New data and tools at TAIR The Arabidopsis

New data and tools at TAIR (The Arabidopsis Information Resource)
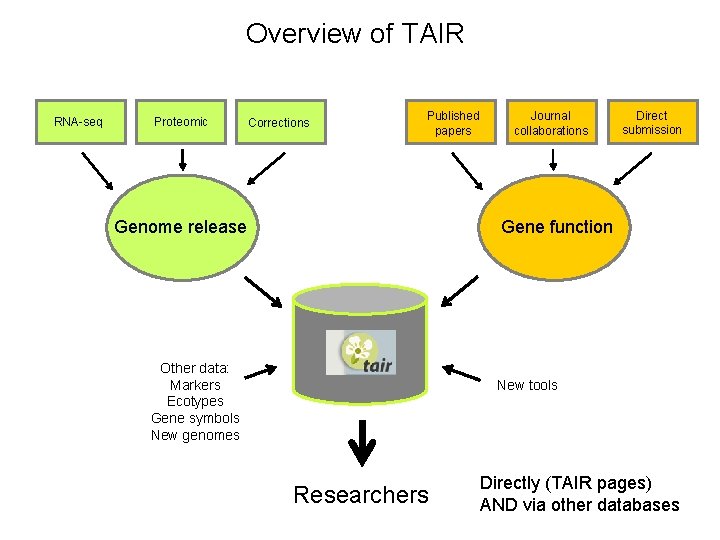
Overview of TAIR RNA-seq Proteomic Corrections Published papers Genome release Journal collaborations Direct submission Gene function Other data: Markers Ecotypes Gene symbols New genomes New tools Researchers Directly (TAIR pages) AND via other databases
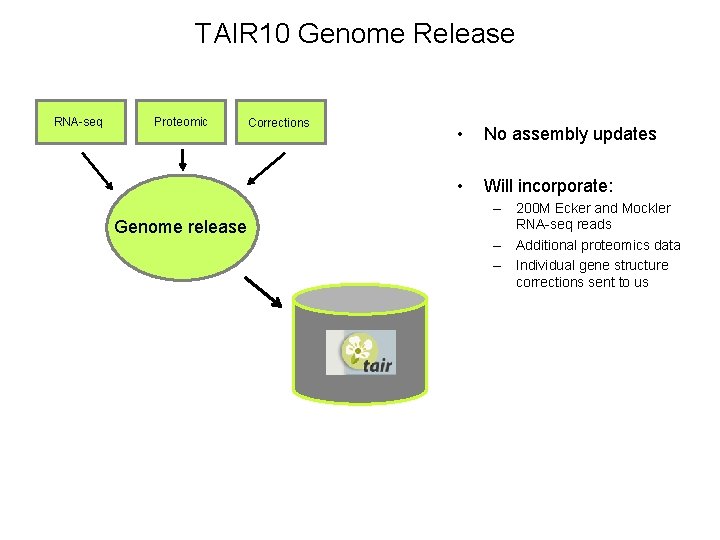
TAIR 10 Genome Release RNA-seq Proteomic Genome release Corrections • No assembly updates • Will incorporate: – 200 M Ecker and Mockler RNA-seq reads – Additional proteomics data – Individual gene structure corrections sent to us

Mapping and Assembly 1. Mapping • • RNA-seq sequences (Tophat (C. Trapnell), Supersplat (T. C. Mockler)) Peptides (6 -frame translation, spliced exon graph) 2. Assembly approaches • Augustus (M. Stanke) o o Uses spliced RNA seq reads, peptides Aim: