NEUTRON CRYSTAL STRUCTURE OF TRIOSEPHOSPHATE ISOMERASE TIM COMPLEXED




- Slides: 13

NEUTRON CRYSTAL STRUCTURE OF TRIOSEPHOSPHATE ISOMERASE (TIM) COMPLEXED WITH ITS INHIBITOR: PGH ESS Science Day – June 14 th Bénédicte LAFUMAT L. mexicana TIM dimer 1/11

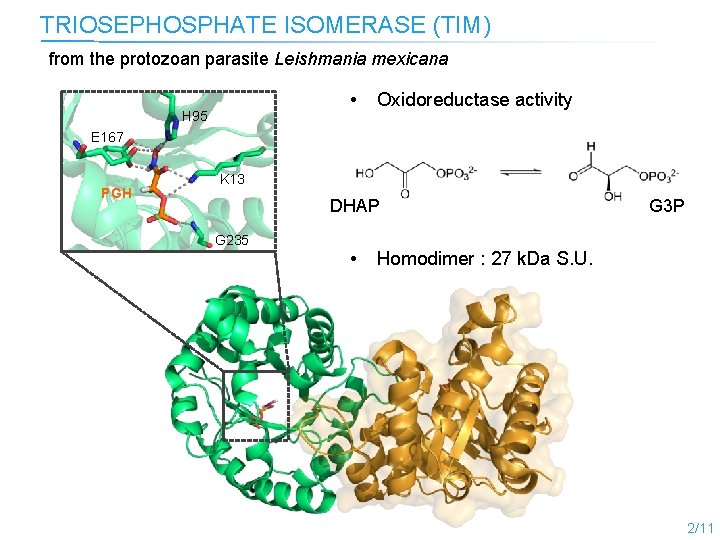
TRIOSEPHOSPHATE ISOMERASE (TIM) from the protozoan parasite Leishmania mexicana • H 95 Oxidoreductase activity E 167 PGH K 13 DHAP G 235 • G 3 P Homodimer : 27 k. Da S. U. 2/11

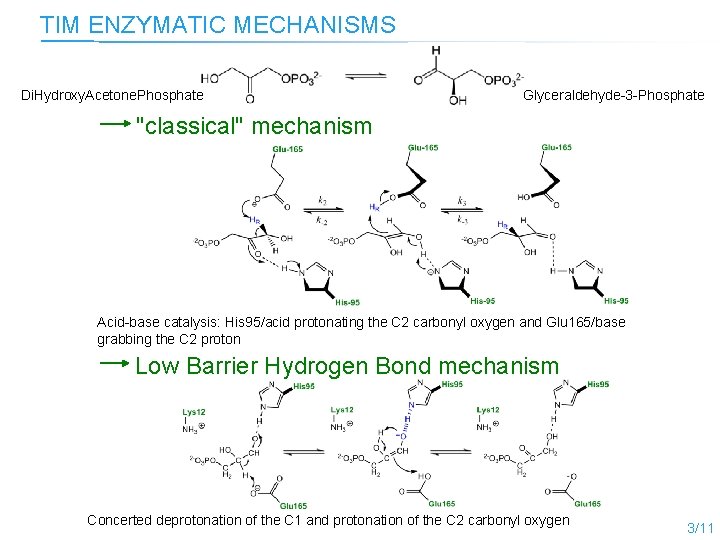
TIM ENZYMATIC MECHANISMS Di. Hydroxy. Acetone. Phosphate Glyceraldehyde-3 -Phosphate "classical" mechanism Acid-base catalysis: His 95/acid protonating the C 2 carbonyl oxygen and Glu 165/base grabbing the C 2 proton Low Barrier Hydrogen Bond mechanism Concerted deprotonation of the C 1 and protonation of the C 2 carbonyl oxygen 3/11

KEY QUESTION H 95 E 167 K 13 PGH G 235 Is TIM processing the “classical” or the “LBHB” enzymatic mechanism? Protonation state of the Glu 235, His 95 and Lys 13 and PGH inhibitor? 4/11

