N Srinivasan Molecular Biophysics Unit Indian Institute of

N. Srinivasan Molecular Biophysics Unit Indian Institute of Science Our Interests MBU Molecular and structural basis of protein action Interactions of proteins in cellular communications URL of our group: http: //pauling. mbu. iisc. ernet. in Support: The Wellcome Trust, London; DBT and CSIR, Delhi

Prediction of protein-protein interactions between a host and a pathogen O. Krishnadev Smita Mohanty K. Anamika Supported by CSIR, DBT and Wellcome Trust

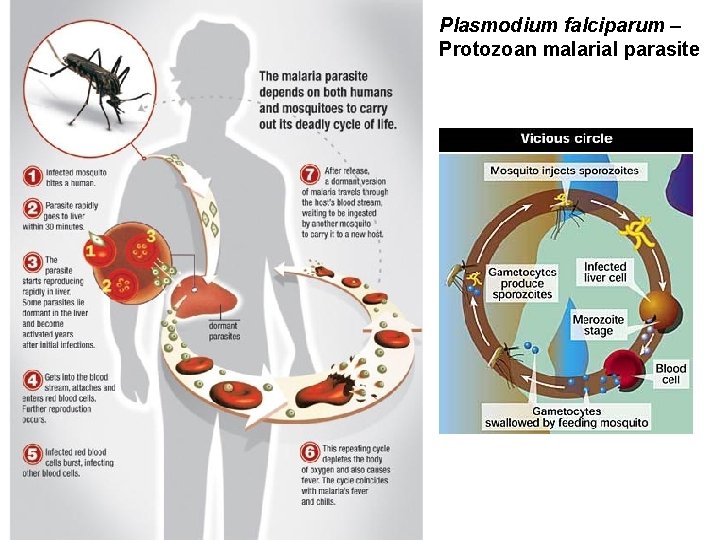
Plasmodium falciparum – Protozoan malarial parasite
Protein-protein interactions network, biological pathways & processes in P. falciparum are dynamic d) e) f) Mohanty, Pandit, Srinivasan (2008) In Protein Interaction Networks and Biological Data Mining (in press) g)
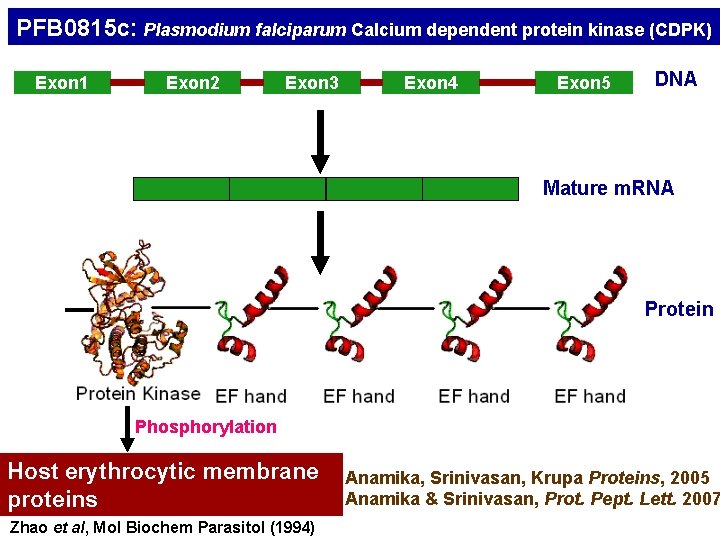
PFB 0815 c: Plasmodium falciparum Calcium dependent protein kinase (CDPK) Exon 1 Exon 2 Exon 3 Exon 4 Exon 5 DNA Mature m. RNA Protein Phosphorylation Host erythrocytic membrane proteins Zhao et al, Mol Biochem Parasitol (1994) Anamika, Srinivasan, Krupa Proteins, 2005 Anamika & Srinivasan, Prot. Pept. Lett. 2007
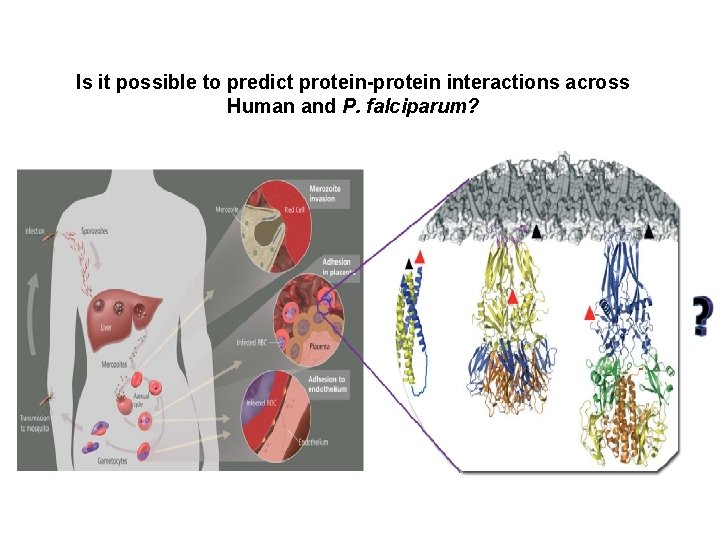
Is it possible to predict protein-protein interactions across Human and P. falciparum?

An Approach for detecting Protein – Protein interactions across host and pathogen organisms Krishnadev & Srinivasan (2008) In Silico Biology
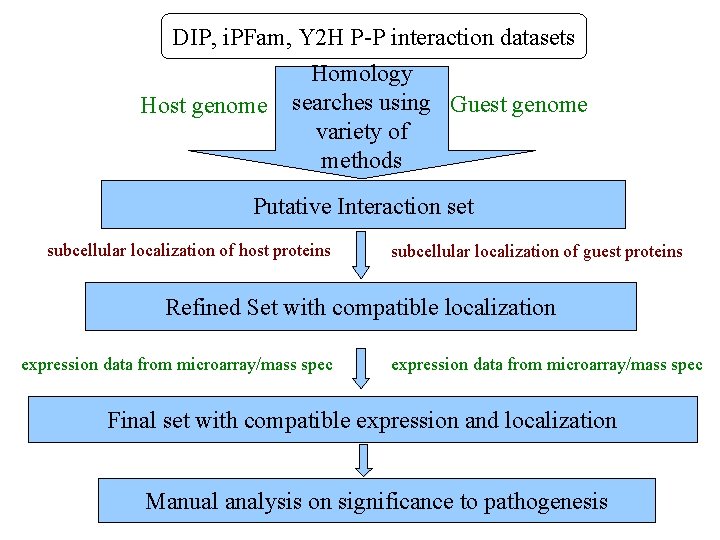
DIP, i. PFam, Y 2 H P-P interaction datasets Host genome Homology searches using Guest genome variety of methods Putative Interaction set subcellular localization of host proteins subcellular localization of guest proteins Refined Set with compatible localization expression data from microarray/mass spec Final set with compatible expression and localization Manual analysis on significance to pathogenesis