Identify additional splice-variants, update existing genes • TAU (T. C. Mockler) o o Uses spliced RNA seq reads Aim: Identify additional splice-variants • Cufflinks (C. Trapnell) o o Uses spliced and unspliced RNA seq data Aim: Identify novel genes

Preliminary Results Augustus/TAU/Cufflinks predicted models are classified into categories: Novel genes Updated genes Splice-variants B-list Rejects 21 812 2134 1586 2318
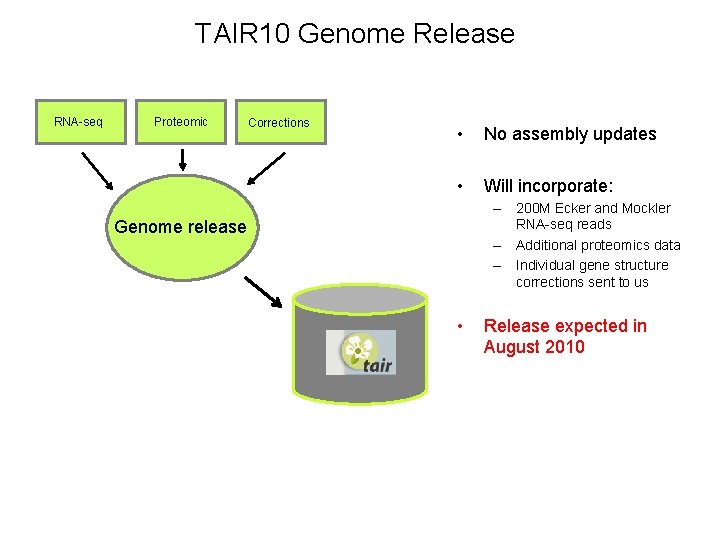
TAIR 10 Genome Release RNA-seq Proteomic Corrections • No assembly updates • Will incorporate: – 200 M Ecker and Mockler RNA-seq reads – Additional proteomics data – Individual gene structure corrections sent to us Genome release • Release expected in August 2010
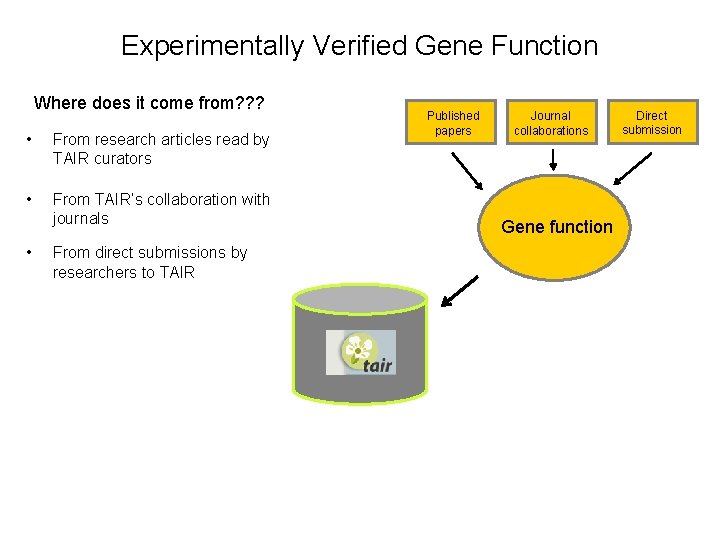
Experimentally Verified Gene Function Where does it come from? ? ? • From research articles read by TAIR curators • From TAIR’s collaboration with journals • From direct submissions by researchers to TAIR Published papers Journal collaborations Gene function Direct submission

