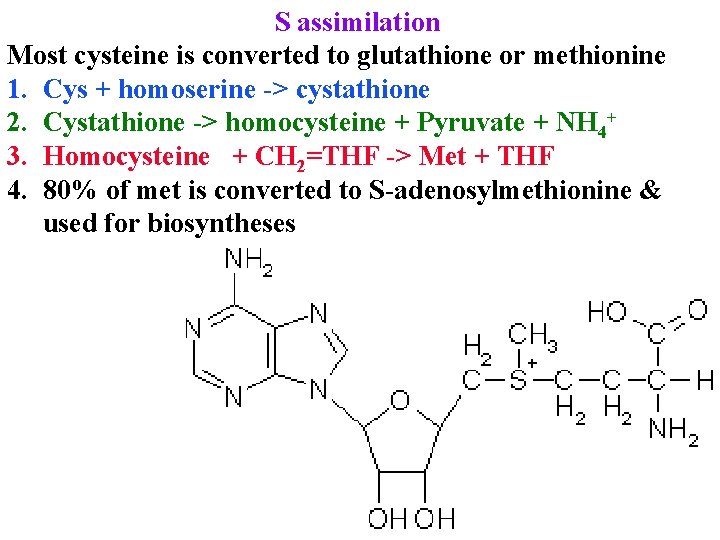
S assimilation Most cysteine is converted to glutathione or methionine 1. Cys + homoserine -> cystathione 2. Cystathione -> homocysteine + Pyruvate + NH 4+ 3. Homocysteine + CH 2=THF -> Met + THF 4. 80% of met is converted to S-adenosylmethionine & used for biosyntheses
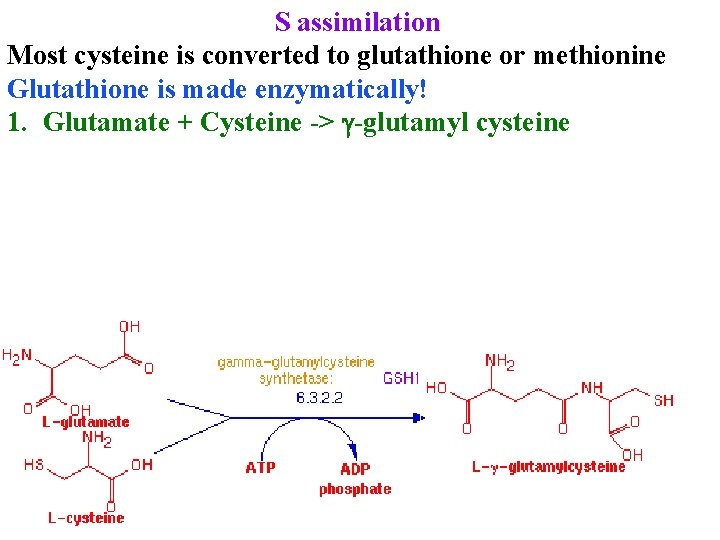
S assimilation Most cysteine is converted to glutathione or methionine Glutathione is made enzymatically! 1. Glutamate + Cysteine -> g-glutamyl cysteine
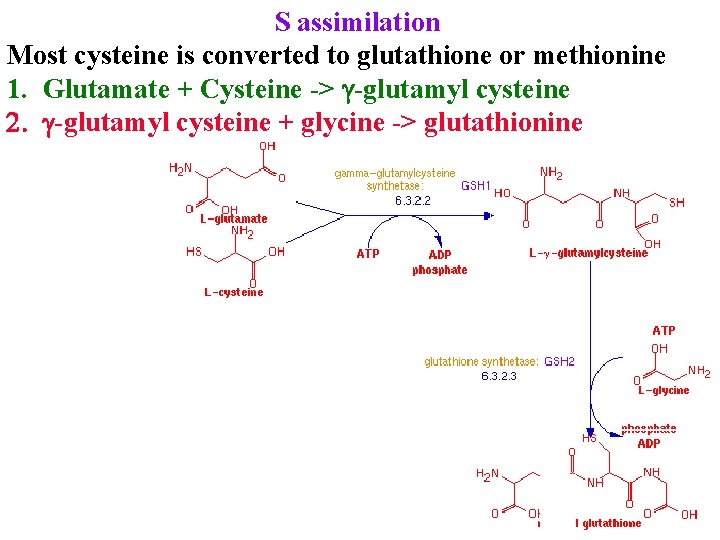
S assimilation Most cysteine is converted to glutathione or methionine 1. Glutamate + Cysteine -> g-glutamyl cysteine 2. g-glutamyl cysteine + glycine -> glutathionine
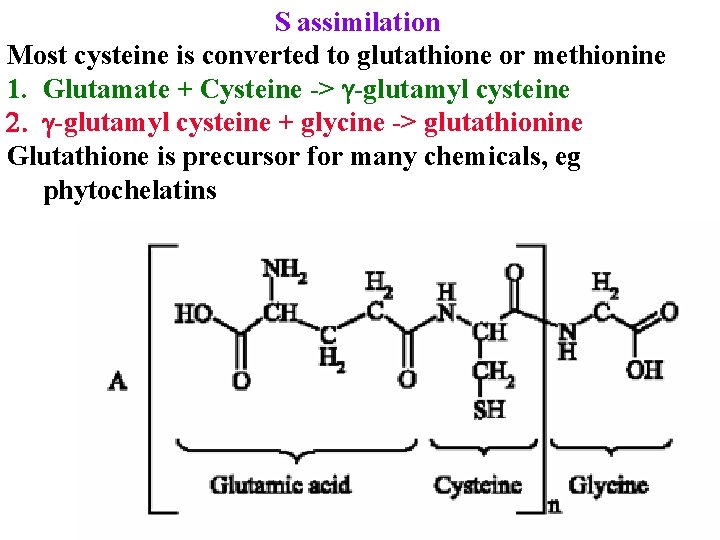
S assimilation Most cysteine is converted to glutathione or methionine 1. Glutamate + Cysteine -> g-glutamyl cysteine 2. g-glutamyl cysteine + glycine -> glutathionine Glutathione is precursor for many chemicals, eg phytochelatins

S assimilation Most cysteine is converted to glutathione or methionine 1. Glutamate + Cysteine -> g-glutamyl cysteine 2. g-glutamyl cysteine + glycine -> glutathionine Glutathione is precursor for many chemicals, eg phytochelatins SAM & glutathione are also precursors for many cell wall components
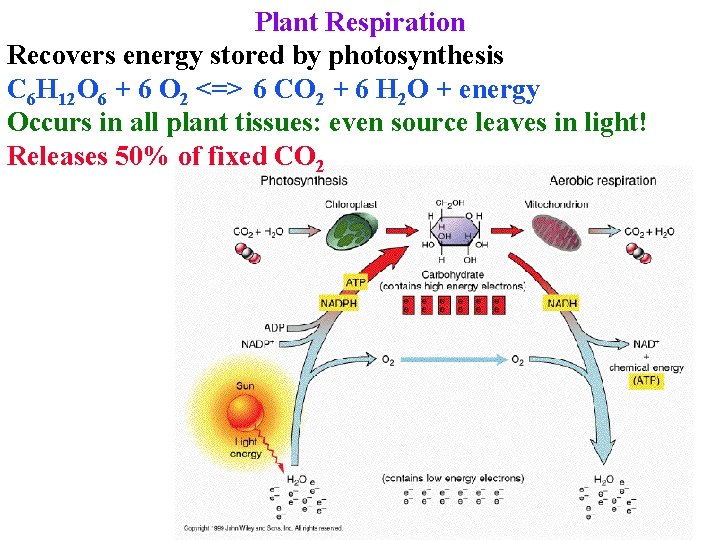
Plant Respiration Recovers energy stored by photosynthesis C 6 H 12 O 6 + 6 O 2 <=> 6 CO 2 + 6 H 2 O + energy Occurs in all plant tissues: even source leaves in light! Releases 50% of fixed CO 2

