Multivariate Genetic Analysis Boulder 2004 Phenotypic Cholesky 1

Multivariate Genetic Analysis Boulder 2004
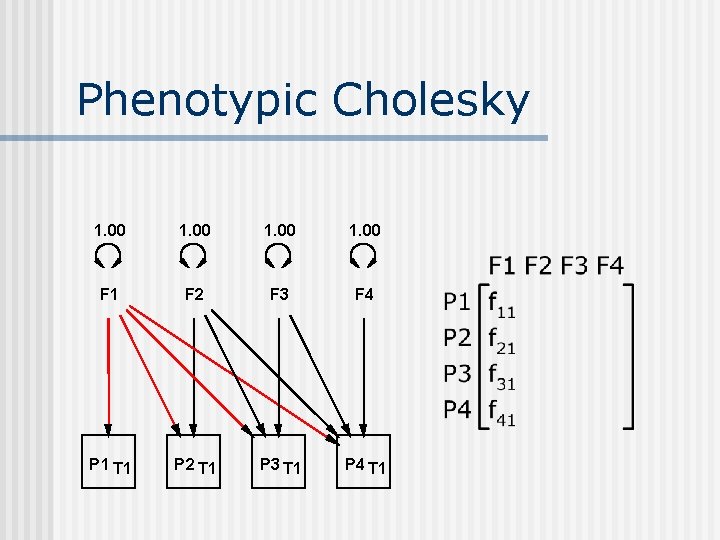
Phenotypic Cholesky 1. 00 F 1 F 2 F 3 F 4 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1
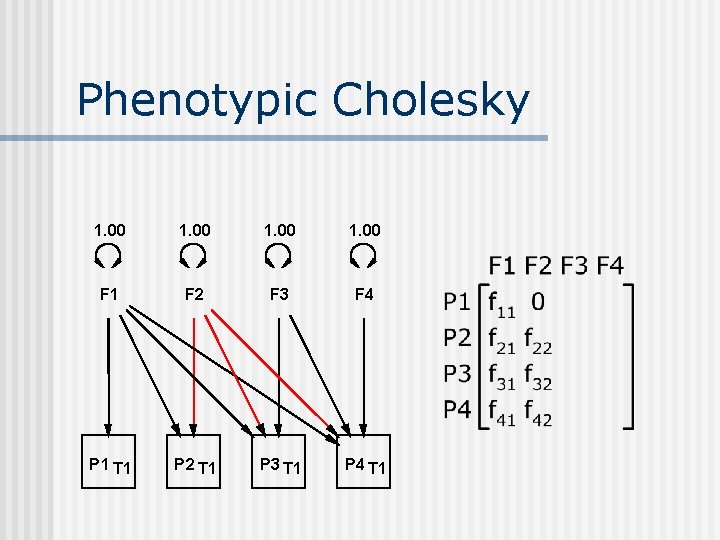
Phenotypic Cholesky 1. 00 F 1 F 2 F 3 F 4 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1
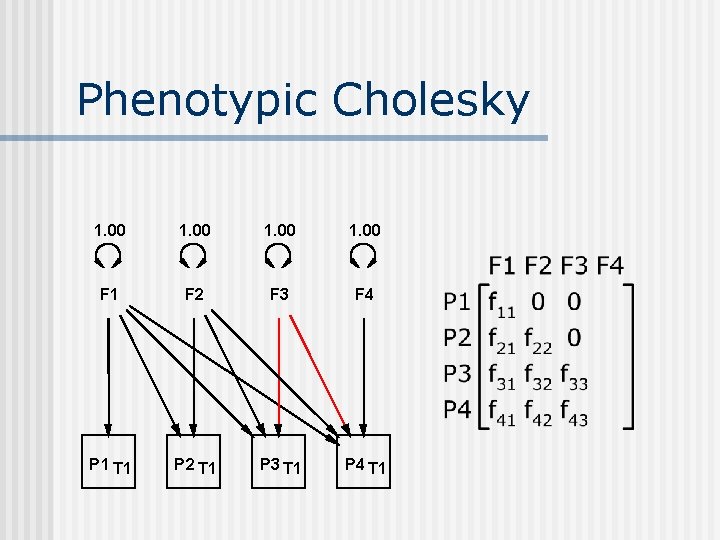
Phenotypic Cholesky 1. 00 F 1 F 2 F 3 F 4 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1
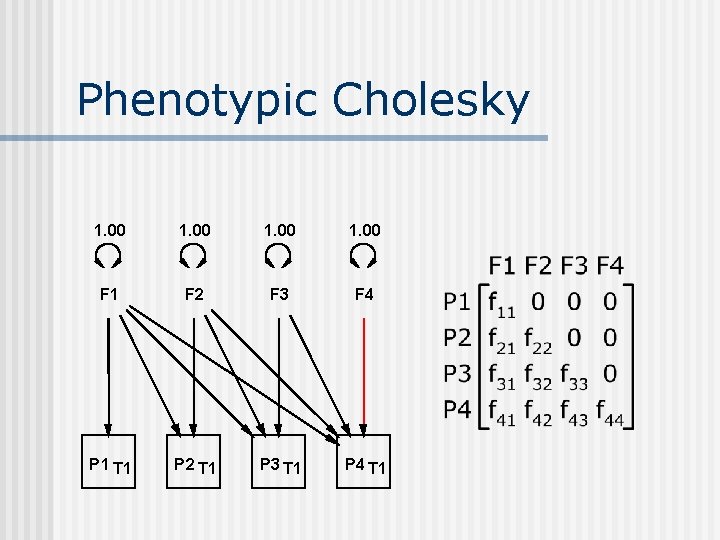
Phenotypic Cholesky 1. 00 F 1 F 2 F 3 F 4 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1
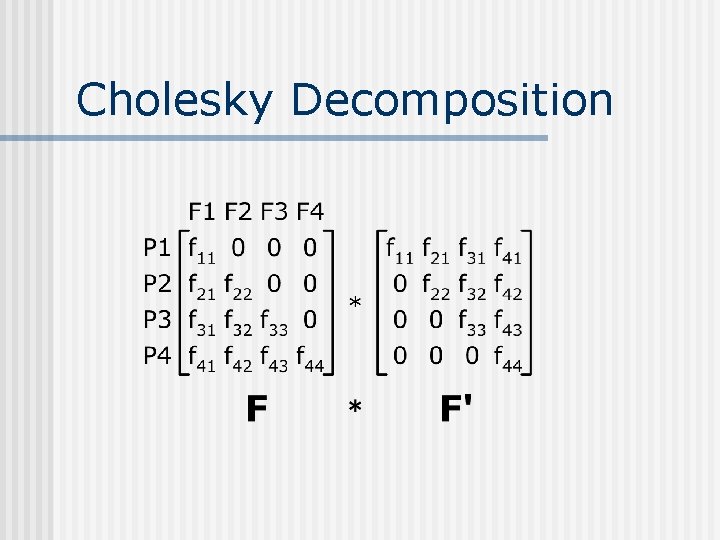
Cholesky Decomposition