Literature Curation • How? – Papers are prioritized according to novelty of gene function results – Highest priority papers are read and gene function is extracted • Why? – A lot of high quality experimental gene function information is only available in the form of articles • How many? – About 1/3 of all new articles containing gene function data are curated at TAIR each year Published papers Gene function
Journal Collaboration • How? Journal collaborations – Author instructions, Excel sheet or online form • Why? – To capture a larger fraction of gene function data – Because publication is the right time to get the data into TAIR • What journals? Gene function

Journal Collaboration
Journal Collaboration • How? Journal collaborations – Author instructions, Excel sheet or online form • Why? Gene function – To capture a larger fraction of gene function data – Because publication is the right time to get the data into TAIR • What journals? Plant Physiology (2008) The Plant Journal (2009) 2010: Journal of Integrative Plant Biology Journal of Experimental Botany Plant Science Environmental Botany Plant Physiology and Biochemistry Plant, Cell and Environment
Direct Submission of Gene Function • Direct submission How? – Excel sheet or online form • Why? – To capture more data with a small curation team – Because researchers are the experts on the genes they study Gene function
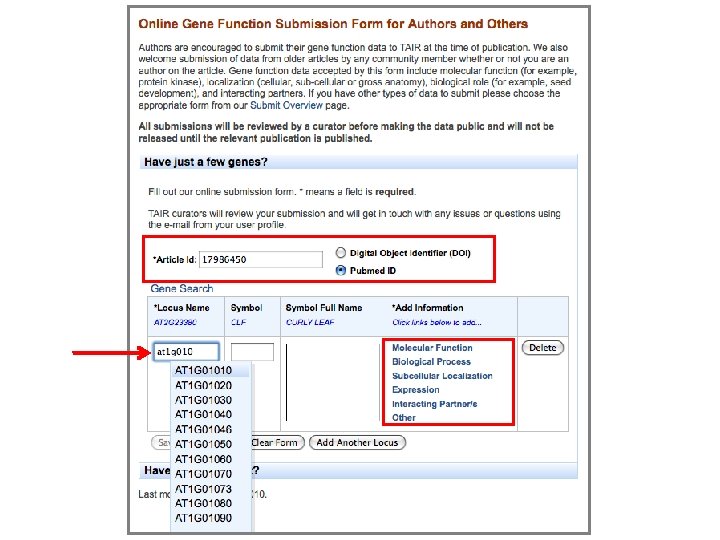
New online submission form 17986450

Why Gene Ontology? • Standardization allows comparison across experiments and species • Hierarchical structure allows high level categorization • Well structured ontology framework facilitates computational analysis • Attached to data source (peer reviewed published research) • Experimental evidence can be distinguished from predictions

