NEW SOLUTIONS FOR HIGHLY MULTIPLEXED PROTEIN ASSAYS VIA
















- Slides: 43

NEW SOLUTIONS FOR HIGHLY MULTIPLEXED PROTEIN ASSAYS VIA MRM Webinar June 04, 2012 1

CAPRION PROTEOMICS • Leading proteomics-based service provider - Biomarker and target discovery - Multiplexed MRM assays - Biomarker verification and validation • Recently acquired Immune. Carta • Multiparametric GLP flow cytometry for immune monitoring • Located in: – Montreal, Canada – Menlo Park, California • Founded in 2000 • 62 employees (55 scientists) • Extensive experience - Over 45 industry and government partners - Over 100 large scale studies completed (clinical and pre-clinical) - All major therapeutic areas and sample types - Expertise in study design, execution, statistical analysis, biological insights and comprehensive reporting 2

TECHNICAL APPROACHES IN MASS SPEC-BASED PROTEOMICS Biomarker Discovery (LC-MS/MS) • Non-hypothesis based discovery approach • Label-free, gel-free quantitative mass spectrometry • Shotgun sequencing • Profile 1000’s of proteins in 100’s samples • Identify differentially expressed proteins as candidate targets and biomarkers Multiplexed Assays (MRM) • Targeted quantification of up to 350 specific proteins • Target list from MS, literature, transcriptomics • Rapid assay development • Ab-free or ab enrichment • Synthetic labeled standards for absolute quantification 3

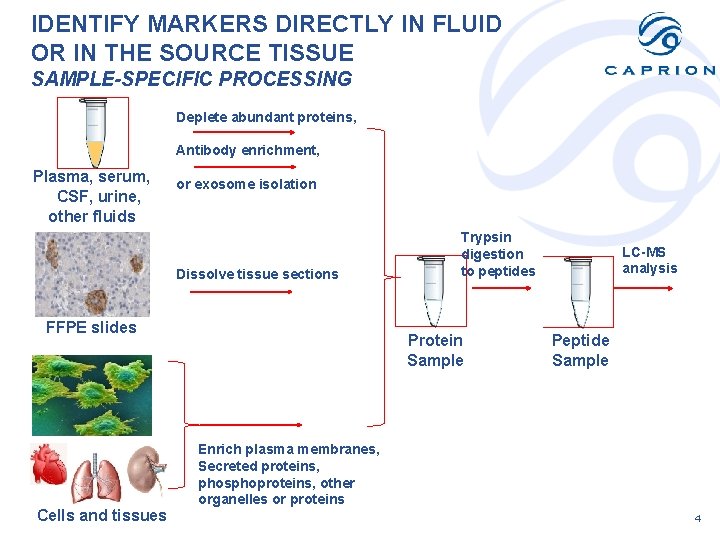
IDENTIFY MARKERS DIRECTLY IN FLUID OR IN THE SOURCE TISSUE SAMPLE-SPECIFIC PROCESSING Deplete abundant proteins, Antibody enrichment, Plasma, serum, CSF, urine, other fluids or exosome isolation Dissolve tissue sections FFPE slides Cells and tissues Trypsin digestion to peptides Protein Sample LC-MS analysis Peptide Sample Enrich plasma membranes, Secreted proteins, phosphoproteins, other organelles or proteins 4

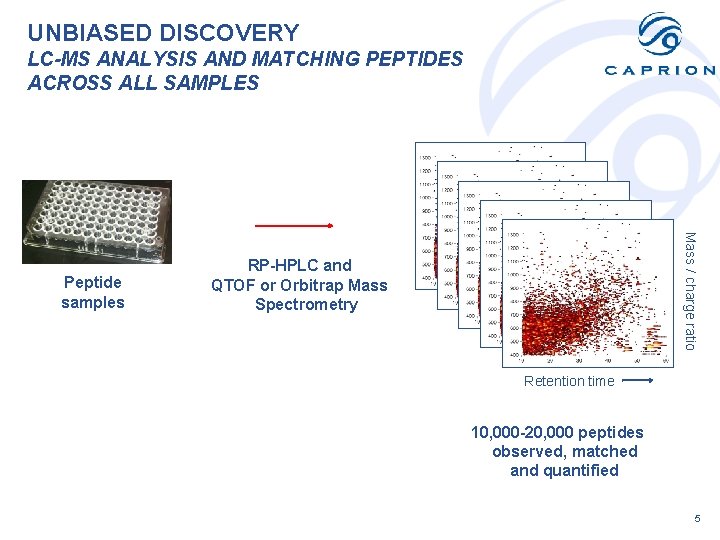
UNBIASED DISCOVERY LC-MS ANALYSIS AND MATCHING PEPTIDES ACROSS ALL SAMPLES Mass / charge ratio Peptide samples RP-HPLC and QTOF or Orbitrap Mass Spectrometry Retention time 10, 000 -20, 000 peptides observed, matched and quantified 5

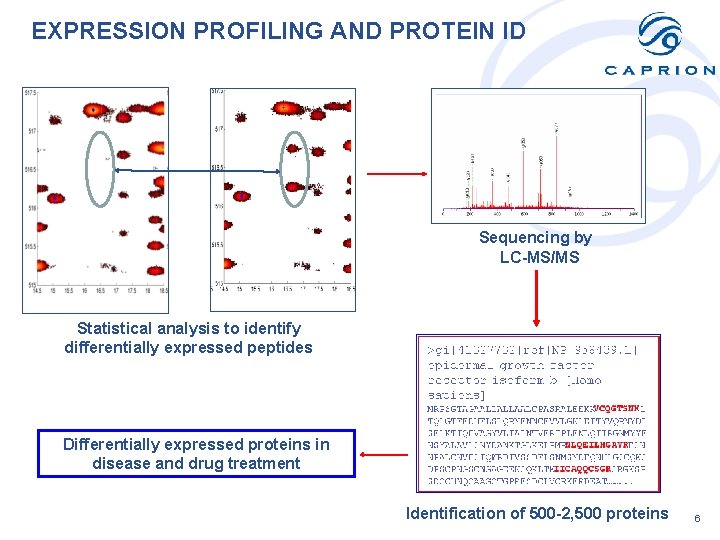
EXPRESSION PROFILING AND PROTEIN ID Sequencing by LC-MS/MS Statistical analysis to identify differentially expressed peptides Differentially expressed proteins in disease and drug treatment Identification of 500 -2, 500 proteins 6