Cholesky Example n n Script: f: hmaesa 17chol. mx 3 MRI measures – 2 IQ subtests: n n n var 1 var 2 var 3 var 4 var 5 = = = cerebellum grey matter white matter calculation letters and numbers Data: f: hmaesa 17mri-iqfactor. rec n n T: hmaes/a 17mri-iq-mz. dat T: hmaesa 17mri-iq-dz. dat

Cholesky #define nvar 5 Begin Matrices; X lower nvar Free ! genetic structure Y lower nvar Free ! shared environment structure Z lower nvar Free ! specific environment structure M full 1 nvar Free ! grand means End Matrices; Begin Algebra; A= X*X'; ! additive genetic covariance C= Y*Y'; ! shared environment covariance E= Z*Z'; ! nonshared environment covariance End Algebra;

Standardization Begin Algebra; R=A+C+E; ! total variance S=(sqrt(I. R))~; ! diagonal matrix of standard deviations P=S*X| S*Y| S*Z; ! standardized estimates End Algebra;
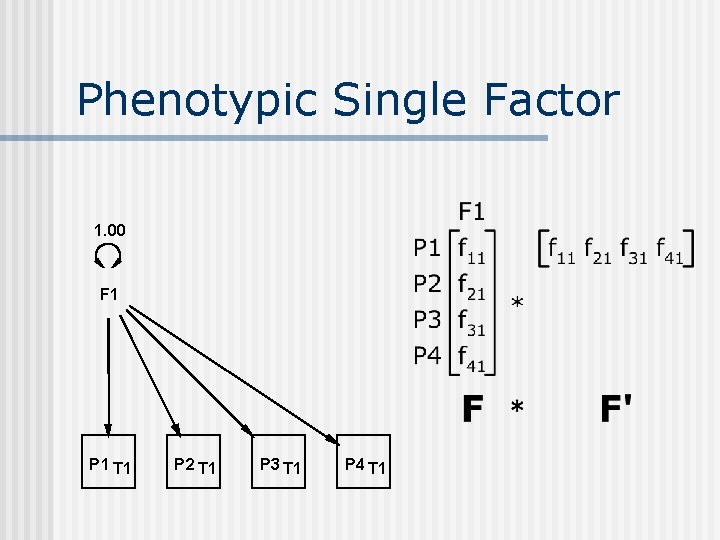
Phenotypic Single Factor 1. 00 F 1 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1
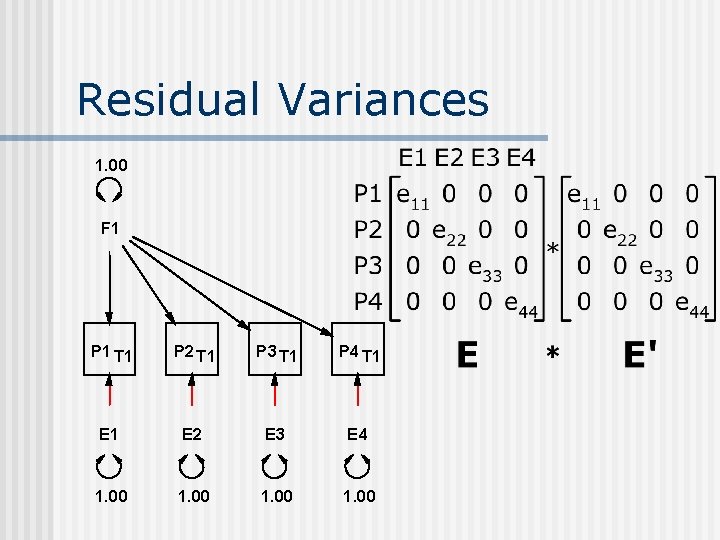
Residual Variances 1. 00 F 1 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1 E 2 E 3 E 4 1. 00
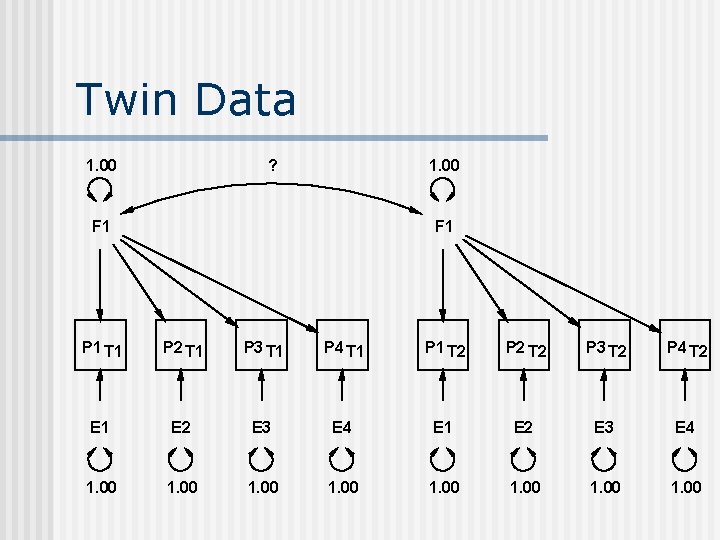
Twin Data 1. 00 ? 1. 00 F 1 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1 P 1 T 2 P 2 T 2 P 3 T 2 P 4 T 2 E 1 E 2 E 3 E 4 1. 00 1. 00
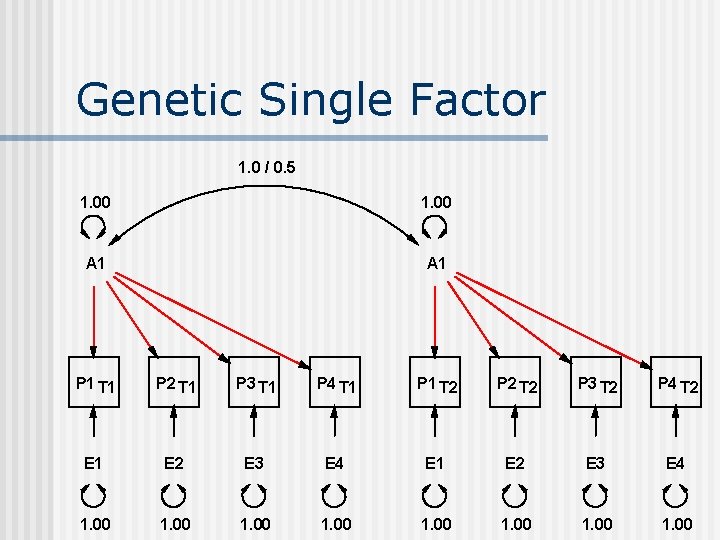
Genetic Single Factor 1. 0 / 0. 5 1. 00 A 1 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1 P 1 T 2 P 2 T 2 P 3 T 2 P 4 T 2 E 1 E 2 E 3 E 4 1. 00 1. 00
![Single [Common] Factor n X: genetic n n Full 4 x 1 Full nvar Single [Common] Factor n X: genetic n n Full 4 x 1 Full nvar](http://slidetodoc.com/presentation_image_h/28ebcd705c33225200a510a0ae30df72/image-14.jpg)