Example Gene Ontology annotations Gene GO term Evidence Reference 3 GO flavors Phot 1 Phototropism Biological process Mutant phenotype Huala et al 1997 Phot 1 Cytoplasm Cellular component Direct assay Sakamoto et al 2002 Phot 1 Serine / threonine kinase activity Molecular function Direct assay Christie et al 1998
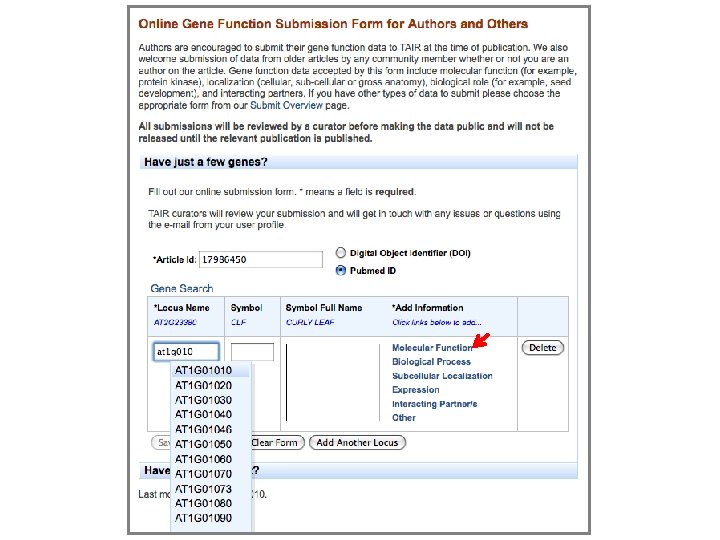
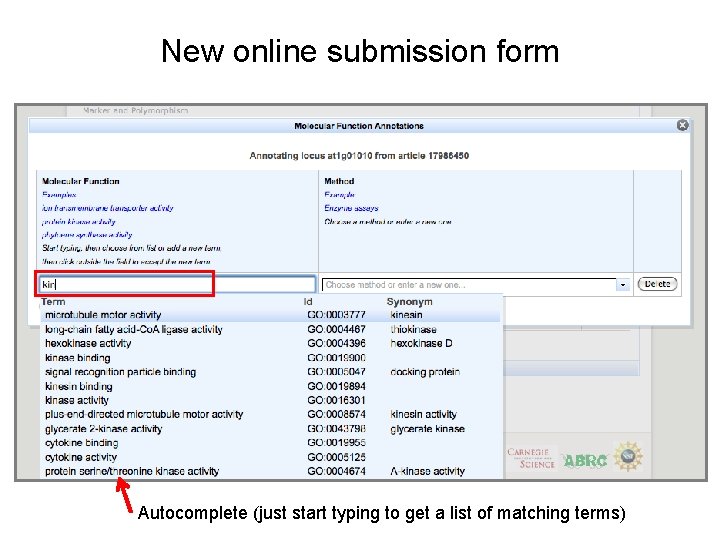
New online submission form Autocomplete (just start typing to get a list of matching terms)
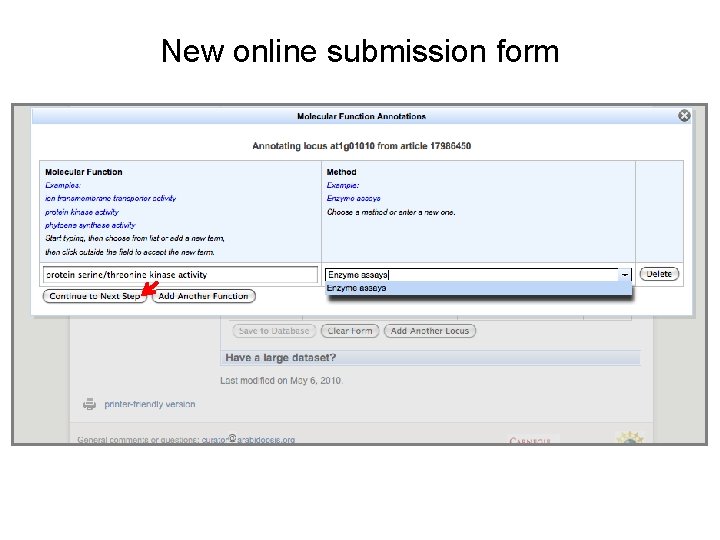
New online submission form
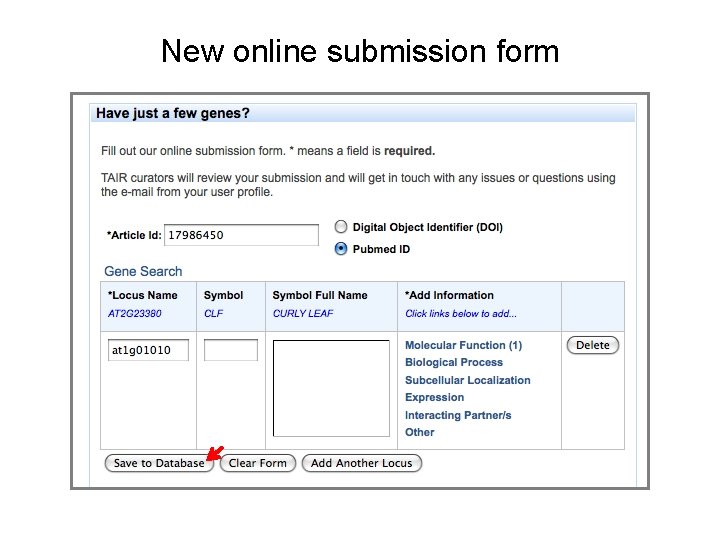
New online submission form
What is the result of TAIR’s effort to capture gene function? Published papers • Journal collaborations How many genes have experimental gene function in TAIR? Gene function Direct submission
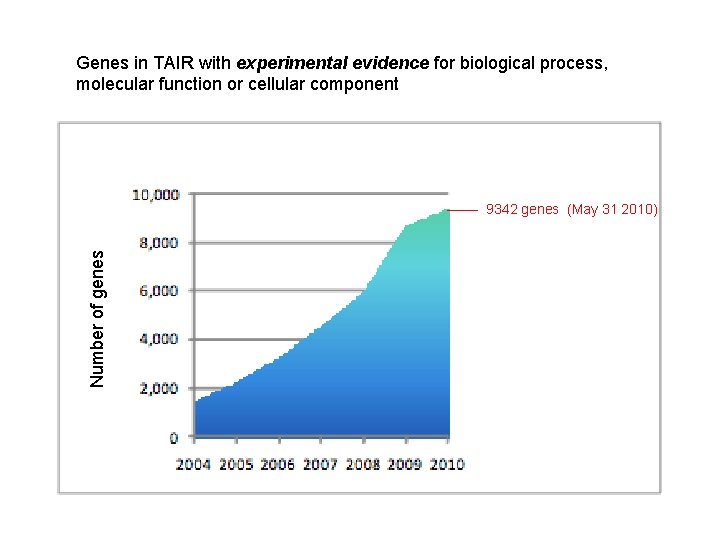
