NCI Cancer Research Data Commons History Vision and















- Slides: 28

NCI Cancer Research Data Commons History, Vision and Progress Tony Kerlavage, Ph. D. Acting Director Center for Biomedical Informatics and Information Technology CRDC Meeting October 12, 2018

1. Program Retrospective Agenda 2. Drivers for a Cancer Research Data Commons 3. Progress & Current Status of the CRDC 2

Program Retrospective The TCGA Challenge: GDC and Cancer Cloud Pilots

4

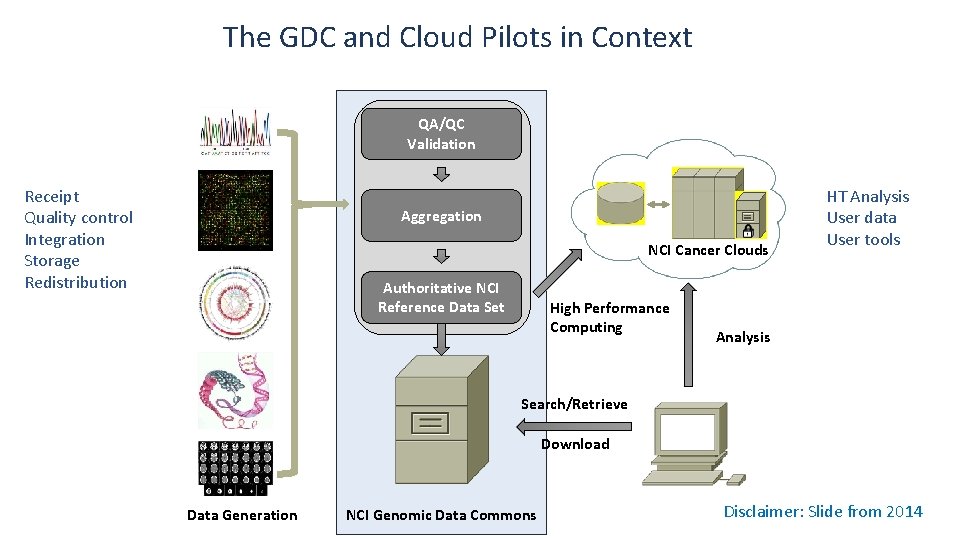
• Unify fragmentary repositories • Support the receipt, quality control, integration, storage, and redistribution of standardized genomic data sets derived from cancer research studies • Harmonization of raw sequence both from existing and new cancer research programs • Application of state-of-the-art methods of generating derived genomic data • Provide the foundation for: • Identification of both high- and low-frequency cancer drivers • Defining genomic determinants of response to therapy • Selecting clinical trial cohorts sharing genetic lesions 5

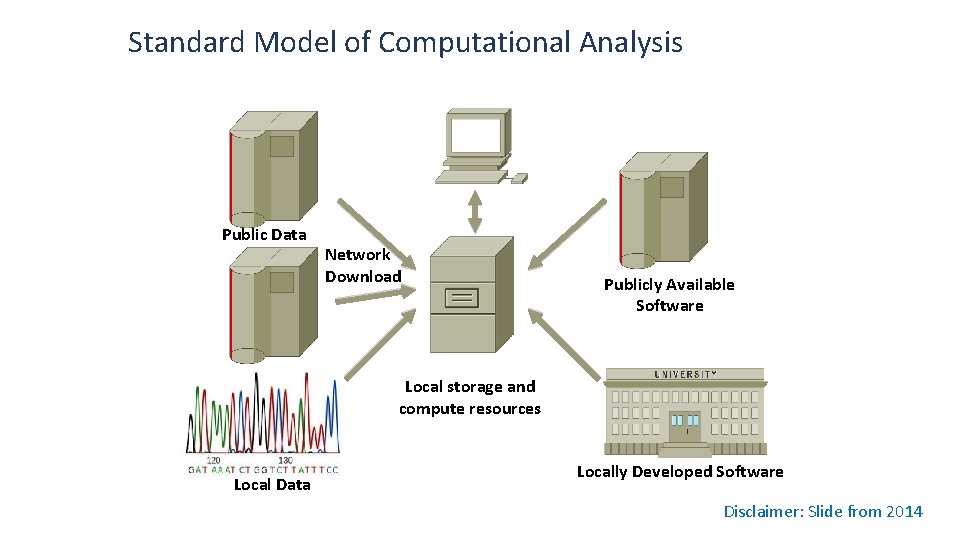
Standard Model of Computational Analysis Public Data Network Download Publicly Available Software Local storage and compute resources Local Data Locally Developed Software Disclaimer: Slide from 2014

