NEMO ERP Analysis Toolkit ERP Pattern Decomposition An






















- Slides: 30

NEMO ERP Analysis Toolkit ERP Pattern Decomposition An Overview

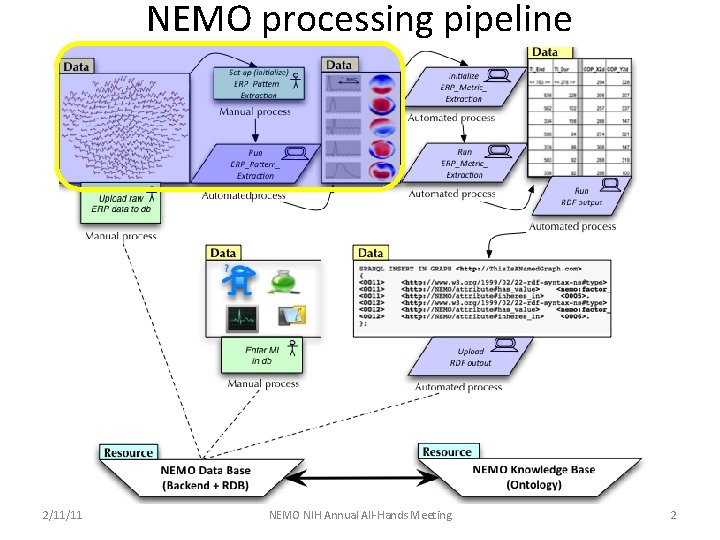
NEMO processing pipeline 2/11/11 NEMO NIH Annual All-Hands Meeting 2

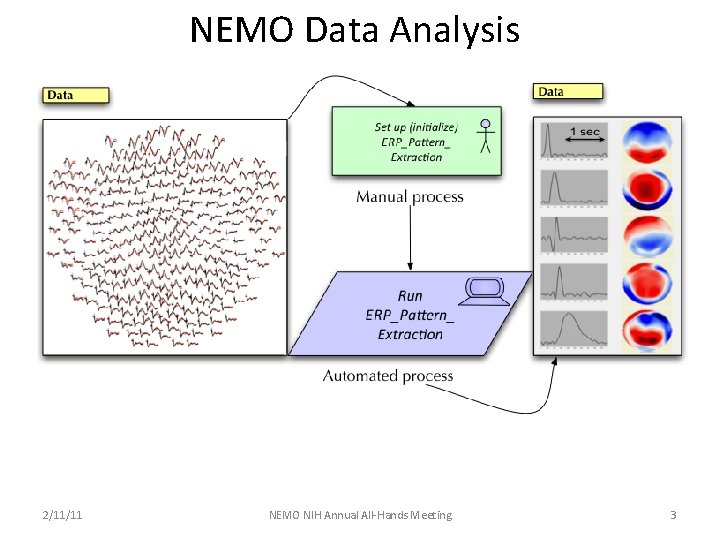
NEMO Data Analysis 2/11/11 NEMO NIH Annual All-Hands Meeting 3

NEMO Information Processing Pipeline ERP Pattern Extraction, Identification and Labeling v Obtain ERP data sets with compatible functional constraints – NEMO consortium data v Decompose / segment ERP data into discrete spatio-temporal patterns – ERP Pattern Decomposition / ERP Pattern Segmentation v Mark-up patterns with their spatial, temporal & functional characteristics – ERP Metric Extraction v Meta-Analysis v Extracted ERP pattern labeling v Extracted ERP pattern clustering v Protocol incorporates and integrates: v ERP pattern extraction v ERP metric extraction/RDF generation v NEMO Data Base (NEMO Portal / NEMO FTP Server) v NEMO Knowledge Base (NEMO Ontology/Query Engine)