Genes in TAIR with experimental evidence for biological process, molecular function or cellular component Number of genes 9342 genes (May 31 2010)
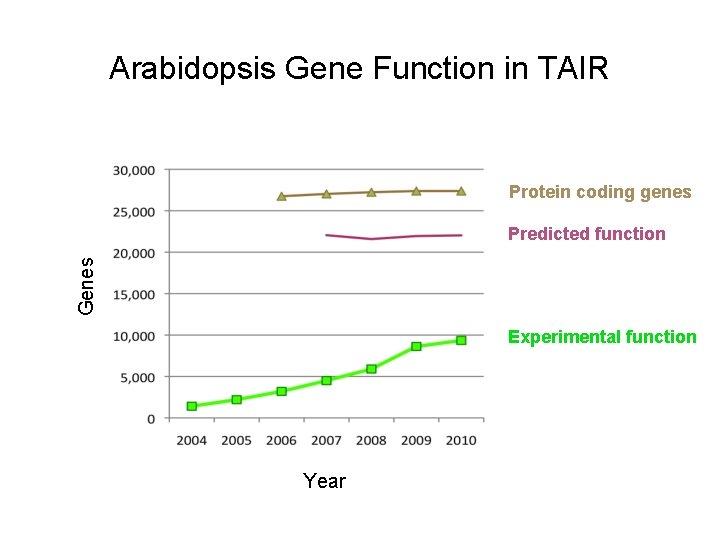
Arabidopsis Gene Function in TAIR Protein coding genes Genes Predicted function Experimental function Year
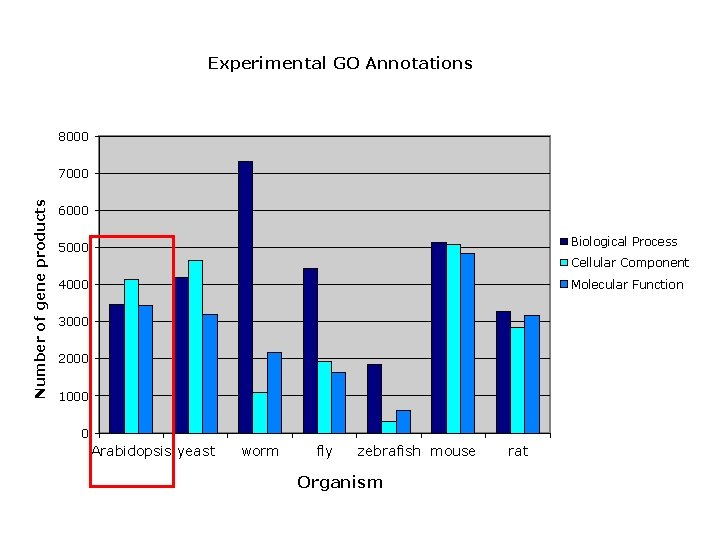
Experimental GO Annotations 8000 Number of gene products 7000 6000 Biological Process 5000 Cellular Component Molecular Function 4000 3000 2000 1000 0 Arabidopsis yeast worm fly zebrafish mouse Organism rat
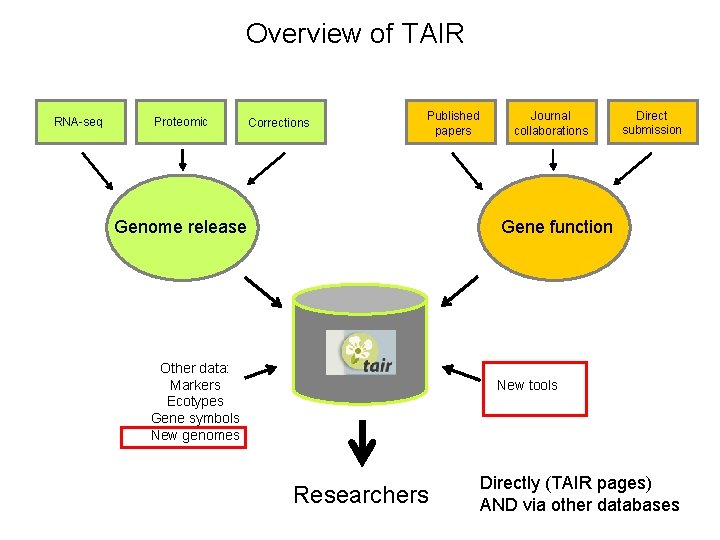
Overview of TAIR RNA-seq Proteomic Corrections Published papers Genome release Journal collaborations Direct submission Gene function Other data: Markers Ecotypes Gene symbols New genomes New tools Researchers Directly (TAIR pages) AND via other databases
GBrowse_syn Tool by Sheldon Mc. Kay, CSHL Alignment data from Pedro Pattyn, Van de Peer lab, U. of Ghent