An approach for remote homology detection and fold recognition from sequence Gowri, Anand, Krishnadev, Sairam Swamy, Gracelin, Smita Mohanty Anand et. al. , Bioinformatics (2005) Gowri et al Nucl. Acids Res. (2006) Gowri et al Proteins (2007)
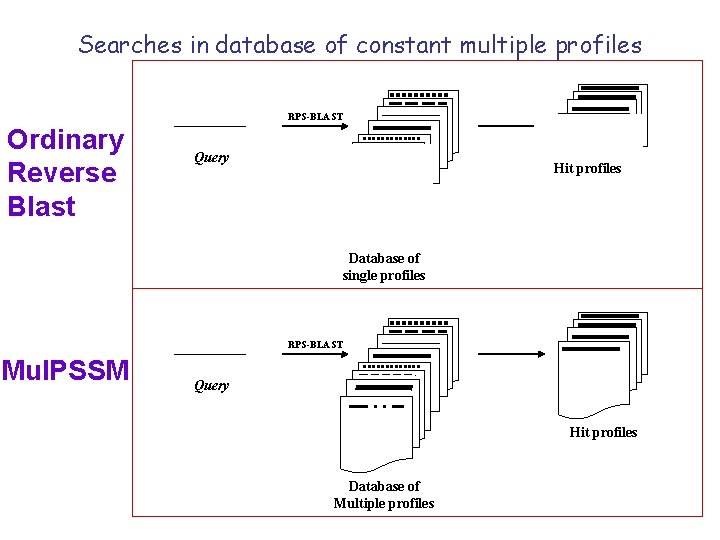
Searches in database of constant multiple profiles RPS-BLAST Ordinary Reverse Blast Query Hit profiles Database of single profiles RPS-BLAST Mul. PSSM Query Hit profiles Database of Multiple profiles

PALI- Database of Phylogeny and ALIgnment of homologous protein structures. S. Balaji, V. S. Gowri, G. Agarwal, S. Pandit, S. Sujatha, C. S. Swamy, O. Krishnadev Balaji et al (2001) Nucl. Acids. Res. Sujatha et al (2001) Bioinformatics Gowri et al (2003) Nucl. Acids. Res. URL: http: //pauling. mbu. iisc. ernet. in/~pali
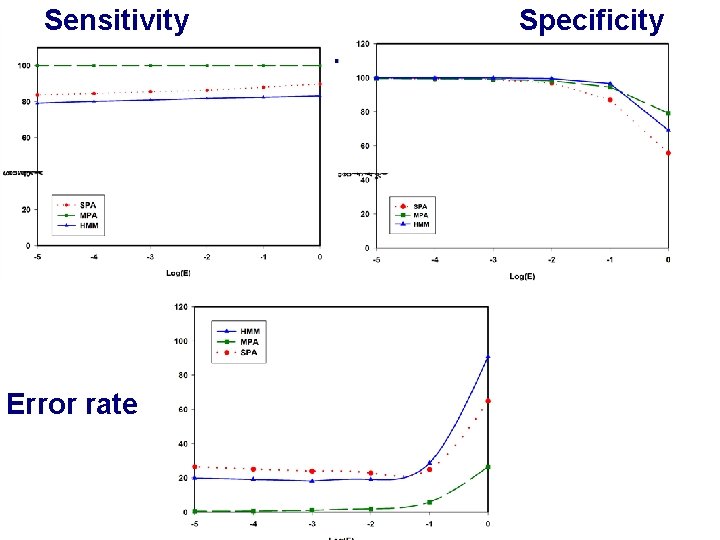
Sensitivity Error rate Specificity
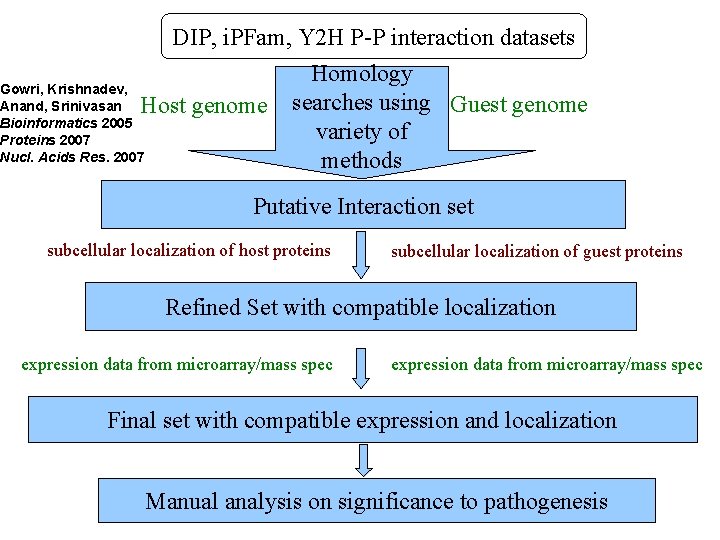
DIP, i. PFam, Y 2 H P-P interaction datasets Gowri, Krishnadev, Anand, Srinivasan Bioinformatics 2005 Proteins 2007 Nucl. Acids Res. 2007 Host genome Homology searches using Guest genome variety of methods Putative Interaction set subcellular localization of host proteins subcellular localization of guest proteins Refined Set with compatible localization expression data from microarray/mass spec Final set with compatible expression and localization Manual analysis on significance to pathogenesis