Plant Respiration Releases 50% of fixed CO 2 Provides energy for all sinks, source leaves at night & helps source during day!
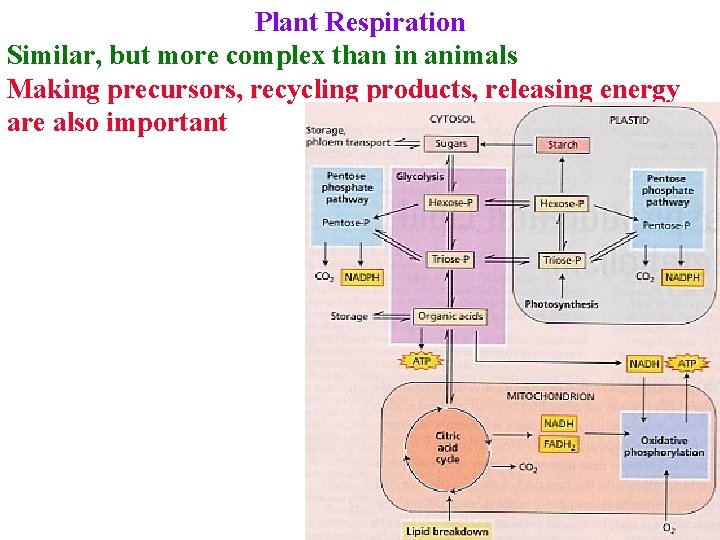
Plant Respiration Similar, but more complex than in animals Making precursors, recycling products, releasing energy are also important
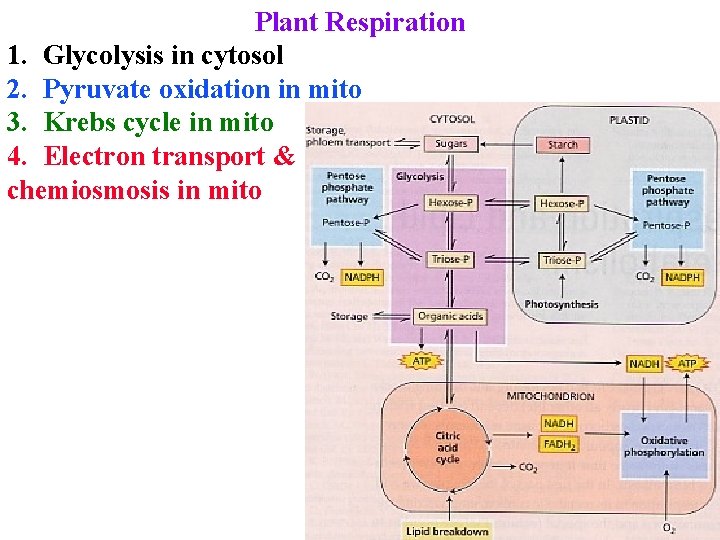
Plant Respiration 1. Glycolysis in cytosol 2. Pyruvate oxidation in mito 3. Krebs cycle in mito 4. Electron transport & chemiosmosis in mito
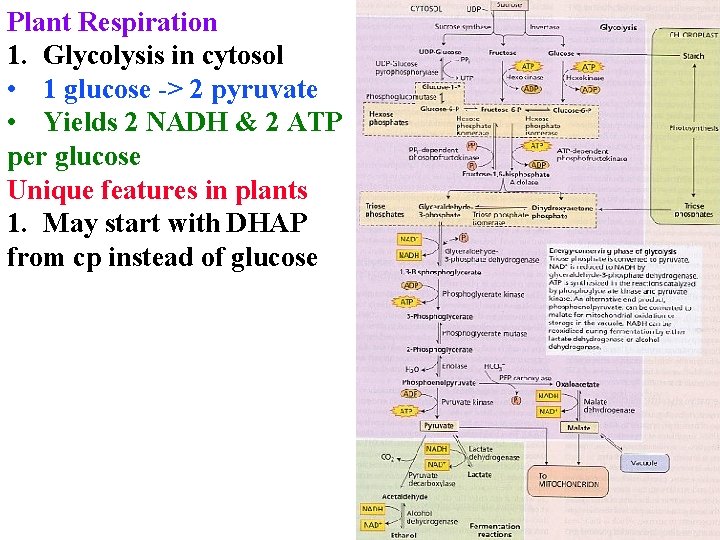
Plant Respiration 1. Glycolysis in cytosol • 1 glucose -> 2 pyruvate • Yields 2 NADH & 2 ATP per glucose Unique features in plants 1. May start with DHAP from cp instead of glucose
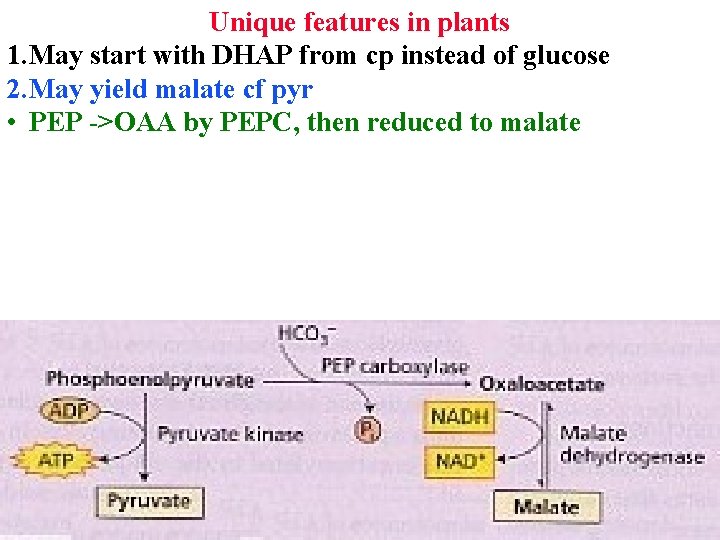
Unique features in plants 1. May start with DHAP from cp instead of glucose 2. May yield malate cf pyr • PEP ->OAA by PEPC, then reduced to malate
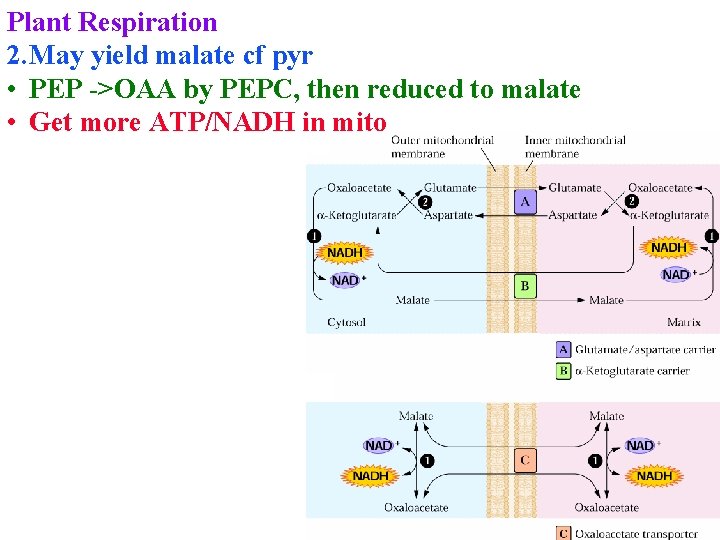
Plant Respiration 2. May yield malate cf pyr • PEP ->OAA by PEPC, then reduced to malate • Get more ATP/NADH in mito
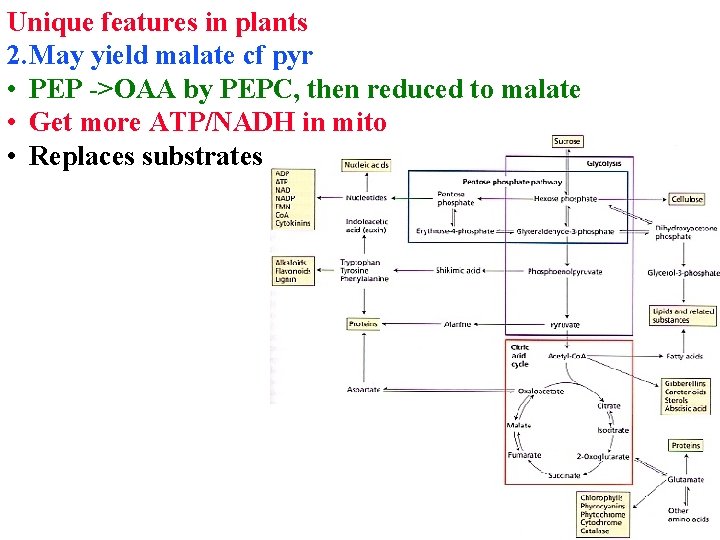
Unique features in plants 2. May yield malate cf pyr • PEP ->OAA by PEPC, then reduced to malate • Get more ATP/NADH in mito • Replaces substrates
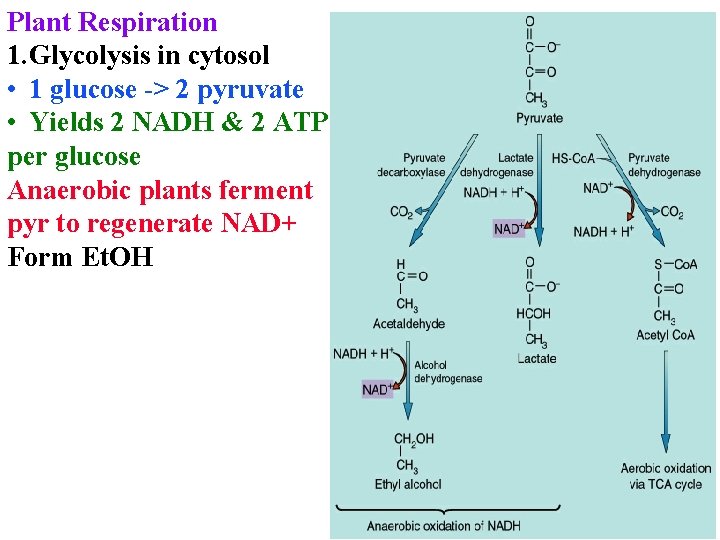
Plant Respiration 1. Glycolysis in cytosol • 1 glucose -> 2 pyruvate • Yields 2 NADH & 2 ATP per glucose Anaerobic plants ferment pyr to regenerate NAD+ Form Et. OH
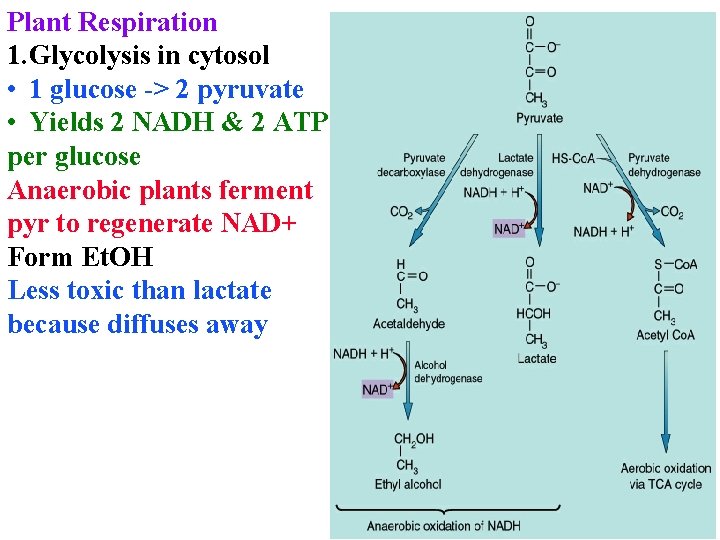
Plant Respiration 1. Glycolysis in cytosol • 1 glucose -> 2 pyruvate • Yields 2 NADH & 2 ATP per glucose Anaerobic plants ferment pyr to regenerate NAD+ Form Et. OH Less toxic than lactate because diffuses away
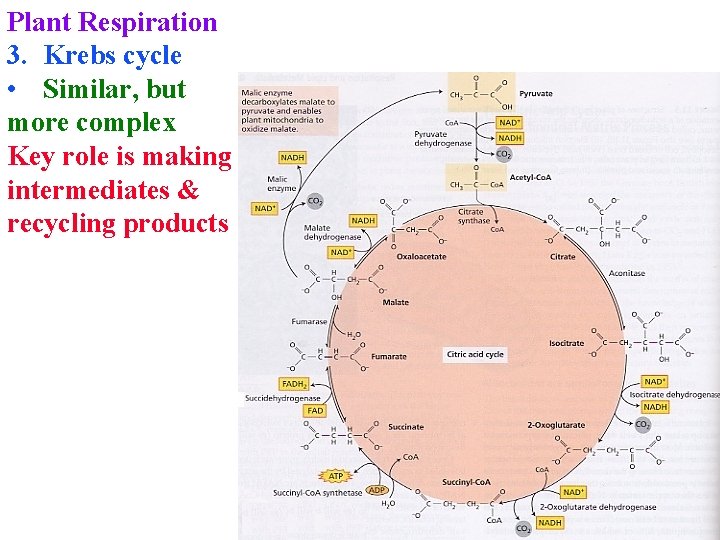
Plant Respiration 3. Krebs cycle • Similar, but more complex Key role is making intermediates & recycling products
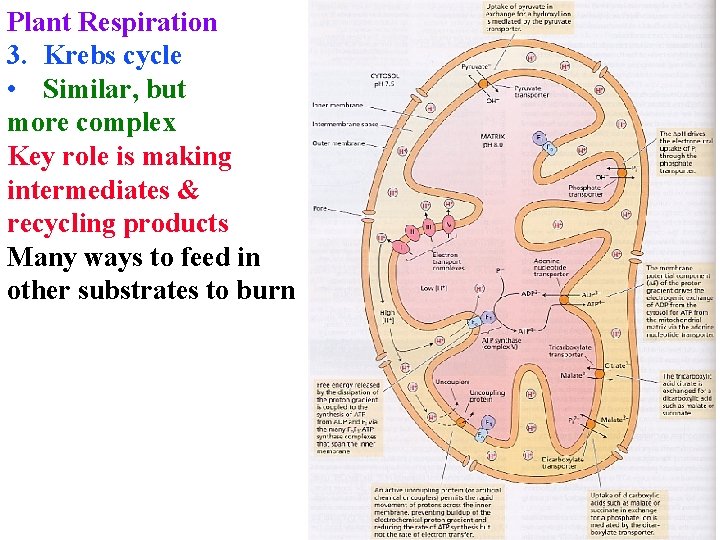
Plant Respiration 3. Krebs cycle • Similar, but more complex Key role is making intermediates & recycling products Many ways to feed in other substrates to burn
