Multiple Sequence Alignment Definition Homology related by descent

Multiple Sequence Alignment

Definition • Homology: related by descent • Homologous sequence positions ATTGCGC ATTGCGC AT-ACGC ATACGC

Reasons for aligning sets of sequences • • Organise data to reflect sequence homology Estimate evolutionary distance Infer phylogenetic trees from homologous sites Highlight conserved sites/regions Highlight variable sites/regions Uncover changes in gene structure Look for evidence of selection Summarise information

Alignments help to Organise Visualise Analyze Sequence Data

The process of aligning sequences is a game involving playing off gaps and mismatches

Ways of aligning multiple sequences • By hand • Automated • Combination

Definition Optimality criteria: some kind rule or scoring scheme to help you to decide what you consider to be the best alignment

Pairwise vs Multiple Sequences • Pairs of sequences typically aligned using exhaustive algorithms (dynamic programming) – complexity of exhaustive methods is O(2 n mn) n = number of sequences m = sequence length • Multiple sequence alignment usually performed using heuristic methods

The Correct Alignment ATTGCGC ATTGCGC AT-ACGC ATACGC ATTGCGC ATA-CGC
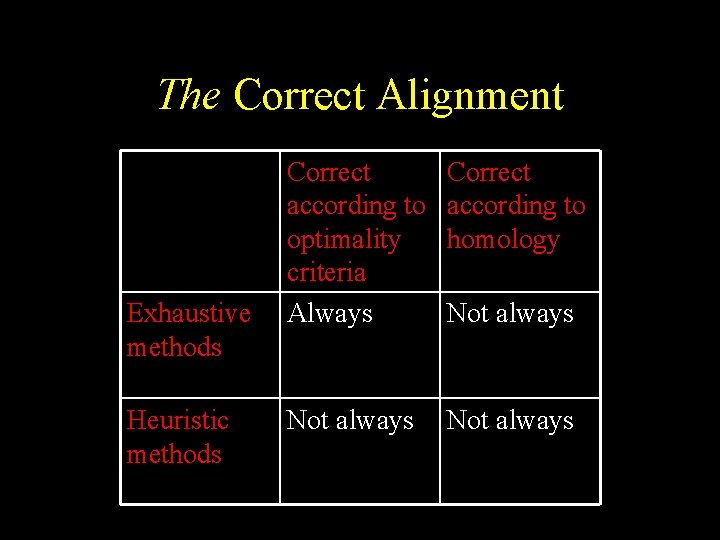
The Correct Alignment Exhaustive methods Heuristic methods Correct according to optimality criteria Always Correct according to homology Not always

• Sequence alignment is easy with sufficiently closely related sequences • Below a certain level of identity sequence alignment may become meaningless – twilight zone for aa sequences ~ 30% • In the twilight zone it is good to make use of additional information if possible (e. g. structure)

Consensus Sequences • Simplest Form: A single sequence which represents the most common amino acid/base in that position Y Y F F Y Y D D E D D G G G G G A I I A A/I V L L V V/L V V E E E Q Q E A A A L L L V L L

Multiple Alignment Formats e. g. Clustal, Phylip, MSF, MEGA etc.

Clustal Format CLUSTAL X (1. 81) multiple sequence alignment CAS 1_BOVIN CAS 1_SHEEP CAS 1_PIG CAS 1_HUMAN CAS 1_RABBIT CAS 1_MOUSE CAS 1_RAT MKLLILTCLVAVALARPKHPIKHQGLPQ----EVLNENMKLLILTCLVAVALARPKHPIKHQGLSP----EVLNENMKLLIFICLAAVALARPKPPLRHQEHLQNEPDSRE-------MRLLILTCLVAVALARPKLPLRYPERLQNPSESSE-------MKLLILTCLVATALARHKFHLGHLKLTQEQPESSEQEILKERK MKLLILTCLVAAAFAMPRLHSRNAVSSQTQ------QQHSSSE MKLLILTCLVAAALALPRAHRRNAVSSQTQ------*: **. *. *: * : . :

Phylip Format (Interleaved) 7 100 SOMA_BOVIN SOMA_SHEEP SOMA_RAT_P SOMA_MOUSE SOMA_RABIT SOMA_PIG_P SOMA_HUMAN MMAAGPRTSL -MAADSQTPW -MATDSRTSW -MAAGSWTAG -MAAGPRTSA -MATGSRTSL LLAFALLCLP LLAFTLLCLP LLTFSLLCLL LLTVSLLCLL LLAFALLCLP LLAFGLLCLP WTQVVGAFPA WPQEAGAFPA WPQEASAFPA WTREVGAFPA WLQEGSAFPT MSLSGLFANA MPLSSLFANA MPLSSLFSNA MPLSSLFANA IPLSRLFDNA VLRAQHLHQL VLRAQHLHQL MLRAHRLHQL AADTFKEFER AADTYKEFER AFDTYQEFEE TYIPEGQRYS AYIPEGQRYS AYIPKEQKYS -IQNTQVAFC -IQNAQAAFC FLQNPQTSLC FSETIPAPTG FSETIPAPTG FSESIPTPSN KNEAQQKSDL KEEAQQRTDM KDEAQQRSDM KDEAQQRSDV REETQQKSNL

