National Cancer Institute The Cancer Biomedical Informatics Grid

National Cancer Institute The Cancer Biomedical Informatics Grid™ (ca. BIG™) 2006 CODATA Conference Beijing, China Mary Jo Deering , Ph. D. Director, Informatics Dissemination NCI Center for Bioinformatics 0

National Cancer Institute Cancer Biomedical Informatics Grid™ (ca. BIGTM) • Common, widely distributed infrastructure permits research community to focus on innovation • Shared vocabulary, data elements, data models facilitate information exchange • Collection of interoperable applications developed to common standards • Raw published cancer research data is available for mining and integration 1

National Cancer Institute Cancer Biomedical Informatics Grid™ (ca. BIGTM) • ca. BIG™ infrastructure and tools are widely applicable outside cancer • ca. BIG™ components may be used by anyone 2

National Cancer Institute ca. BIG™ principles • Open source • Open access • Open development • Federated 3

National Cancer Institute ca. BIG™’s Informatics Core 4
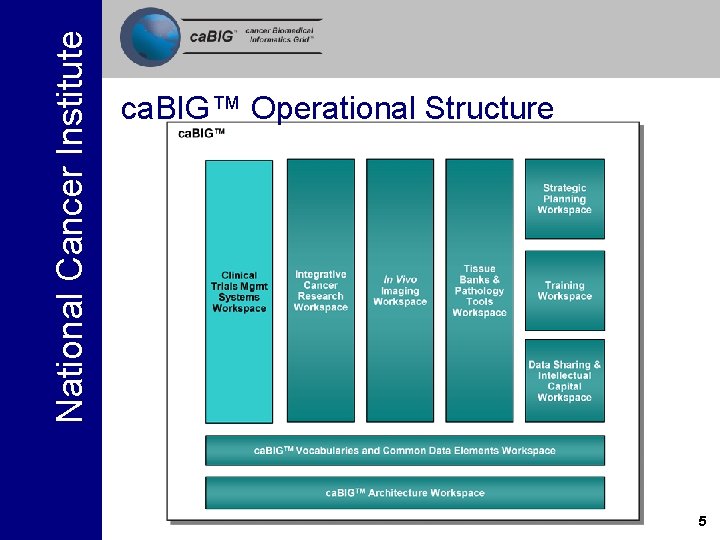
National Cancer Institute ca. BIG™ Operational Structure 5

National Cancer Institute 2006 Clinical Trial Tools Development Activities • ca. AERS • Patient Study Calendar • Lab Data Hub • Making other CTMS systems ca. BIG compatible 6
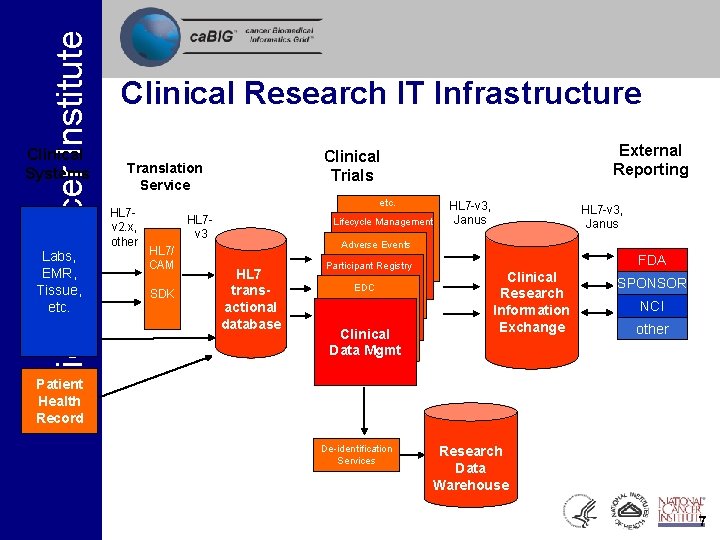
National Cancer Institute Clinical Systems Labs, EMR, Tissue, etc. Clinical Research IT Infrastructure Translation Service HL 7 v 2. x, other External Reporting Clinical Trials etc. HL 7 v 3 HL 7/ CAM SDK Lifecycle Management HL 7 -v 3, Janus Adverse Events HL 7 transactional database Participant Registry EDC Clinical Data Mgmt FDA Clinical Research Information Exchange SPONSOR NCI other Patient Health Record De-identification Services Research Data Warehouse 7