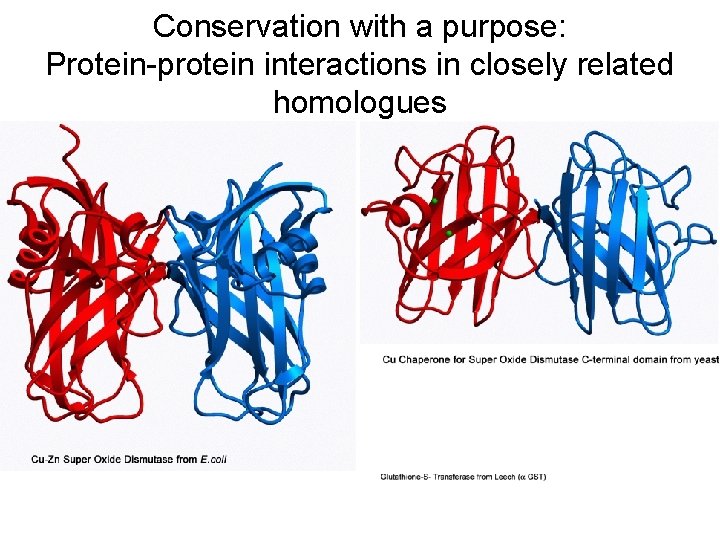
Conservation with a purpose: Protein-protein interactions in closely related homologues
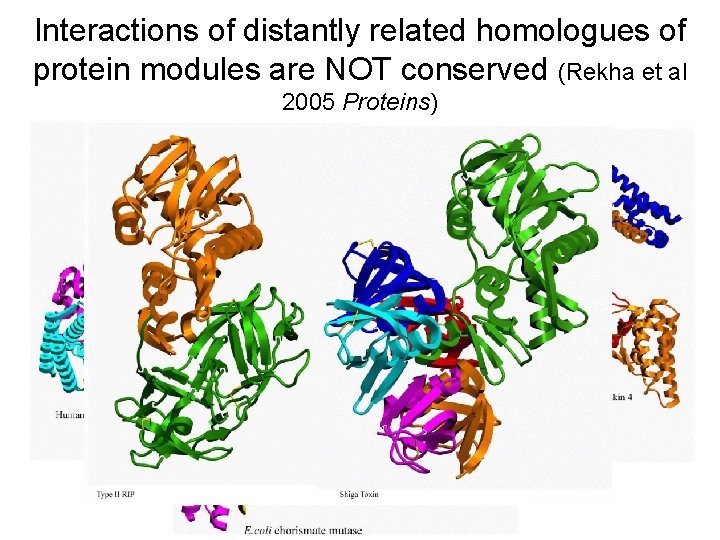
Interactions of distantly related homologues of protein modules are NOT conserved (Rekha et al 2005 Proteins)
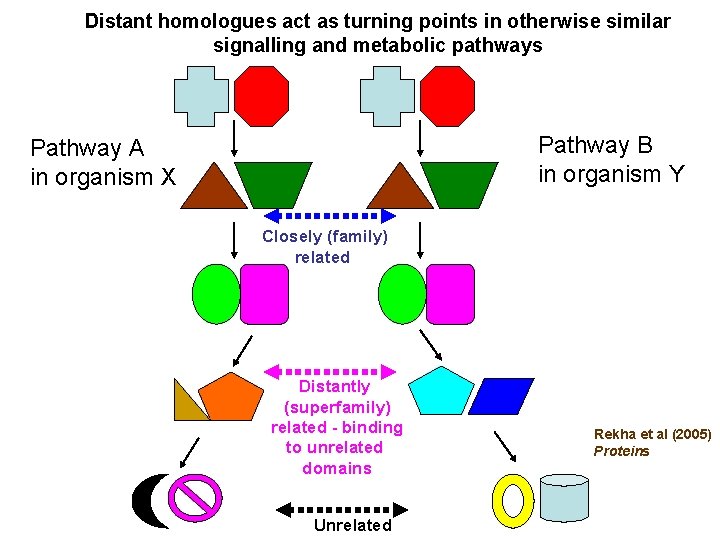
Distant homologues act as turning points in otherwise similar signalling and metabolic pathways Pathway B in organism Y Pathway A in organism X Closely (family) related Distantly (superfamily) related - binding to unrelated domains Unrelated Rekha et al (2005) Proteins
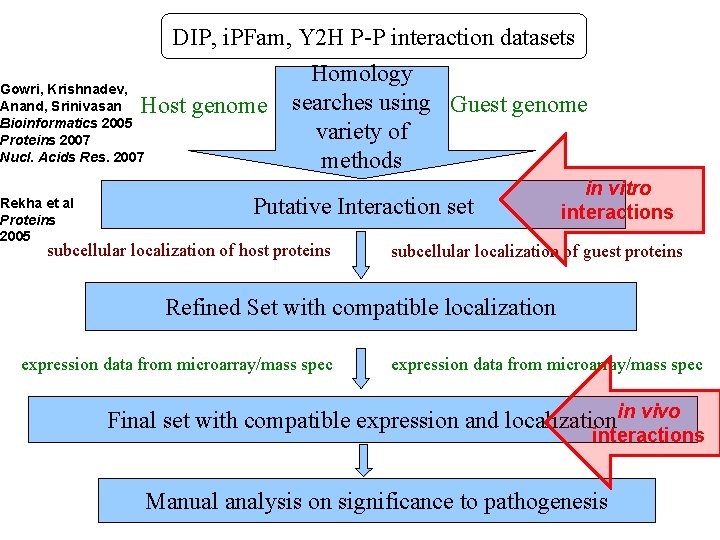
DIP, i. PFam, Y 2 H P-P interaction datasets Gowri, Krishnadev, Anand, Srinivasan Bioinformatics 2005 Proteins 2007 Nucl. Acids Res. 2007 Host genome Rekha et al Proteins 2005 Homology searches using Guest genome variety of methods Putative Interaction set subcellular localization of host proteins in vitro interactions subcellular localization of guest proteins Refined Set with compatible localization expression data from microarray/mass spec Final set with compatible expression and localizationin vivo interactions Manual analysis on significance to pathogenesis