ERP Pattern Decomposition Tool MATLAB and Directory Configuration v Get Latest Toolkit Version (NEMO Wiki : Screencasts : Versions ) – Update your local (working) copy of the NEMO Sourceforge Repository v Configure MATLAB (NEMO Wiki : Screencasts : NEMO ERP Analysis Toolkit I) – MATLAB R 2010 a / R 2010 b, Optimization and Statistics Toolboxes – Add to the MATLAB path, with subfolders: § NEMO_ERP_Dataset_Import / NEMO_ERP_Dataset_Information § NEMO_ERP_Metric_Extraction / NEMO_ERP_Pattern_Decomposition / NEMO_ERP_Pattern_Segmentation v Configure Experiment Folder (NEMO Wiki : Screencasts : NEMO ERP Analysis Toolkit I & II) – Create an experiment-specific parent folder containing Data, Metric Extraction, Pattern Decomposition and Pattern Segmentation subfolders – Copy the metric extraction, decomposition and segmentation script templates from your NEMO Sourceforge Repository working copy to their respective script subfolders – Add the experiment-specific parent folder, with its subfolders, to the MATLAB path

ERP Pattern Decomposition Tool Metascript Configuration – Step 1 of 7: Data Parameters v File_Name v Electrode_Montage_ID v Cell_Index v Factor_Index v ERP_Onset_Latency v ERP_Offset_Latency v ERP_Baseline_Latency

ERP Pattern Decomposition Tool Metascript Configuration – Step 1 of 7: Data Parameters v File_Name – Name of an EGI segmented simple binary file, as a single-quoted string § Example: ‘Sim. Erp. Data. raw’ § At present, Metric Extraction only accepts factor files from the Pattern Decomposition tool v Electrode_Montage_ID – Name of an EGI/Biosemi electrode montage file, as a single-quoted string § Valid montage strings: ‘GSN-128’, ‘GSN-256’, ‘HCGSN-128’, ‘HCGSN-256’, ‘Biosemi-64+5 exg’, ‘Biosemi-64 -sans. NZ_LPA_RPA’ § The NEMO ERP Analysis Toolkit will require EEGLAB channel location file (. ced) format for all proprietary, user-specified, montages v Cell_Index – Indices of cells / conditions to import, as a MATLAB vector § Indices correspond to the ordering of cells in the data file § See Metric_obj. Dataset. Metadata. Src. File. Info. Cellcode for the ordered list of conditions v Factor_Index – Indices of PCA factors to import, as a MATLAB vector § Indices correspond to the ordering of factors in the data file

ERP Pattern Decomposition Tool Metascript Configuration – Step 1 of 7: Data Parameters v ERP_Onset_Latency – Time, in milliseconds, of the first ERP sample point to import, as a MATLAB scalar § 0 ms = stimulus onset § Positive values specify post-stimulus time points, negative values pre-stimulus time points § All latencies must be in integer multiples of the sampling interval (for example, +’ve / -’ve multiples of 4 ms @ 250 Hz) v ERP_Offset_Latency – Time, in milliseconds, of the last ERP sample point to import, as a MATLAB scalar § 0 ms = stimulus onset § Positive values specify post-stimulus time points, and must be greater than the ERP_Onset_Latency § ERP_Offset_Latency must not exceed the final data sample point (for example, a 1000 ms ERP with a 200 ms baseline: maximum 800 ms ERP_Offset_Latency) v ERP_Baseline_Latency – Time, in negative milliseconds, of the pre-stimulus ERP sample points to exclude from import, as a MATLAB scalar § ERP_Baseline_Latency = 0 no baseline § To import pre-stimulus sample points, specify ERP_Baseline_Latency < ERP_Onset_Latency < 0 § All latencies must be within the data range (for example, a 1000 ms ERP with a 200 ms baseline: ERP_Baseline_Latency = -200 ms, ERP_Onset_Latency = 0 ms and ERP_Offset_Latency = 800 ms imports the 800 ms post-stimulus interval, including stimulus onset)

ERP Pattern Decomposition Tool Metascript Configuration – Step 2 of 7: Experiment Parameters (Required) v Lab_ID v Experiment_ID v Session_ID v Subject_Group_ID v Subject_ID v Experiment_Info