Limitations of the Standard Model for Large Data • Assuming the 2. 5 PB TCGA data set • Storage and data protection cost are ~$2 M /year • Downloading TCGA data at 10 Gb/sec would take ~23 days • Only large institutions have the ability to utilize this data • These data will continue to grow at an increasing rate

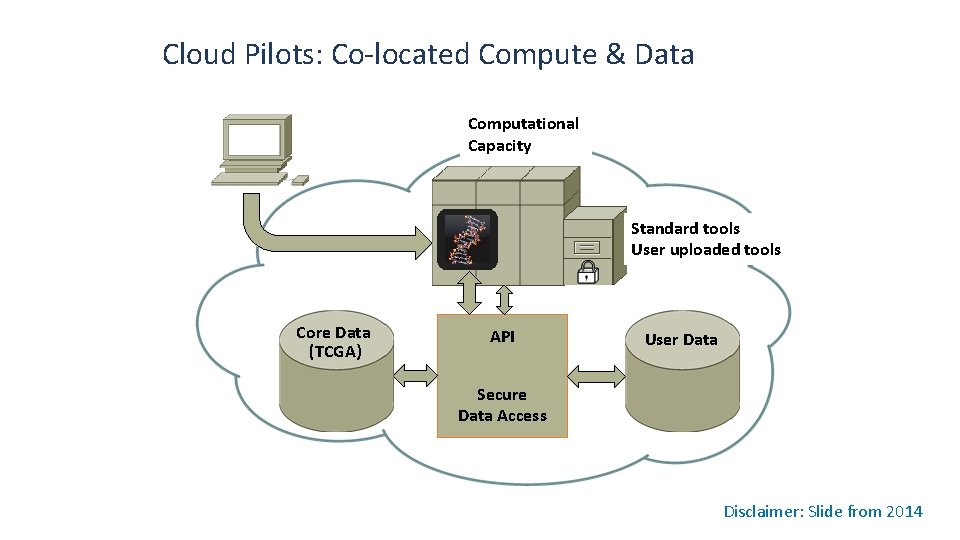
Cloud Pilots: Co-located Compute & Data Computational Capacity Standard tools User uploaded tools Core Data (TCGA) API User Data Secure Data Access Disclaimer: Slide from 2014

The GDC and Cloud Pilots in Context QA/QC Validation Receipt Quality control Integration Storage Redistribution Aggregation NCI Cancer Clouds Authoritative NCI Reference Data Set High Performance Computing HT Analysis User data User tools Analysis Search/Retrieve Download Data Generation NCI Genomic Data Commons Disclaimer: Slide from 2014

Cloud Pilot Project Structure • Goal was to democratize access to NCI genomic and associated data • Managed through CBIIT in partnership with the Center for Cancer Genomics (CCG) – Coordinating with the Genomic Data Commons (GDC) • Three contracts awarded to: – Broad Institute – Institute for Systems Biology – Seven Bridges Genomics • Period of performance: Sept 2014 – Sept 2016 – https: //cbiit. nci. nih. gov/ncip/nci-cancer-genomics-cloud-pilots – Anticipated go-live date: January 2016

Cloud Pilot Project Considerations • Innovation! • Open Design • Designs required to be released under a non-viral, open source license • Build for Extensibility & Sustainability • Initial clouds focused on a set of “core datatypes” • Extend to additional datatypes without major refactoring of the existing system • Cost assessments for operating at current scale and at 10/100 fold increases in storage, compute and usage • Data Security • First human genomic data in the cloud! • Manage Open vs. Controlled Access data • FISMA moderate system, Fed. RAMP certified cloud provider, NCI ATO, Trusted Partnership

NCI Cancer Genomics Cloud Pilots provide: • Access to large genomic data sets without need to download Use emerging GA 4 GH standards • Access to popular pipelines and visualization tools • Ability for researchers to bring their own tools and pipelines to the data • Ability for researchers to bring their own data and analyze in combination with existing genomic data • Workspaces, for researchers to save and share their data and results of analyses Democratize access to NCI-generated genomic and related data, and to create a cost-effective way to provide scalable computational capacity to the cancer research community. 12