Single [Common] Factor n X: genetic n n Full 4 x 1 Full nvar x nfac Y: shared environmental Z: specific environmental
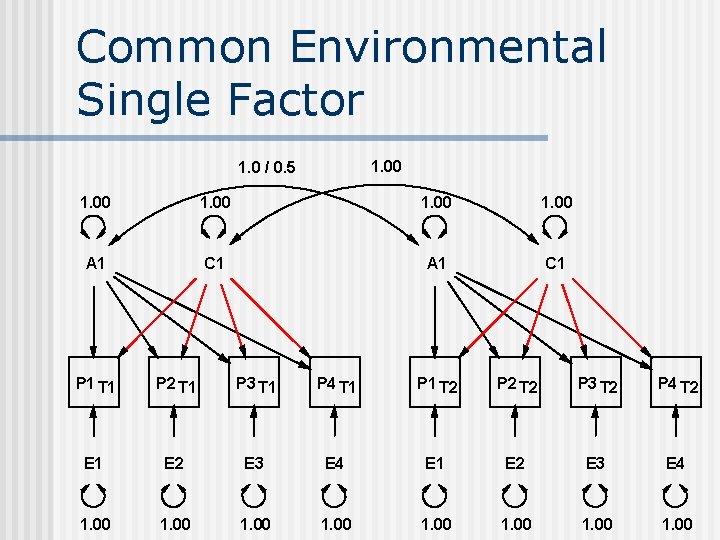
Common Environmental Single Factor 1. 00 1. 0 / 0. 5 1. 00 A 1 C 1 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1 P 1 T 2 P 2 T 2 P 3 T 2 P 4 T 2 E 1 E 2 E 3 E 4 1. 00 1. 00
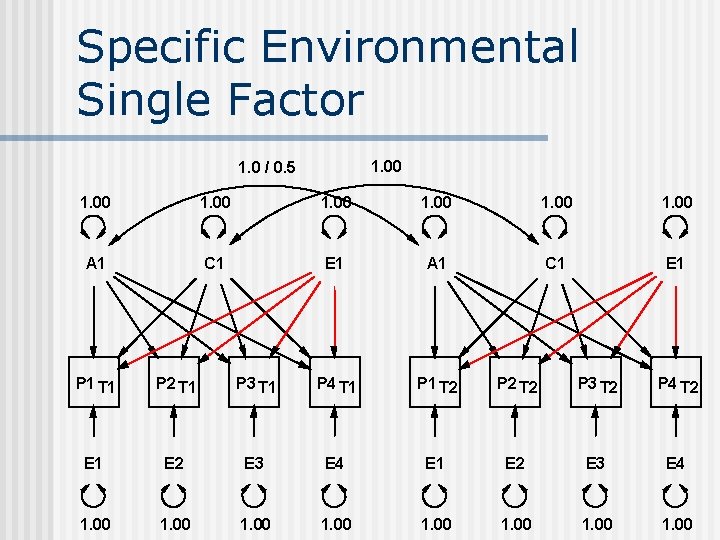
Specific Environmental Single Factor 1. 00 1. 0 / 0. 5 1. 00 A 1 C 1 E 1 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1 P 1 T 2 P 2 T 2 P 3 T 2 P 4 T 2 E 1 E 2 E 3 E 4 1. 00 1. 00
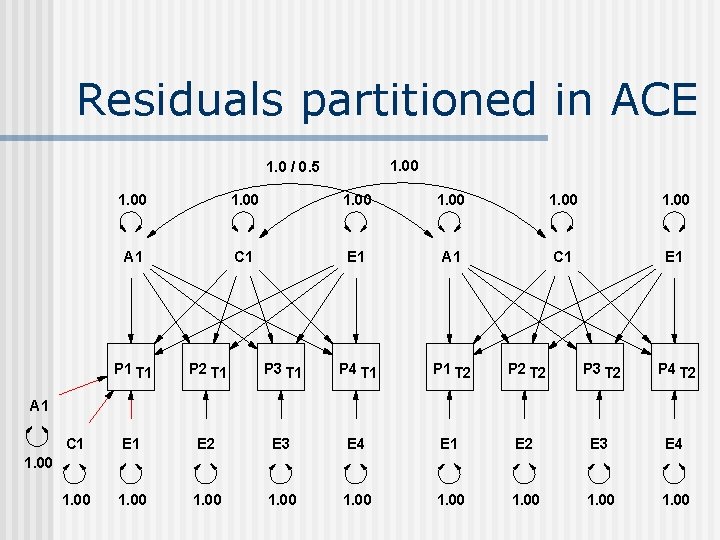
Residuals partitioned in ACE 1. 00 1. 0 / 0. 5 1. 00 A 1 C 1 E 1 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1 P 1 T 2 P 2 T 2 P 3 T 2 P 4 T 2 C 1 E 2 E 3 E 4 E 1 E 2 E 3 E 4 1. 00 1. 00 A 1 1. 00