National Cancer Institute Integrated Cancer Research • Microarray Repositories • Data Analysis & Statistics • Informatics for Proteomics • Genome Annotation • Pathways Tools • Translational Tools • Population Sciences and Cancer Control 8
9 National Cancer Institute
10 National Cancer Institute

11 National Cancer Institute

National Cancer Institute Tissue Banks and Pathology Tools • ca. TISSUE Core (WU) – Core specimen handling and tracking functions • ca. TISSUE Clinical Annotation Engine (UPMC) - Annotation of specimens with clinical data • ca. TIES (UPMC) - Text extraction and de-identification of surgical pathology reports 12
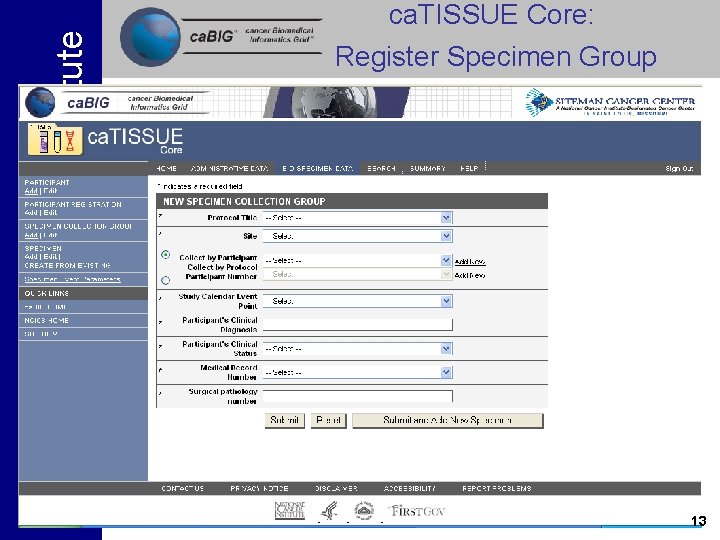
National Cancer Institute ca. TISSUE Core: Register Specimen Group 13
National Cancer Institute ca. IMAGE – Cancer Images Database • ca. IMAGE allows researchers to submit and retrieve images and annotations. • Images are streamed for efficient access. • Researchers can search images based on tissue and diagnosis and experiment information. • Use of common terminology originating from the NCI Enterprise Vocabulary Server (EVS). 14
15 National Cancer Institute

National Cancer Institute ca. BIG™ Compatibility • ca. BIG™ is all about Interoperability • Extensible infrastructure • Ensures partnerships • Evolving • Compatibility Guidelines at https: //cabig. nci. nih. gov/guidelines_documentation – – Key is to create tools for sharing information Expandable and modular software to plug into existing systems so current development efforts are not wasted Encourages relationships between academic, government and industry Compatibility guidelines are being translated into certification procedures 16

National Cancer Institute Interoperability of a system to access and use the parts or equipment of another system Syntactic interoperability Semantic interoperability 17
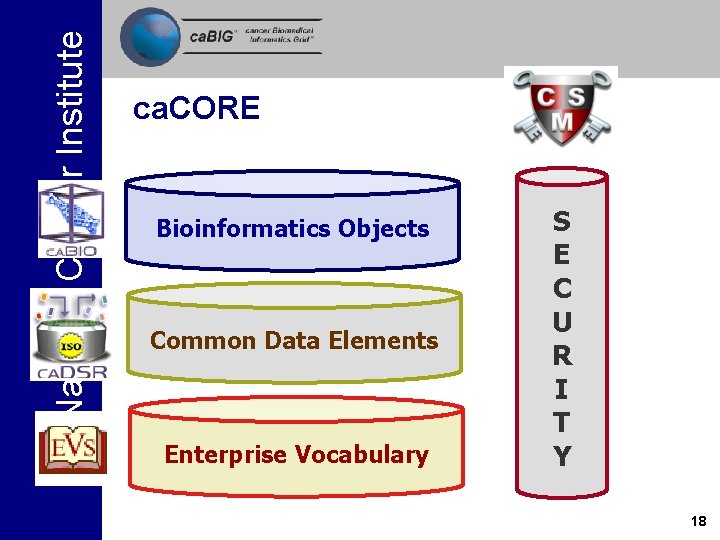
National Cancer Institute ca. CORE Bioinformatics Objects Common Data Elements Enterprise Vocabulary S E C U R I T Y 18