Drivers for a Cancer Research Data Commons Precision Medicine in oncology is a Grand Challenge

Precision Medicine is a Grand Challenge Requires: • Deep biological understanding • Advances in scientific methods • Advances in instrumentation • Advances in technology • Advances in data management and computation Cancer Research and Care generate detailed data that are critical to create a learning health system for cancer 14

National Cancer Data Ecosystem for Sharing and Analysis Overall goal: “Enable all participants across the cancer research and care continuum to contribute, access, combine and analyze diverse data that will enable new discoveries and lead to lowering the burden of cancer. ” Overarching goals Recommendations • Accelerate progress in cancer, including prevention & screening • Build a National Cancer Data Ecosystem • From cutting edge basic research to wider uptake of standard of care • Encourage greater cooperation and collaboration • Within and between academia, government, and private sector • Enhanced cloud-computing platforms • Services that link disparate information, including clinical, image, and molecular data • Essential underlying data science infrastructure, standards, methods, and portals for the Cancer Data Ecosystem • Enhance data sharing 15

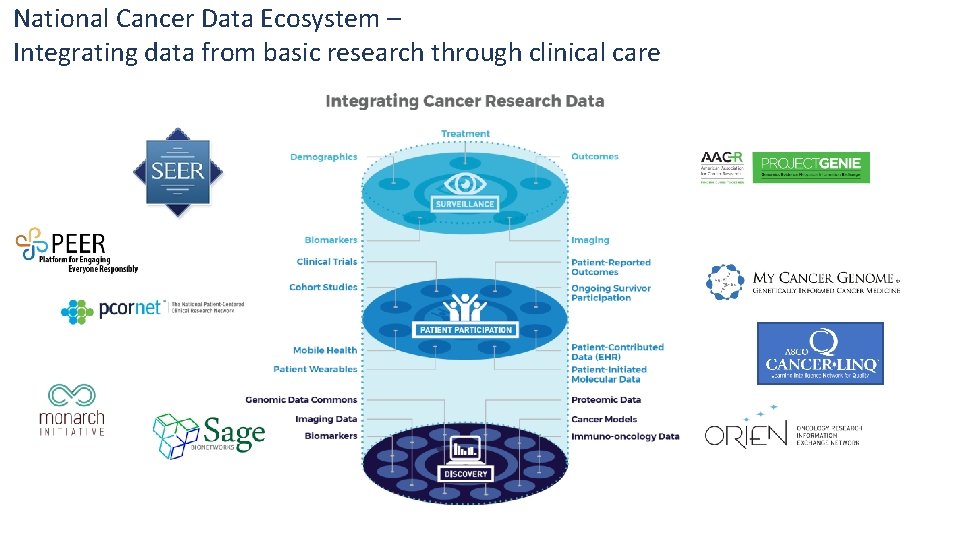
National Cancer Data Ecosystem – Integrating data from basic research through clinical care

Many NCI Programs Generating Multimodal Data Clinical Proteomics Tumor Analysis Consortium* TCIA The Cancer Imaging Archive*

The Cancer Research Data Commons Progress and Status

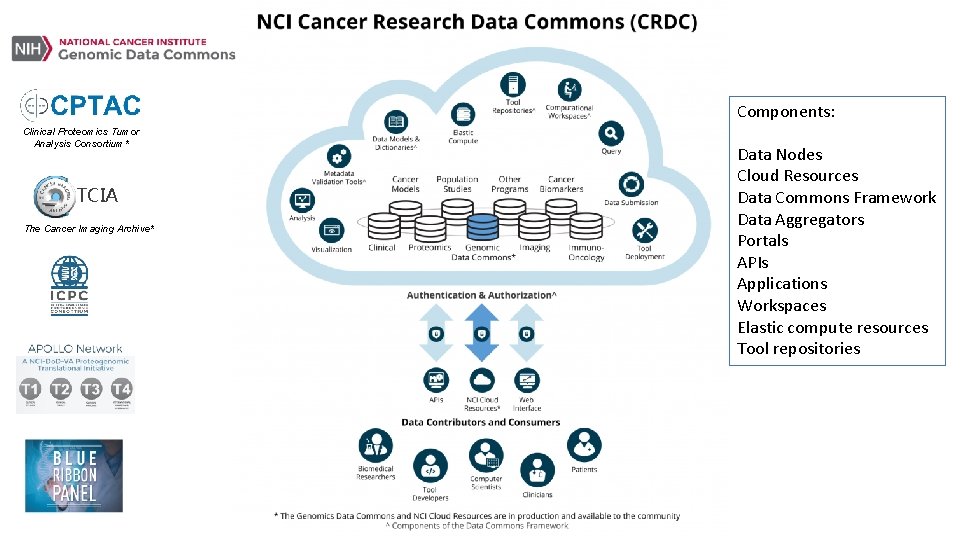
Components: Clinical Proteomics Tumor Analysis Consortium* TCIA The Cancer Imaging Archive* Data Nodes Cloud Resources Data Commons Framework Data Aggregators Portals APIs Applications Workspaces Elastic compute resources Tool repositories