Residual Factors n n n T: genetic U: shared environmental V: specific environmental n n Diag 4 x 4 Diag nvar x nvar
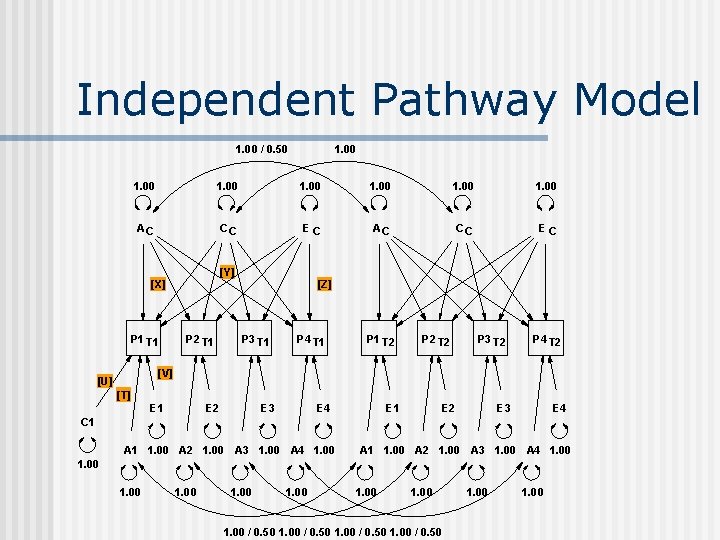
Independent Pathway Model 1. 00 / 0. 50 1. 00 1. 00 AC CC EC [Y] [X] P 1 T 1 P 2 T 1 [Z] P 3 T 1 P 4 T 1 P 1 T 2 P 2 T 2 P 3 T 2 P 4 T 2 [V] [U] [T] E 1 E 2 E 3 E 4 C 1 A 1 1. 00 A 2 1. 00 A 3 1. 00 A 4 1. 00 1. 00 / 0. 50 1. 00