National Cancer Institute Professional Documentation 19

National Cancer Institute ca. CORE Software Development Kit Components • UML Modeling Tool (any with XMI export) • Semantic Connector (concept binding utility) • UML Loader (model registration in ca. DSR) • Codegen (middleware code generator) • Security Adaptor (Common Security Module) • ca. CORE SDK generates a ca. BIG-Silver compliant system 20
21 National Cancer Institute

National Cancer Institute Grid Technology in ca. BIGTM • What is a ‘Grid’ – “A Grid is a system that coordinates resources that are not subject to centralized control using standard, open, general-purpose protocols and interfaces to deliver nontrivial qualities of service. ” - Ian Foster Grid Today, July 20, 2002 • Grid Technology supplies two useful components to a network of computers: – Advertising: Inform the network about the capabilities of new systems – Discovery: Allow users to find resources that meet their needs. • The ca. Grid project is the ‘Grid in ca. BIGTM’; the actual infrastructure that data and analytical services will use to interoperate. • The current ca. Grid is version 0. 5; ca. Grid 1. 0 in December. • The combination of data and analytical service nodes in ca. BIGTM produced a design that utilizes a variety of standard Grid technologies including the Globus Toolkit and OGSA-DAI, DQP, GRAM, etc. 22
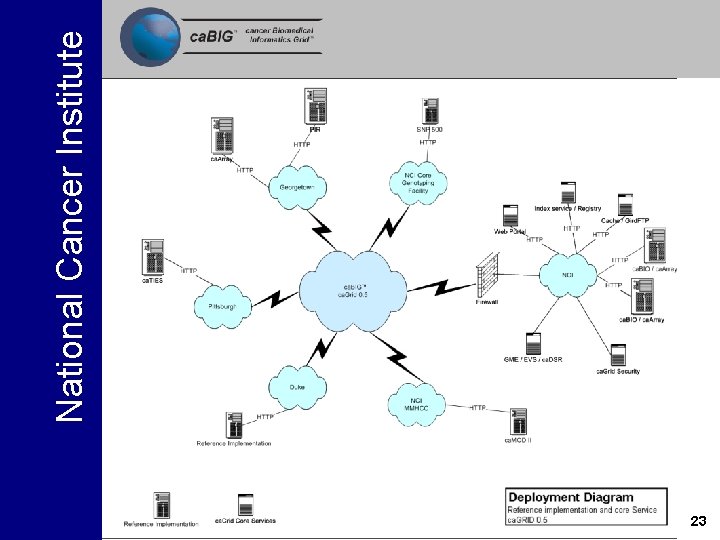
National Cancer Institute Test bed Infrastructure ca. Grid 0. 5 Test Bed 23

National Cancer Institute Cancer Biomedical Informatics Grid™ (ca. BIGTM) • ca. BIG™ infrastructure and tools are widely applicable outside cancer • ca. BIG™ components may be used by anyone 24

National Cancer Institute Contact Information Mary Jo Deering, Ph. D Director for Informatics Dissemination NCI Center for Bioinformatics National Cancer Institute National Institutes of Health, USDHHS 6116 Executive Blvd. - #403 Rockville, MD 20852 (o) 301 -496 -3458 (f) 301 -480 -4222 deeringm@mail. nih. gov 25

National Cancer Institute Additional Background and Detail • The following slides were not included in the presentation. 26

National Cancer Institute Current ca. BIG™ community • NCI-designated Cancer Centers (50) – Academic Centers (integrated into broader biomedical infrastructure) – Stand-alone (community leaders) – Community outreach • NCI Divisions and Programs • National Institutes of Health • Other Government Agencies • Industry • International Groups – Standards development organizations – U. K. ’s National Cancer Research Institute • ~900 active participants 27