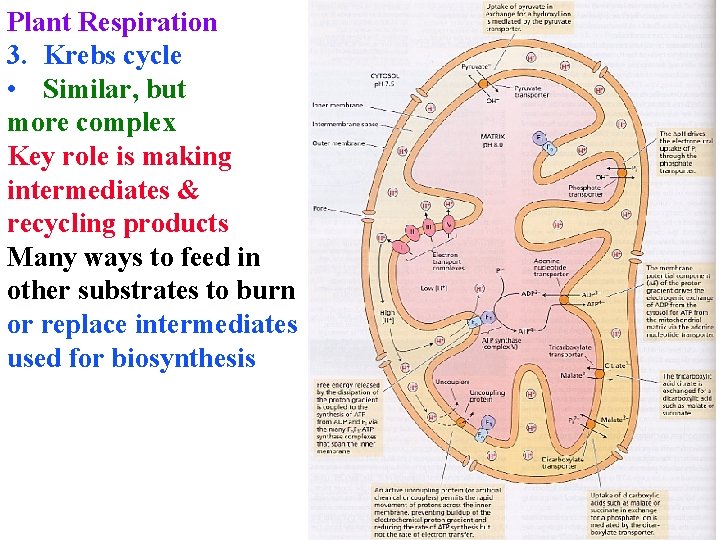
Plant Respiration 3. Krebs cycle • Similar, but more complex Key role is making intermediates & recycling products Many ways to feed in other substrates to burn or replace intermediates used for biosynthesis
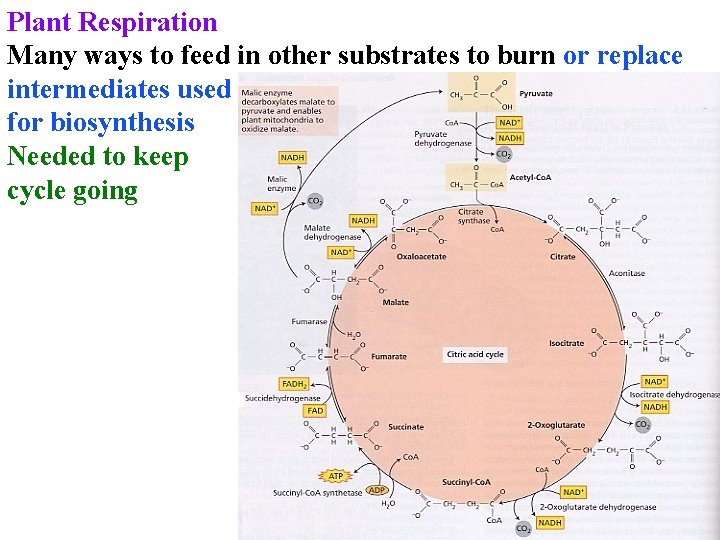
Plant Respiration Many ways to feed in other substrates to burn or replace intermediates used for biosynthesis Needed to keep cycle going
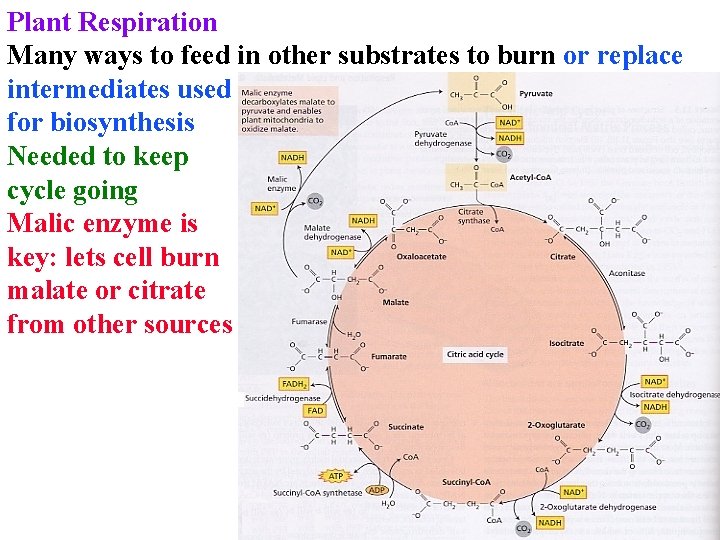
Plant Respiration Many ways to feed in other substrates to burn or replace intermediates used for biosynthesis Needed to keep cycle going Malic enzyme is key: lets cell burn malate or citrate from other sources
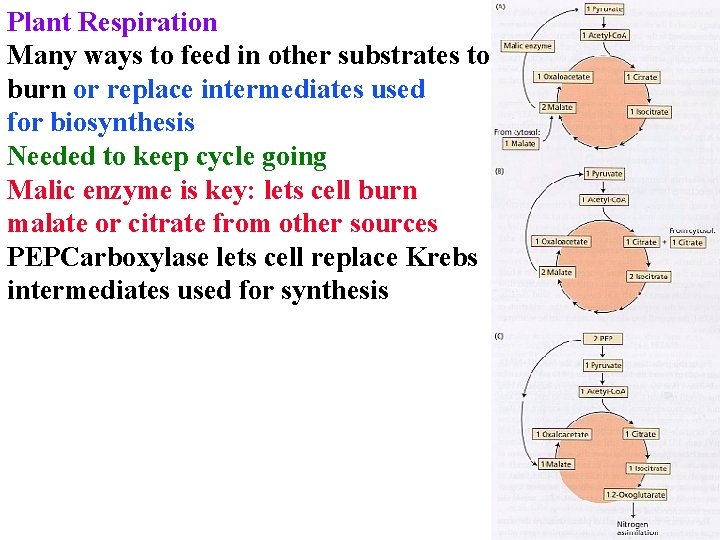
Plant Respiration Many ways to feed in other substrates to burn or replace intermediates used for biosynthesis Needed to keep cycle going Malic enzyme is key: lets cell burn malate or citrate from other sources PEPCarboxylase lets cell replace Krebs intermediates used for synthesis
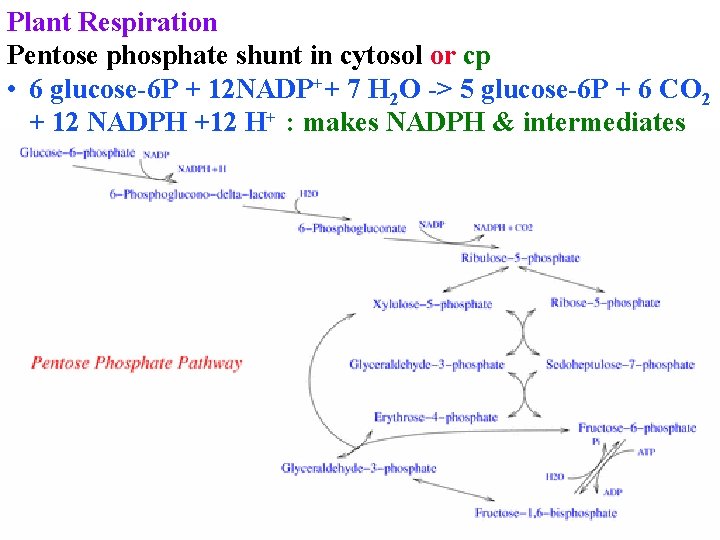
Plant Respiration Pentose phosphate shunt in cytosol or cp • 6 glucose-6 P + 12 NADP++ 7 H 2 O -> 5 glucose-6 P + 6 CO 2 + 12 NADPH +12 H+ : makes NADPH & intermediates
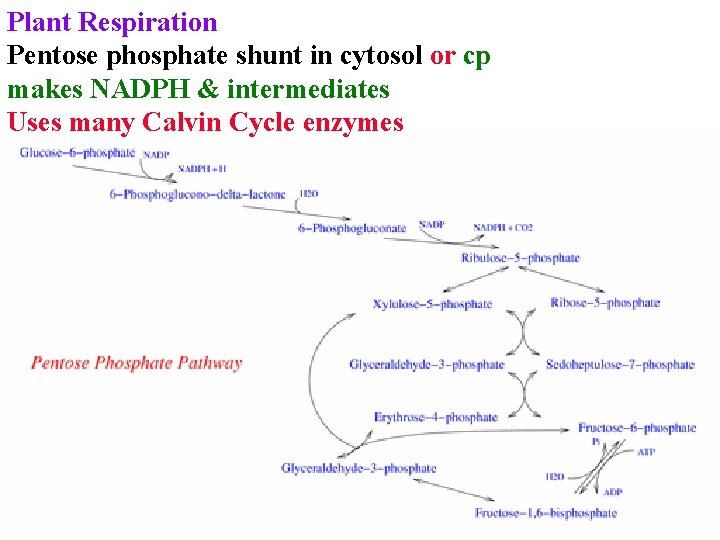
Plant Respiration Pentose phosphate shunt in cytosol or cp makes NADPH & intermediates Uses many Calvin Cycle enzymes
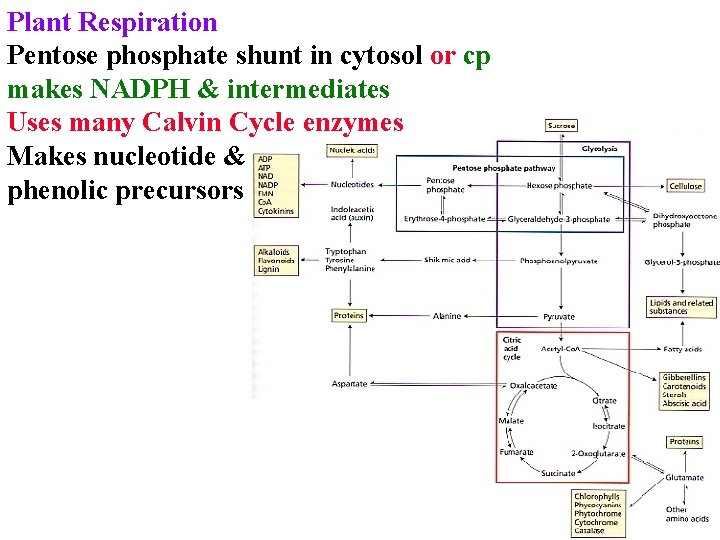
Plant Respiration Pentose phosphate shunt in cytosol or cp makes NADPH & intermediates Uses many Calvin Cycle enzymes Makes nucleotide & phenolic precursors
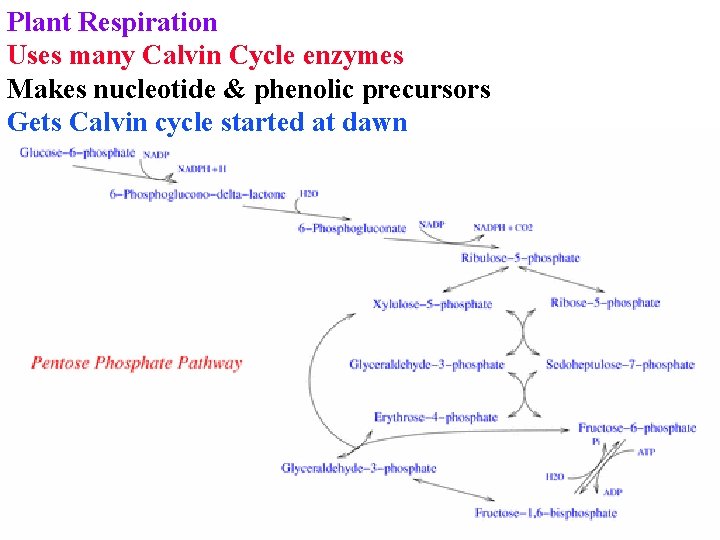
Plant Respiration Uses many Calvin Cycle enzymes Makes nucleotide & phenolic precursors Gets Calvin cycle started at dawn
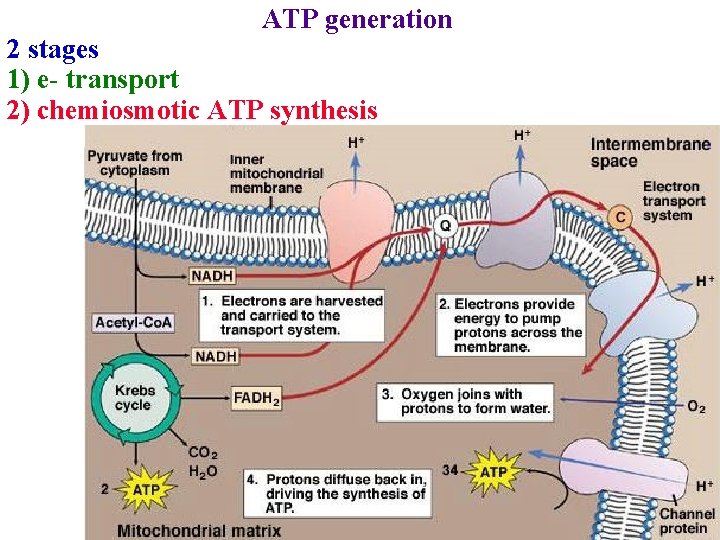
ATP generation 2 stages 1) e- transport 2) chemiosmotic ATP synthesis
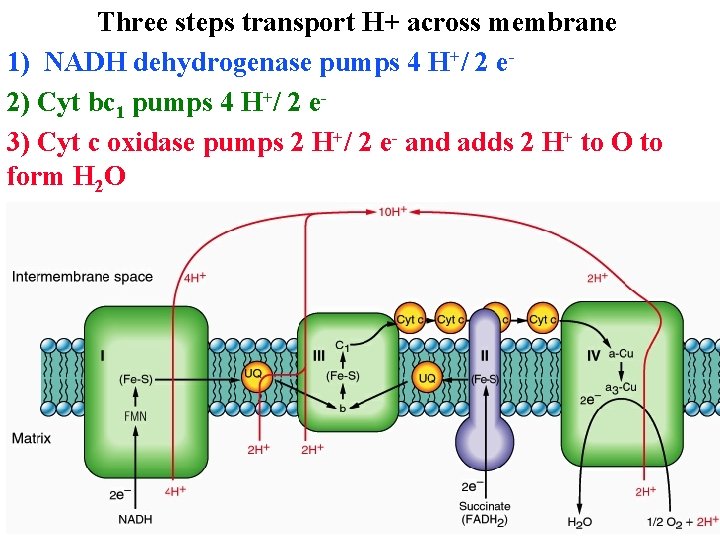
Three steps transport H+ across membrane 1) NADH dehydrogenase pumps 4 H+/ 2 e 2) Cyt bc 1 pumps 4 H+/ 2 e 3) Cyt c oxidase pumps 2 H+/ 2 e- and adds 2 H+ to O to form H 2 O