Data Commons Framework – What Is It? Reusable, expandable framework for a Data Commons Core principles and structures for a Data Commons Set of modular components that can be leveraged across Data Commons Modular Components Secure user authentication and authorization Digital ID / Metadata services Domain-specific, extensible data models and dictionaries API and container environments for tools and pipelines Access to computational workspaces for storing data, tools, and results 20

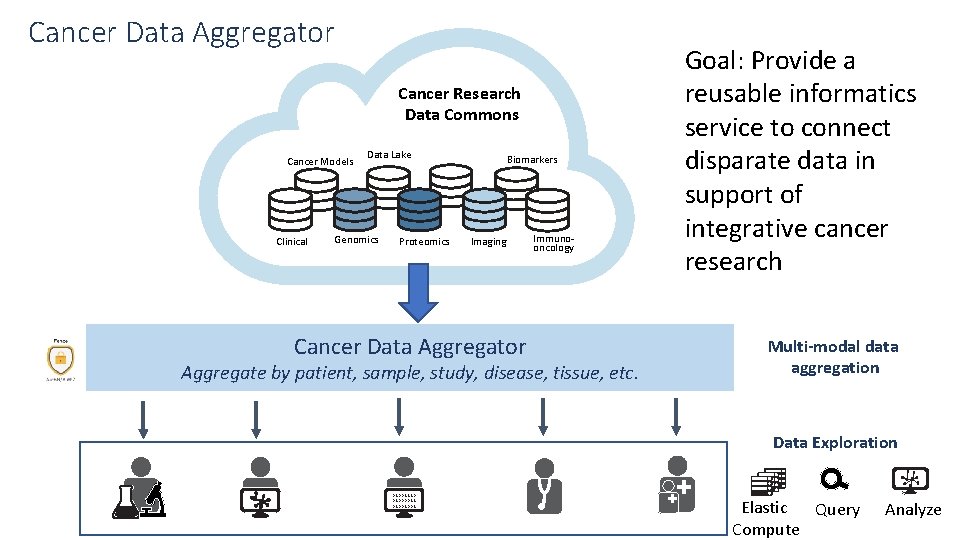
Cancer Data Aggregator Cancer Research Data Commons Cancer Models Clinical Data Lake Genomics Proteomics Biomarkers Imaging Cancer Data Aggregator Immunooncology Aggregate by patient, sample, study, disease, tissue, etc. Goal: Provide a reusable informatics service to connect disparate data in support of integrative cancer research Multi-modal data aggregation Data Exploration 01001110 01000011 01001001 Elastic Query Compute Analyze

Cancer Data Aggregator Driving Projects • Human Tumor Atlas • Will generate a significant volume of disparate primary data and metadata, including: • Single-cell and bulk –omics data sets (genomic, transcriptomic, epigenomic, proteomic, etc. ) • 2 D and 3 D molecular imaging • Clinical pathology and radiomics • Programs such as APOLLO and CPTAC 3 that are collecting multi-modal data • Aggregation of clinical, epidemiology, and exposure information