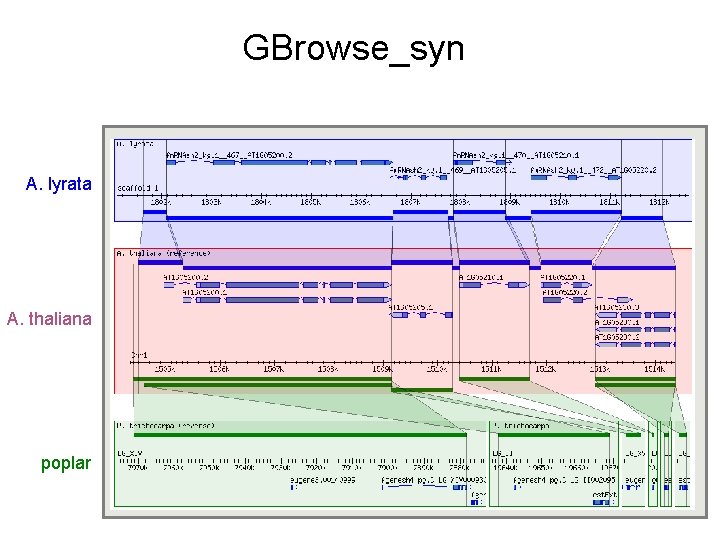
GBrowse_syn A. lyrata A. thaliana poplar
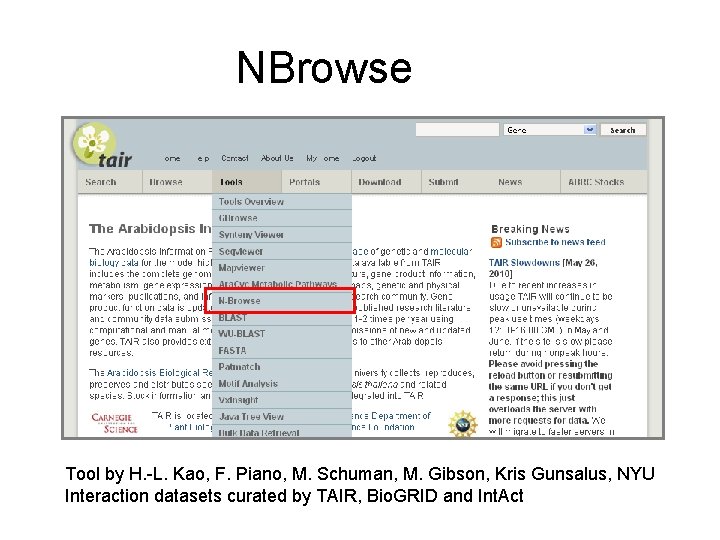
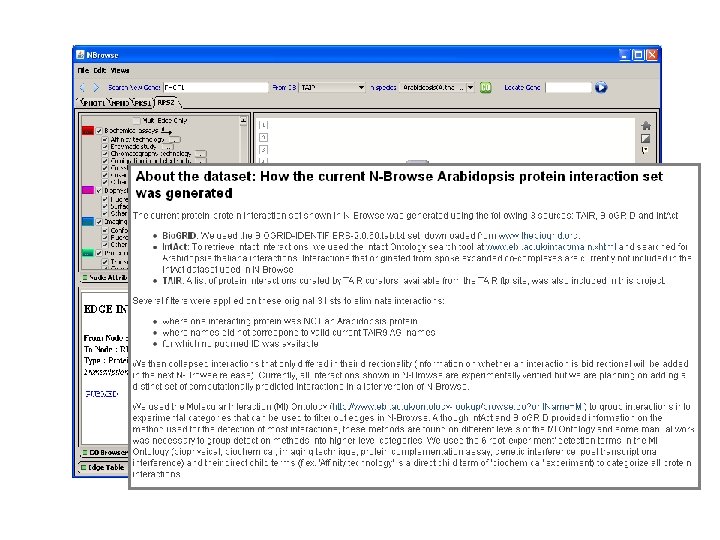
NBrowse Tool by H. -L. Kao, F. Piano, M. Schuman, M. Gibson, Kris Gunsalus, NYU Interaction datasets curated by TAIR, Bio. GRID and Int. Act
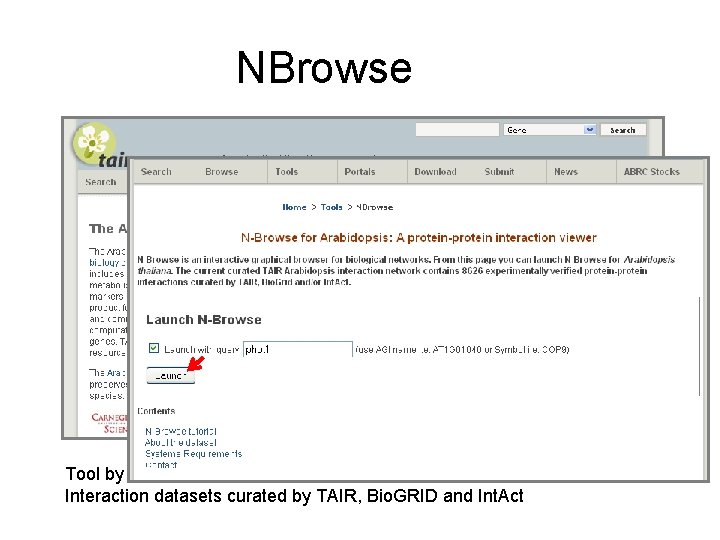
NBrowse Tool by H. -L. Kao, F. Piano, M. Schuman, M. Gibson, Kris Gunsalus, NYU Interaction datasets curated by TAIR, Bio. GRID and Int. Act
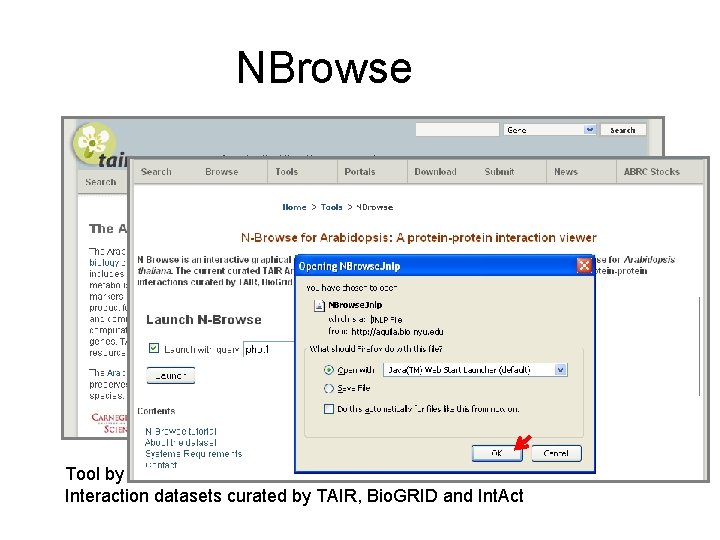
NBrowse Tool by H. -L. Kao, F. Piano, M. Schuman, M. Gibson, Kris Gunsalus, NYU Interaction datasets curated by TAIR, Bio. GRID and Int. Act
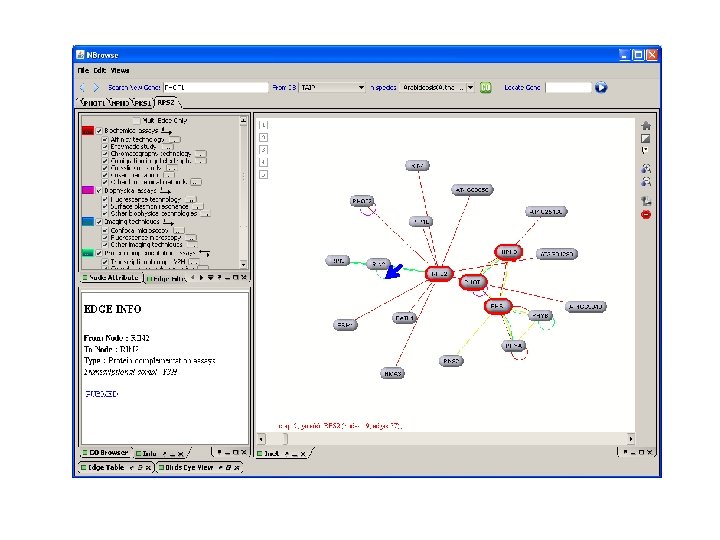

Arabidopsis lyrata Genes have been loaded Working on adding some gene function information and improving searching
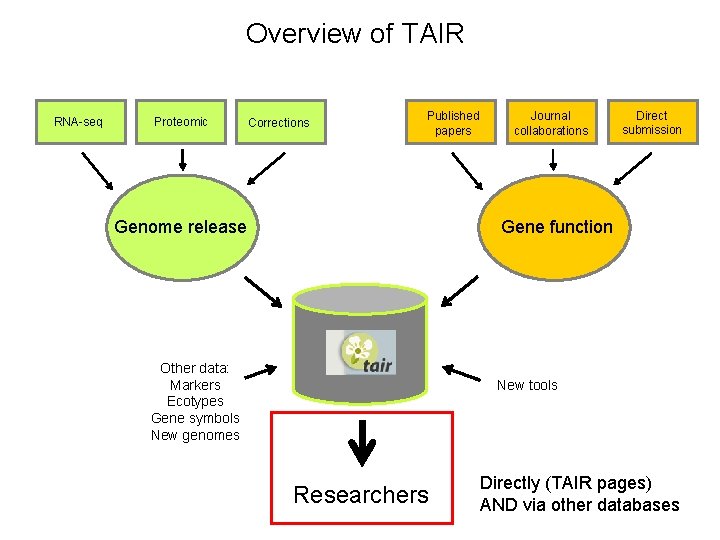