Host – Pathogen pairs studied Phage - Bacteria Herpes Virus - Human M. tuberculosis – Human M. leprae – Human P. falciparum – Human C. diptheriae - Human

Our approach has a good success rate Example, herpes virus and human P-P interactions • 12 experimentally studied interactions using two-hybrid and co-immunoprecipitation. • 11 predicted’ by our method. • Novel predictions • Viral ribonucleotide reductase small subunit with human p 53 protein. This can point to a mechanism of regulating apoptosis which occurs upon viral infection. • Known PPI in other organisms • Viral uracil DNA glycosylase with human PCNA important for replication. Viral UDG shown to localize to DNA replisome.
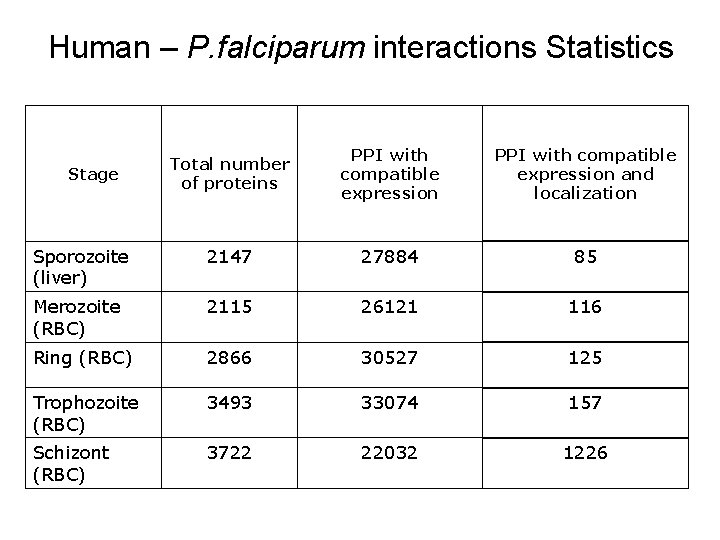
Human – P. falciparum interactions Statistics Total number of proteins PPI with compatible expression and localization Sporozoite (liver) 2147 27884 85 Merozoite (RBC) 2115 26121 116 Ring (RBC) 2866 30527 125 Trophozoite (RBC) 3493 33074 157 Schizont (RBC) 3722 22032 1226 Stage
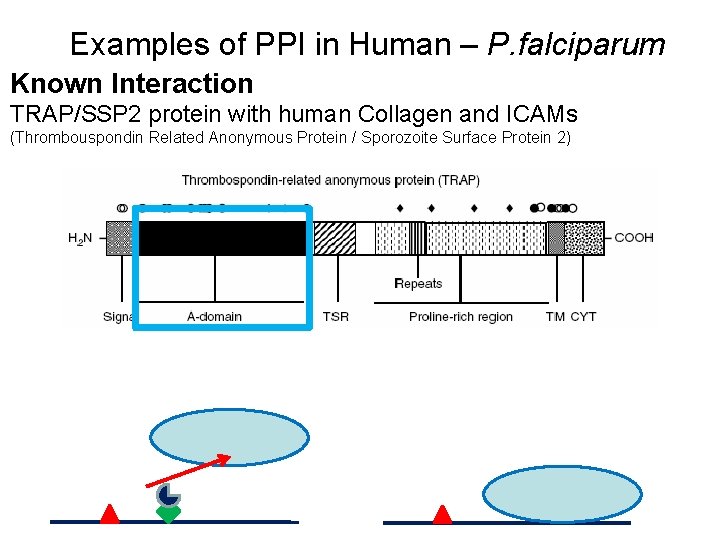
Examples of PPI in Human – P. falciparum Known Interaction TRAP/SSP 2 protein with human Collagen and ICAMs (Thrombouspondin Related Anonymous Protein / Sporozoite Surface Protein 2)

