Neurosciences Research Building Mass Spectrometry FacilityDepartment of Pharmaceutical

蛋白质代谢的信息处理进展 Neurosciences Research Building 关慎恒 Mass Spectrometry Facility/Department of Pharmaceutical Chemistry, Institute for Neurodegenerative Diseases and Department of Neurology, University of California, San Francisco 第二届中国计算蛋白质组学研讨会
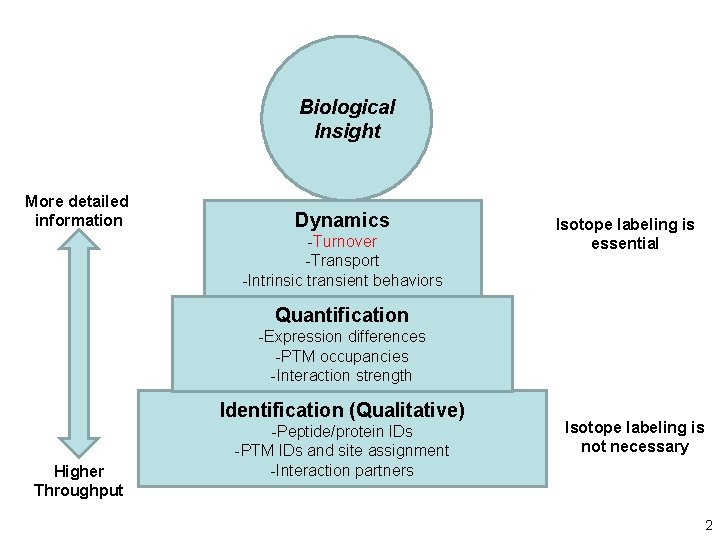
Biological Insight More detailed information Dynamics -Turnover -Transport -Intrinsic transient behaviors Isotope labeling is essential Quantification -Expression differences -PTM occupancies -Interaction strength Identification (Qualitative) Higher Throughput -Peptide/protein IDs -PTM IDs and site assignment -Interaction partners Isotope labeling is not necessary 2
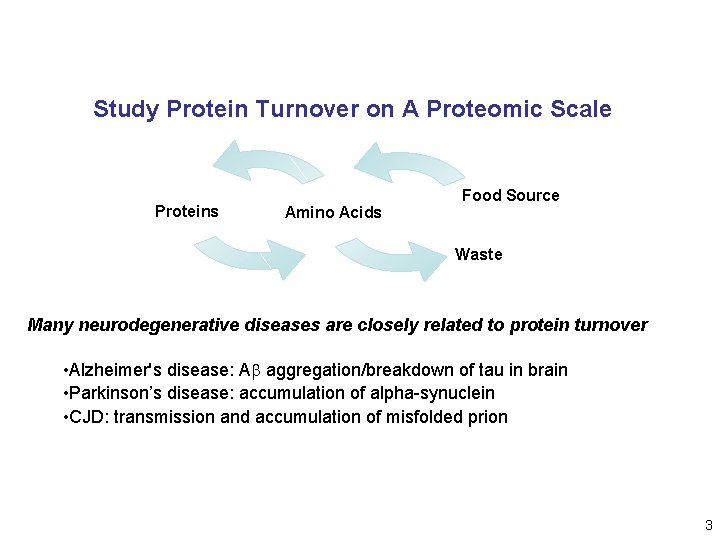