Independent Pathway Example n n Script: f: hmaesa 17ind 3 f. mx 3 MRI measures – 2 IQ subtests: n n n var 1 var 2 var 3 var 4 var 5 = = = cerebellum grey matter white matter calculation letters and numbers Data: f: hmaesa 17mri-iqfactor. rec n n T: hmaes/a 17mri-iq-mz. dat T: hmaesa 17mri-iq-dz. dat

Independent Pathway #define nvar 5 #define nfac 1 G 1: Define matrices Calculation Begin Matrices; X full nvar 3 Y full nvar nfac Z full nvar nfac Free T diag nvar Free U diag nvar V diag nvar Free M full 1 nvar Free End Matrices; ! common factor genetic paths ! common factor shared env paths ! common factor specific env paths ! variable specific genetic paths ! variable specific shared env paths ! variable specific residual paths ! grand means

Three Genetic Factors Specify X ! declare free parameters for 3 genetic common factors ! G 1 G 2 G 3 100 110 0 101 111 0 102 112 0 103 0 120 104 0 121 Drop T 1 5 5 ! fix genetic specific for last variable for identification
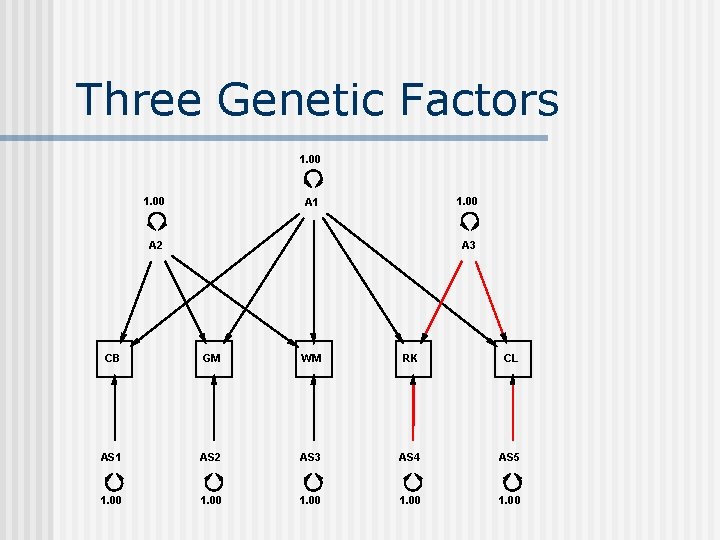
Three Genetic Factors 1. 00 A 1 A 2 A 3 CB GM WM RK CL AS 1 AS 2 AS 3 AS 4 AS 5 1. 00

Independent II Begin Algebra; A= X*X' + T*T'; C= Y*Y' + U*U'; E= Z*Z' + V*V'; End Algebra; ! additive genetic covariance ! shared environment covariance ! specific environment covariance Begin Algebra; R=A+C+E; ! total variance S=(sqrt(I. R))~; ! diagonal matrix of standard deviations P=S*X| S*Y| S*Z| d 2 v(S*T)'| d 2 v(S*U)'| d 2 v(S*V)'; ! standardized estimates for common and specific factors End Algebra; d 2 v : diagonal to vector

Path Diagram to Matrices Variance Component a 2 c 2 e 2 Common Factors [X]full 5 x 1 [Y]full 5 x 1 [Z]full 5 x 1 Residual Factors [T]diag 5 x 5 [U]diag 5 x 5 [V]diag 5 x 5 #define nvar 5 #define nfac 1