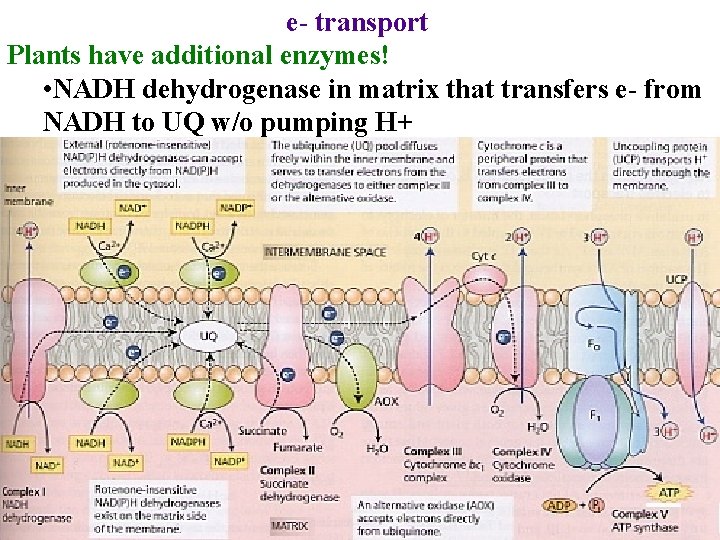
e- transport Plants have additional enzymes! • NADH dehydrogenase in matrix that transfers e- from NADH to UQ w/o pumping H+
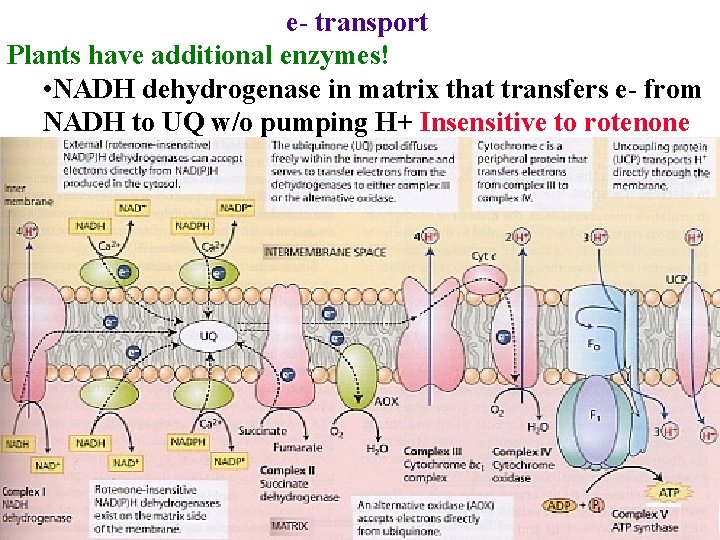
e- transport Plants have additional enzymes! • NADH dehydrogenase in matrix that transfers e- from NADH to UQ w/o pumping H+ Insensitive to rotenone
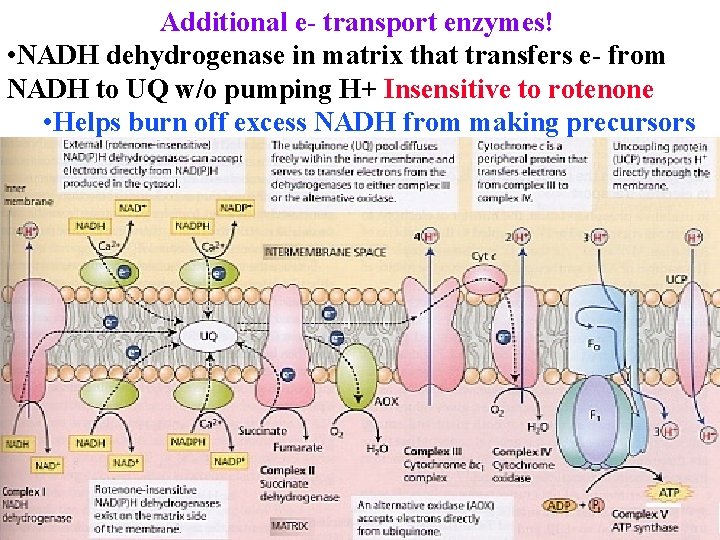
Additional e- transport enzymes! • NADH dehydrogenase in matrix that transfers e- from NADH to UQ w/o pumping H+ Insensitive to rotenone • Helps burn off excess NADH from making precursors
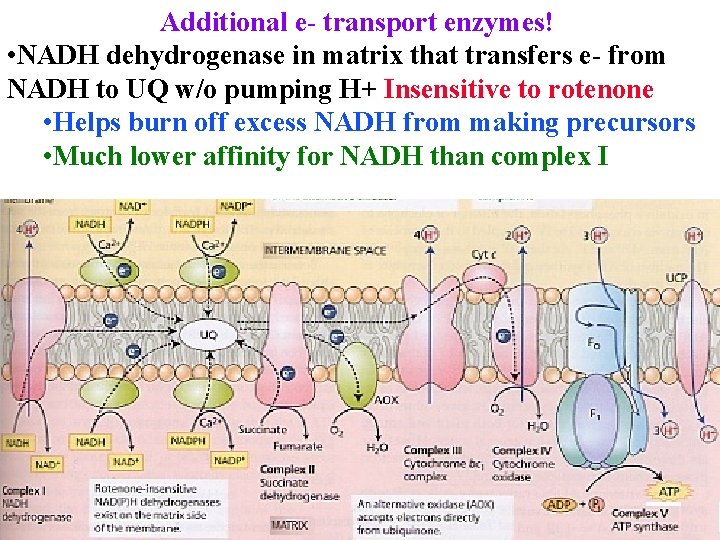
Additional e- transport enzymes! • NADH dehydrogenase in matrix that transfers e- from NADH to UQ w/o pumping H+ Insensitive to rotenone • Helps burn off excess NADH from making precursors • Much lower affinity for NADH than complex I
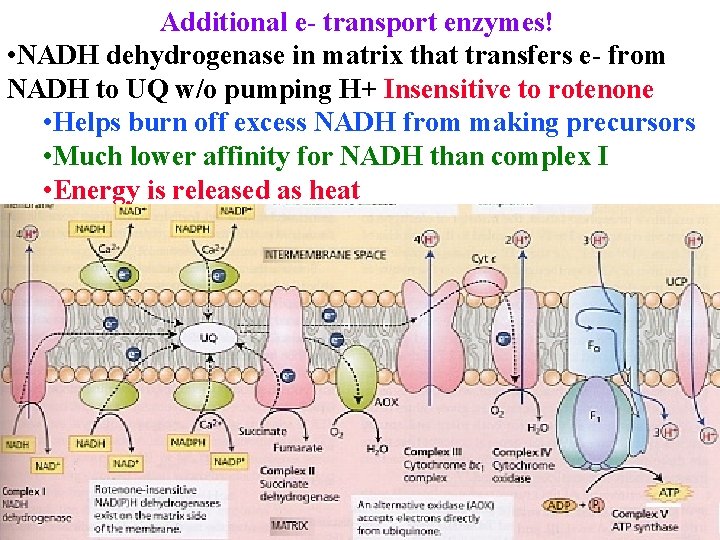
Additional e- transport enzymes! • NADH dehydrogenase in matrix that transfers e- from NADH to UQ w/o pumping H+ Insensitive to rotenone • Helps burn off excess NADH from making precursors • Much lower affinity for NADH than complex I • Energy is released as heat
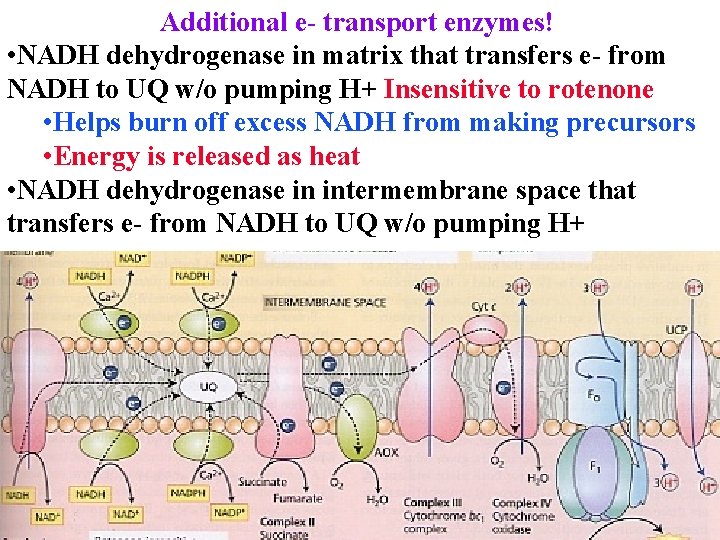
Additional e- transport enzymes! • NADH dehydrogenase in matrix that transfers e- from NADH to UQ w/o pumping H+ Insensitive to rotenone • Helps burn off excess NADH from making precursors • Energy is released as heat • NADH dehydrogenase in intermembrane space that transfers e- from NADH to UQ w/o pumping H+
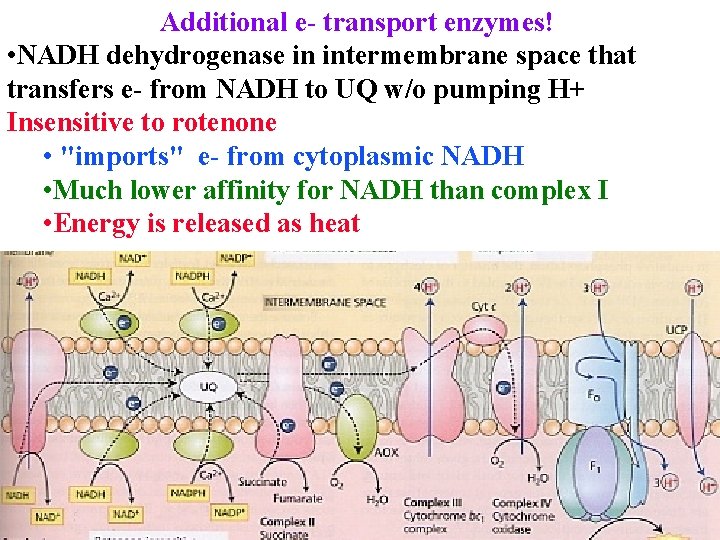
Additional e- transport enzymes! • NADH dehydrogenase in intermembrane space that transfers e- from NADH to UQ w/o pumping H+ Insensitive to rotenone • "imports" e- from cytoplasmic NADH • Much lower affinity for NADH than complex I • Energy is released as heat
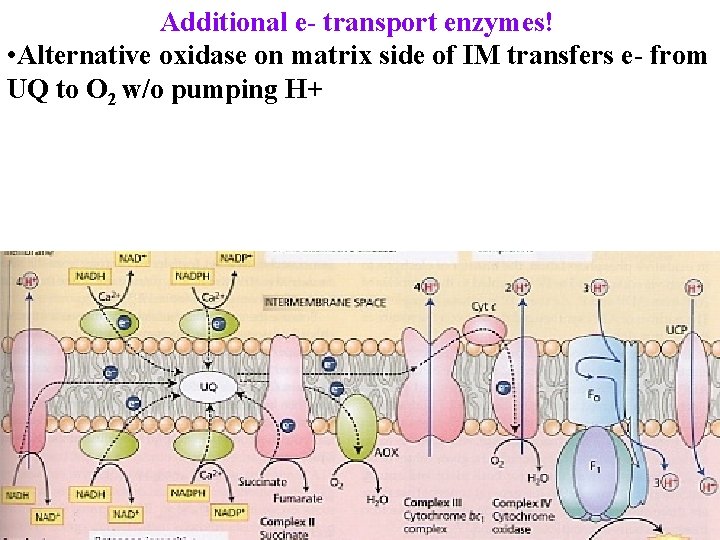
Additional e- transport enzymes! • Alternative oxidase on matrix side of IM transfers e- from UQ to O 2 w/o pumping H+