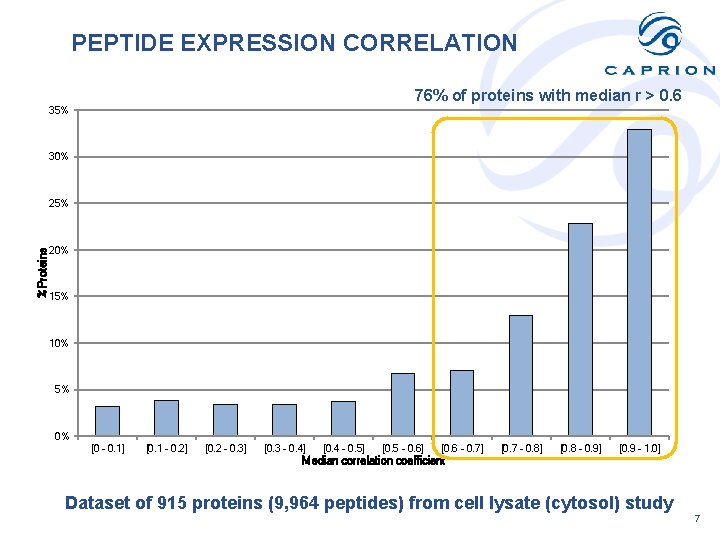
PEPTIDE EXPRESSION CORRELATION 76% of proteins with median r > 0. 6 35% 30% %Proteins 25% 20% 15% 10% 5% 0% [0 - 0. 1] [0. 1 - 0. 2] [0. 2 - 0. 3] [0. 3 - 0. 4] [0. 4 - 0. 5] [0. 5 - 0. 6] [0. 6 - 0. 7] [0. 7 - 0. 8] [0. 8 - 0. 9] [0. 9 - 1. 0] Median correlation coefficient Dataset of 915 proteins (9, 964 peptides) from cell lysate (cytosol) study 7

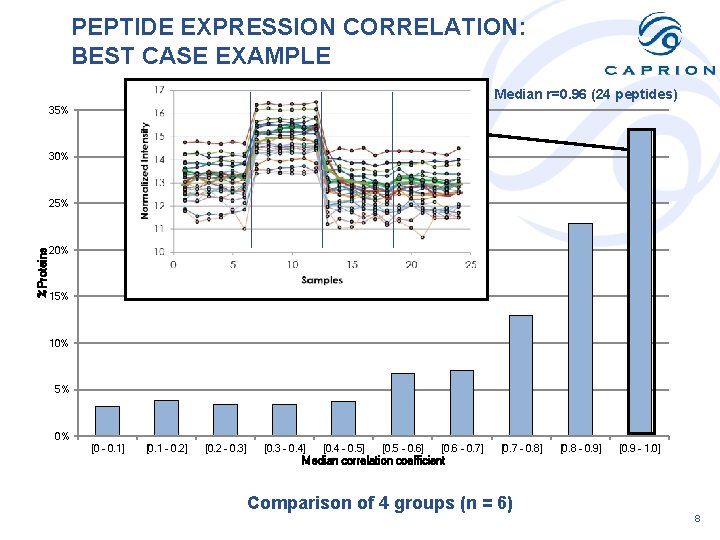
PEPTIDE EXPRESSION CORRELATION: BEST CASE EXAMPLE Median r=0. 96 (24 peptides) 35% 30% %Proteins 25% 20% 15% 10% 5% 0% [0 - 0. 1] [0. 1 - 0. 2] [0. 2 - 0. 3] [0. 3 - 0. 4] [0. 4 - 0. 5] [0. 5 - 0. 6] [0. 6 - 0. 7] [0. 7 - 0. 8] [0. 8 - 0. 9] [0. 9 - 1. 0] Median correlation coefficient Comparison of 4 groups (n = 6) 8

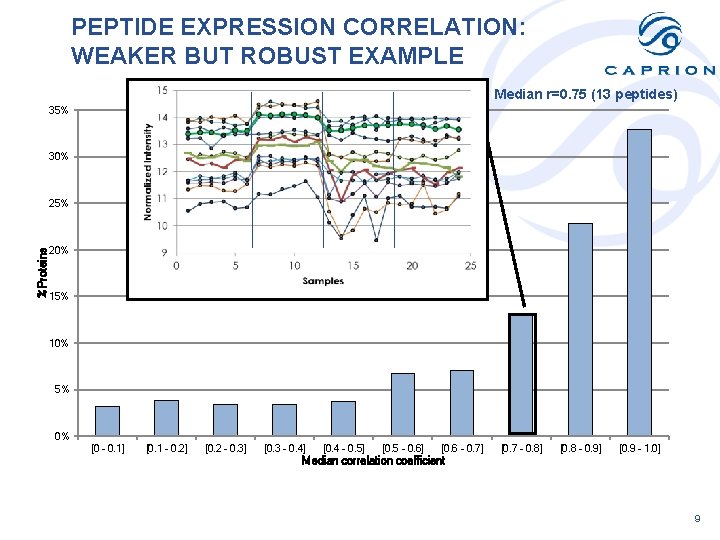
PEPTIDE EXPRESSION CORRELATION: WEAKER BUT ROBUST EXAMPLE Median r=0. 75 (13 peptides) 35% 30% %Proteins 25% 20% 15% 10% 5% 0% [0 - 0. 1] [0. 1 - 0. 2] [0. 2 - 0. 3] [0. 3 - 0. 4] [0. 4 - 0. 5] [0. 5 - 0. 6] [0. 6 - 0. 7] [0. 7 - 0. 8] [0. 8 - 0. 9] [0. 9 - 1. 0] Median correlation coefficient 9