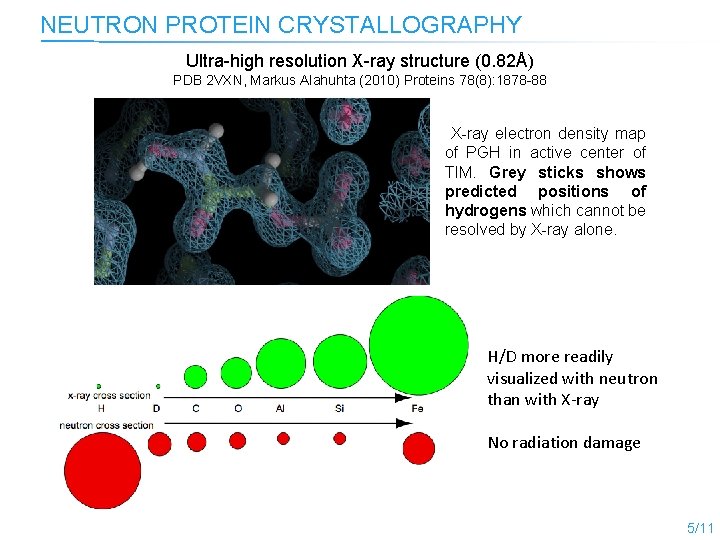
NEUTRON PROTEIN CRYSTALLOGRAPHY Ultra-high resolution X-ray structure (0. 82Å) PDB 2 VXN, Markus Alahuhta (2010) Proteins 78(8): 1878 -88 X-ray electron density map of PGH in active center of TIM. Grey sticks shows predicted positions of hydrogens which cannot be resolved by X-ray alone. H/D more readily visualized with neutron than with X-ray No radiation damage 5/11

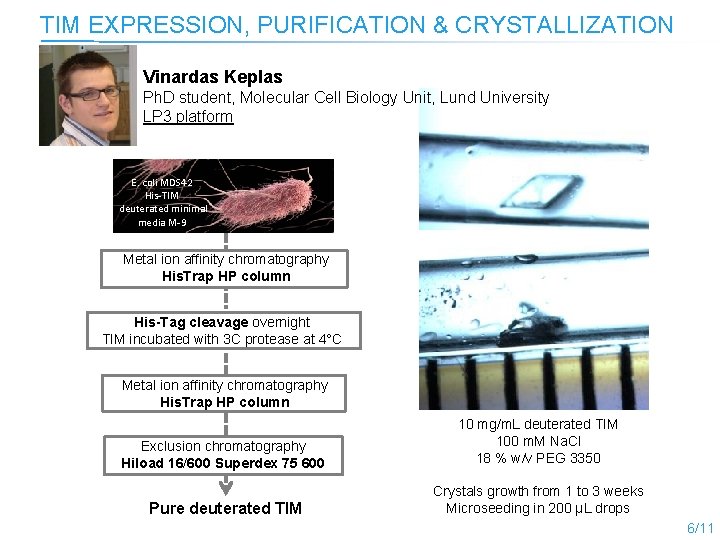
TIM EXPRESSION, PURIFICATION & CRYSTALLIZATION Vinardas Keplas Ph. D student, Molecular Cell Biology Unit, Lund University LP 3 platform E. coli MDS 42 His-TIM deuterated minimal media M-9 Metal ion affinity chromatography His. Trap HP column His-Tag cleavage overnight TIM incubated with 3 C protease at 4°C Metal ion affinity chromatography His. Trap HP column Exclusion chromatography Hiload 16/600 Superdex 75 600 10 mg/m. L deuterated TIM 100 m. M Na. Cl 18 % w/v PEG 3350 Pure deuterated TIM Crystals growth from 1 to 3 weeks Microseeding in 200 µL drops 6/11

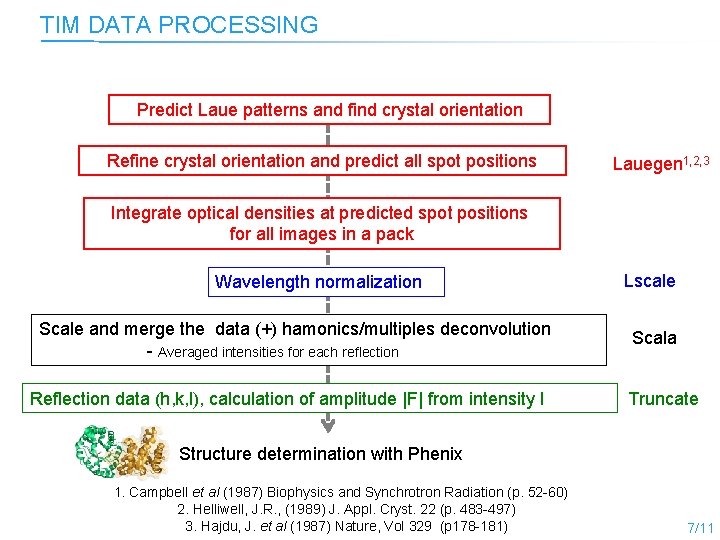
TIM DATA PROCESSING Predict Laue patterns and find crystal orientation Refine crystal orientation and predict all spot positions Lauegen 1, 2, 3 Integrate optical densities at predicted spot positions for all images in a pack Wavelength normalization Scale and merge the data (+) hamonics/multiples deconvolution - Averaged intensities for each reflection Reflection data (h, k, l), calculation of amplitude |F| from intensity I Lscale Scala Truncate Structure determination with Phenix 1. Campbell et al (1987) Biophysics and Synchrotron Radiation (p. 52 -60) 2. Helliwell, J. R. , (1989) J. Appl. Cryst. 22 (p. 483 -497) 3. Hajdu, J. et al (1987) Nature, Vol 329 (p 178 -181) 7/11