National Cancer Institute Four Domain Workspaces and two Cross Cutting Workspaces have been launched DOMAIN WORKSPACE 1 Clinical Trial Management Systems Addresses the need for consistent, open and comprehensive tools for clinical trials management. DOMAIN WORKSPACE 2 Integrative Cancer Research Provides tools and systems to enable integration and sharing of information. DOMAIN WORKSPACE 3 Tissue Banks & Pathology Tools Provides for the integration, development, and implementation of tissue and pathology tools. DOMAIN WORKSPACE 4 Imaging Provides for the sharing and analysis of in vivo imaging data. Responsible for evaluating, developing, and integrating CROSS CUTTING WORKSPACE 1 systems for vocabulary and ontology content, Vocabularies & Common standards, and software systems for content delivery. Data Elements Developing architectural standards and architecture necessary for other workspaces. CROSS CUTTING WORKSPACE 2 Architecture 28

National Cancer Institute Strategic Level Workspaces Data Sharing and Intellectual Capital Training Strategic Planning Addresses issues related to the sharing of data, applications and infrastructure both within the consortium and in the larger cancer research community. Developing strategies for providing training in the use of the ca. BIG developed resources including on-line tutorials, workshops, and training programs. Assists in identifying strategic priorities for the development and evolution of the ca. BIGTM effort. 29

National Cancer Institute REMBRANDT: Building a robust translational research framework for brain tumor studies REpository of Molecular BRAin Neoplasia Da. Ta http: //rembrandt. nci. nih. gov 30
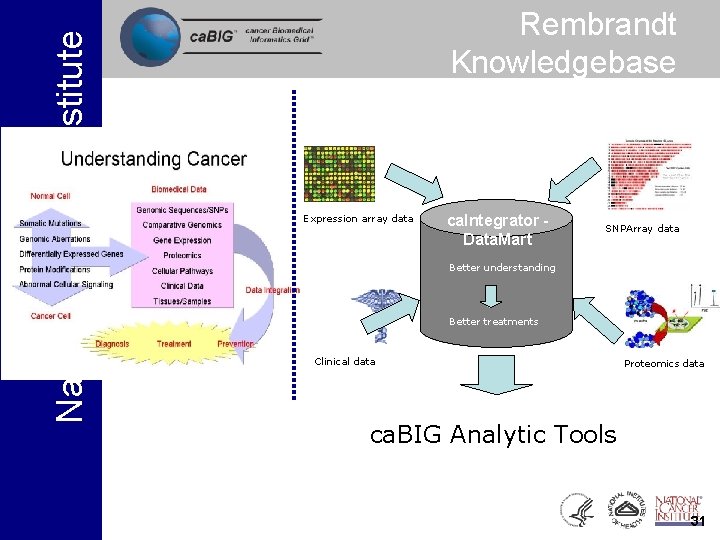
National Cancer Institute Rembrandt Knowledgebase Expression array data ca. Integrator Data. Mart SNPArray data Better understanding Better treatments Clinical data Proteomics data ca. BIG Analytic Tools 31

National Cancer Institute ca. BIGTM Compatibility Guidelines • The ca. BIGTM compatibility guidelines are designed to insure that systems designed in a Federated environment are still interoperable on the ca. BIGTM Grid, both syntactically and semantically • Since achieving interoperability is a process, ca. BIGTM recognizes four levels of compatibility, starting from Legacy (not interoperable) through Bronze, Silver and Gold (fully interoperable) • ca. BIGTM compatibility is all about interfaces rather than the scientific content of the system 32
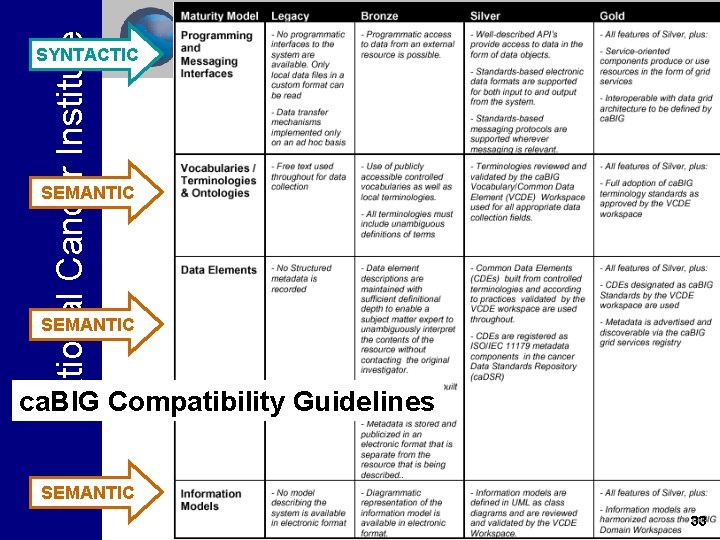
National Cancer Institute SYNTACTIC SEMANTIC ca. BIG Compatibility Guidelines SEMANTIC 33