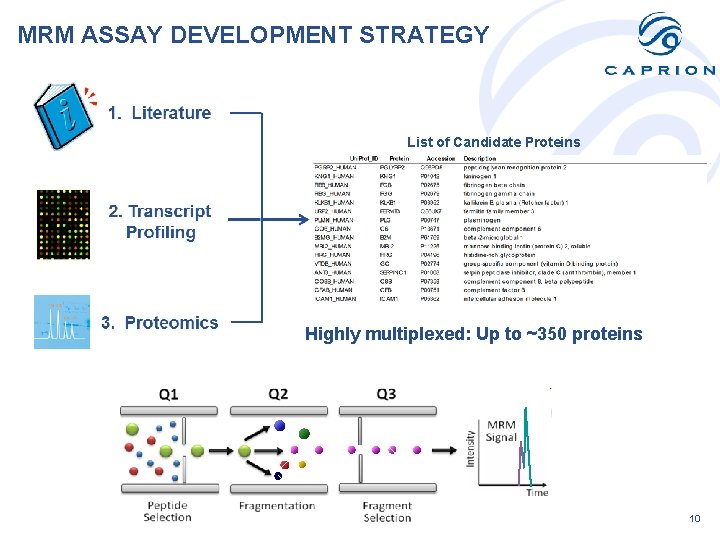
MRM ASSAY DEVELOPMENT STRATEGY List of Candidate Proteins Highly multiplexed: Up to ~350 proteins 10

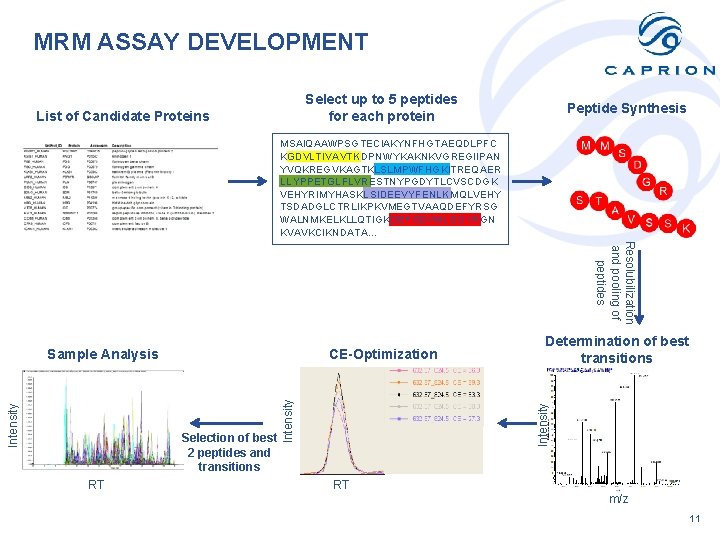
MRM ASSAY DEVELOPMENT Select up to 5 peptides for each protein List of Candidate Proteins Peptide Synthesis MSAIQAAWPSGTECIAKYNFHGTAEQDLPFC KGDVLTIVAVTKDPNWYKAKNKVGREGIIPAN YVQKREGVKAGTKLSLMPWFHGKITREQAER LLYPPETGLFLVRESTNYPGDYTLCVSCDGK VEHYRIMYHASKLSIDEEVYFENLKMQLVEHY TSDADGLCTRLIKPKVMEGTVAAQDEFYRSG WALNMKELKLLQTIGKGEFGDVMLGDYRGN KVAVKCIKNDATA… Resolubilization and pooling of peptides CE-Optimization RT Determination of best transitions Intensity Selection of best 2 peptides and transitions Intensity Sample Analysis RT m/z 11

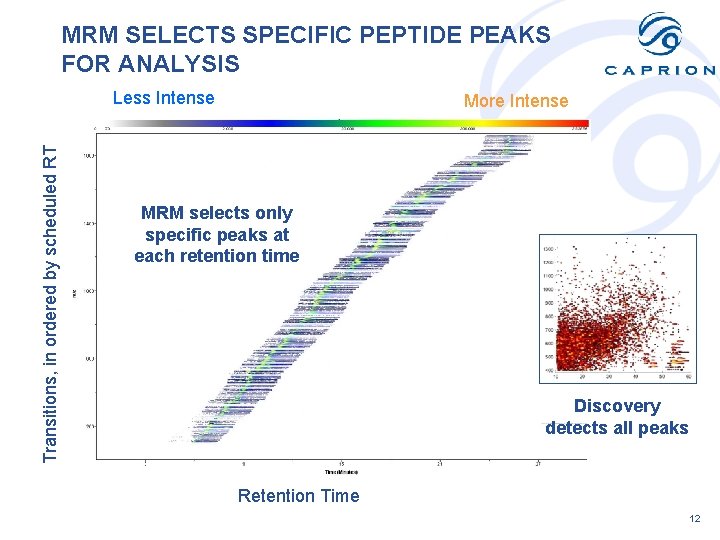
MRM SELECTS SPECIFIC PEPTIDE PEAKS FOR ANALYSIS Transitions, in ordered by scheduled RT Less Intense More Intense MRM selects only specific peaks at each retention time Discovery detects all peaks Retention Time 12

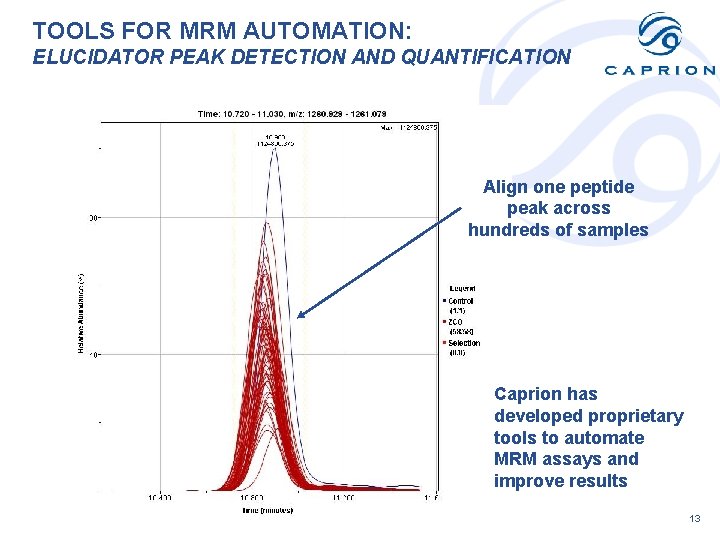
TOOLS FOR MRM AUTOMATION: ELUCIDATOR PEAK DETECTION AND QUANTIFICATION Align one peptide peak across hundreds of samples Caprion has developed proprietary tools to automate MRM assays and improve results 13