ERP Pattern Decomposition Tool Metascript Configuration – Step 2 of 7: Experiment Parameters (Required) v Lab_ID – Laboratory identification label, as a single-quoted string § Example: ‘My Simulated Lab’ v Experiment_ID – Experiment identification label, as a single-quoted string § Example: ‘My Simulated Experiment’ v Session_ID – Session identification label, as a single-quoted string § Example: ‘My Simulated Session’ v Subject_Group_ID – Subject group identification label, as a single-quoted string § Example: ‘My Simulated Subject Group’ v Subject_ID – Subject identification label, as a single-quoted string § Example: ‘My Simulated Subject # 1’ v Experiment_Info – Experiment note, as a single-quoted string § Example: ‘t. PCA with Infomax rotation’

ERP Pattern Decomposition Tool Metascript Configuration – Step 3 of 7: Experiment Parameters (Optional) v Event_Type_Label v Stimulus_Modality_Label v Cell_Label_Descriptor

ERP Pattern Decomposition Tool Metascript Configuration – Step 3 of 7: Experiment Parameters (Optional) v Event_Type_Label – MATLAB cell array of cell/condition event type labels § One label per cell/condition, as a single-quoted string § Example: {‘Sim. Event. Type 1’, ‘Sim. Event. Type 2’, ‘Sim. Event. Type 3’} v Stimulus_Type_Label – MATLAB cell array of cell/condition stimulus type labels § One label per cell/condition, as a single-quoted string § Example: {‘Sim. Stimulus. Type 1’, ‘Sim. Stimulus. Type 2’, ‘Sim. Stimulus. Type 3’} v Stimulus_Modality_Label – MATLAB cell array of cell/condition stimulus modality labels § One label per cell/condition, as a single-quoted string § Example: {‘Sim. Stimulus. Modality 1’, ‘Sim. Stimulus. Modality 2’, ‘Sim. Stimulus. Modality 3’} v Cell_Label_Descriptor – MATLAB cell array of cell/condition description labels § One label per cell/condition, as a single-quoted string § Optional Labels: E-prime assigned cell codes imported from input data file § Example: {‘Sim. Condition. Description 1’, ‘Sim. Condition. Description 2’, ‘Sim. Condition. Description 3’}

ERP Pattern Decomposition Tool Metascript Configuration – Step 4 of 7: Stage 1 Component Decomposition Parameters Stage 1 t. PCA v PCAmode v MAT_TYPE v ROTATION v LOADING v NUM_FAC v SORTOPT v GAVE

ERP Pattern Decomposition Tool Metascript Configuration – Step 4 of 7: Stage 1 Component Decomposition Parameters v PCAmode – Specifies the PCA mode, as a single-quoted string § ‘temp’: Temporal PCA, in which time points are variables § ‘spat’: Spatial PCA, in which channel voltages are variables v MAT_TYPE – Specifies the PCA eigenvector/relationship matrix, as a single-quoted string § ‘COV’: Covariance matrix (mean correction) § ‘COR’: Correlation matrix (mean + variance correction) § ‘SCP’: Sum of squares cross product (no mean/variance correction) v ROTATION – Specifies the PCA factor rotation type, as a single-quoted string § ‘IMAX’: Infomax - ”Statistically Independent” factor loadings via high-order statistics § ‘VMAX’: Varimax - Maximal variance factor loadings subject to orthogonality constraint § ‘PMAX’: Promax - Relaxes factor orthogonality constraint of relationship matrix eigenvectors Promax rotation is automatically applied subsequent to Varimax rotation when ROTATION = ‘VMAX’

ERP Pattern Decomposition Tool Metascript Configuration – Step 4 of 7: Stage 1 Component Decomposition Parameters v LOADING – Specifies factor loading type, the rotated factor loading scaling transform, as a single-quoted string § ‘N’: None § ‘K’: Kaiser § ‘C’: Covariance § ‘W’: Cureton-Mulaik v NUM_FAC – Specifies the number of PCA factors to rotate, as a MATLAB scalar § For s. PCA: 1. LE. NUM_FAC. LE. number of electrode channels § For t. PCA: 1. LE. NUM_FAC. LE. number of imported ERP time points v SORTOPT – Specifies the ordering (sort) of post-rotation PCA factors, as a single quoted string § ‘Pre. Rot’: Sort in order of decreasing pre-rotation (eigenvector) factor variance § ‘Fac. Var’: Sort in order of decreasing post-rotation factor variance, via Fac. Var parameter v GAVE – Optionally perform analysis on grand average data § ‘N’: Perform analysis on subject average data only § ‘Y’: Perform analysis on grand average data; convert factor scores to subject average form for export