Study Protein Turnover on A Proteomic Scale Proteins Amino Acids Food Source Waste Many neurodegenerative diseases are closely related to protein turnover • Alzheimer's disease: Ab aggregation/breakdown of tau in brain • Parkinson’s disease: accumulation of alpha-synuclein • CJD: transmission and accumulation of misfolded prion 3
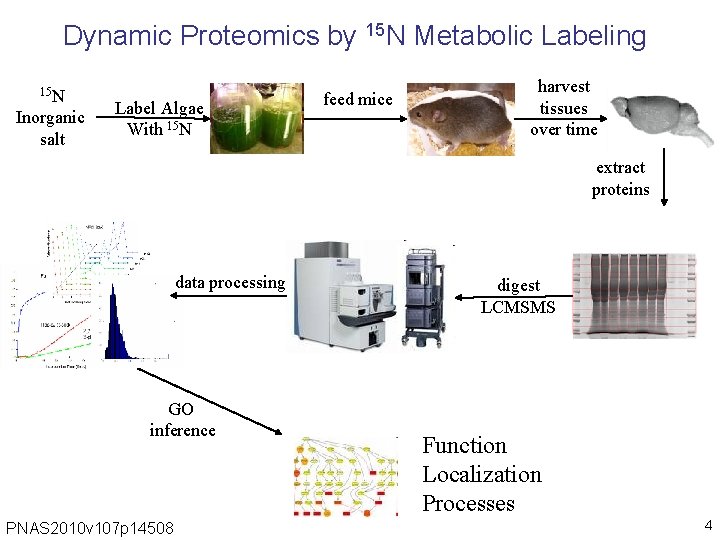
Dynamic Proteomics by 15 N Metabolic Labeling 15 N Inorganic salt Label Algae With 15 N feed mice harvest tissues over time extract proteins data processing GO inference PNAS 2010 v 107 p 14508 digest LCMSMS Function Localization Processes 4
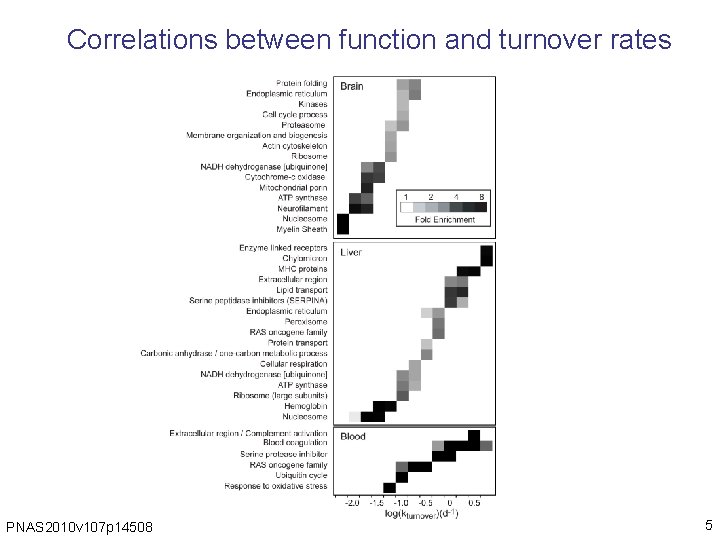
Correlations between function and turnover rates PNAS 2010 v 107 p 14508 5
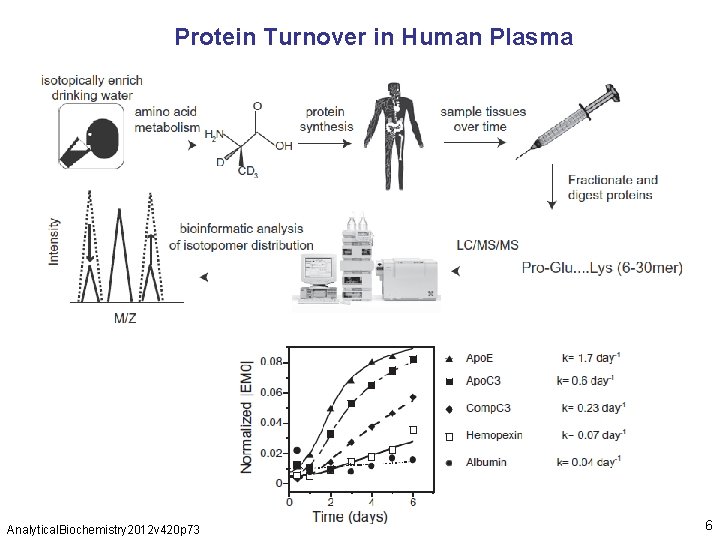
Protein Turnover in Human Plasma Analytical. Biochemistry 2012 v 420 p 73 6
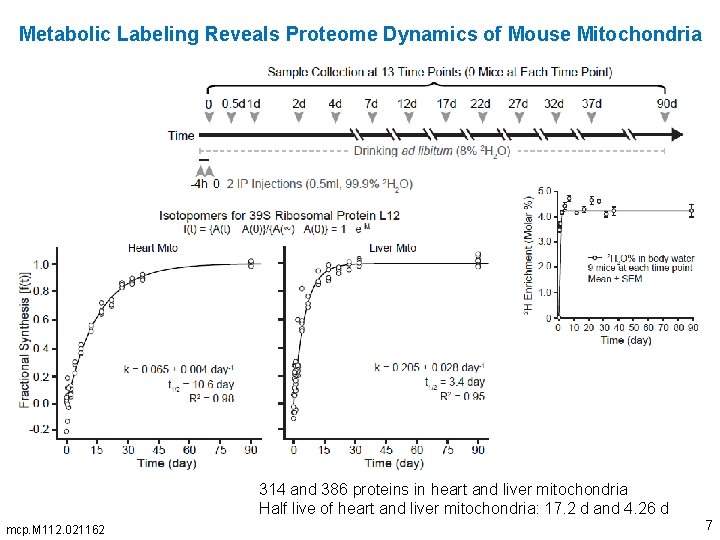
Metabolic Labeling Reveals Proteome Dynamics of Mouse Mitochondria 314 and 386 proteins in heart and liver mitochondria Half live of heart and liver mitochondria: 17. 2 d and 4. 26 d mcp. M 112. 021162 7