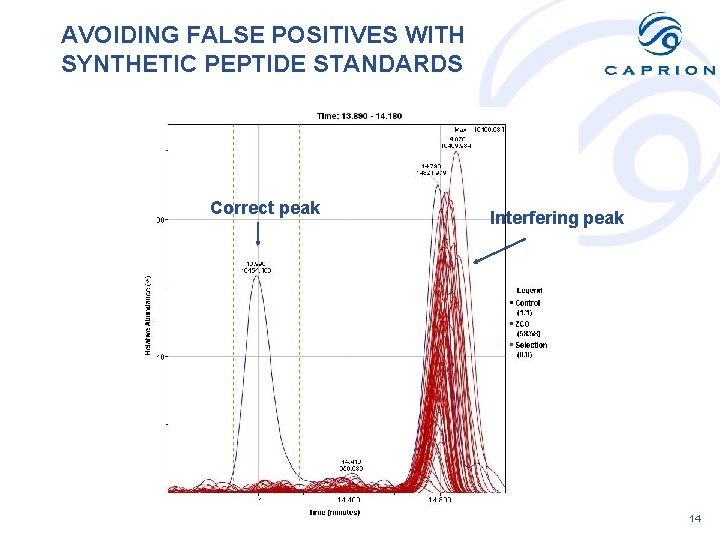
AVOIDING FALSE POSITIVES WITH SYNTHETIC PEPTIDE STANDARDS Correct peak Interfering peak 14

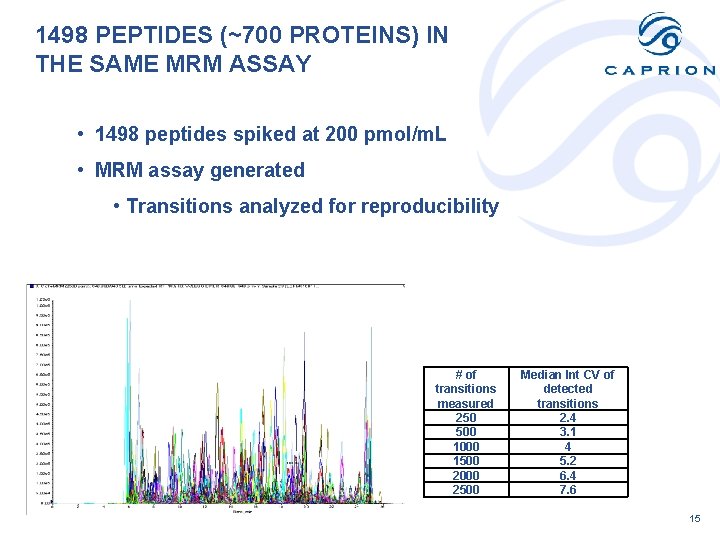
1498 PEPTIDES (~700 PROTEINS) IN THE SAME MRM ASSAY • 1498 peptides spiked at 200 pmol/m. L • MRM assay generated • Transitions analyzed for reproducibility # of transitions measured 250 500 1000 1500 2000 2500 Median Int CV of detected transitions 2. 4 3. 1 4 5. 2 6. 4 7. 6 15

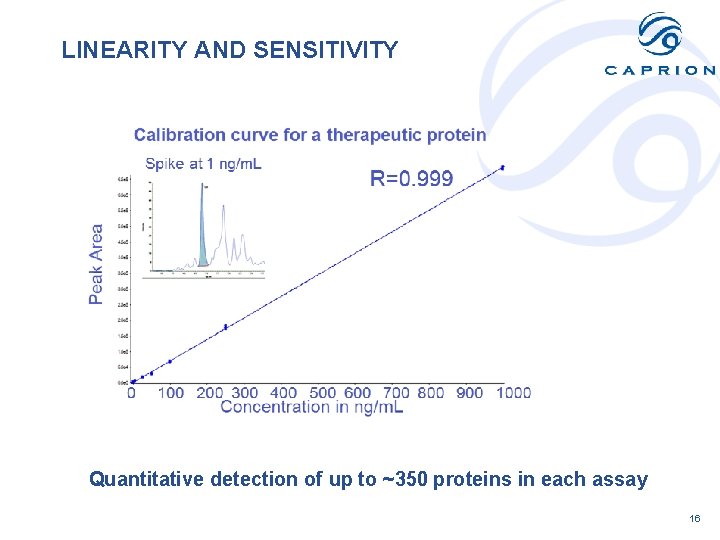
LINEARITY AND SENSITIVITY Quantitative detection of up to ~350 proteins in each assay 16

Options for MRM assays 17

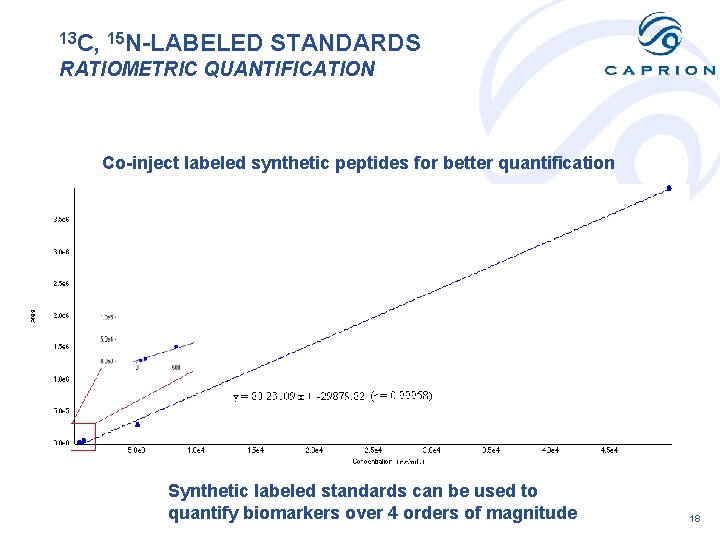
13 C, 15 N-LABELED STANDARDS RATIOMETRIC QUANTIFICATION Co-inject labeled synthetic peptides for better quantification Synthetic labeled standards can be used to quantify biomarkers over 4 orders of magnitude 18