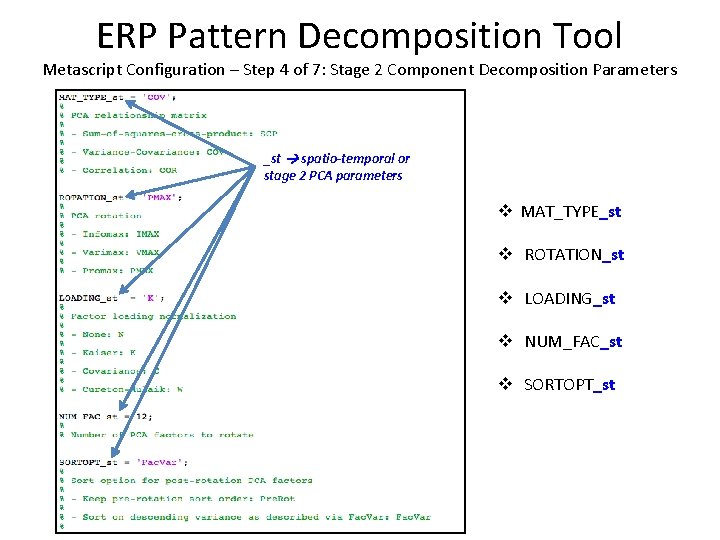
ERP Pattern Decomposition Tool Metascript Configuration – Step 4 of 7: Stage 2 Component Decomposition Parameters Stage 1 t. PCA _st spatio-temporal or stage 2 PCA parameters v MAT_TYPE_st v ROTATION_st v LOADING_st v NUM_FAC_st v SORTOPT_st

ERP Pattern Decomposition Tool Metascript Configuration – Step 4 of 7: Stage 2 Component Decomposition Parameters v PCAmode – Specifies the PCA mode, as a single-quoted string § ‘temp’: Temporal PCA, in which time points are variables § ‘spat’: Spatial PCA, in which channel voltages are variables Stage 1 t. PCA Stage 2 s. PCA Stage 1 s. PCA Stage 2 t. PCA v MAT_TYPE_st – Specifies the PCA eigenvector/relationship matrix, as a single-quoted string § ‘COV’: Covariance matrix (mean correction) § ‘COR’: Correlation matrix (mean + variance correction) § ‘SCP’: Sum of squares cross product (no mean/variance correction) v ROTATION_st – Specifies the PCA factor rotation type, as a single-quoted string § ‘IMAX’: Infomax - ”Statistically Independent” factor loadings via high-order statistics § ‘VMAX’: Varimax - Maximal variance factor loadings subject to orthogonality constraint § ‘PMAX’: Promax - Relaxes factor orthogonality constraint of relationship matrix eigenvectors Promax rotation is automatically applied subsequent to Varimax rotation when ROTATION = ‘VMAX’

ERP Pattern Decomposition Tool Metascript Configuration – Step 4 of 7: Stage 2 Component Decomposition Parameters v LOADING_st – Specifies factor loading type, the rotated factor loading scaling transform, as a single-quoted string § ‘N’: None § ‘K’: Kaiser § ‘C’: Covariance § ‘W’: Cureton-Mulaik v NUM_FAC_st – Specifies the number of PCA factors to rotate, as a MATLAB scalar § 1. LE. NUM_FAC_st. LE. NUM_FAC (Number of stage 1 factors to rotate) v SORTOPT_st – Specifies the ordering (sort) of post-rotation PCA factors, as a single quoted string § ‘Pre. Rot’: Sort in order of decreasing pre-rotation (eigenvector) factor variance § ‘Fac. Var’: Sort in order of decreasing post-rotation factor variance, via Fac. Var parameter v GAVE – Optionally perform analysis on grand average data Specified in Stage 1 § ‘N’: Perform analysis on subject average data only § ‘Y’: Perform analysis on grand average data; convert factor scores to subject average form for export