Additional e- transport enzymes! • Alternative oxidase on matrix side of IM transfers e- from UQ to O 2 w/o pumping H+ • Insensitive to Cyanide, Azide or CO

Additional e- transport enzymes! • Alternative oxidase on matrix side of IM transfers e- from UQ to O 2 w/o pumping H+ • Insensitive to Cyanide, Azide or CO • Sensitive to SHAM (salicylhydroxamic acid)
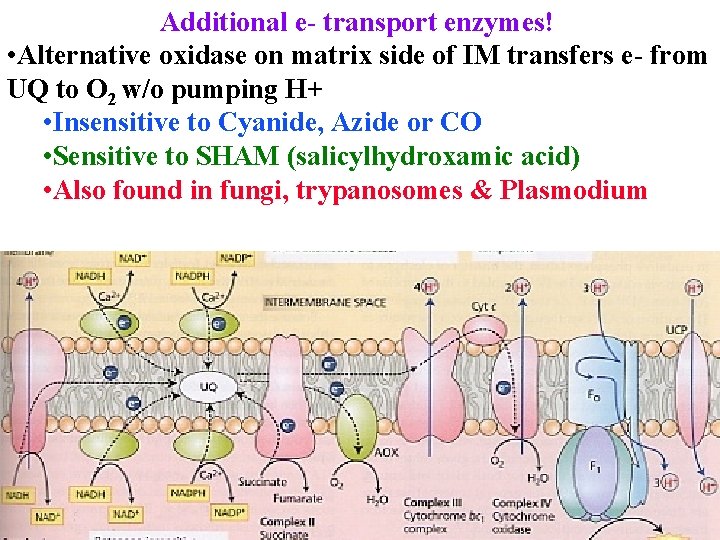
Additional e- transport enzymes! • Alternative oxidase on matrix side of IM transfers e- from UQ to O 2 w/o pumping H+ • Insensitive to Cyanide, Azide or CO • Sensitive to SHAM (salicylhydroxamic acid) • Also found in fungi, trypanosomes & Plasmodium
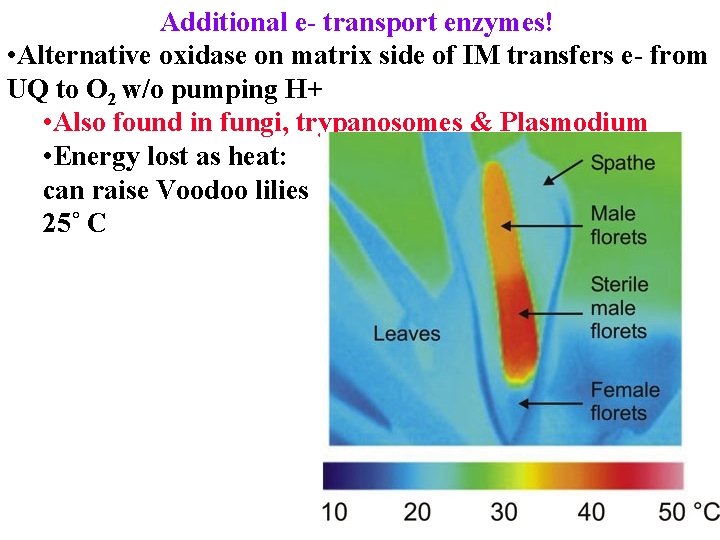
Additional e- transport enzymes! • Alternative oxidase on matrix side of IM transfers e- from UQ to O 2 w/o pumping H+ • Also found in fungi, trypanosomes & Plasmodium • Energy lost as heat: can raise Voodoo lilies 25˚ C
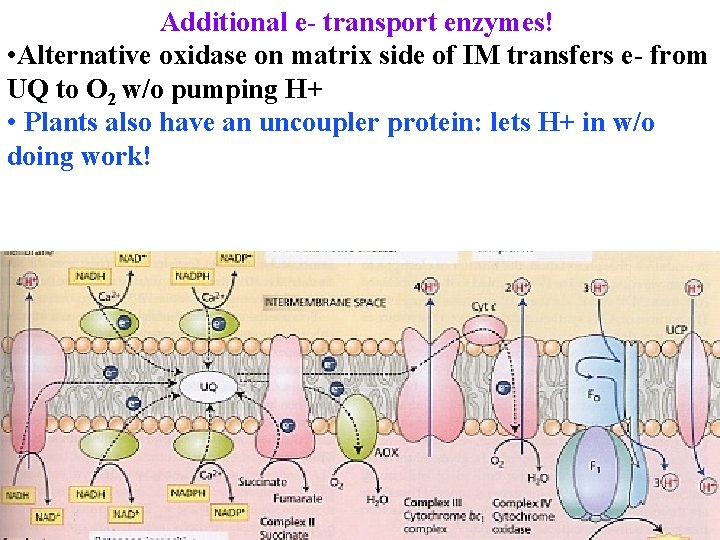
Additional e- transport enzymes! • Alternative oxidase on matrix side of IM transfers e- from UQ to O 2 w/o pumping H+ • Plants also have an uncoupler protein: lets H+ in w/o doing work!
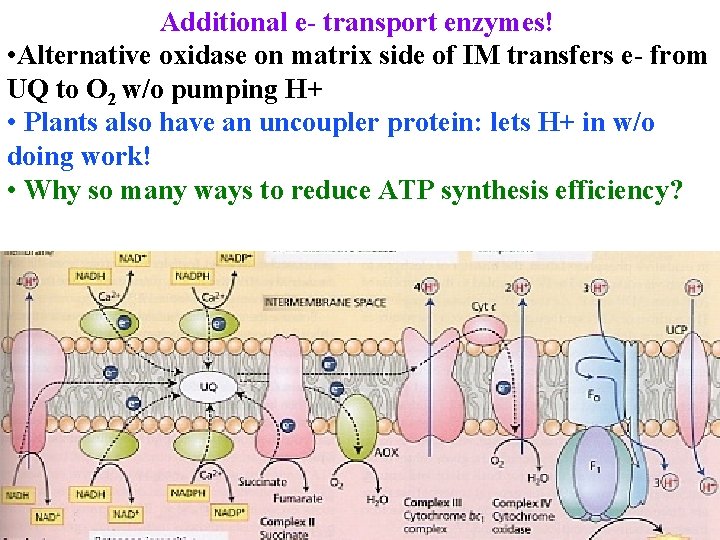
Additional e- transport enzymes! • Alternative oxidase on matrix side of IM transfers e- from UQ to O 2 w/o pumping H+ • Plants also have an uncoupler protein: lets H+ in w/o doing work! • Why so many ways to reduce ATP synthesis efficiency?
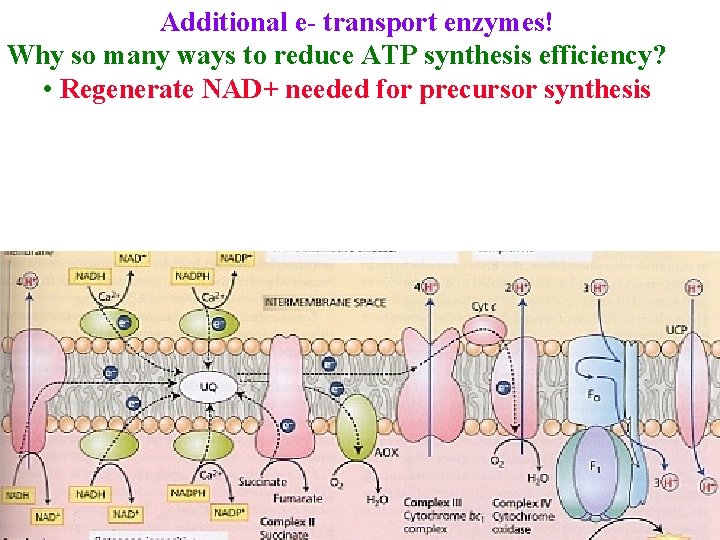
Additional e- transport enzymes! Why so many ways to reduce ATP synthesis efficiency? • Regenerate NAD+ needed for precursor synthesis
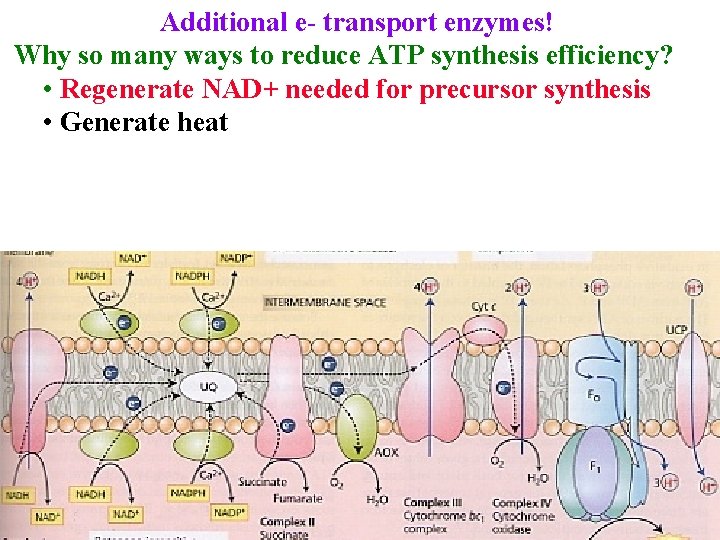
Additional e- transport enzymes! Why so many ways to reduce ATP synthesis efficiency? • Regenerate NAD+ needed for precursor synthesis • Generate heat
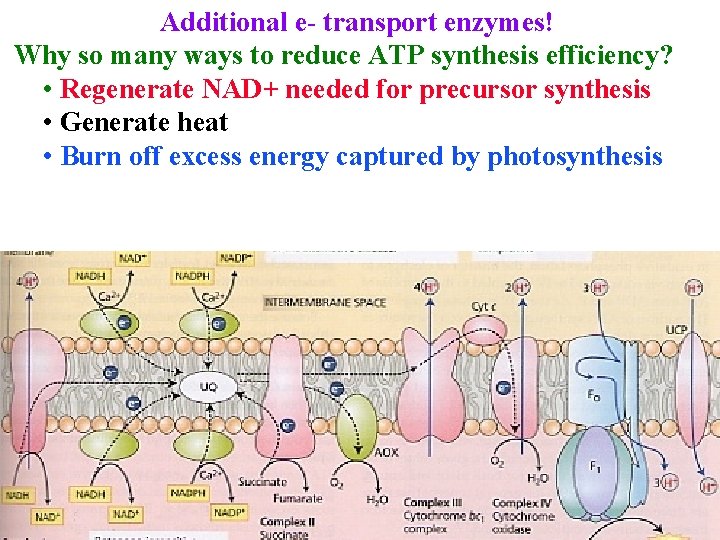
Additional e- transport enzymes! Why so many ways to reduce ATP synthesis efficiency? • Regenerate NAD+ needed for precursor synthesis • Generate heat • Burn off excess energy captured by photosynthesis
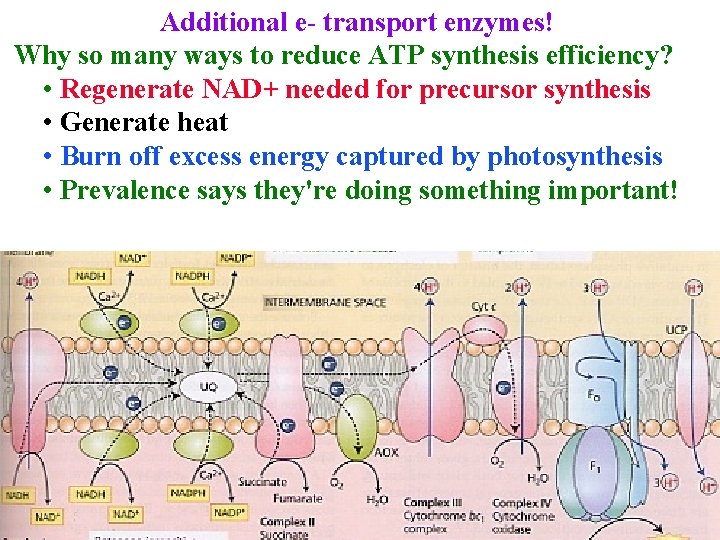
Additional e- transport enzymes! Why so many ways to reduce ATP synthesis efficiency? • Regenerate NAD+ needed for precursor synthesis • Generate heat • Burn off excess energy captured by photosynthesis • Prevalence says they're doing something important!
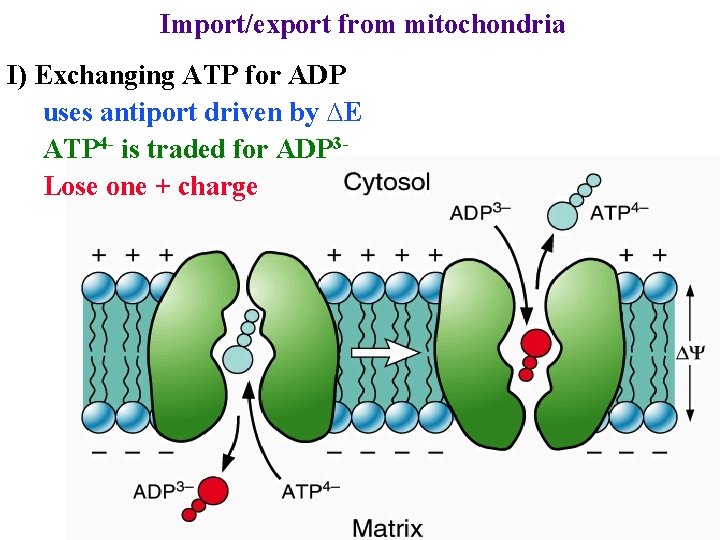
Import/export from mitochondria I) Exchanging ATP for ADP uses antiport driven by ∆E ATP 4 - is traded for ADP 3 Lose one + charge
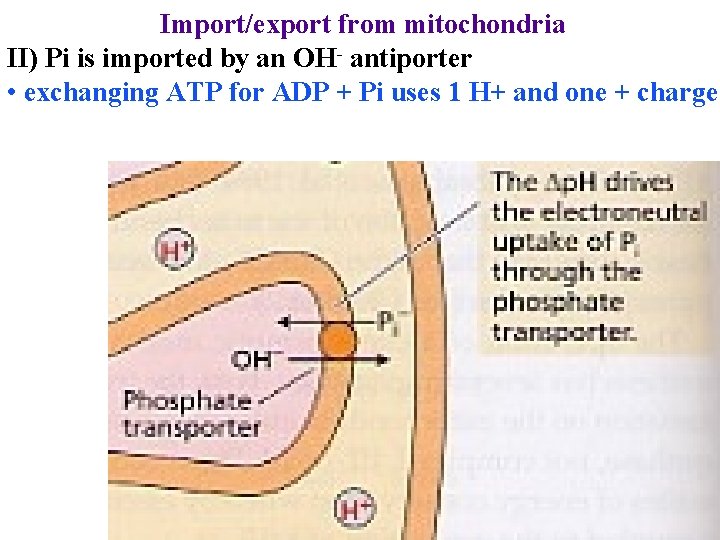
Import/export from mitochondria II) Pi is imported by an OH- antiporter • exchanging ATP for ADP + Pi uses 1 H+ and one + charge