National Cancer Institute Common Data Elements • What do all those data classes and attributes actually mean, anyway? • Data descriptors or “semantic metadata” required • Computable, commonly structured, reusable units of metadata are “Common Data Elements” or CDEs. • NCI uses the ISO/IEC 11179 standard for metadata structure and registration • Semantics all drawn from Enterprise Vocabulary Service resources 34

National Cancer Institute Cancer Data Standards Repository (ca. DSR) • Basic ca. DSR unit of metadata information to describe a datum is a Common Data Element or CDE • Enterprise-class system for storing metadata, with APIs that give runtime access to both metadata and semantics • Implements the ISO 11179 standard, a flexible model for describing arbitrary metadata • Used to describe metadata associated with clinical case report forms and UML Models 35

National Cancer Institute Enterprise Vocabulary Services • Controlled vocabulary resources for ca. CORE and the cancer research community • Vocabulary Products and Services – NCI Thesaurus – NCI Metathesaurus – External vocabularies • NCI Thesaurus - controlled vocabulary source for metadata – Has excellent coverage of cancer terminology – Expands based on needs for additional terminology – Based on concepts rather than terms – Each concept has a unique identifier or CUI with definitions and synonym 36

National Cancer Institute Data Standards in ca. BIG™ • The V/CDE workspace is responsible for facilitating the development and ratification of Data Standards for ca. BIG™ • Data Standards can be Vocabularies or Common Data Elements (CDEs) with their associated controlled terminology • A ca. BIG™ Data Standard is, in effect, a ‘pre-approved’ mechanism for semantically modeling an attribute or series of attributes in a data object. Ideally, having a standard available shortens development time for other projects that need to present such data • Whenever possible, ca. BIG™ adopts standards that are derived from other standards bodies (HL 7, ISO, USPS, UPU, W 3 C, etc. ) and in general use within our community • In the last year, the V/CDE workspace has developed a consensus driven mechanism for approving Data Standards and applied it to an increasing number of CDEs 37
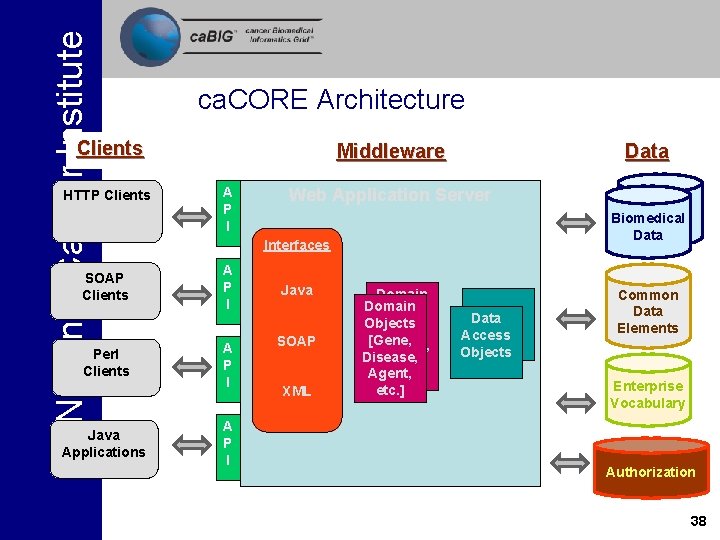
National Cancer Institute ca. CORE Architecture Clients HTTP Clients SOAP Clients Perl Clients Java Applications Middleware A P I Data Web Application Server Biomedical Data Interfaces A P I Java SOAP XML Domain Objects [Gene, Disease, etc. ] Agent, etc. ] Data Access Objects Common Data Elements Enterprise Vocabulary Authorization 38