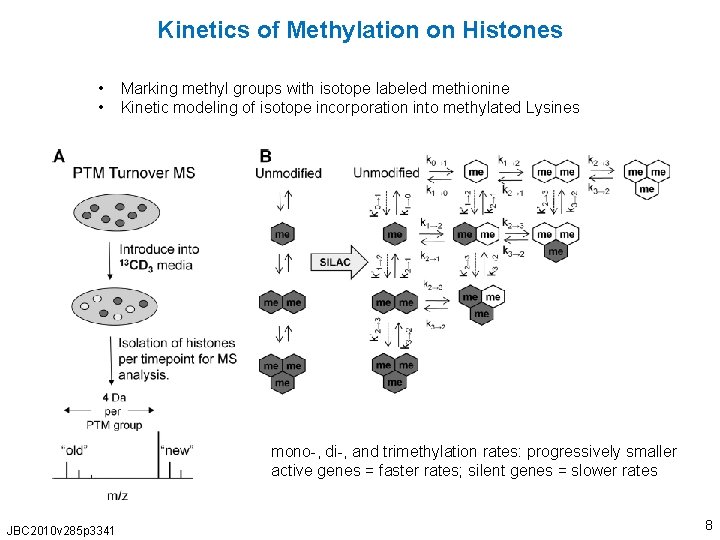
Kinetics of Methylation on Histones • • Marking methyl groups with isotope labeled methionine Kinetic modeling of isotope incorporation into methylated Lysines mono-, di-, and trimethylation rates: progressively smaller active genes = faster rates; silent genes = slower rates JBC 2010 v 285 p 3341 8
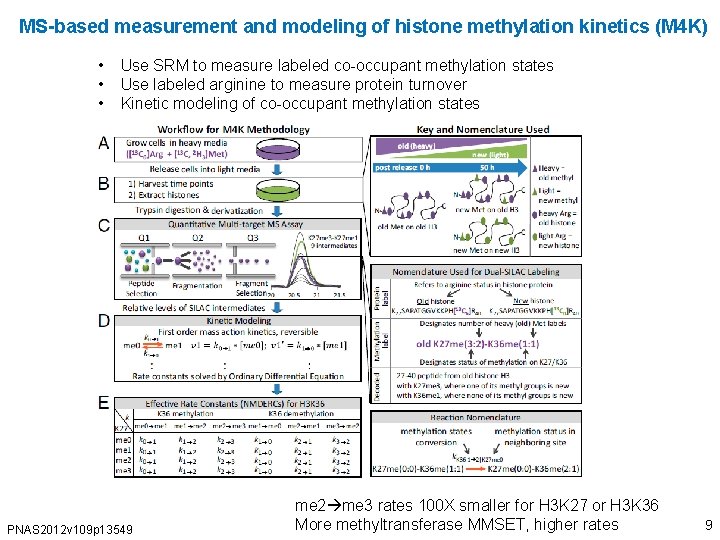
MS-based measurement and modeling of histone methylation kinetics (M 4 K) • • • Use SRM to measure labeled co-occupant methylation states Use labeled arginine to measure protein turnover Kinetic modeling of co-occupant methylation states PNAS 2012 v 109 p 13549 me 2 me 3 rates 100 X smaller for H 3 K 27 or H 3 K 36 More methyltransferase MMSET, higher rates 9
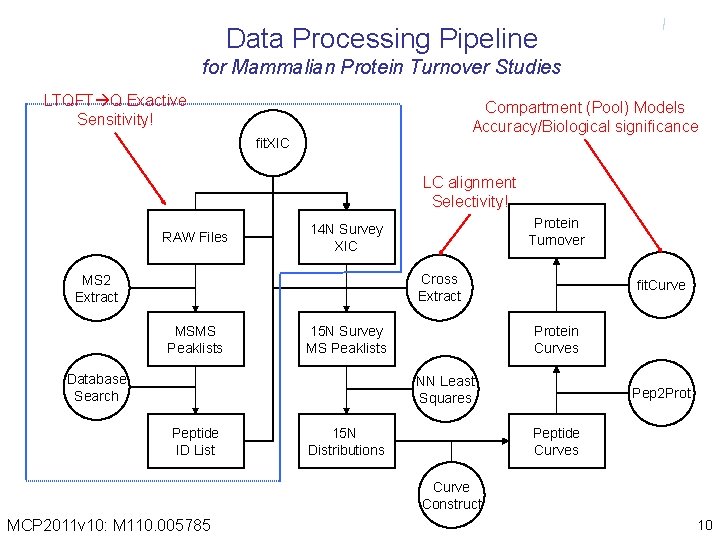