Recognition of Interactions TRAP/SSP 2 protein with human Collagen and ICAMs • TRAP/SSP 2 activity disruption leads to a marked decrease in the ability to invade the human hepatocytes. • TRAP/SSP 2 shown to be present on the sporozoite membrane. • Experimental evidence points out a mechanism of gliding motility in apicomplexans which involves non-specific binding of pathogen surface proteins to human surface proteins. • Presence of A-domain which is known to interact with different human cell surface proteins
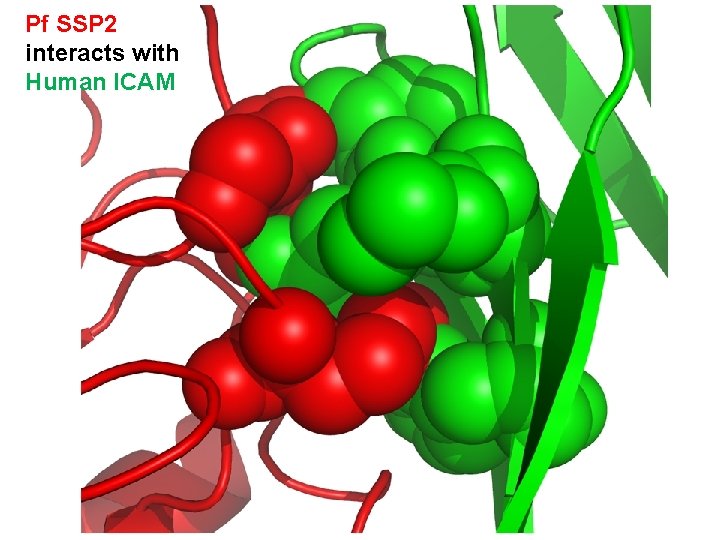
Pf SSP 2 interacts with Human ICAM
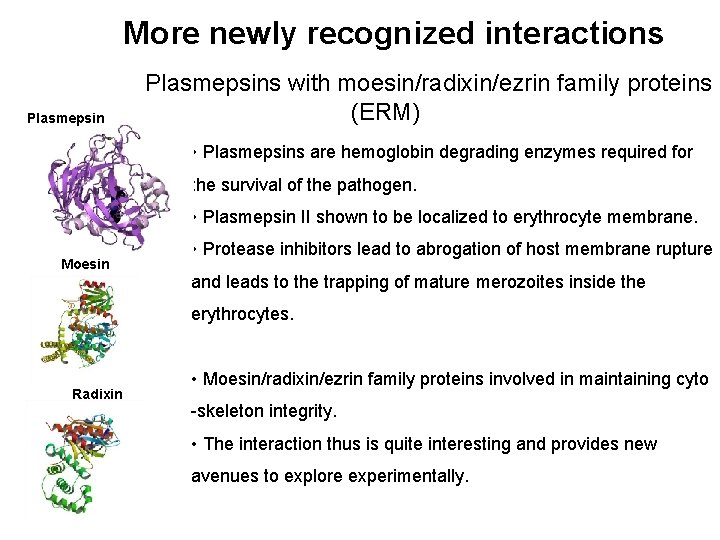
More newly recognized interactions Plasmepsins with moesin/radixin/ezrin family proteins (ERM) • Plasmepsins are hemoglobin degrading enzymes required for the survival of the pathogen. • Plasmepsin II shown to be localized to erythrocyte membrane. Moesin • Protease inhibitors lead to abrogation of host membrane rupture and leads to the trapping of mature merozoites inside the erythrocytes. Radixin • Moesin/radixin/ezrin family proteins involved in maintaining cyto -skeleton integrity. • The interaction thus is quite interesting and provides new avenues to explore experimentally.
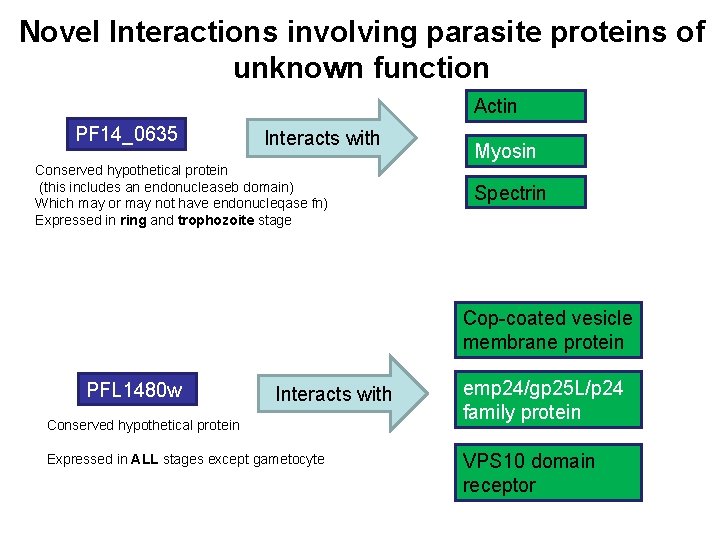
Novel Interactions involving parasite proteins of unknown function Actin PF 14_0635 Interacts with Conserved hypothetical protein (this includes an endonucleaseb domain) Which may or may not have endonucleqase fn) Expressed in ring and trophozoite stage Myosin Spectrin Cop-coated vesicle membrane protein PFL 1480 w Interacts with Conserved hypothetical protein Expressed in ALL stages except gametocyte emp 24/gp 25 L/p 24 family protein VPS 10 domain receptor
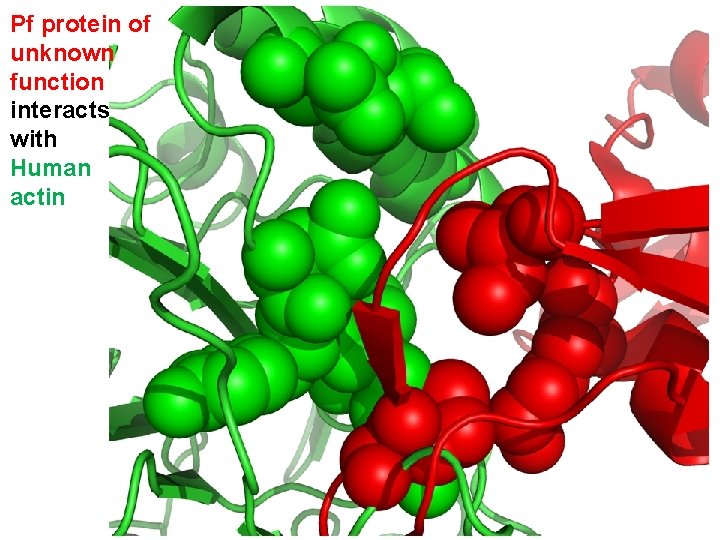
Pf protein of unknown function interacts with Human actin
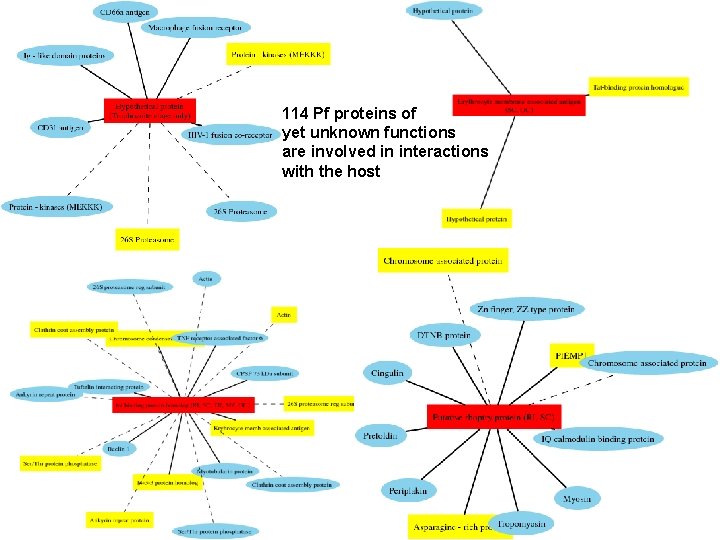
114 Pf proteins of yet unknown functions are involved in interactions with the host
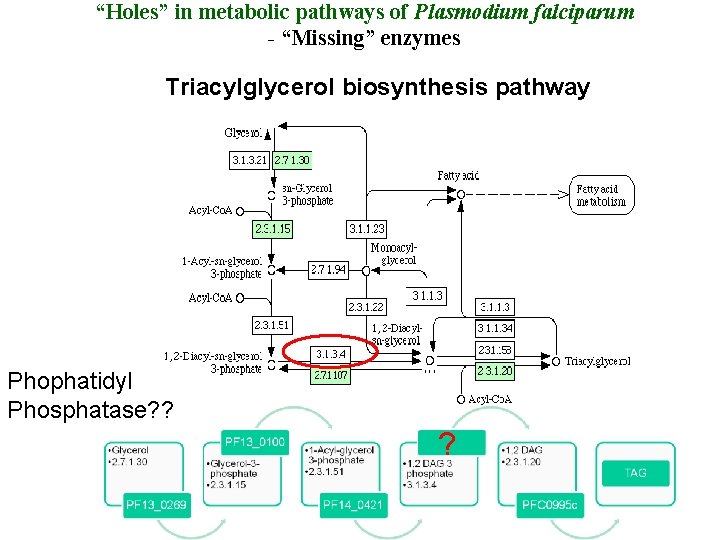
“Holes” in metabolic pathways of Plasmodium falciparum - “Missing” enzymes Triacylglycerol biosynthesis pathway Phophatidyl Phosphatase? ? ?