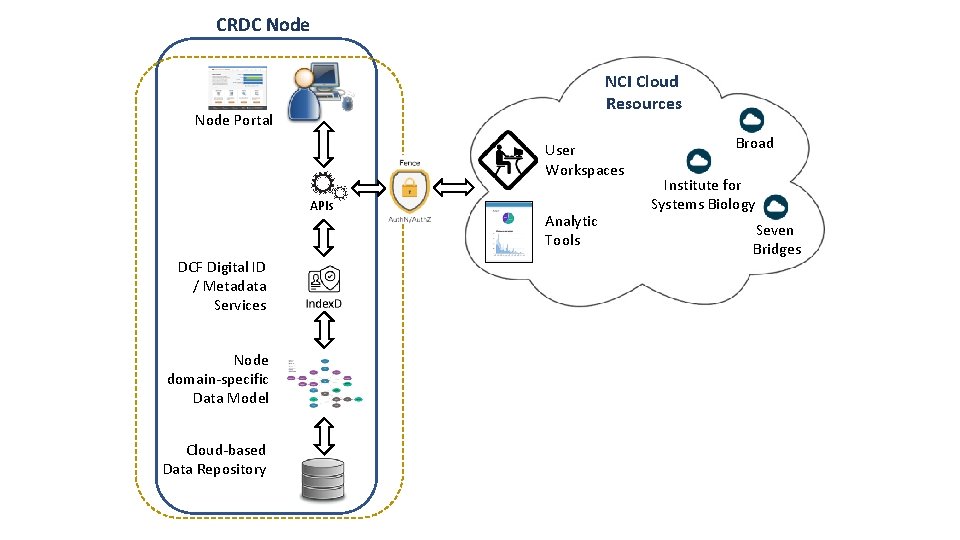
CRDC Node NCI Cloud Resources Node Portal User Workspaces APIs DCF Digital ID / Metadata Services Node domain-specific Data Model Cloud-based Data Repository Analytic Tools Broad Institute for Systems Biology Seven Bridges

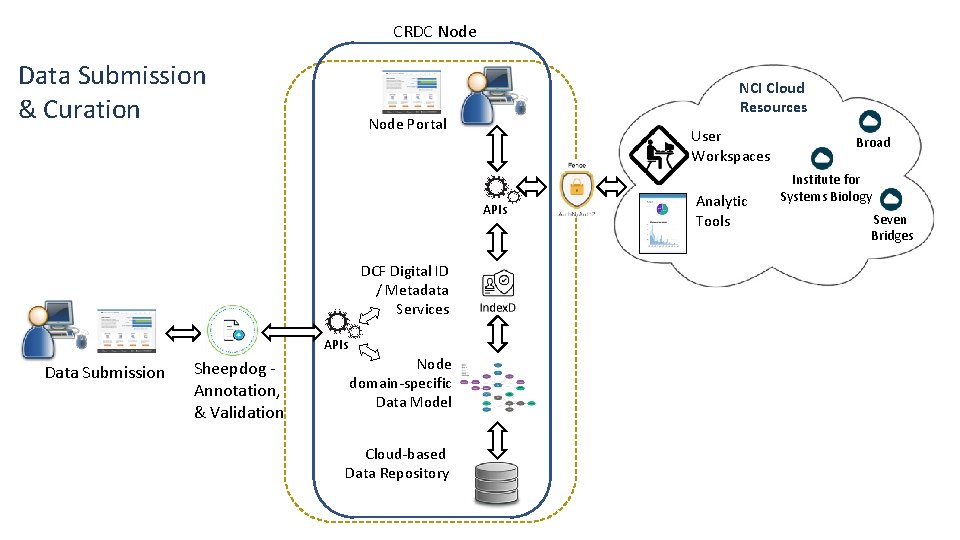
CRDC Node Data Submission & Curation NCI Cloud Resources Node Portal User Workspaces APIs DCF Digital ID / Metadata Services APIs Data Submission Sheepdog Annotation, & Validation Node domain-specific Data Model Cloud-based Data Repository Analytic Tools Broad Institute for Systems Biology Seven Bridges