Data Processing Pipeline for Mammalian Protein Turnover Studies LTQFT Q Exactive Sensitivity! Compartment (Pool) Models Accuracy/Biological significance fit. XIC LC alignment Selectivity! RAW Files Protein Turnover 14 N Survey XIC Cross Extract MS 2 Extract MSMS Peaklists 15 N Survey MS Peaklists Database Search fit. Curve Protein Curves NN Least Squares Peptide ID List 15 N Distributions Pep 2 Prot Peptide Curves Curve Construct MCP 2011 v 10: M 110. 005785 10
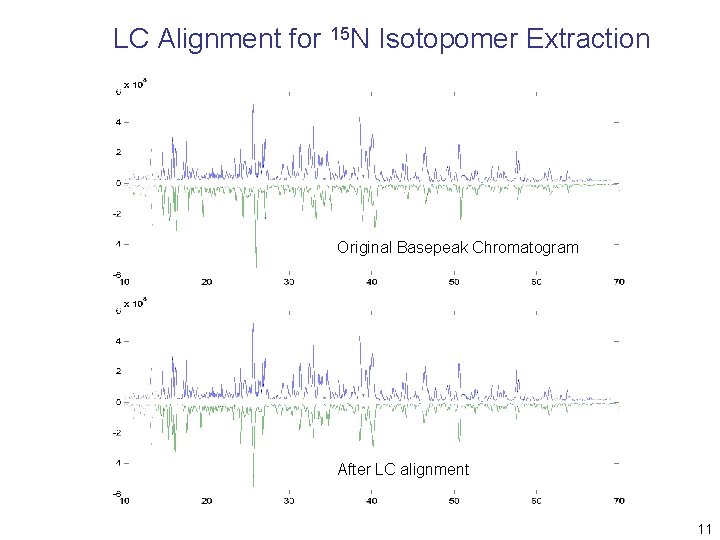
LC Alignment for 15 N Isotopomer Extraction Original Basepeak Chromatogram After LC alignment 11
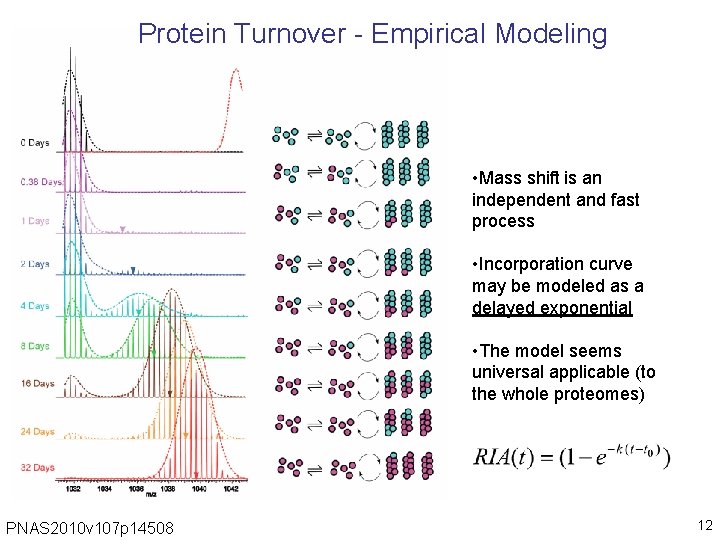
Protein Turnover - Empirical Modeling • Mass shift is an independent and fast process • Incorporation curve may be modeled as a delayed exponential • The model seems universal applicable (to the whole proteomes) PNAS 2010 v 107 p 14508 12
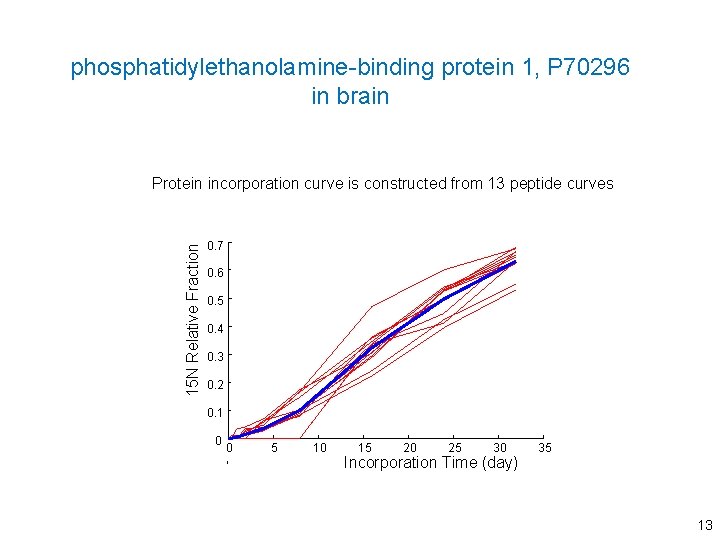