P. falciparum genome sequence reveals a total of 5363 proteins. Nature 419: 498 -511 Non-enzymes • 75 different physiological processes are characterized encompassing 696 proteins. • Several enzymes involved in metabolic pathways are not yet identified. • Approximately 57 enzymatic steps are still undetected
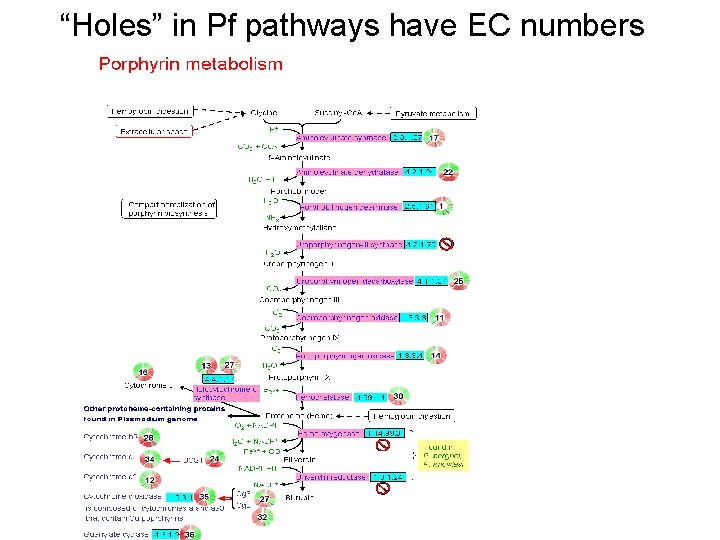
“Holes” in Pf pathways have EC numbers
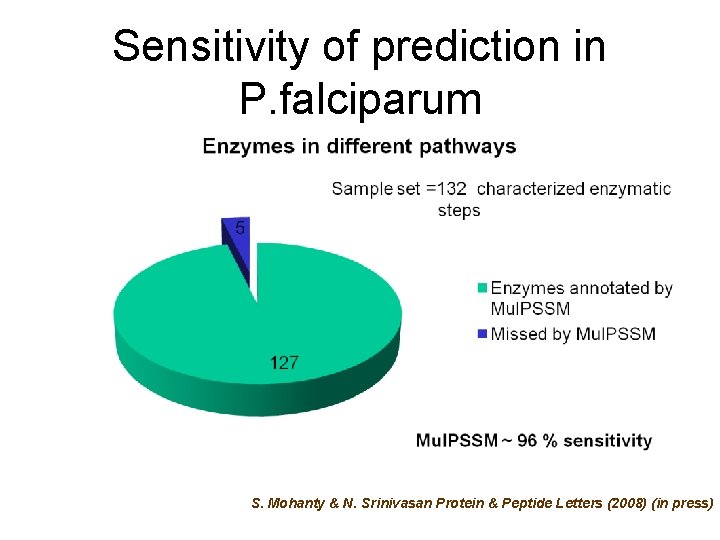
Sensitivity of prediction in P. falciparum S. Mohanty & N. Srinivasan Protein & Peptide Letters (2008) (in press)
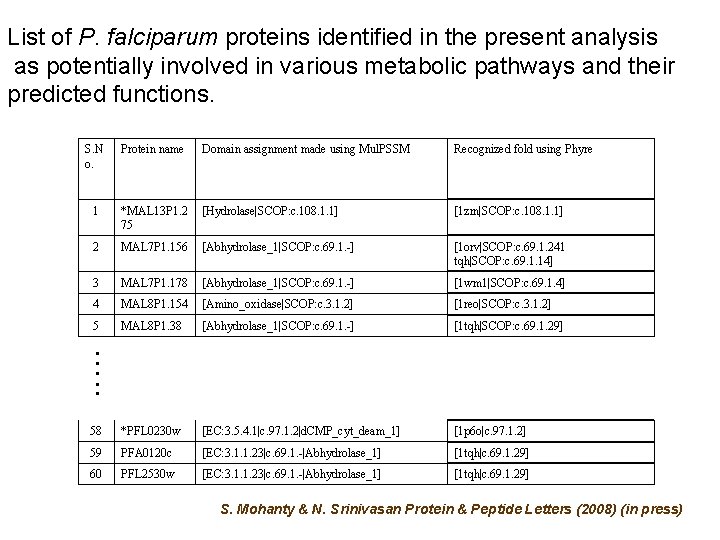
List of P. falciparum proteins identified in the present analysis as potentially involved in various metabolic pathways and their predicted functions. S. N o. Protein name Domain assignment made using Mul. PSSM Recognized fold using Phyre 1 *MAL 13 P 1. 2 75 [Hydrolase|SCOP: c. 108. 1. 1] [1 zrn|SCOP: c. 108. 1. 1] 2 MAL 7 P 1. 156 [Abhydrolase_1|SCOP: c. 69. 1. -] [1 orv|SCOP: c. 69. 1. 241 tqh|SCOP: c. 69. 1. 14] 3 MAL 7 P 1. 178 [Abhydrolase_1|SCOP: c. 69. 1. -] [1 wm 1|SCOP: c. 69. 1. 4] 4 MAL 8 P 1. 154 [Amino_oxidase|SCOP: c. 3. 1. 2] [1 reo|SCOP: c. 3. 1. 2] 5 MAL 8 P 1. 38 [Abhydrolase_1|SCOP: c. 69. 1. -] [1 tqh|SCOP: c. 69. 1. 29] …. . 58 *PFL 0230 w [EC: 3. 5. 4. 1|c. 97. 1. 2|d. CMP_cyt_deam_1] [1 p 6 o|c. 97. 1. 2] 59 PFA 0120 c [EC: 3. 1. 1. 23|c. 69. 1. -|Abhydrolase_1] [1 tqh|c. 69. 1. 29] 60 PFL 2530 w [EC: 3. 1. 1. 23|c. 69. 1. -|Abhydrolase_1] [1 tqh|c. 69. 1. 29] S. Mohanty & N. Srinivasan Protein & Peptide Letters (2008) (in press)
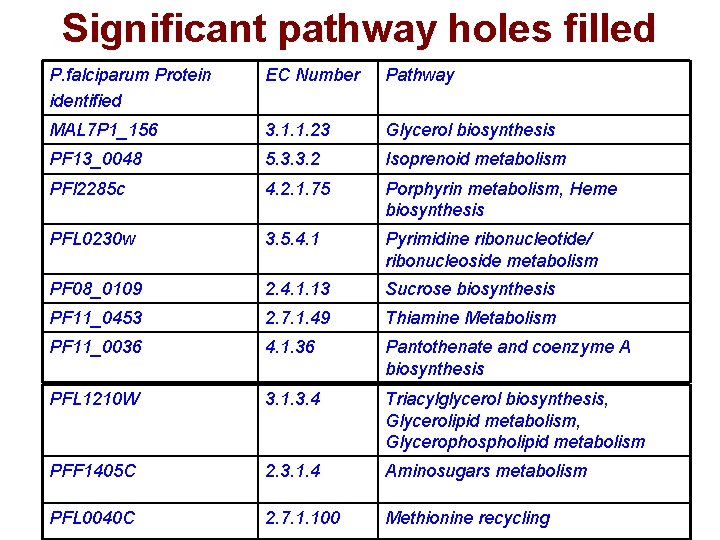
Significant pathway holes filled P. falciparum Protein identified EC Number Pathway MAL 7 P 1_156 3. 1. 1. 23 Glycerol biosynthesis PF 13_0048 5. 3. 3. 2 Isoprenoid metabolism PFl 2285 c 4. 2. 1. 75 Porphyrin metabolism, Heme biosynthesis PFL 0230 w 3. 5. 4. 1 Pyrimidine ribonucleotide/ ribonucleoside metabolism PF 08_0109 2. 4. 1. 13 Sucrose biosynthesis PF 11_0453 2. 7. 1. 49 Thiamine Metabolism PF 11_0036 4. 1. 36 Pantothenate and coenzyme A biosynthesis PFL 1210 W 3. 1. 3. 4 Triacylglycerol biosynthesis, Glycerolipid metabolism, Glycerophospholipid metabolism PFF 1405 C 2. 3. 1. 4 Aminosugars metabolism PFL 0040 C 2. 7. 1. 100 Methionine recycling
![Triacylglycerol biosynthesis ? Phophatidyl phosphatase [E. C 3. 1. 3. 4] Triacylglycerol biosynthesis ? Phophatidyl phosphatase [E. C 3. 1. 3. 4]](http://slidetodoc.com/presentation_image/cda1e1ee4b1cd84d32fe560fa22120bb/image-35.jpg)
Triacylglycerol biosynthesis ? Phophatidyl phosphatase [E. C 3. 1. 3. 4]
![Phophatidyl phosphatase [E. C 3. 1. 3. 4] • Catalyzes hydrolysis of L-phosphatidate to Phophatidyl phosphatase [E. C 3. 1. 3. 4] • Catalyzes hydrolysis of L-phosphatidate to](http://slidetodoc.com/presentation_image/cda1e1ee4b1cd84d32fe560fa22120bb/image-36.jpg)
Phophatidyl phosphatase [E. C 3. 1. 3. 4] • Catalyzes hydrolysis of L-phosphatidate to 1, 2 -diacylglycerol and phosphate. • Involved in: • Glycerophospholipid metabolism • Glycerolipid metabolism • Triacylglycerol biosyntheisis • Essential for membrane biogenesis and also in production of energy. • Identified PAP 2 superfamily related protein (PFL 1210 w, Id=15%, E-value=: 2 e-08 in Mul. PSSM search ). Members known to act as phophatidyl phosphatases
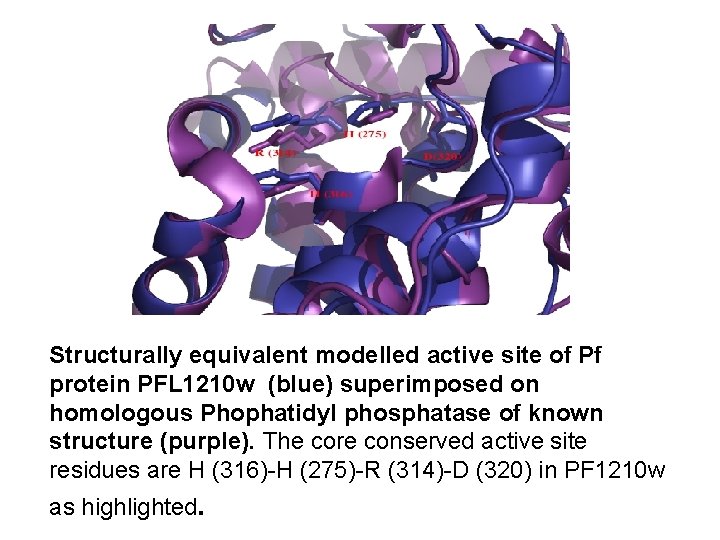
Structurally equivalent modelled active site of Pf protein PFL 1210 w (blue) superimposed on homologous Phophatidyl phosphatase of known structure (purple). The core conserved active site residues are H (316)-H (275)-R (314)-D (320) in PF 1210 w as highlighted.