Import/export from mitochondria II) Pi is imported by an OH- antiporter • exchanging ATP for ADP + Pi uses 1 H+ and one + charge • Use ∆Pi for other import

Import/export from mitochondria II) Pi is imported by an OH- antiporter III) Pyruvate is imported by an OH- antiporter
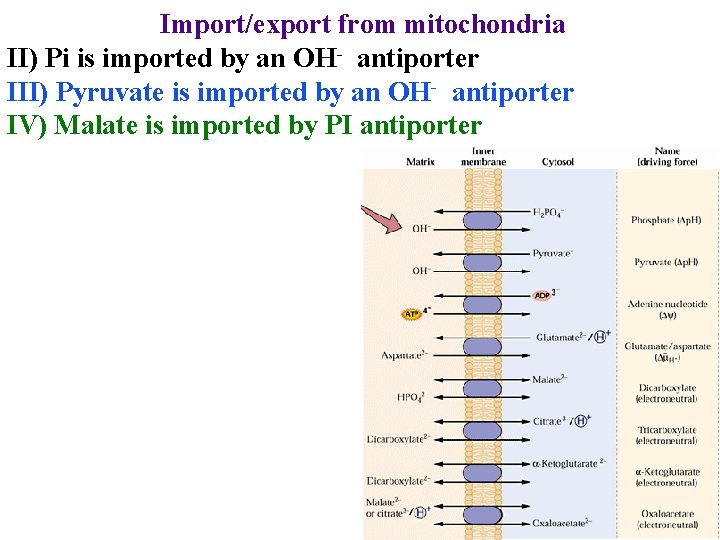
Import/export from mitochondria II) Pi is imported by an OH- antiporter III) Pyruvate is imported by an OH- antiporter IV) Malate is imported by PI antiporter
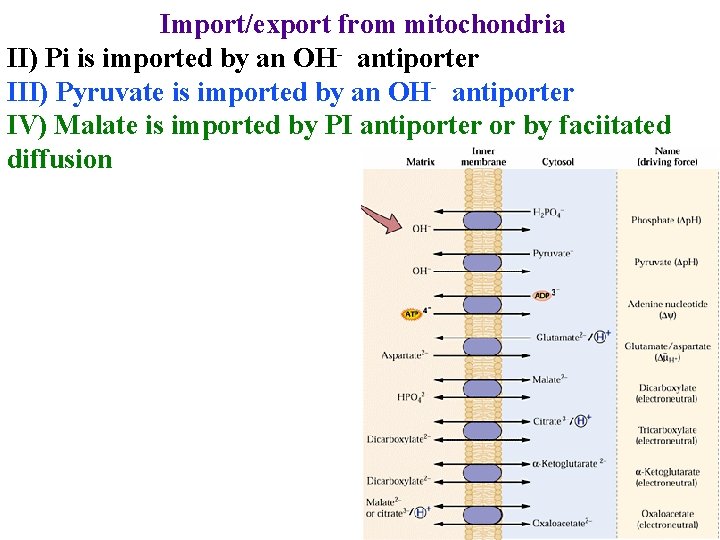
Import/export from mitochondria II) Pi is imported by an OH- antiporter III) Pyruvate is imported by an OH- antiporter IV) Malate is imported by PI antiporter or by faciitated diffusion
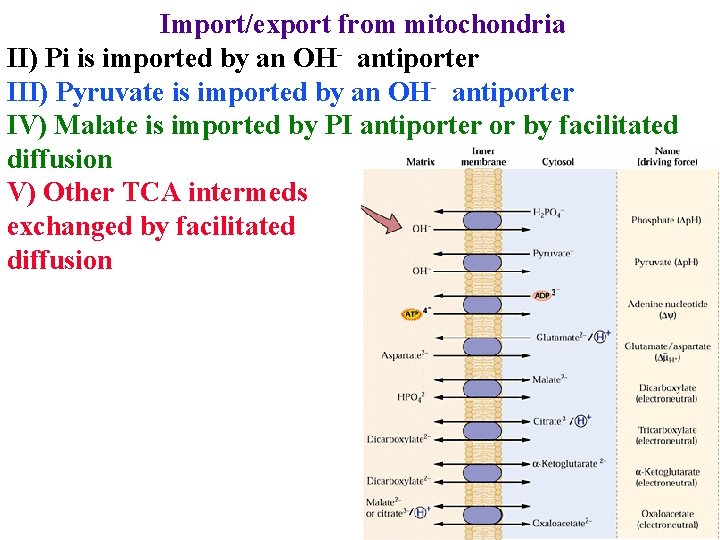
Import/export from mitochondria II) Pi is imported by an OH- antiporter III) Pyruvate is imported by an OH- antiporter IV) Malate is imported by PI antiporter or by facilitated diffusion V) Other TCA intermeds exchanged by facilitated diffusion

Import/export from mitochondria VI) Mito DNA encodes ~ 35 proteins, also r. RNA & t. RNA • subunits of ATP synthase & complexes I, III & IV

Import/export from mitochondria VI) Mito DNA encodes ~ 35 proteins, also r. RNA & t. RNA • subunits of ATP synthase & complexes I, III & IV • some m. RNA are trans-spliced from 2 diff transcripts!

Import/export from mitochondria VI) Mito DNA encodes ~ 35 proteins, also r. RNA & t. RNA • subunits of ATP synthase & complexes I, III & IV • some m. RNA are trans-spliced from 2 diff transcripts! • some m. RNA are edited: bases changed after synthesis!

Import/export from mitochondria VI) Mito DNA encodes ~ 35 proteins, also r. RNA & t. RNA • subunits of ATP synthase & complexes I, III & IV • some m. RNA are trans-spliced from 2 diff transcripts! • some m. RNA are edited: bases changed after synthesis! • mt. DNA recombines to form new genes, some poison pollen development to create cytoplasmic male sterility

Import/export from mitochondria VI) Mito DNA encodes ~ 35 proteins, also r. RNA & t. RNA • subunits of ATP synthase & complexes I, III & IV • some m. RNA are trans-spliced from 2 diff transcripts! • some m. RNA are edited: bases changed after synthesis! • mt. DNA recombines to form new genes, some poison pollen development to create cytoplasmic male sterility • Pollen don't transmit mito!

Import/export from mitochondria VI) Mito DNA encodes ~ 35 proteins, also r. RNA & t. RNA • subunits of ATP synthase & complexes I, III & IV • some m. RNA are trans-spliced from 2 diff transcripts! • some m. RNA are edited: bases changed after synthesis! • mt. DNA recombines to form new genes, some poison pollen development to create cytoplasmic male sterility • Pollen don't transmit mito! • Other 2000 proteins are imported from cytosol posttranslationally • made with N-terminal presequence targeting them to mito