phosphatidylethanolamine-binding protein 1, P 70296 in brain 15 N Relative Fraction Protein incorporation curve is constructed from 13 peptide curves 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 0. 1 0 0 5 10 15 20 25 30 Incorporation Time (day) 35 13

JBiol. Chem 1939 v 130 p 703 14
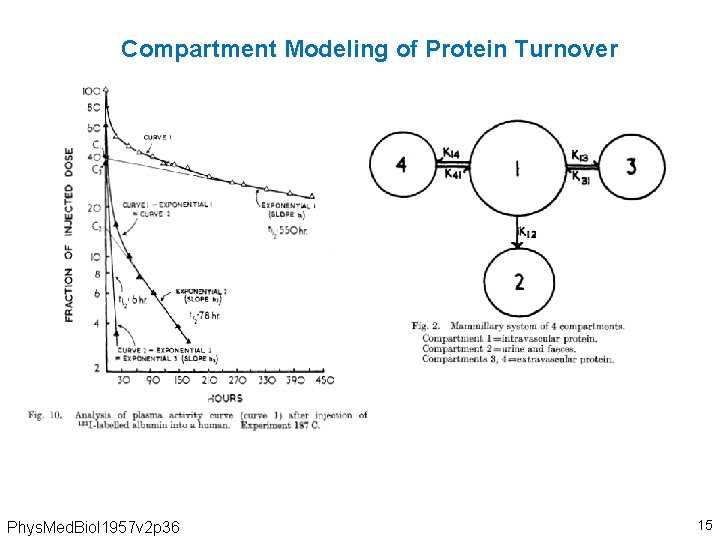
Compartment Modeling of Protein Turnover Phys. Med. Biol 1957 v 2 p 36 15
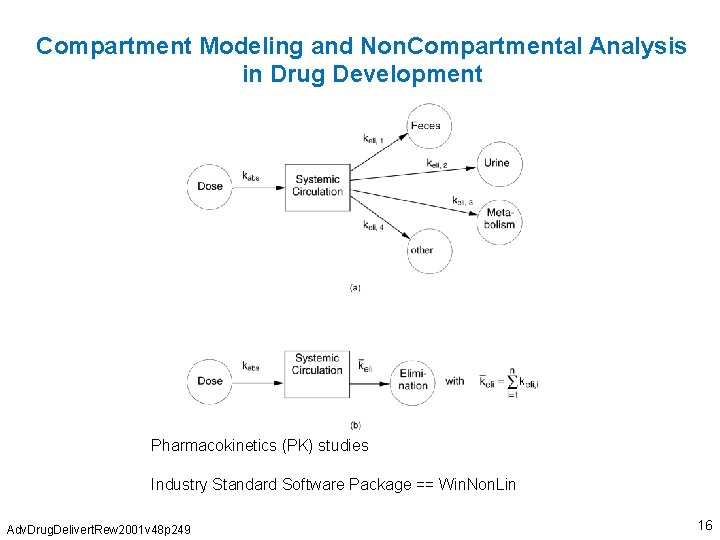
Compartment Modeling and Non. Compartmental Analysis in Drug Development Pharmacokinetics (PK) studies Industry Standard Software Package == Win. Non. Lin Adv. Drug. Delivert. Rew 2001 v 48 p 249 16
![Compartment (Pool) Modeling “分池模型” RA[A]T V RA[A] Anal. Chem 2012 v 84 p 4014 Compartment (Pool) Modeling “分池模型” RA[A]T V RA[A] Anal. Chem 2012 v 84 p 4014](http://slidetodoc.com/presentation_image/29523b65ea66d542d5af57d9db5a9ddb/image-17.jpg)
Compartment (Pool) Modeling “分池模型” RA[A]T V RA[A] Anal. Chem 2012 v 84 p 4014 17
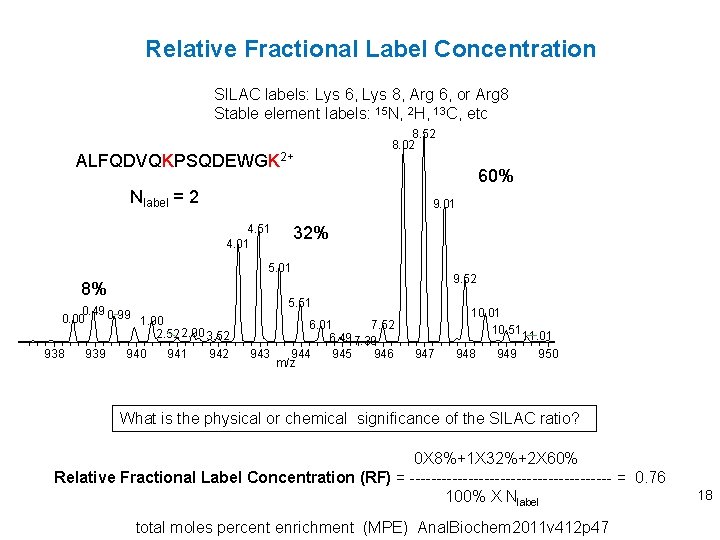