ERP Pattern Decomposition Tool Metascript Configuration – Step 5 of 7: Export to EGI Simple Binary Parameters Stage 1 v Num_Fac_Export Stage 2 v Num_Fac_Export_st v Cell_IO_Rule v Output_File_Type v Grand_Avg_Add v Exclude_Channel

ERP Pattern Decomposition Tool Metascript Configuration – Step 5 of 7: Export to EGI Simple Binary Parameters v Num_Fac_Export / Num_Fac_Export_st – Specifies the number of stage 1 / stage 2 PCA factors to export, as a MATLAB scalar § 1. LE. Num_Fac_Export. LE. NUM_FAC (# of stage 1 PCA factors to rotate) § 1. LE. Num_Fac_Export_st. LE. NUM_FAC_st (# of stage 2 PCA factors to rotate) v Cell_IO_Rule – Specifies the input cell to output cell rule, as a 2 D MATLAB array § Output cell x input cell logical indexing matrix § Type <My. Pattern. Decomposition. Object>. Help. Topic(‘PCAto. Egi. Sbin’) For Detail v Output_File_Type – Specifies the output PCA factor file type, as a single quoted string § ‘G’: Grand average factor file (Average across subject factors for each cell type | 1 file) § ‘S’: Subject average factor file (Subject-specific factors for each cell type | 1 file per subject) v Grand_Avg_Add – Specifies option to add grand average to factor reconstructions § ‘N’: Do not add grand average to factor reconstructions § ‘Y’: Add grand average to factor reconstructions v Exclude_Channel – List of peri-ocular or midline channels to omit in ANOVA (N/A = []), as a MATLAB vector

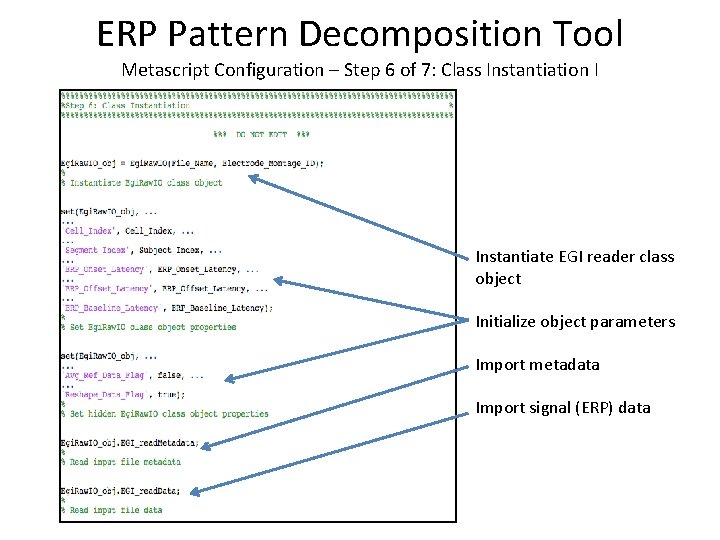
ERP Pattern Decomposition Tool Metascript Configuration – Step 6 of 7: Class Instantiation I Instantiate EGI reader class object Initialize object parameters Import metadata Import signal (ERP) data

ERP Pattern Decomposition Tool Metascript Configuration – Step 6 of 7: Class Instantiation I (EP Toolkit) Instantiate EGI reader class object Initialize object parameters Import metadata and signal (ERP) data via EPToolkit’s ep_read. Data

ERP Pattern Decomposition Tool Metascript Configuration – Step 6 of 7: Class Instantiation II Instantiate Pattern Decomposition class object Initialize object parameters