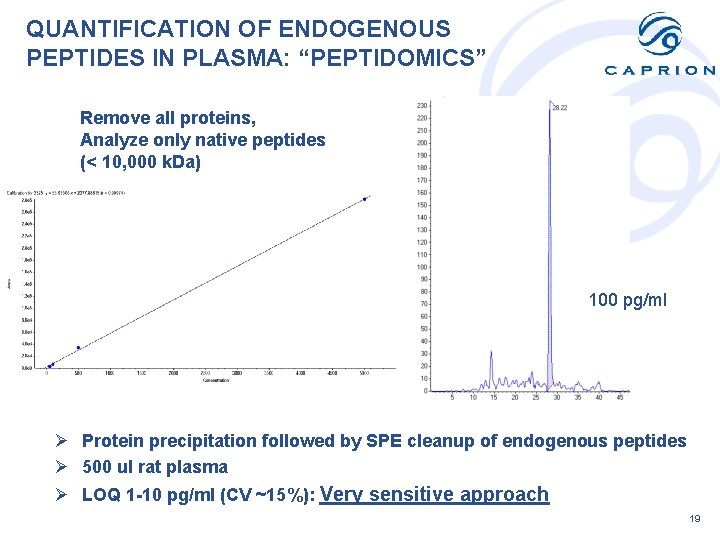
QUANTIFICATION OF ENDOGENOUS PEPTIDES IN PLASMA: “PEPTIDOMICS” Remove all proteins, Analyze only native peptides (< 10, 000 k. Da) 100 pg/ml Ø Protein precipitation followed by SPE cleanup of endogenous peptides Ø 500 ul rat plasma Ø LOQ 1 -10 pg/ml (CV ~15%): Very sensitive approach 19

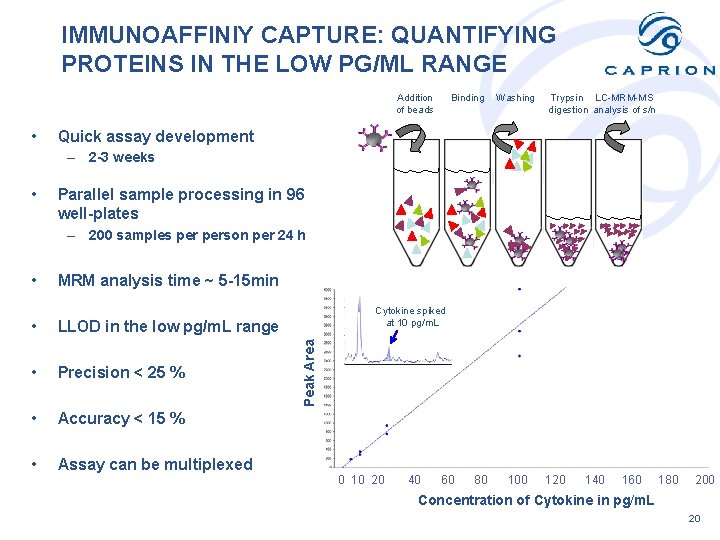
IMMUNOAFFINIY CAPTURE: QUANTIFYING PROTEINS IN THE LOW PG/ML RANGE Addition of beads • Binding Washing Trypsin LC-MRM-MS digestion analysis of s/n Quick assay development – 2 -3 weeks • Parallel sample processing in 96 well-plates – 200 samples person per 24 h MRM analysis time ~ 5 -15 min • LLOD in the low pg/m. L range • Precision < 25 % • Accuracy < 15 % • Assay can be multiplexed Cytokine spiked at 10 pg/m. L Peak Area • 0 10 20 40 60 80 100 120 140 160 180 200 Concentration of Cytokine in pg/m. L 20

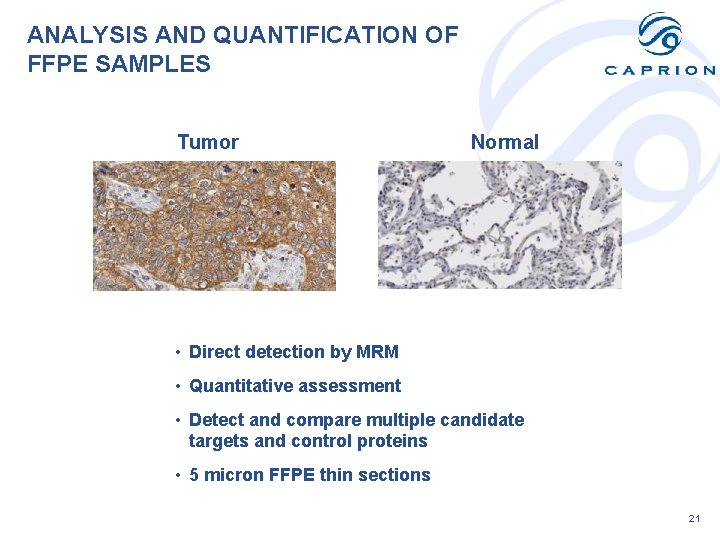
ANALYSIS AND QUANTIFICATION OF FFPE SAMPLES Tumor Normal • Direct detection by MRM • Quantitative assessment • Detect and compare multiple candidate targets and control proteins • 5 micron FFPE thin sections 21

Case Study 1: Non-Targeted Discovery in Plasma Followed by MRM Markers of active TB in plasma and serum 22