Relative Fractional Label Concentration SILAC labels: Lys 6, Lys 8, Arg 6, or Arg 8 Stable element labels: 15 N, 2 H, 13 C, etc ALFQDVQKPSQDEWGK 2+ 8. 52 8. 02 Nlabel = 2 60% 9. 01 4. 51 4. 01 32% 5. 01 8% 0. 49 0. 99 0. 00 1. 90 2. 52 2. 90 3. 52 938 939 940 941 942 9. 52 5. 51 7. 52 6. 01 6. 49 7. 39 943 944 945 946 m/z 10. 01 10. 51 11. 01 947 948 949 950 What is the physical or chemical significance of the SILAC ratio? 0 X 8%+1 X 32%+2 X 60% Relative Fractional Label Concentration (RF) = ------------------- = 0. 76 100% X Nlabel total moles percent enrichment (MPE) Anal. Biochem 2011 v 412 p 47 18
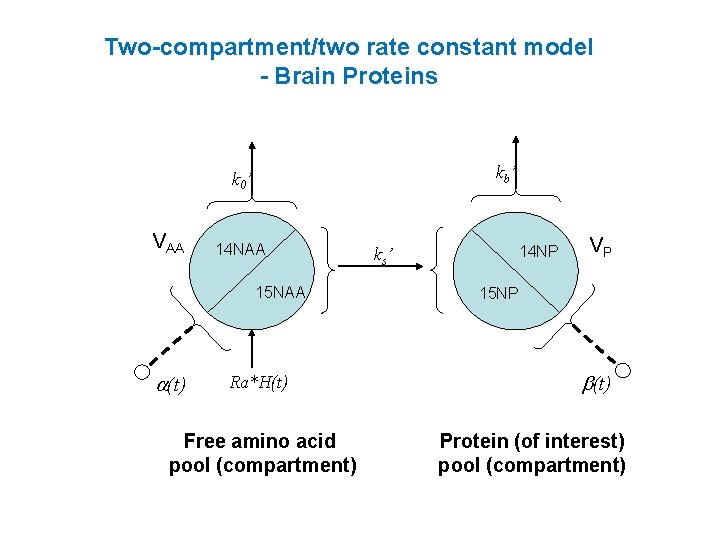
Two-compartment/two rate constant model - Brain Proteins kb ’ k 0 ’ VAA 14 NAA 15 NAA a(t) Ra*H(t) Free amino acid pool (compartment) 14 NP k s’ VP 15 NP b(t) Protein (of interest) pool (compartment)

Two-compartment/two rate constant model 2