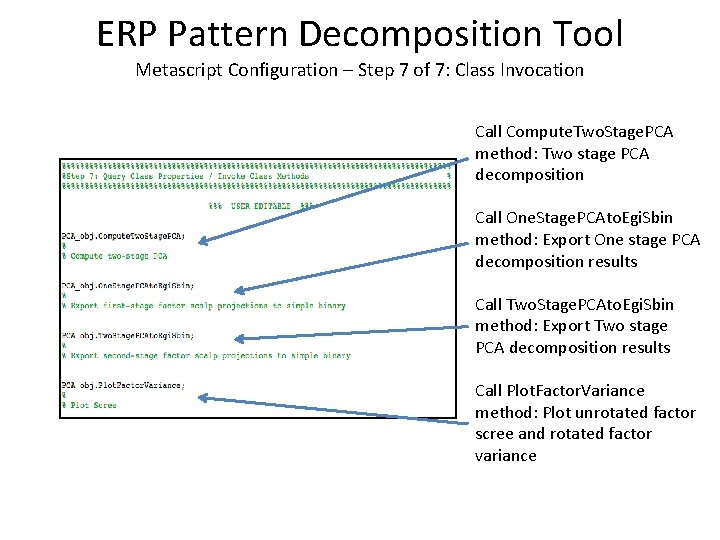
ERP Pattern Decomposition Tool Metascript Configuration – Step 7 of 7: Class Invocation Call Compute. Two. Stage. PCA method: Two stage PCA decomposition Call One. Stage. PCAto. Egi. Sbin method: Export One stage PCA decomposition results Call Two. Stage. PCAto. Egi. Sbin method: Export Two stage PCA decomposition results Call Plot. Factor. Variance method: Plot unrotated factor scree and rotated factor variance

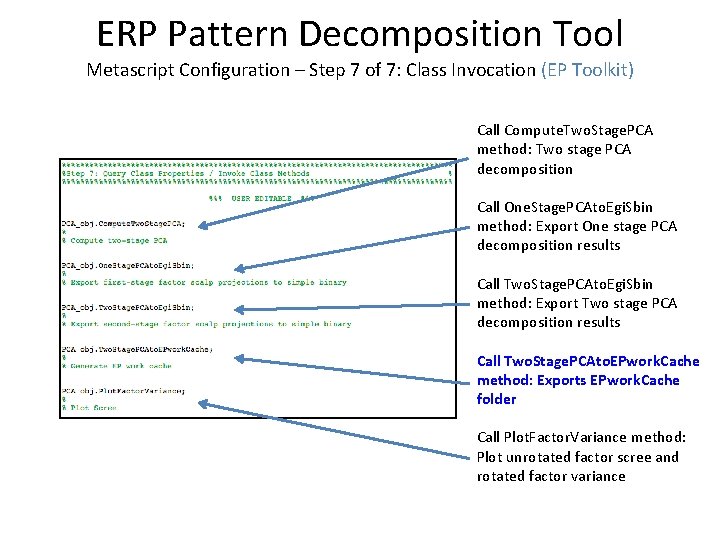
ERP Pattern Decomposition Tool Metascript Configuration – Step 7 of 7: Class Invocation (EP Toolkit) Call Compute. Two. Stage. PCA method: Two stage PCA decomposition Call One. Stage. PCAto. Egi. Sbin method: Export One stage PCA decomposition results Call Two. Stage. PCAto. Egi. Sbin method: Export Two stage PCA decomposition results Call Two. Stage. PCAto. EPwork. Cache method: Exports EPwork. Cache folder Call Plot. Factor. Variance method: Plot unrotated factor scree and rotated factor variance

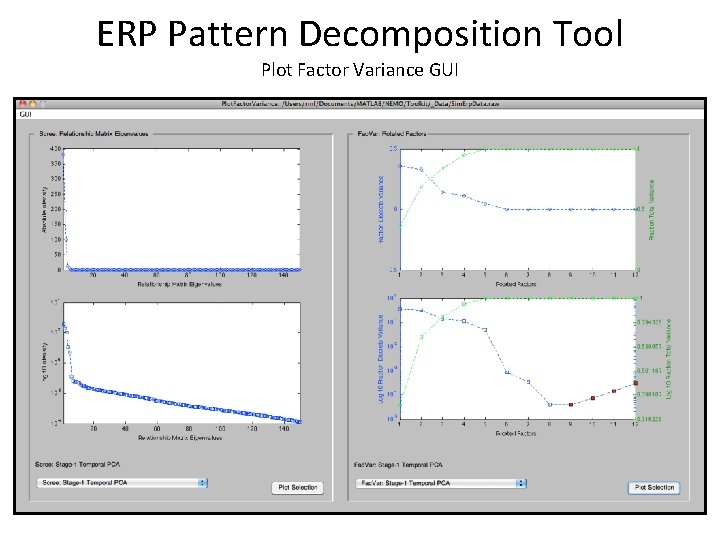
ERP Pattern Decomposition Tool Plot Factor Variance GUI