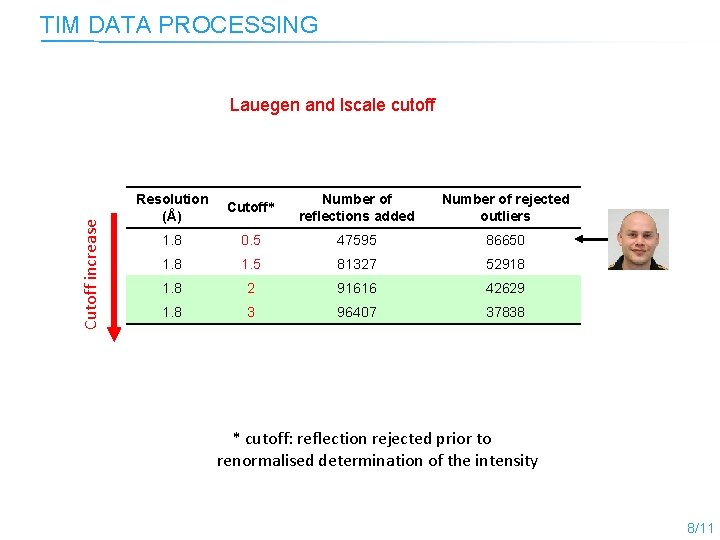
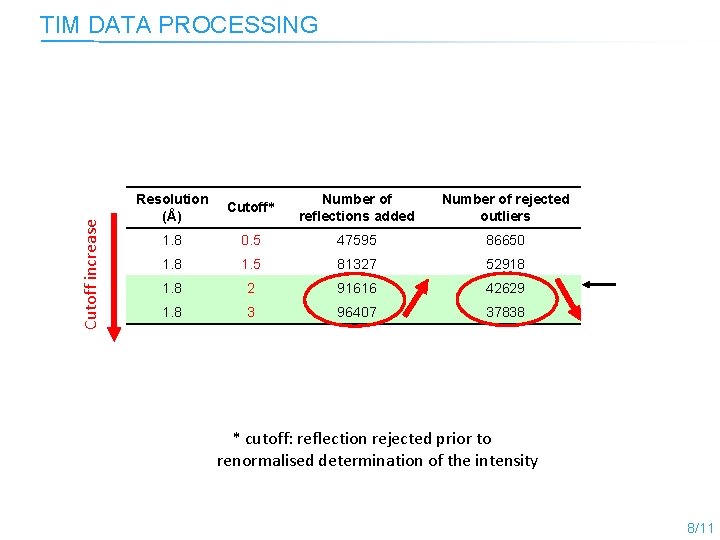
TIM DATA PROCESSING Cutoff increase Lauegen and lscale cutoff Resolution (Å) Cutoff* Number of reflections added Number of rejected outliers 1. 8 0. 5 47595 86650 1. 8 1. 5 81327 52918 1. 8 2 91616 42629 1. 8 3 96407 37838 * cutoff: reflection rejected prior to renormalised determination of the intensity 8/11

Cutoff increase TIM DATA PROCESSING Resolution (Å) Cutoff* Number of reflections added Number of rejected outliers 1. 8 0. 5 47595 86650 1. 8 1. 5 81327 52918 1. 8 2 91616 42629 1. 8 3 96407 37838 * cutoff: reflection rejected prior to renormalised determination of the intensity 8/11