Overview of TAIR RNA-seq Proteomic Corrections Published papers Genome release Journal collaborations Direct submission Gene function Other data: Markers Ecotypes Gene symbols New genomes New tools Researchers Directly (TAIR pages) AND via other databases
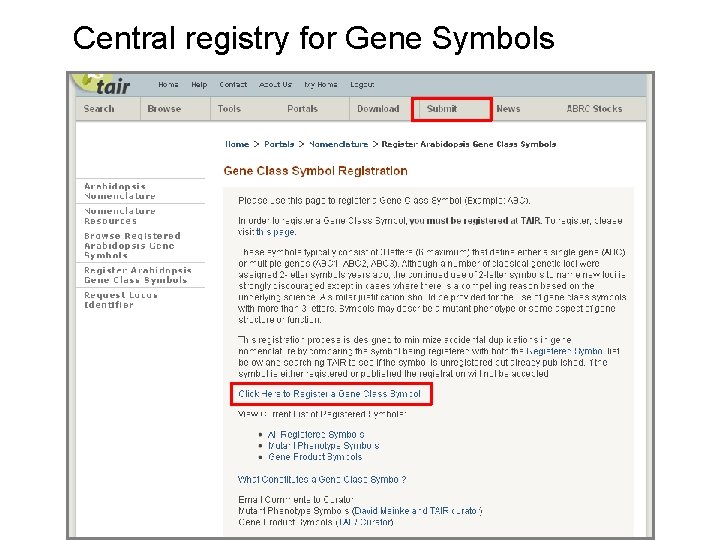
Central registry for Gene Symbols
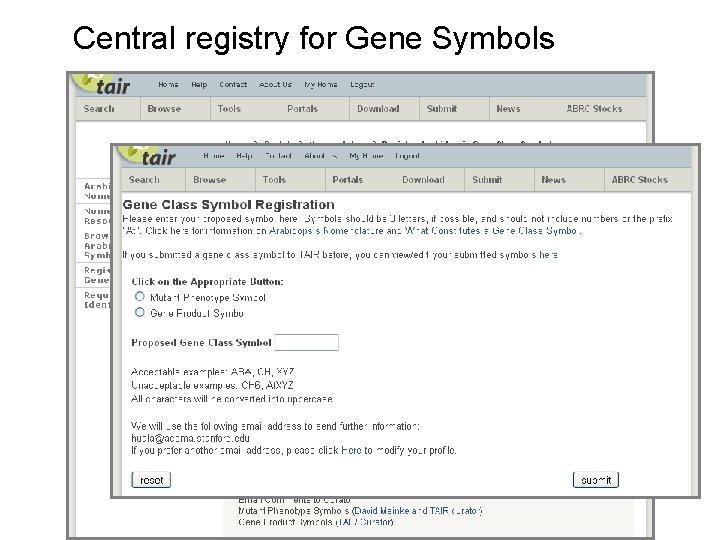
Central registry for Gene Symbols
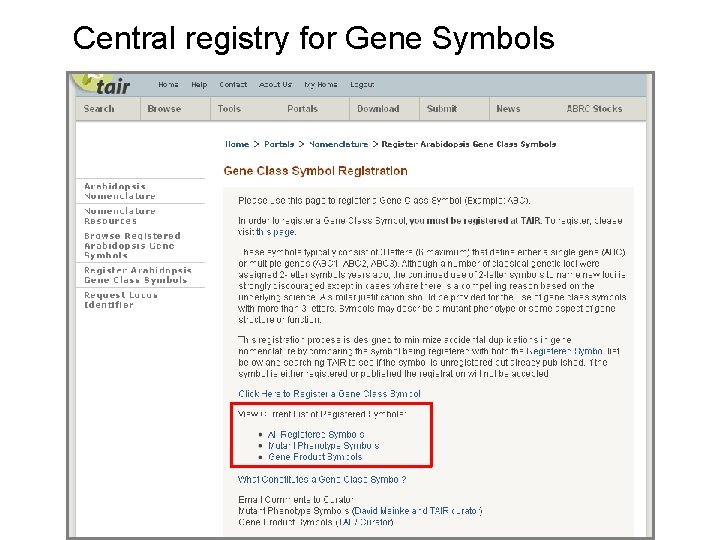
Central registry for Gene Symbols
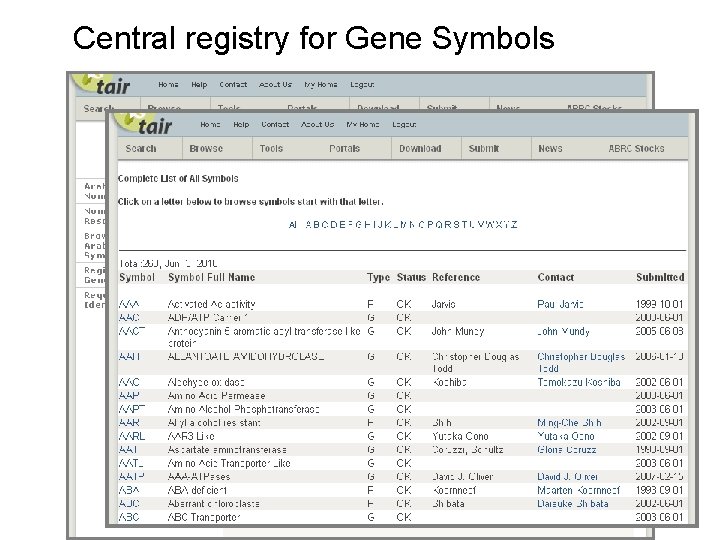
Central registry for Gene Symbols
Helpdesk
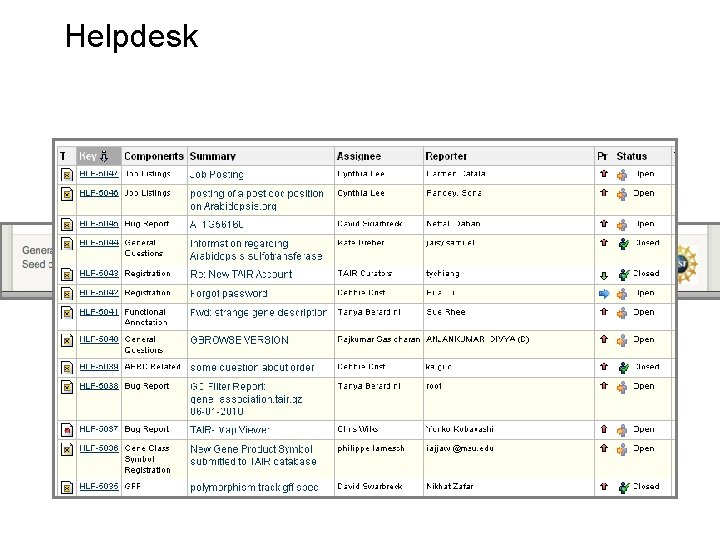
Helpdesk
Helpdesk
RSS news feed
RSS news feed

TAIR Facebook Page
TAIR Twitter Feed

TAIR Staff Genome Annotation: Gene Function/GO: ? David Swarbreck Philippe Lamesch Rajkumar Sasidharan Tanya Berardini Donghui Li Tech Team: Bob Muller Larry Ploetz Shanker Singh Chris Wilks (50%) Cynthia Lee
Host Institution: Funding Agencies: TAIR Sponsors: Partner:
- Slides: 47