National Cancer Institute Use cases for ca. Grid • Advertisement – Service Provider composes service metadata describing the service and publishes it to grid. • Discovery – Researcher (or application developer) specifies search criteria describing a service of interest – The research submits the discovery request to a discovery service, which identifies a list of services matching the criteria, and returns the list. • Invocation – Researcher (or application developer) instantiates the grid service and access its resources 39

National Cancer Institute ca. Grid 0. 5 Services • Data Services – ca. BIO: Gene-centric bioinformatics objects • NCICB-Rockville, MD – ca. Array: MAGE-OM compliant microarray repository • NCICB-Rockville, MD • Lombardi Cancer Center-Georgetown, DC – grid. PIR: Protein Information Resource • Lombardi Cancer Center-Georgetown, DC – ca. TIES: Text Information Extraction System for pathology reports • UPMC-Pittsburgh, PA – SNP 500: Polymorphism database with population frequencies • NCI Core Genotyping Facility-Gaithersburg, MD – ca. MOD II: Cancer Model Organism Database • NCI Mouse Models of Human Cancer Consortium (MMHCC) • Analytical Service – RProteomics: Statistical analysis of proteomics data • Duke-Durham, NC 40
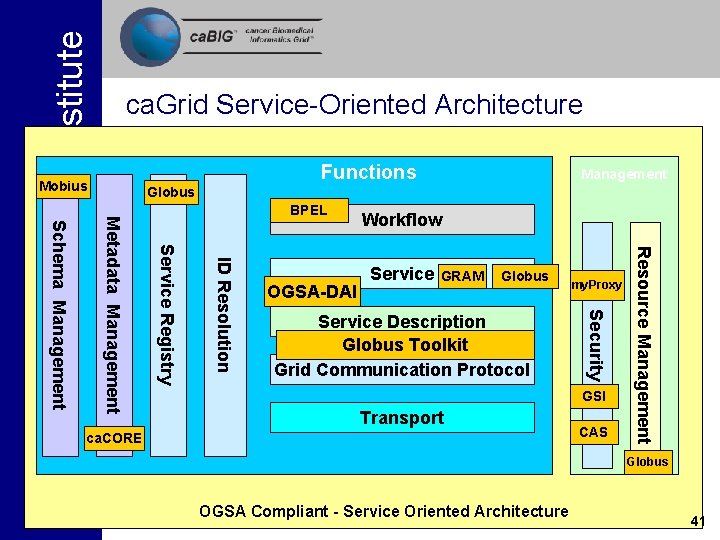
Functions Mobius Management Globus Service GRAM Globus Service Description Globus Toolkit Grid Communication Protocol my. Proxy GSI Transport ca. CORE CAS Resource Management OGSA-DAI Workflow Security ID Resolution Schema Management Metadata Management BPEL Service Registry National Cancer Institute ca. Grid Service-Oriented Architecture Globus OGSA Compliant - Service Oriented Architecture 41

National Cancer Institute Enabling Technology • The NCI provides freely available enabling technology for ca. BIGTM compatibility • These technologies are distributed under a ‘non-viral’ open source license. • ca. CORE – Enterprise Vocabulary Services (EVS) – Cancer Data Standards Repository (ca. DSR) • ca. CORE Software Development Kit – When complete process is followed, the outcome is a ca. BIG ‘Silver’ compliant data system. 42

National Cancer Institute How can my research benefit from ca. BIG™ Tools? • Everything developed by the program is open source and freely available • Training is available at https: //cabig. nci. nih. gov/training • The latest versions of all the software developed as part of the project can be obtained from the ca. BIG™ project gforge site: – http: //gforge. nci. nih. gov 43

National Cancer Institute ca. BIG™: Getting Involved • To get involved with ca. BIG™: – Track ca. BIG™ activities on the NCI’s ca. BIG™ website, https: //cabig. nci. nih. gov/ – Attend ca. BIG™ Annual Meeting, February 5 -7, 2007, Wardman Park Marriott, Washington, DC – Learn about the existing bioinformatics infrastructure, ca. CORE, at https: //ncicb. nci. nih. gov/core – Download currently available ca. BIG™ tools from the ca. BIG™ website at https: //cabig. nci. nih. gov/inventory – Sign up for the ca. BIG™ mailing list at http: //list. nih. gov/archives/cabig_announce. html • Please visit the main ca. BIG™ website for more information: https: //cabig. nci. nih. gov/ 44
- Slides: 45