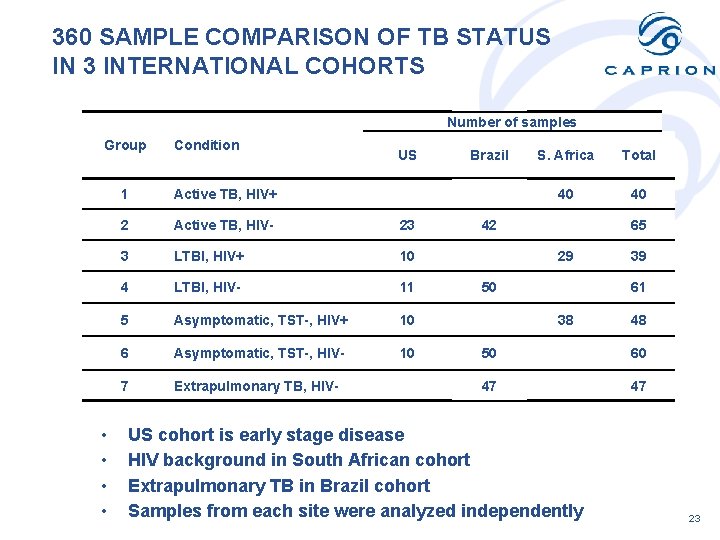
360 SAMPLE COMPARISON OF TB STATUS IN 3 INTERNATIONAL COHORTS Number of samples Group • • Condition US Brazil S. Africa Total 1 Active TB, HIV+ 40 40 2 Active TB, HIV- 23 42 65 3 LTBI, HIV+ 10 29 39 4 LTBI, HIV- 11 50 61 5 Asymptomatic, TST-, HIV+ 10 38 48 6 Asymptomatic, TST-, HIV- 10 50 60 7 Extrapulmonary TB, HIV- 47 47 US cohort is early stage disease HIV background in South African cohort Extrapulmonary TB in Brazil cohort Samples from each site were analyzed independently 23

SUMMARY OF DISCOVERY RESULTS • High abundance proteins depleted with Ig. Y 14/Supermix resin • Single LC-MS analysis per sample • Identified and quantified ~450 proteins • Compared active TB to other cohorts • Number of differentially expressed proteins: US: 105 Brazil: 146 S. Africa: 49 Non-redundant total: 200 • Most differentially expressed proteins assigned to defined processes: Immune response ~50% Lipid metabolism ~20% Lung damage & wound healing ~20% 24

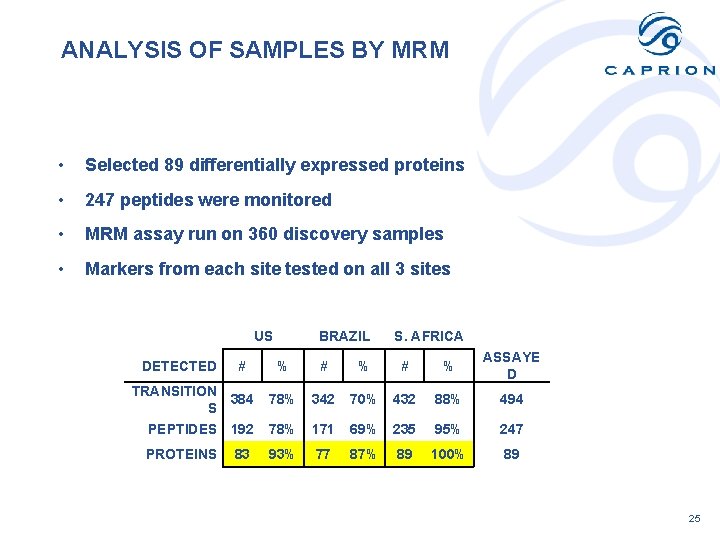
ANALYSIS OF SAMPLES BY MRM • Selected 89 differentially expressed proteins • 247 peptides were monitored • MRM assay run on 360 discovery samples • Markers from each site tested on all 3 sites US BRAZIL S. AFRICA % # % ASSAYE D TRANSITION 384 S 78% 342 70% 432 88% 494 PEPTIDES 192 78% 171 69% 235 95% 247 PROTEINS 93% 77 87% 89 100% 89 DETECTED # 83 25

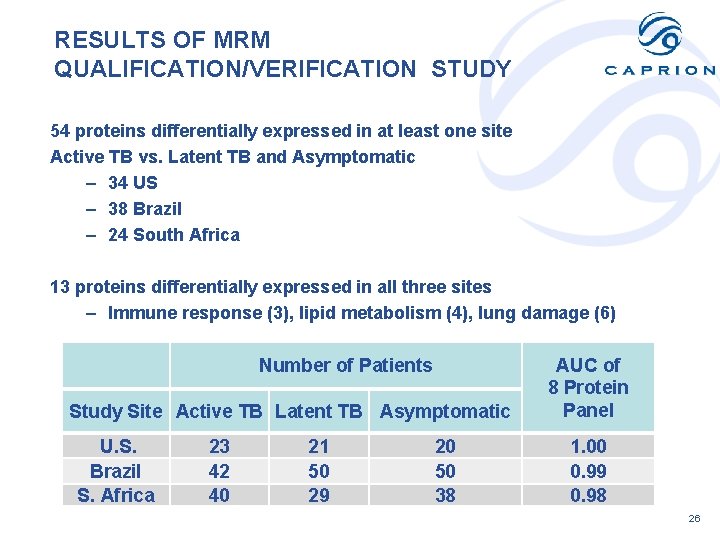
RESULTS OF MRM QUALIFICATION/VERIFICATION STUDY 54 proteins differentially expressed in at least one site Active TB vs. Latent TB and Asymptomatic – 34 US – 38 Brazil – 24 South Africa 13 proteins differentially expressed in all three sites – Immune response (3), lipid metabolism (4), lung damage (6) Number of Patients Study Site Active TB Latent TB Asymptomatic U. S. Brazil S. Africa 23 42 40 21 50 29 20 50 38 AUC of 8 Protein Panel 1. 00 0. 99 0. 98 26

Case Study 2: Targeted Discovery of Biomarkers by MRM in Plasma Markers of H 5 N 1 vaccine response 27

DISCOVERY STUDY DESIGN: SAMPLES NIAID-funded clinical trial involving 451 healthy adults • 291 samples from 97 patients receiving the highest dose (90 µg) of inactivated subvirion influenza A (H 5 N 1) vaccine: 1. Baseline (pre-vaccination) 2. Samples taken 28 days post-dose 1 3. Samples taken 28 days post-dose 2 • Immunogenicity determined by hemagglutination inhibition (HAI) and microneutralization (MN) • Response: increase in HAI or MN titer by 4 -fold or more between the baseline and post-dose 2 values GENDER Female Male SEASONAL FLU VACCINE No No Yes Yes RESPONSE NON-RESPONDER RESPONDER NUMBER OF PATIENTS 7 22 11 11 10 13 13 10 28