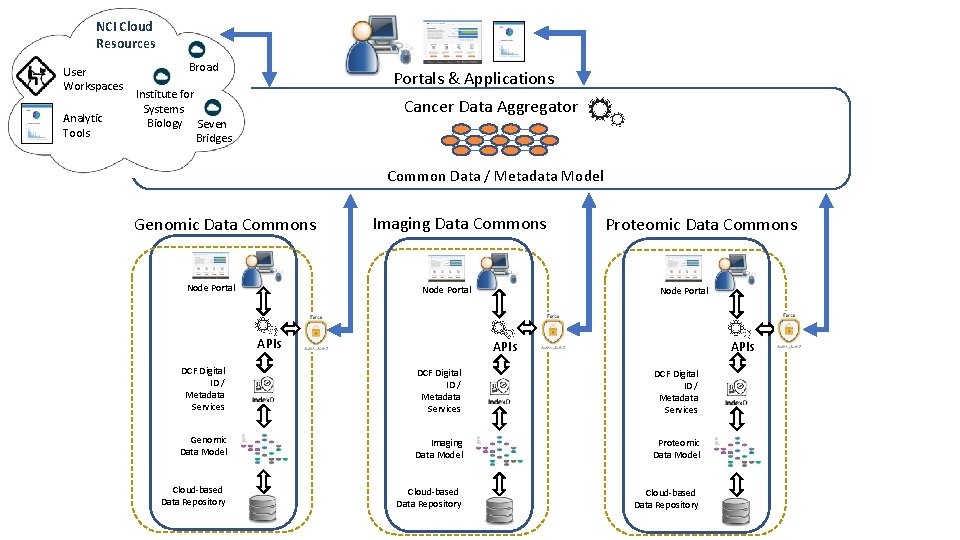
NCI Cloud Resources User Workspaces Analytic Tools Broad Portals & Applications Institute for Systems Biology Seven Bridges Cancer Data Aggregator Common Data / Metadata Model Genomic Data Commons Node Portal Imaging Data Commons Node Portal APIs Proteomic Data Commons Node Portal APIs DCF Digital ID / Metadata Services Genomic Data Model Imaging Data Model Proteomic Data Model Cloud-based Data Repository

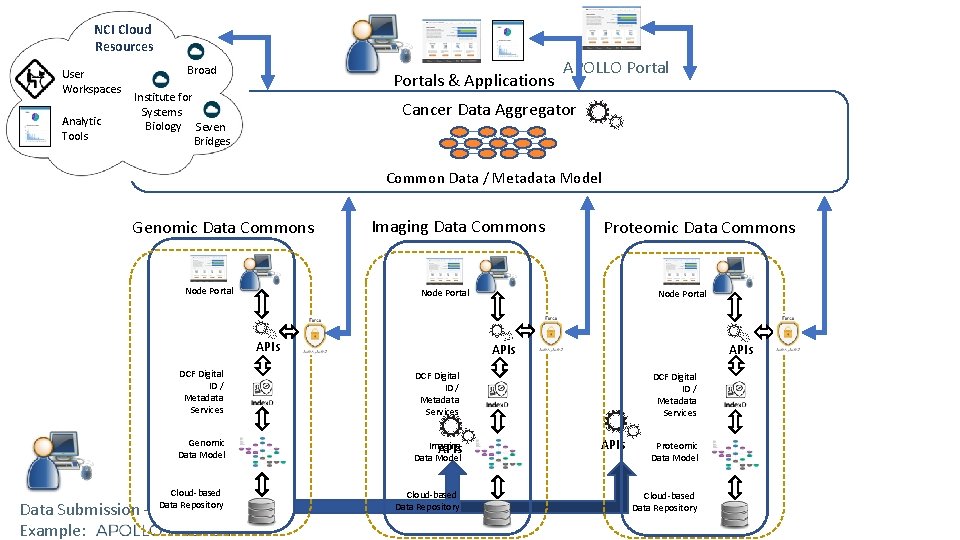
NCI Cloud Resources User Workspaces Analytic Tools Broad Portals & Applications Institute for Systems Biology Seven Bridges APOLLO Portal Cancer Data Aggregator Common Data / Metadata Model Genomic Data Commons Node Portal Imaging Data Commons Node Portal APIs DCF Digital ID / Metadata Services Genomic Data Model Data Submission – Example: Cloud-based Data Repository Proteomic Data Commons Node Portal APIs DCF Digital ID / Metadata Services APIs Imaging APIs Data Model Cloud-based Data Repository DCF Digital ID / Metadata Services APIs Proteomic Data Model Cloud-based Data Repository

Status of CRDC Components • Data Commons Framework • Fence, Index. D in use by Cloud Resources and some Data Nodes • Other components developed and ready for use by Data Nodes • Proteomic Data Commons (PDC) • Contract awarded September 2017 • Limited Pilot launched October 2018; Production version within 12 months • Imaging Data Commons (IDC) • RFP to be issued by December 1 • Award by February 2019; Pilot by February 2020 • Cancer Immuno-oncology Data Commons (CIDC) • Awarded September 2017 • Launch for data collection late 2018 • Integrated Canine Data Commons (ICDC) • Awarded September 2018 • Pilot by or before September 2020 • DCEG Population Cohort • Concept phase

www. cancer. gov/espanol