ERP Pattern Decomposition Tool Folder Output for Sim. Erp. Data. raw v Pattern Decomposition output folder contents – RAW files • t. PCA: Input. Data. File_t. PCA_GAV/AVG. raw • s. PCA: Input. Data. File_s. PCA_GAV/AVG. raw • st. PCA/ts. PCA: Input. Data. File_st. PCA/ts. PCA_GAV/AVG. raw – Epwork Folder: EP Toolkit integration folder (if used EPT_read. Data) – Nemo. Erp. Pattern. Decompostion workspace object in MATLAB (. mat) format Input data file Time stamp

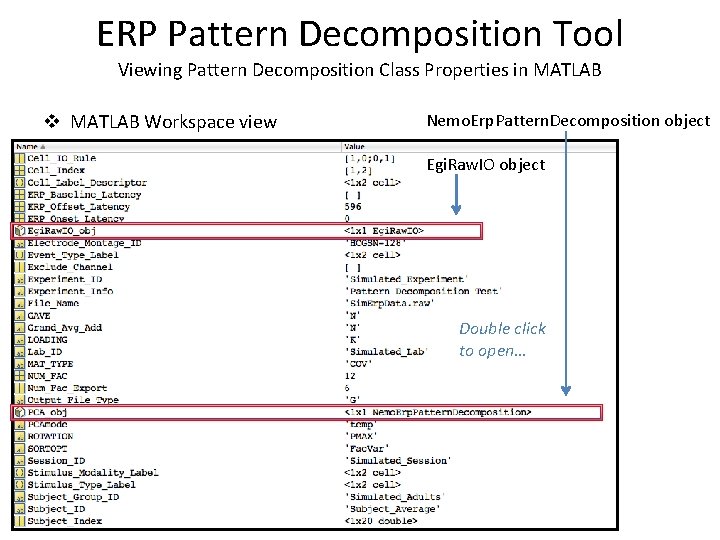
ERP Pattern Decomposition Tool Viewing Pattern Decomposition Class Properties in MATLAB v MATLAB Workspace view Nemo. Erp. Pattern. Decomposition object Egi. Raw. IO object Double click to open…

ERP Pattern Decomposition Tool Viewing Pattern Decomposition Class Properties in MATLAB v MATLAB Workspace view Keep on double clicking … v EPread. Data. Input: MATLAB structure of input parameters to ep_read. Data v Epdata: MATLAB structure of output data and metadata from ep_read. Data v EGIread. Data. Input: MATLAB structure of (optional) input parameters to EGI_read. Data and EGI_read. Meta. Data v Metadata: MATLAB structure of output metadata from EGI_read. Metadata v Data: MATLAB structure of output data from EGI_read. Data

ERP Pattern Decomposition Tool Viewing Pattern Decomposition Class Properties in MATLAB v MATLAB Workspace view Keep on double clicking … v EPdo. PCAInput: MATLAB structure of input parameters to ep_do. PCA v Factor. Results: MATLAB structure of output factor decomposition and metadata from ep_do. PCA v EPdo. PCAst. Input: MATLAB structure of input parameters to second PCA step (ep_do. PCAst) v Factor. Results. ST: MATLAB structure of output factor decomposition and metadata from second PCA step (ep_do. PCAst) v PCAto. Egi. Sbin: MATLAB structure of input parameters to One. Stage. PCAto. Egi. Sbin / Two. Stage. PCAto. Egi. Sbin