Import/export from mitochondria VI) Mito genome encodes ~ 35 proteins, remainder are imported from cytosol post-translationally • made with N-terminal presequence targeting them to mito
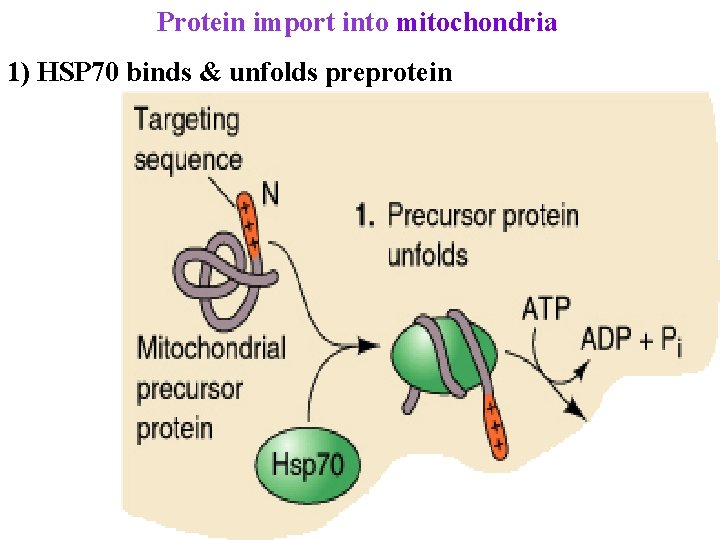
Protein import into mitochondria 1) HSP 70 binds & unfolds preprotein
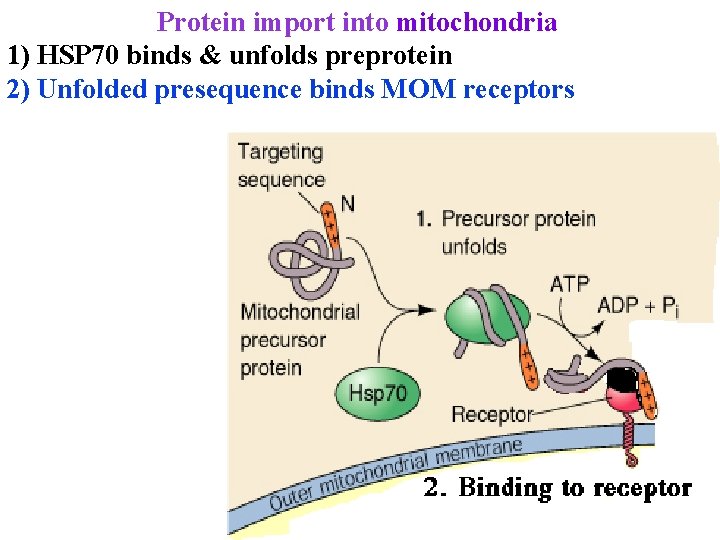
Protein import into mitochondria 1) HSP 70 binds & unfolds preprotein 2) Unfolded presequence binds MOM receptors
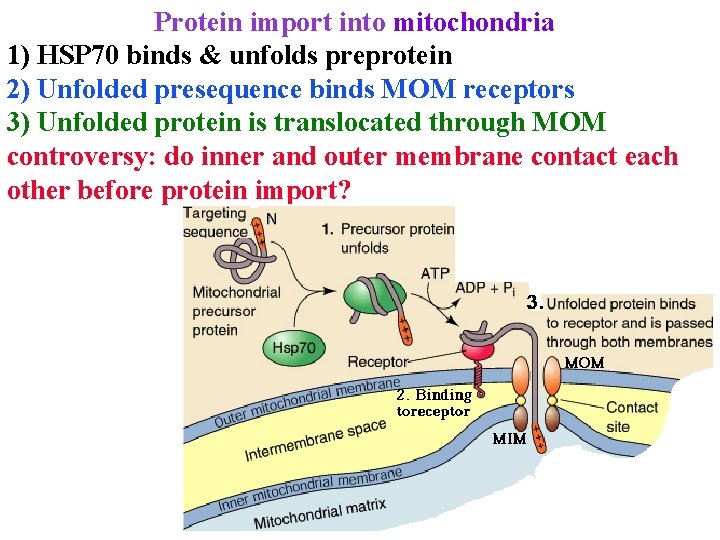
Protein import into mitochondria 1) HSP 70 binds & unfolds preprotein 2) Unfolded presequence binds MOM receptors 3) Unfolded protein is translocated through MOM controversy: do inner and outer membrane contact each other before protein import?
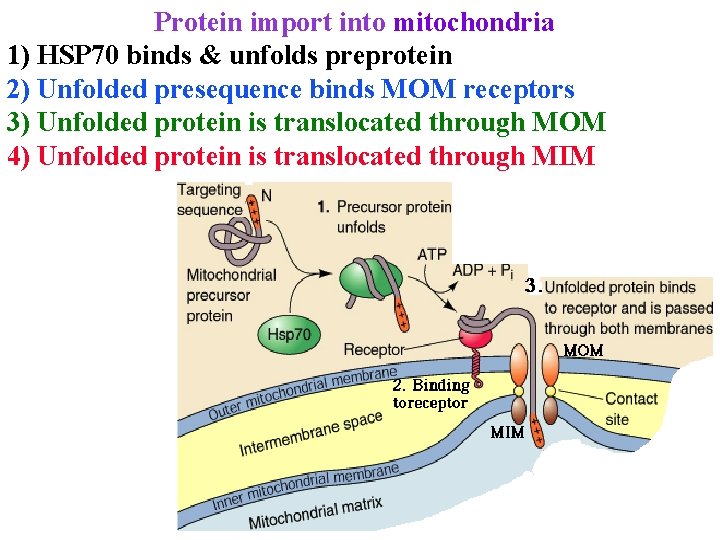
Protein import into mitochondria 1) HSP 70 binds & unfolds preprotein 2) Unfolded presequence binds MOM receptors 3) Unfolded protein is translocated through MOM 4) Unfolded protein is translocated through MIM
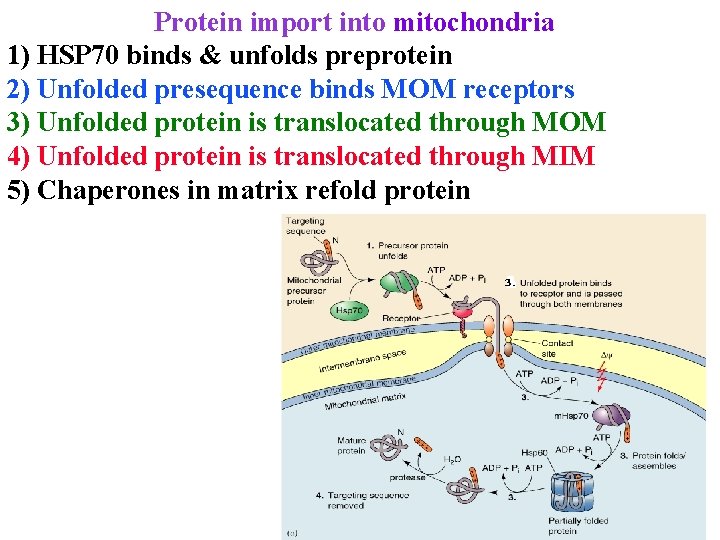
Protein import into mitochondria 1) HSP 70 binds & unfolds preprotein 2) Unfolded presequence binds MOM receptors 3) Unfolded protein is translocated through MOM 4) Unfolded protein is translocated through MIM 5) Chaperones in matrix refold protein
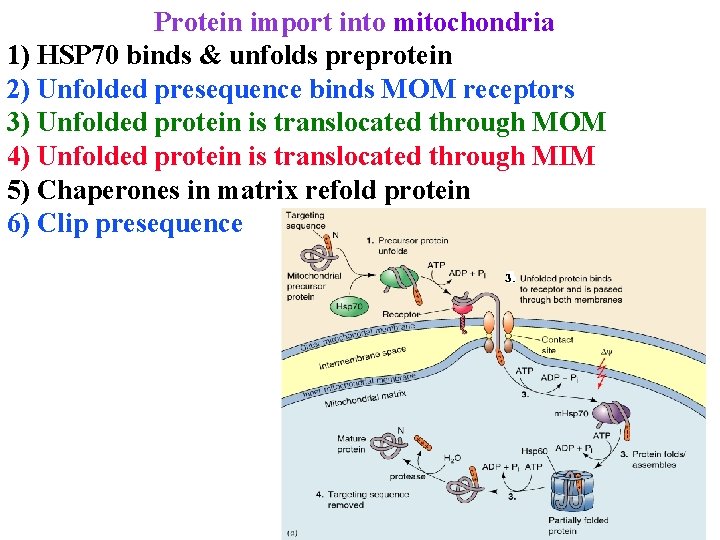
Protein import into mitochondria 1) HSP 70 binds & unfolds preprotein 2) Unfolded presequence binds MOM receptors 3) Unfolded protein is translocated through MOM 4) Unfolded protein is translocated through MIM 5) Chaperones in matrix refold protein 6) Clip presequence
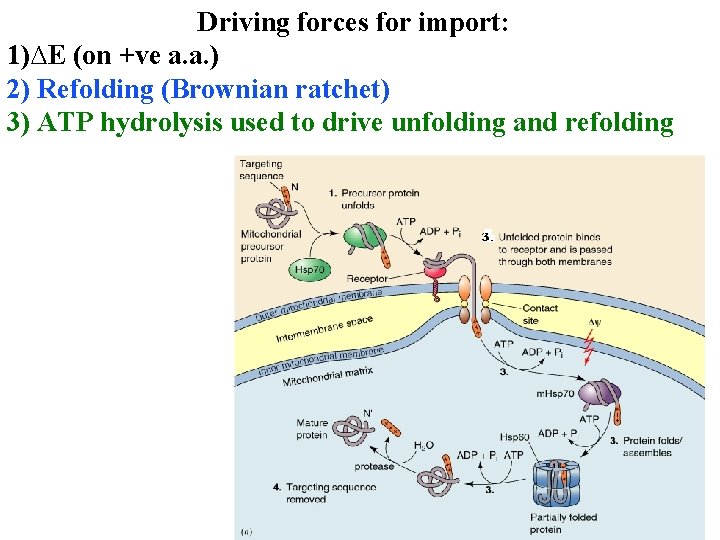
Driving forces for import: 1)∆E (on +ve a. a. ) 2) Refolding (Brownian ratchet) 3) ATP hydrolysis used to drive unfolding and refolding
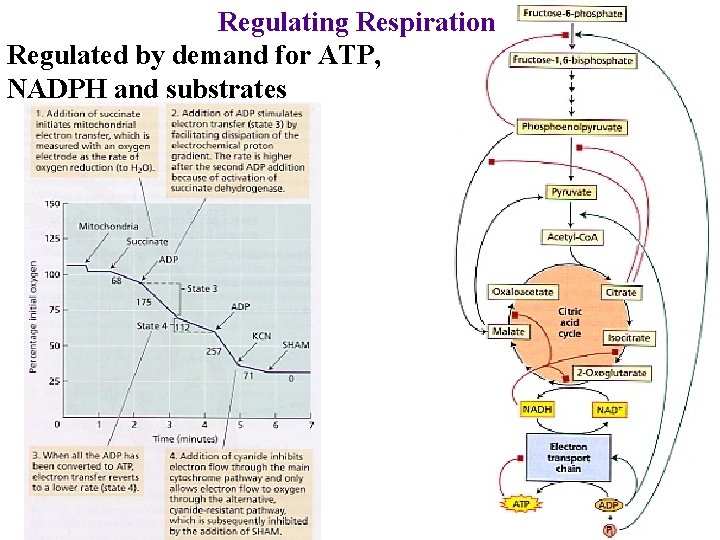
Regulating Respiration Regulated by demand for ATP, NADPH and substrates
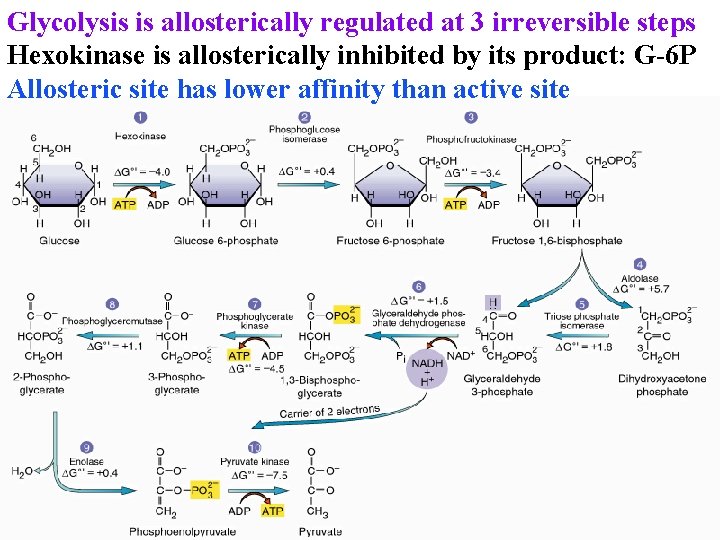
Glycolysis is allosterically regulated at 3 irreversible steps Hexokinase is allosterically inhibited by its product: G-6 P Allosteric site has lower affinity than active site
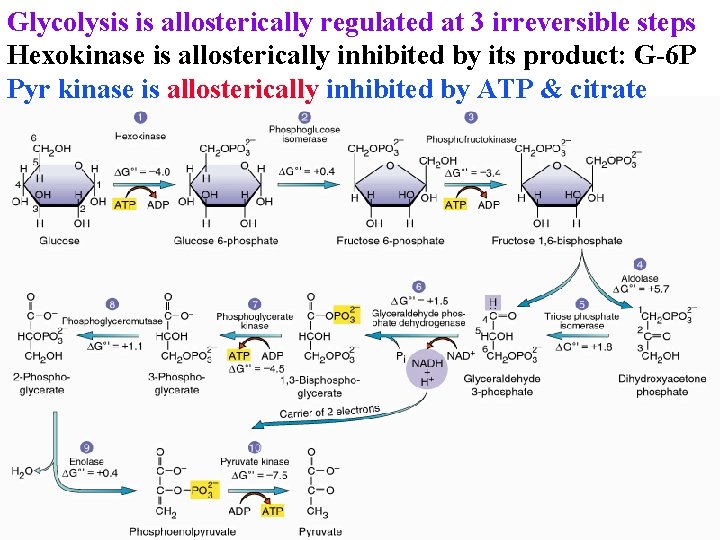
Glycolysis is allosterically regulated at 3 irreversible steps Hexokinase is allosterically inhibited by its product: G-6 P Pyr kinase is allosterically inhibited by ATP & citrate
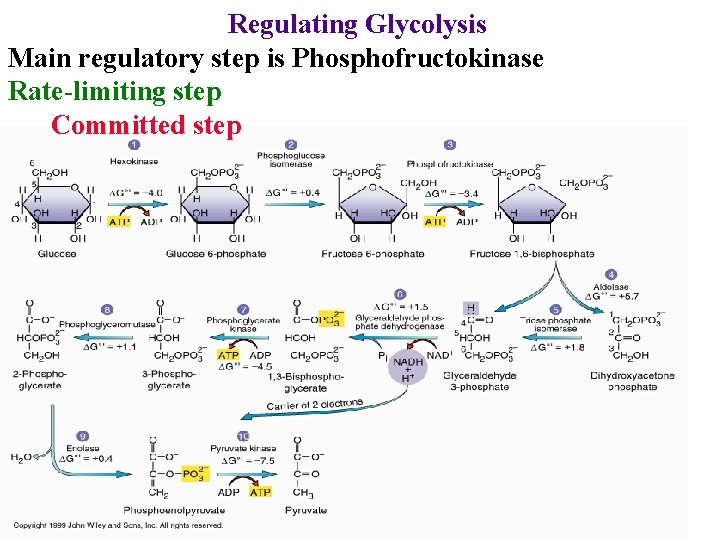
Regulating Glycolysis Main regulatory step is Phosphofructokinase Rate-limiting step Committed step