Path Diagram to Matrices Variance Component a 2 c 2 e 2 Common Factors [X]full 5 x 3 [Y] [Z]full 5 x 1 Residual Factors [T]diag 5 x 5 [U] [V]diag 5 x 5 #define nvar 5 #define nfac 1
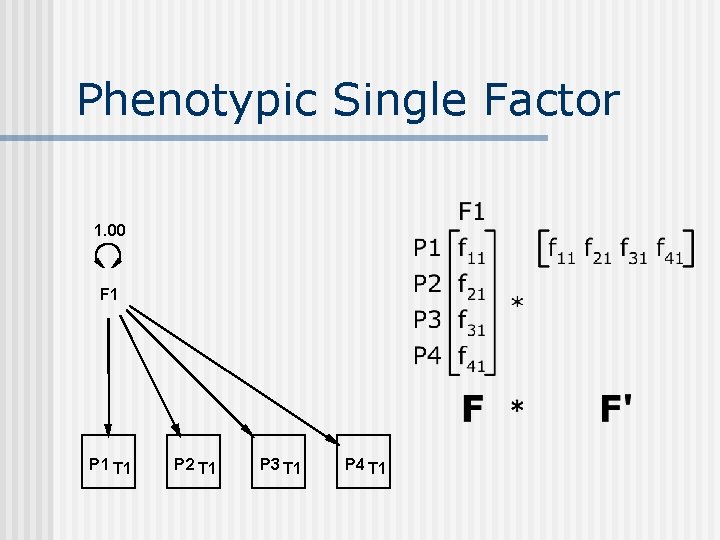
Phenotypic Single Factor 1. 00 F 1 P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1
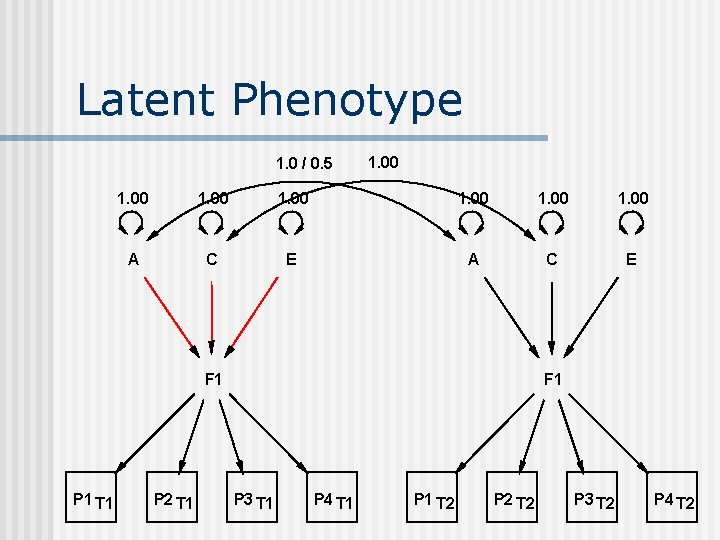
Latent Phenotype 1. 0 / 0. 5 1. 00 1. 00 A C E F 1 P 1 T 1 P 2 T 1 F 1 P 3 T 1 P 4 T 1 P 1 T 2 P 2 T 2 P 3 T 2 P 4 T 2
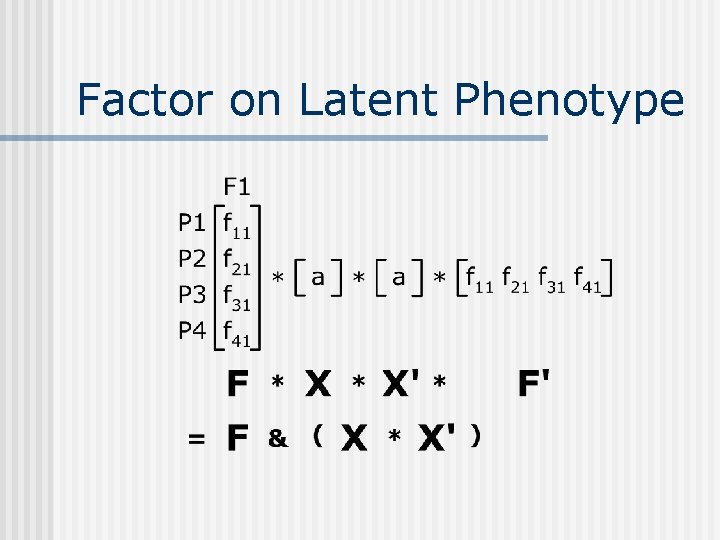
Factor on Latent Phenotype
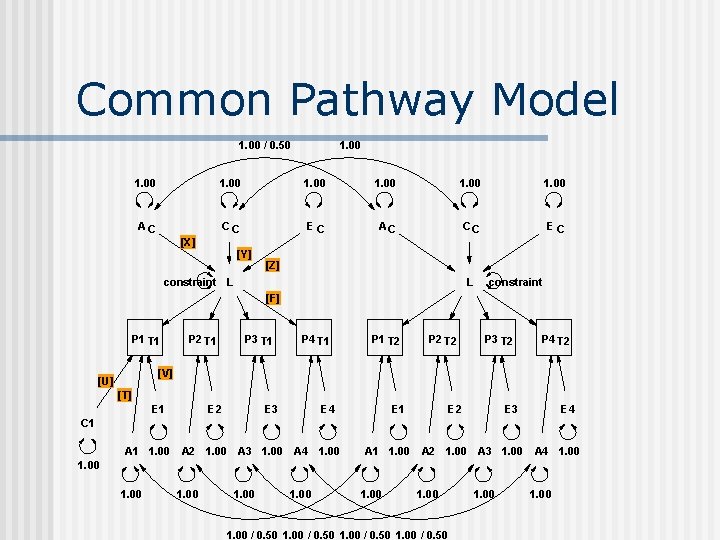
Common Pathway Model 1. 00 / 0. 50 1. 00 1. 00 AC CC EC [X] [Y] [Z] constraint L L constraint [F] P 1 T 1 P 2 T 1 P 3 T 1 P 4 T 1 P 1 T 2 P 2 T 2 P 3 T 2 P 4 T 2 [V] [U] [T] E 1 E 2 E 3 E 4 C 1 A 1 1. 00 A 2 1. 00 A 3 1. 00 A 4 1. 00 1. 00 / 0. 50 1. 00