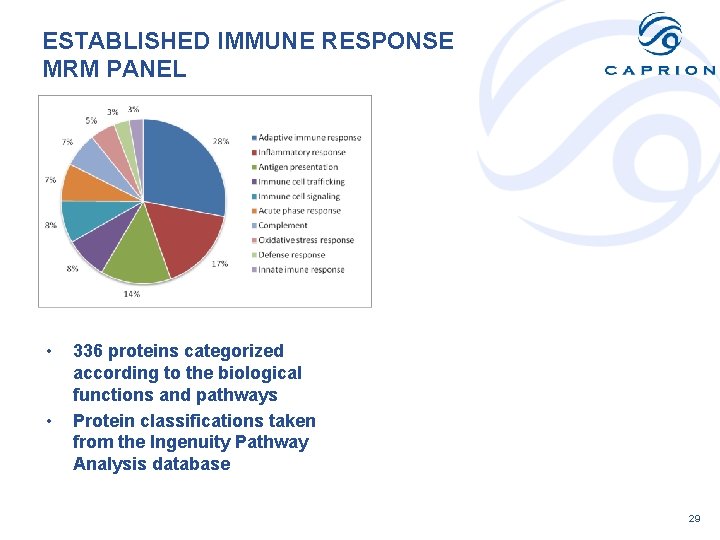
ESTABLISHED IMMUNE RESPONSE MRM PANEL • • 336 proteins categorized according to the biological functions and pathways Protein classifications taken from the Ingenuity Pathway Analysis database 29

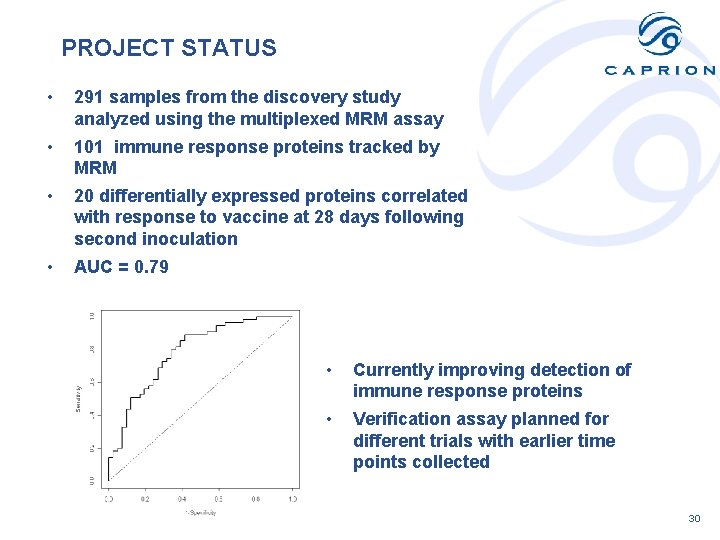
PROJECT STATUS • 291 samples from the discovery study analyzed using the multiplexed MRM assay • 101 immune response proteins tracked by MRM • 20 differentially expressed proteins correlated with response to vaccine at 28 days following second inoculation • AUC = 0. 79 • Currently improving detection of immune response proteins • Verification assay planned for different trials with earlier time points collected 30

Case Study 3: Discovery in the Source Tissue Followed by MRM in Plasma a) Identify initial candidates in secretome of NSCLC tumors b) Detect and quantify circulating markers in plasma 31

DISTINGUISHING BENIGN FROM MALIGNANT LUNG NODULES Problem: • 150, 000 lung nodules detected in the US each year – 60% are benign and 40% are malignant • Standard of care is CT scans performed every 6 months • If the nodule enlarges, a biopsy may be ordered Goal: • Develop blood-based test for early detection of small, malignant nodules to be used in conjunction with imaging Approach: • Identify 347 candidate markers for testing in patient plasma – Lung cancer tissue-based discovery studies – plasma membrane – secretory vesicles – Transcriptomics – Literature review • Establish MRM assay and screen patient plasma 32

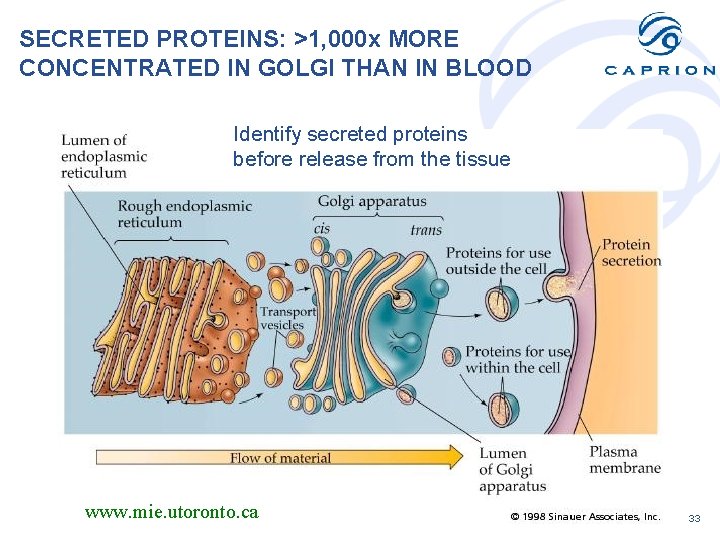
SECRETED PROTEINS: >1, 000 x MORE CONCENTRATED IN GOLGI THAN IN BLOOD Identify secreted proteins before release from the tissue www. mie. utoronto. ca 33

LUNG TUMOR SECRETED PROTEINS • Study Design – Collaboration with local hospitals – IRB approved, patient consented – 24 patients – Comparison of tumors and adjacent normal • Results – 1196 proteins identified in lung secretome – 200 up-regulated in tumor 34