Increase in PLP, DAG and TAG in Merozoites stage, essential for parasitophorous vacuolar membrane synthesis. Microbiol Rev, 1979, 43, 453 -495 Gene (Function) PF 13_0269 (EC 2. 7. 1. 30) PF 13_0100 (EC 2. 3. 1. 15) PF 14_0421 (EC 3. 1. 3. 4) PFL 1210 w (EC 3. 1. 3. 4) PF 14_0681 (EC 2. 7. 1. 107) PFC 0995 c (EC 2. 3. 1. 20) SPO RIN TRO SCH MER GAM

Application of prediction of hostpathogen protein-protein interaction to Mycobacterial organisms
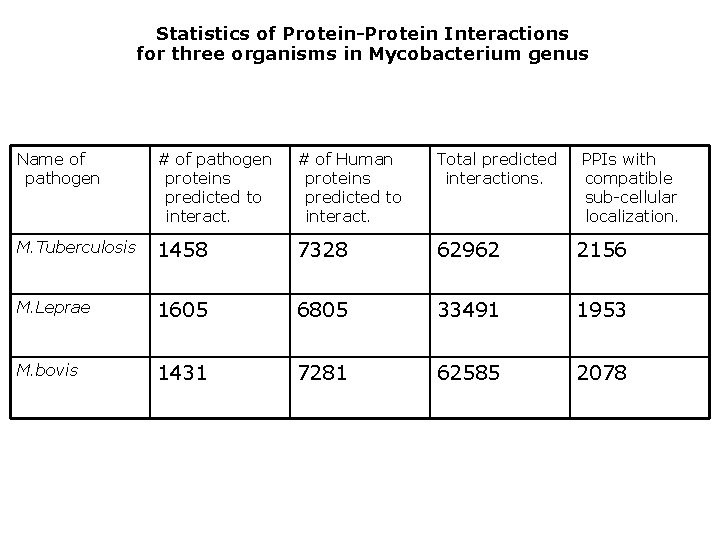
Statistics of Protein-Protein Interactions for three organisms in Mycobacterium genus Name of pathogen # of pathogen proteins predicted to interact. # of Human proteins predicted to interact. Total predicted interactions. PPIs with compatible sub-cellular localization. M. Tuberculosis 1458 7328 62962 2156 M. Leprae 1605 6805 33491 1953 M. bovis 1431 7281 62585 2078
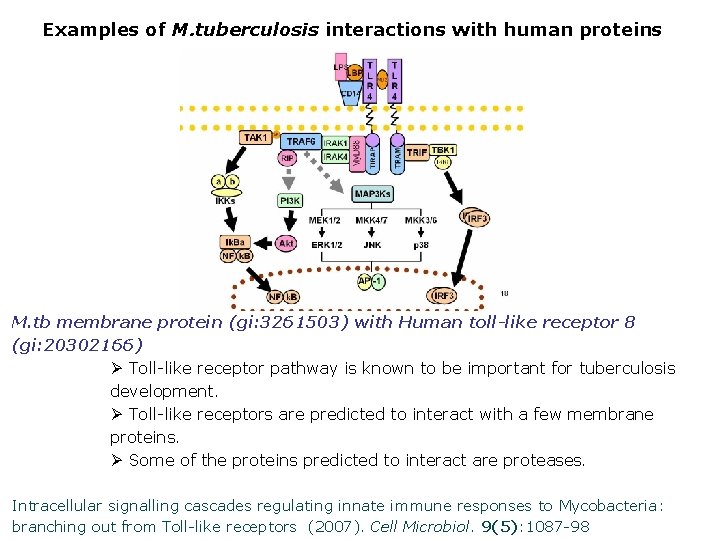
Examples of M. tuberculosis interactions with human proteins M. tb membrane protein (gi: 3261503) with Human toll-like receptor 8 (gi: 20302166) Ø Toll-like receptor pathway is known to be important for tuberculosis development. Ø Toll-like receptors are predicted to interact with a few membrane proteins. Ø Some of the proteins predicted to interact are proteases. Intracellular signalling cascades regulating innate immune responses to Mycobacteria: branching out from Toll-like receptors (2007). Cell Microbiol. 9(5): 1087 -98

Examples of M. tuberculosis interactions with human proteins M. tb 10 k. Da antigen (gi: 1449369) with Human TNF receptor-associated protein 1 (gi: 7706485) Ø 10 k. Da antigen known to be important for pathogenesis from various experiments. Ø M. tuberculosis interferes with TNF signalling which is important for immune evasion. Ø Interaction predicted using DIP interaction between two proteins from Yeast. Ø Predicted interaction can help in understanding TNF signalling in infected macrophages. Effect of Mycobacterium tuberculosis-Specific 10 -Kilodalton Antigen on Macrophage Release of Tumor Necrosis Factor Alpha and Nitric Oxide. (2002) Infection and Immunity 70(12): 6558 -6566