Solution of two-compartment/two-rate constant model b t
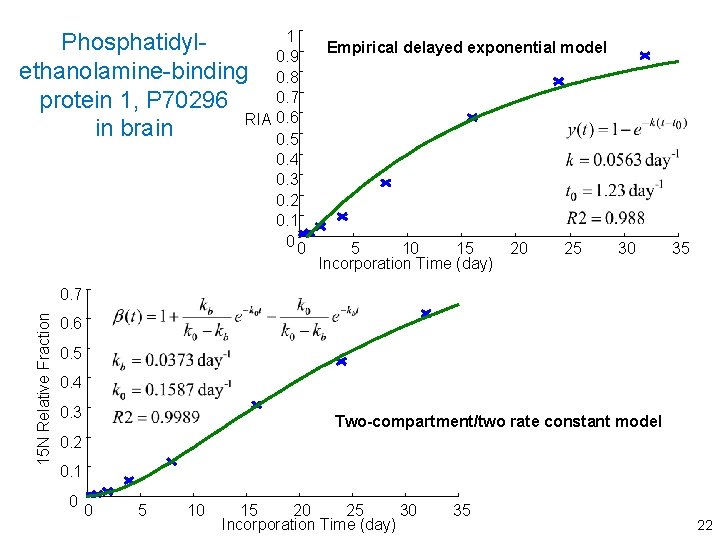
1 Phosphatidyl 0. 9 ethanolamine-binding 0. 8 0. 7 protein 1, P 70296 RIA 0. 6 in brain 0. 5 0. 4 0. 3 0. 2 0. 1 00 Empirical delayed exponential model 5 10 15 Incorporation Time (day) 20 25 30 35 15 N Relative Fraction 0. 7 0. 6 0. 5 0. 4 0. 3 Two-compartment/two rate constant model 0. 2 0. 1 0 0 5 10 15 20 25 30 Incorporation Time (day) 35 22
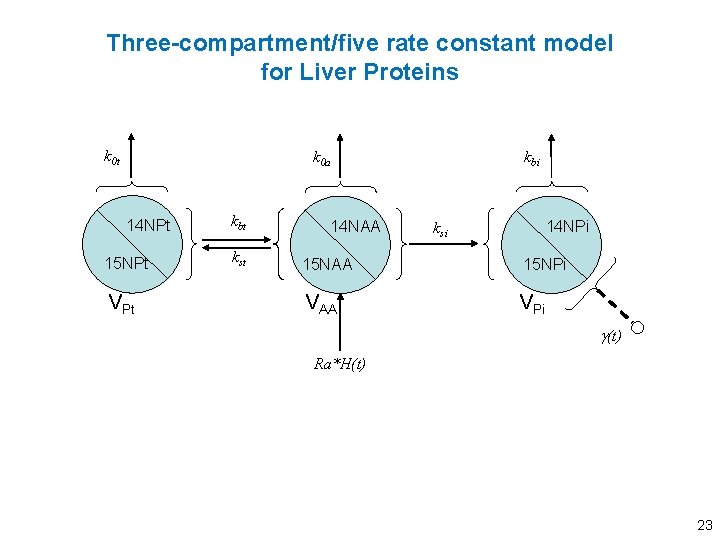
Three-compartment/five rate constant model for Liver Proteins k 0 t kbi k 0 a 14 NPt 15 NPt VPt kbt kst 14 NAA 14 NPi ksi 15 NAA 15 NPi VAA VPi g(t) Ra*H(t) 23
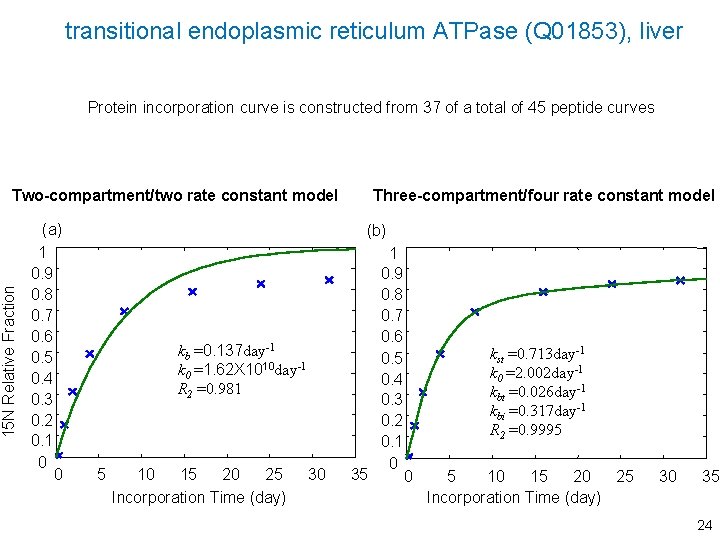
transitional endoplasmic reticulum ATPase (Q 01853), liver Protein incorporation curve is constructed from 37 of a total of 45 peptide curves 15 N Relative Fraction Two-compartment/two rate constant model (a) 1 0. 9 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 0. 1 0 0 Three-compartment/four rate constant model (b) kb =0. 137 day-1 k 0 =1. 62 X 1010 day-1 R 2 =0. 981 5 10 15 20 25 Incorporation Time (day) 30 35 1 0. 9 0. 8 0. 7 0. 6 0. 5 0. 4 0. 3 0. 2 0. 1 0 kst =0. 713 day-1 k 0 =2. 002 day-1 kbt =0. 026 day-1 kbi =0. 317 day-1 R 2 =0. 9995 0 5 10 15 20 25 Incorporation Time (day) 30 35 24

Compartment Modeling 1. Fit better to experimental data with a minimal number of parameters 2. Fitting parameters have biological significance 3. Individual rate constants are determined 25

Studies Cellular models – on going Aging models – on going Disease models: Prion infected - planned Technical improvement High sensitivity Instrument - installed LC alignment – implemented Processing speed and QC – on going 26

神经退化性疾病研究所 John C. Price Shigenari Hayashi Alma L. Burlingame Sina Ghaemmaghami Stanley B. Prusiner 药化系质谱中心 27
- Slides: 27