Common Pathway Example n n Script: f: hmaesa 17comp. mx 3 MRI measures – 2 IQ subtests: n n n var 1 var 2 var 3 var 4 var 5 = = = cerebellum grey matter white matter calculation letters and numbers Data: f: hmaesa 17mri-iqfactor. rec n n T: hmaes/a 17mri-iq-mz. dat T: hmaesa 17mri-iq-dz. dat

Compath Pathway #define nvar 5 #define nfac 1 Begin Matrices; X full nfac Free Y full nfac Z full nfac Free T diag nvar Free U diag nvar V diag nvar Free F full nvar nfac Free M full 1 nvar Free End Matrices; ! latent factor genetic paths ! latent factor shared env paths ! latent factor specific env paths ! variable specific genetic paths ! variable specific shared env paths ! variable specific residual paths ! loadings on latent factor ! grand means

Compath II Begin Algebra; A= F&(X*X') + T*T'; ! genetic variance components C= F&(Y*Y') + U*U'; ! shared environment covariance E= F&(Z*Z') + V*V'; ! specific environment covariance L= X*X' + Y*Y' + Z*Z'; ! variance of latent factor End Algebra; Begin Algebra; R=A+C+E; ! total variance S=(sqrt(D. R))~; ! diagonal matrix of standard deviations P=S*F| d 2 v(S*T)'| d 2 v(S*U)'| d 2 v(S*V)'; ! standardized estimates for loadings and specific factors W= X*X' | Y*Y'| Z*Z'; ! standardized estimates for latent phenotype End Algebra;

Path Diagram to Matrices Variance a 2 Component c 2 e 2 Common Factor [X]full 1 x 1 [Y]full 1 x 1 [Z]full 1 x 1 Residual Factors [T]diag [U]diag [V]diag 5 x 5 #define nvar 5 #define nfac 1 [F]full 5 x 1

Path Diagram to Matrices Variance a 2 Component c 2 e 2 Common Factor [X]full 1 x 1 [Y] [Z]full 1 x 1 Residual Factors [T]diag [U] 5 x 5 #define nvar 5 #define nfac 1 [V]diag 5 x 5 [F]full 5 x 1

Summary n Independent Pathway Model Biometric Factors Model n Loadings differ for genetic and environmental common factors n n Common Pathway Model Psychometric Factors Model n Loadings equal for genetic and environmental common factor n
- Slides: 36