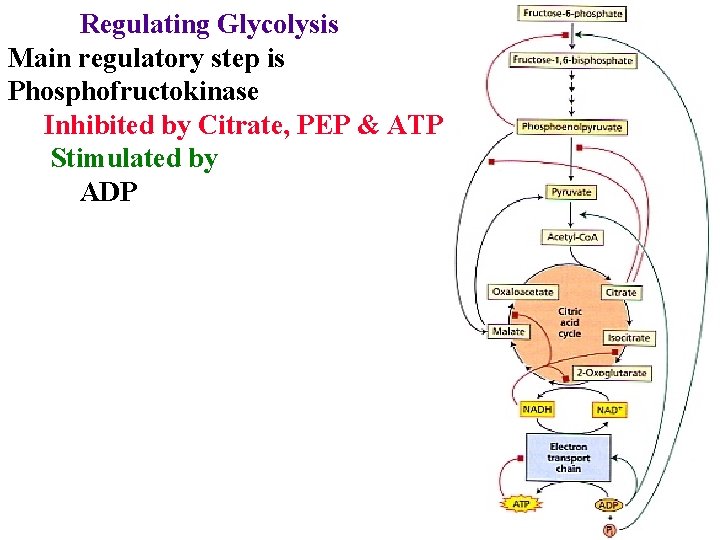
Regulating Glycolysis Main regulatory step is Phosphofructokinase Inhibited by Citrate, PEP & ATP Stimulated by ADP
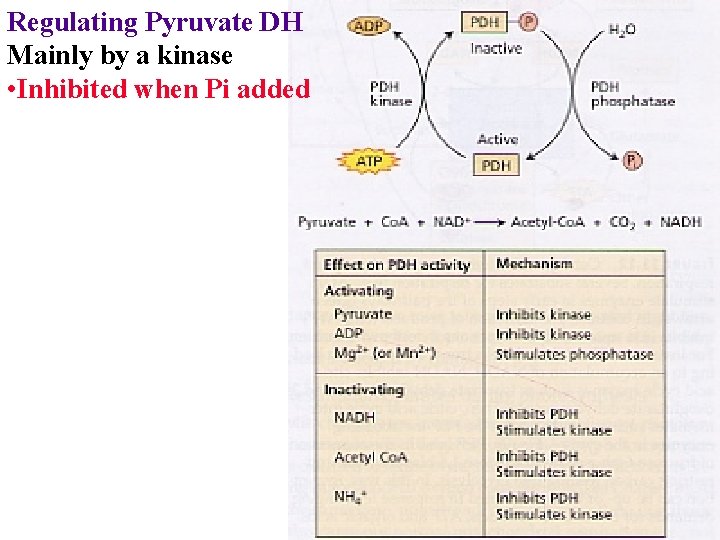
Regulating Pyruvate DH Mainly by a kinase • Inhibited when Pi added
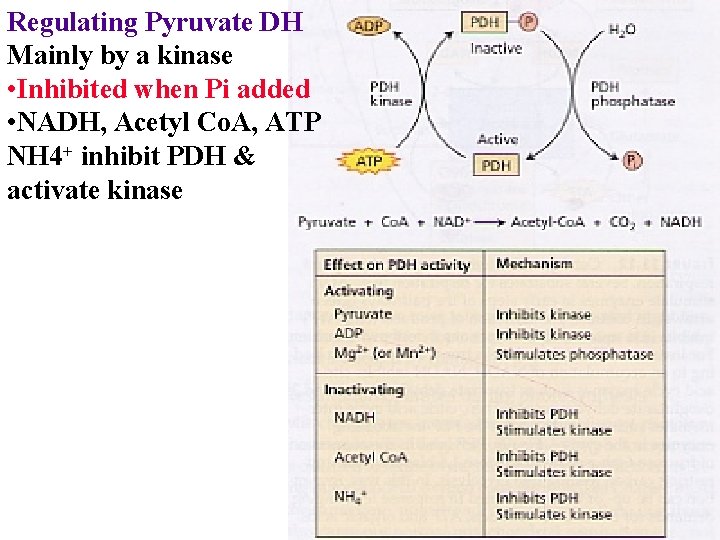
Regulating Pyruvate DH Mainly by a kinase • Inhibited when Pi added • NADH, Acetyl Co. A, ATP NH 4+ inhibit PDH & activate kinase
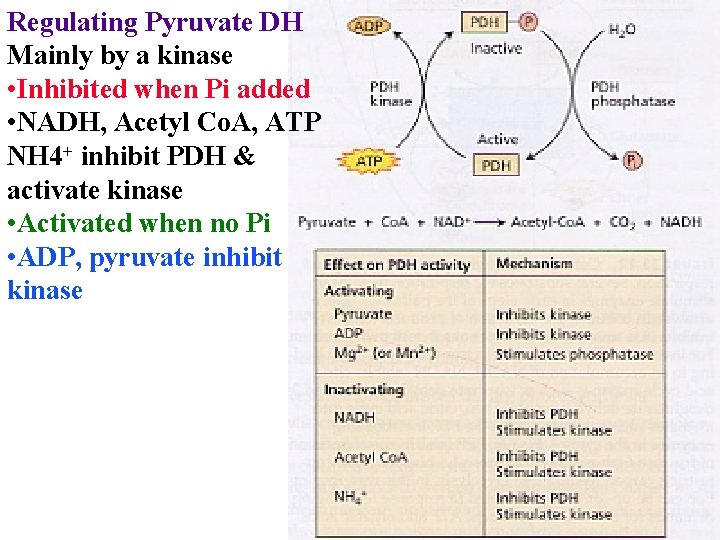
Regulating Pyruvate DH Mainly by a kinase • Inhibited when Pi added • NADH, Acetyl Co. A, ATP NH 4+ inhibit PDH & activate kinase • Activated when no Pi • ADP, pyruvate inhibit kinase
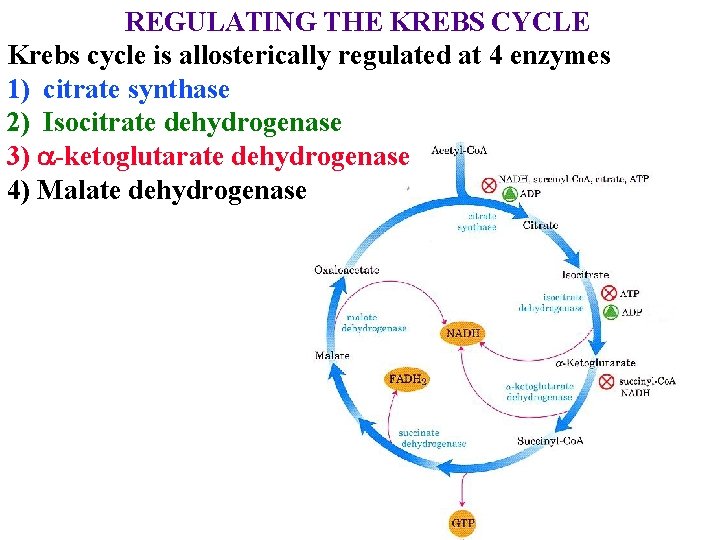
REGULATING THE KREBS CYCLE Krebs cycle is allosterically regulated at 4 enzymes 1) citrate synthase 2) Isocitrate dehydrogenase 3) a-ketoglutarate dehydrogenase 4) Malate dehydrogenase
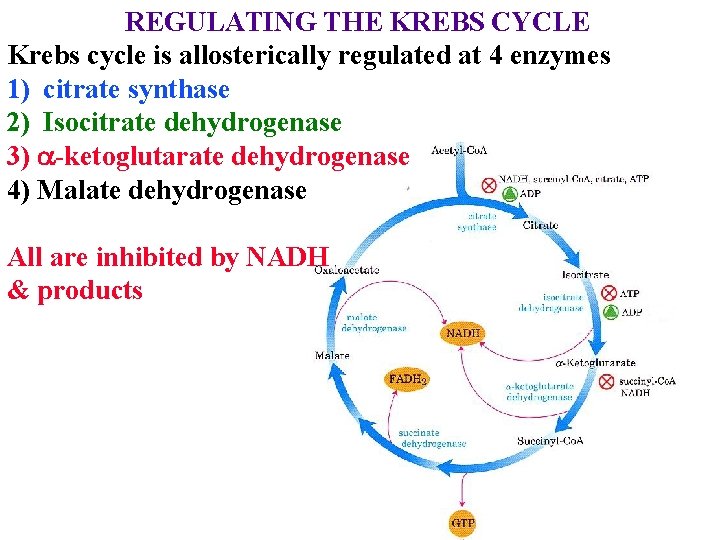
REGULATING THE KREBS CYCLE Krebs cycle is allosterically regulated at 4 enzymes 1) citrate synthase 2) Isocitrate dehydrogenase 3) a-ketoglutarate dehydrogenase 4) Malate dehydrogenase All are inhibited by NADH & products
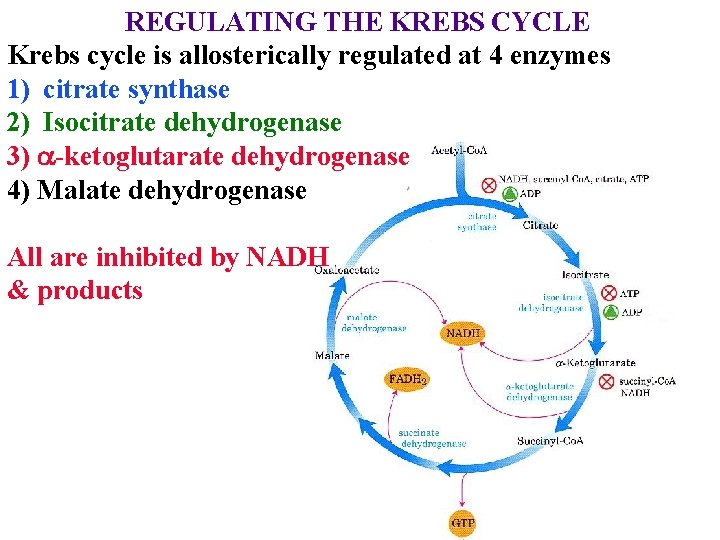
REGULATING THE KREBS CYCLE Krebs cycle is allosterically regulated at 4 enzymes 1) citrate synthase 2) Isocitrate dehydrogenase 3) a-ketoglutarate dehydrogenase 4) Malate dehydrogenase All are inhibited by NADH & products

Environmental factors 1) Temperature • Rate ~ doubles for each 10˚ C increase up to ~ 40˚ • At higher T start to denature

Environmental factors 1) Temperature • Rate ~ doubles for each 10˚ C increase up to ~ 40˚ • At higher T start to denature 2) p. O 2 • Respiration declines if p. O 2 <5%

Environmental factors 1) Temperature • Rate ~ doubles for each 10˚ C increase up to ~ 40˚ • At higher T start to denature 2) p. O 2 • Respiration declines if p. O 2 <5% • Problem for flooded roots

Environmental factors 1) Temperature • Rate ~ doubles for each 10˚ C increase up to ~ 40˚ • At higher T start to denature 2) p. O 2 • Respiration declines if p. O 2 <5% • Problem for flooded roots 3) p. CO 2 • Inhibits respiration at 3%

Environmental factors 1) Temperature • Rate ~ doubles for each 10˚ C increase up to ~ 40˚ • At higher T start to denature 2) p. O 2 • Respiration declines if p. O 2 <5% • Problem for flooded roots 3) p. CO 2 • Inhibits respiration at 3% • No obvious effects at 700 ppm, yet biomass reduced
- Slides: 158