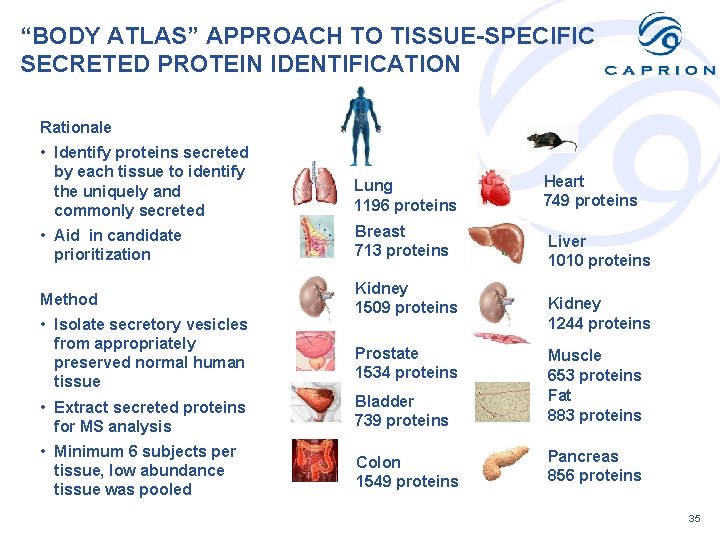
“BODY ATLAS” APPROACH TO TISSUE-SPECIFIC SECRETED PROTEIN IDENTIFICATION Rationale • Identify proteins secreted by each tissue to identify the uniquely and commonly secreted Lung 1196 proteins • Aid in candidate prioritization Breast 713 proteins Method Kidney 1509 proteins • Isolate secretory vesicles from appropriately preserved normal human tissue Prostate 1534 proteins Heart 749 proteins Liver 1010 proteins Kidney 1244 proteins • Extract secreted proteins for MS analysis Bladder 739 proteins Muscle 653 proteins Fat 883 proteins • Minimum 6 subjects per tissue, low abundance tissue was pooled Colon 1549 proteins Pancreas 856 proteins 35

90 -PROTEIN LUNG CANCER MRM PANEL • Development of MRM assay – Predict best 10 peptides/protein, synthesize – Determine empirically the best 2, monitor 2 transitions each • Pre-verification study – Depleted plasma from 20 cancer and 20 control subjects • Results – 64 targeted proteins (71%) successfully detected in cancer and control samples – Candidate classifiers identified 36

CANDIDATE CLASSIFIERS IDENTIFIED Current study: Full set of 347 candidate markers and more samples (>600) from multiple sources 37

Current Studies: Integrated Diagnostics Alzheimer’s Panel AD Biomarker Discovery in Plasma 38

ALZHEIMER’S DISEASE DIAGNOSTIC DISCOVERY • Initial study incorporated 50 candidate markers, tested on 60 samples – AD patients – healthy controls • Currently testing an expanded panel of 350 proteins on 300 samples – – AD patients MCI healthy controls other dementias) 39

MRM PANEL FROM INITIAL STUDY ACHIEVES 80% SENSITIVITY AND SPECIFICITY 40

Off the Shelf Assays: Plasma Proteins CSF Proteins Secreted Proteins, Various Tissues Rat Toxicology 41

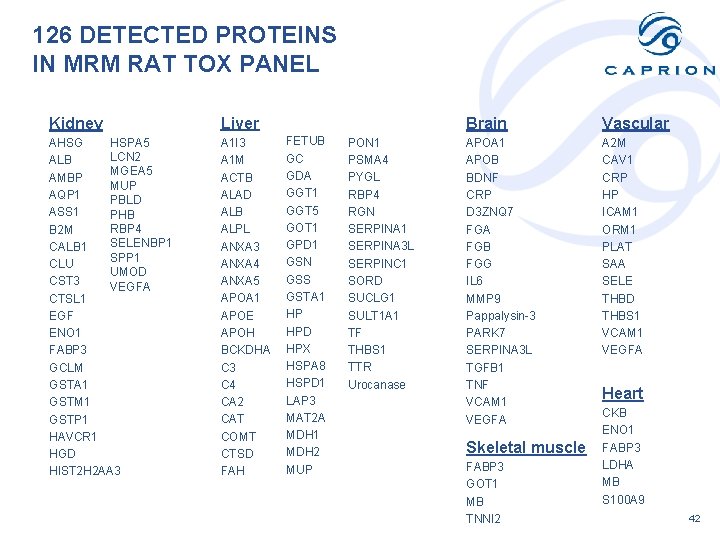
126 DETECTED PROTEINS IN MRM RAT TOX PANEL Kidney Liver HSPA 5 AHSG LCN 2 ALB MGEA 5 AMBP MUP AQP 1 PBLD ASS 1 PHB RBP 4 B 2 M SELENBP 1 CALB 1 SPP 1 CLU UMOD CST 3 VEGFA CTSL 1 EGF ENO 1 FABP 3 GCLM GSTA 1 GSTM 1 GSTP 1 HAVCR 1 HGD HIST 2 H 2 AA 3 A 1 I 3 A 1 M ACTB ALAD ALB ALPL ANXA 3 ANXA 4 ANXA 5 APOA 1 APOE APOH BCKDHA C 3 C 4 CA 2 CAT COMT CTSD FAH FETUB GC GDA GGT 1 GGT 5 GOT 1 GPD 1 GSN GSS GSTA 1 HP HPD HPX HSPA 8 HSPD 1 LAP 3 MAT 2 A MDH 1 MDH 2 MUP PON 1 PSMA 4 PYGL RBP 4 RGN SERPINA 1 SERPINA 3 L SERPINC 1 SORD SUCLG 1 SULT 1 A 1 TF THBS 1 TTR Urocanase Brain Vascular APOA 1 APOB BDNF CRP D 3 ZNQ 7 FGA FGB FGG IL 6 MMP 9 Pappalysin-3 PARK 7 SERPINA 3 L TGFB 1 TNF VCAM 1 VEGFA A 2 M CAV 1 CRP HP ICAM 1 ORM 1 PLAT SAA SELE THBD THBS 1 VCAM 1 VEGFA Skeletal muscle FABP 3 GOT 1 MB TNNI 2 Heart CKB ENO 1 FABP 3 LDHA MB S 100 A 9 42