Examples of M. tuberculosis interactions with human proteins M. tb Probable respiratory nitrogen reductase (nar. G, gi: 2117199) with Human apoptosis-inducing factor (gi : 21389617) ØM. tb infected macrophages show abnormal apoptosis. ØThe mechanisms by which M. tb triggers this transformation is unknown. Ønar. G known to be important for survival under in vitro hypoxic conditions. ØReports suggest nar. G deletion mutants can survive in vivo conditions. ØPredicted interaction can point to a novel function for nar. G in the pathogen. ØIs nar. G involved in the apoptosis regulation in infected macrophages ? Apoptosis-Inducing Factor Participation in Bovine Macrophage Mycobacterium bovis. Induced Caspase-Independent Cell Death. (2007) Infect Immun. 75(3): 1223– 1228.
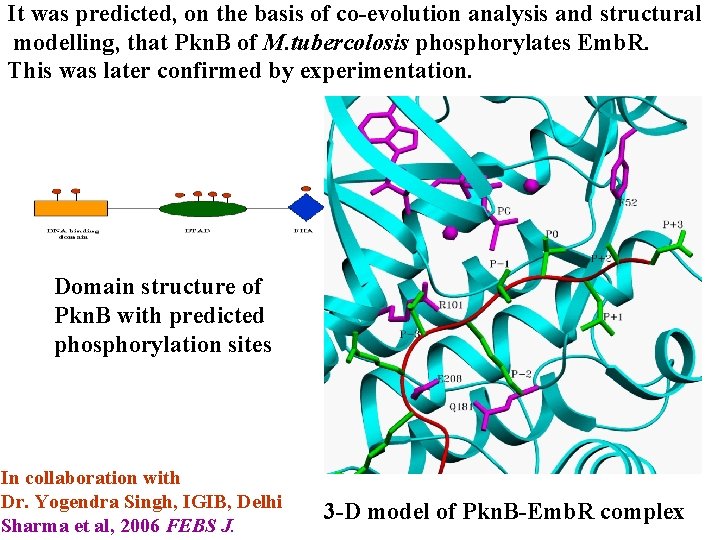
It was predicted, on the basis of co-evolution analysis and structural modelling, that Pkn. B of M. tubercolosis phosphorylates Emb. R. This was later confirmed by experimentation. Domain structure of Pkn. B with predicted phosphorylation sites In collaboration with Dr. Yogendra Singh, IGIB, Delhi Sharma et al, 2006 FEBS J. 3 -D model of Pkn. B-Emb. R complex
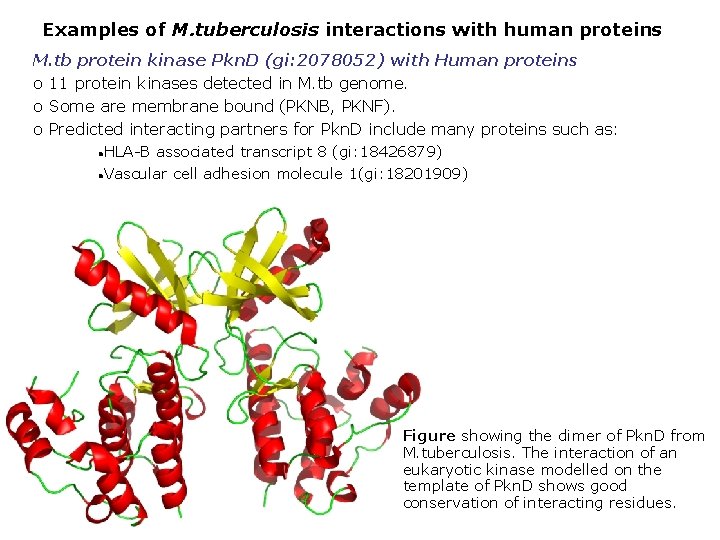
Examples of M. tuberculosis interactions with human proteins M. tb protein kinase Pkn. D (gi: 2078052) with Human proteins o 11 protein kinases detected in M. tb genome. o Some are membrane bound (PKNB, PKNF). o Predicted interacting partners for Pkn. D include many proteins such as: HLA-B associated transcript 8 (gi: 18426879) Vascular cell adhesion molecule 1(gi: 18201909) Figure showing the dimer of Pkn. D from M. tuberculosis. The interaction of an eukaryotic kinase modelled on the template of Pkn. D shows good conservation of interacting residues.

M. tuberculosis interactions with human proteins – Novel examples M. tb probable export protein (gi: 1314027) with Human proteins • Many pathogen proteins are exported to host cytoplasm. • Exported proteins play an important role in pathophysiology. • Function of many of the proteins exported is not known. • Predicted interaction between an export protein and actin is thus interesting. • Predicted interactions include many phosphatases in human. M. tb conserved hypothetical protein (gi: 2104343) with Bcl-XL interacting protein (gi: 20070384) • M. tb arrests the apoptosis of infected macriopahges. • Mechanism is not completely understood. • Bcl-XL protein accessory protein involved in apoptosis. • Predicted interaction could thus help us in understanding apoptosis signalling in TB. Mycobacterium tuberculosis Promotes Apoptosis in Human Neutrophils by Activating Caspase-3 and Altering Expression of Bax/Bcl-x. L Via an Oxygen-Dependent Pathway (2002) The Journal of Immunology 168: 6358 -6365.
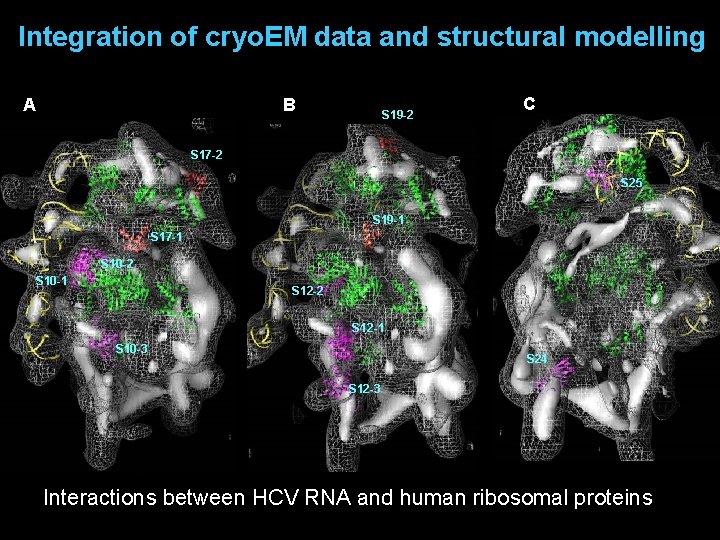
Integration of cryo. EM data and structural modelling A B S 19 -2 C S 17 -2 S 25 S 19 -1 S 17 -1 S 10 -2 S 10 -1 S 12 -2 S 12 -1 S 10 -3 S 24 S 12 -3 Interactions between HCV RNA and human ribosomal proteins

Conclusions • Within organism interactions can be used to detect across organisms interactions. • Combining physiological data from different sources leads to recognition of potential interactions during pathogenesis. • Previously unknown human - parasite protein-protein interactions were predicted involving parasite proteins known to be important for pathogenesis of P. falciparum and M. tubercolosis • We could detect interactions involving parasite proteins of yet unknown function. This could improve our understanding of the pathogenesis process • Our approach is general so it could be applied to any pair of host-guest organisms. This is helpful as currently no large-scale experimental approach is available for identification of comprehensive list of hostpathogen protein-protein interactions.

Acknowledgements Smita Mohanty K. Anamika (P. falciparum pathways) (P. falciparum kinomics) V. S. Gowri Nidhi Tyagi (Pf hypothetical proteins) O. Krishnadev (Host-pathogen interactions) Mul. PSSM team V. S. Gowri B. Anand Tina Graceline O. Krishnadev Smita Mohanty (Pf-specific protein families) Supported by The Wellcome Trust, London Department of Biotechnology, New Delhi Council of Scientific and Industrial Research, New Delhi

THANK YOU
- Slides: 50