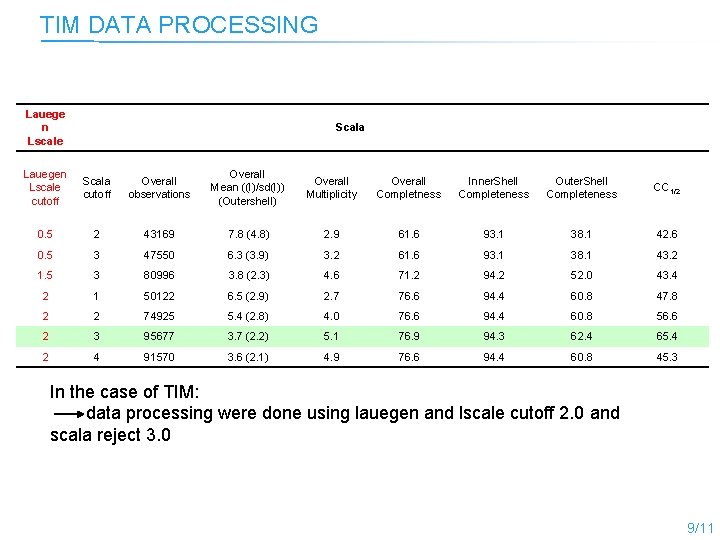
TIM DATA PROCESSING Lauege n Lscale Scala Lauegen Lscale cutoff Scala cutoff Overall observations Overall Mean ((I)/sd(I)) (Outershell) Overall Multiplicity Overall Completness Inner. Shell Completeness Outer. Shell Completeness CC 1/2 0. 5 2 43169 7. 8 (4. 8) 2. 9 61. 6 93. 1 38. 1 42. 6 0. 5 3 47550 6. 3 (3. 9) 3. 2 61. 6 93. 1 38. 1 43. 2 1. 5 3 80996 3. 8 (2. 3) 4. 6 71. 2 94. 2 52. 0 43. 4 2 1 50122 6. 5 (2. 9) 2. 7 76. 6 94. 4 60. 8 47. 8 2 2 74925 5. 4 (2. 8) 4. 0 76. 6 94. 4 60. 8 56. 6 2 3 95677 3. 7 (2. 2) 5. 1 76. 9 94. 3 62. 4 65. 4 2 4 91570 3. 6 (2. 1) 4. 9 76. 6 94. 4 60. 8 45. 3 In the case of TIM: data processing were done using lauegen and lscale cutoff 2. 0 and scala reject 3. 0 9/11

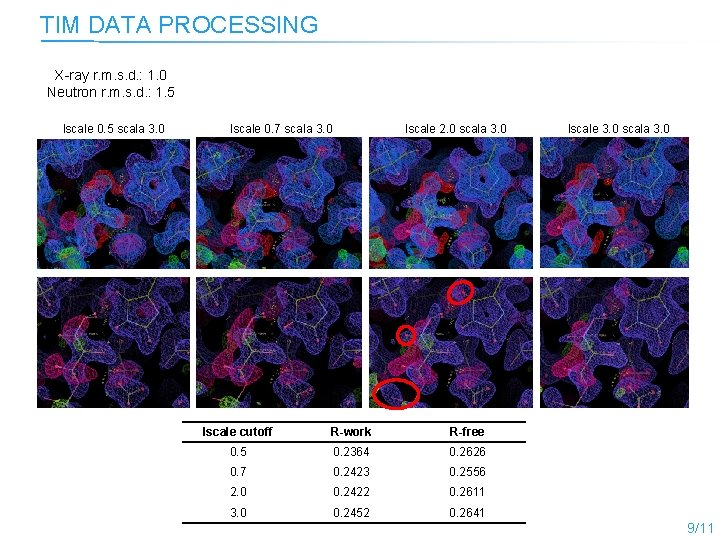
TIM DATA PROCESSING X-ray r. m. s. d. : 1. 0 Neutron r. m. s. d. : 1. 5 lscale 0. 5 scala 3. 0 lscale 0. 7 scala 3. 0 lscale 2. 0 scala 3. 0 lscale cutoff R-work R-free 0. 5 0. 2364 0. 2626 0. 7 0. 2423 0. 2556 2. 0 0. 2422 0. 2611 3. 0 0. 2452 0. 2641 lscale 3. 0 scala 3. 0 9/11

CONCLUSION & PERSPECTIVES 1. Improvement of the data treatment: Lauegen, lscale and scala cutoff 2. Need of new data processing software developements to: Initially better orient the crystal Nicolas Coquelle: https: //github. com/coquellen Find the best compromise to solve the structure Rejection cutoff of reflections 3. TIM structure present possible tautomerization of PGH inhibitor 10/11

ACKNOWLEDGEMENTS Esko Oksanen NMX Scientific Project Leader, ESS Vinardas Keplas Ph. D student, Molecular Cell Biology Unit, Lund University Claes von Wachenfeldt Senior lecturer, Molecular Cell Biology Unit, Lund University Valeria Bugris Research Associate, BRC (Szeged, HU) Matthew Blakeley Nicolas Coquelle LADI-III responsible, ILL (Grenoble, FR) LADI-III co-responsible, ILL (Grenoble, FR) Zoë Fisher Head of Deuteration Macromolecular Crystallization (DEMAX), ESS All the NMX, DEMAX and LP 3 group Wolfgang Knecht, LP 3 Manager Maria Gourdon, LP 3 scientist Annika Rogstam, LP 3 scientist Katarina Koruza, Ph. D student 11/11