Phylip Format (Sequential) 3 100 Rat ATGGTGCACCTGATGCTGAGAAGGCTGCTGTTAATGGCCG TGGTGGCTGGAGTGGCCAGTGCCCTGGCTCACAAGTACCACTAA Mouse ATGGTGCACCTGATGCTGAGAAGGCTGCTGTCTCTTGCCT TGGGGAAAGGTGAACTCCGATGAAGTTGGTGGTGAGGCCCTGGG Rabbit ATGGTGCATCTGTCCAGT---GAGGAGAAGTCTGCGGTCACTGC TGGGGCAAGGTGAATGTGGAAGAAGTTGGTGGTGAGGCCCTGGG

Mega Format #mega TITLE: No title #Rat #Mouse #Rabbit #Human #Oppossum #Chicken #Frog ATGGTGCACCTGACTGATGCTGAGAAGGCTGCTGT ATGGTGCATCTGTCCAGT---GAGGAGAAGTCTGC ATGGTGCACCTGACTCCT---GAGGAGAAGTCTGC ATGGTGCACTTGACTTTT---GAGGAGAAGAACTG ATGGTGCACTGGACTGCT---GAGGAGAAGCAGCT ---ATGGGTTTGACAGCACATGATCGT---CAGCT

Progressive Multiple Alignment • Heuristic • Perform pairwise alignments • Align sequences to alignments or alignments to existing alignments (profile alignments • Do the alignments in some sensible order

Progressive versus Simultaneous • speed versus accuracy • simultaneous methods are capable of working out an ‘exact’ solution to the problem of multiple sequence alignment (e. g. NCBI’s MSA – user interface QAlign)

Iterative methods • Several progressive alignment methods can be iterated – e. g. Barton-Sternberg, Clustal. X

Clustal. X Algorithm • Perform pairwise alignments and calculate distances for all pairs of sequences • Construct guide tree (dendrogram) joining the most similar sequences using Neighbour Joining • Align sequences, starting at the leaves of the guide tree. This involves the pair-wise comparisons as well as comparison of single sequence with a group of seqs (Profile)

• Clustal. X is not optimal • There are known areas in which Clustal. X performs badly e. g. – errors introduced early cannot be corrected by subsequent information – alignments of sequences of differing lengths cause strange guide trees and unpredictable effects – edges: Clustal. X does not penalise gaps at edges • There alternatives to Clustal. X available

T-Coffee • JMB 2000 • Also a progressive alignment method • Designed to solve some of the problems with clustal (in particular the problem of clustals inability to correct errors that appear early in the process of alignment) • Can consider global and local pair-wise alignments

Using Clustal. X • Start with sequences in FASTA format (or an existing alignment in Clustal format • [Do Alignment] on the alignment menu


Clustal. X Parameters • • • Scoring Matrix Gap opening penalty Gap extension penalty Protein gap parameters Additional algorithm parameters Secondary structure penalties

Score Matrices • Pairwise matrices and multiple alignment matrix series • PAM (Dayhoff), BLOSUM (Hennikof), GONNET (default), user defined • Transition (A<->G)/Transversion (C<-T) ratio – low for distantly related sequences

Gap Penalties • Linear gap penalties – Affine gap penalties p = (o + l. e) • Gap opening • Gap extension • Protein specific penalties (on by default) – Increase the probability of gaps associated with certain residues – Increase the chances of gaps in loop regions (> 5 hydrophilic residues)

Algorithm parameters • • Slow-accurate pair-wise alignment Do alignment from guide tree Reset gaps before aligning (iteration) Delay Divergent sequences (%)

Additional displays • Column Scores • Low quality regions • Exceptional residues

Multiple Alignment Tips • Align pairs of sequences using an optimal method • Progressive alignment programs such as Clustal. X for multiple alignment • Choose representative sequences to align carefully • Choose sequences of comparable lengths • Progressive alignment programs may be combined • Review alignment by eye and edit • If you have a choice align amino acid sequences rather than nucleotides

Alignment of coding regions • Nucleotide sequences much harder to align accurately than proteins • Protein coding sequences can be aligned using the protein sequences – e. g. Bio. Edit: toggle translation to amino acid, call clustalw to align, edit alignment by hand, toggle back to nucleotide • In-frame nucleotide alignments can be used, e. g. to determine non-synonymous and synonymous distances separately

Multiple Alignments and Phylogenetic Trees – You can make a more accurate multiple sequence alignment if you know the tree already – A phylogenetic tree is only as good as the alignment from which it was produced – The process of constructing a multiple alignment (unlike pair-wise) needs to take account of phylogenetic relationships

Editing a multiple sequence alignment • It is NOT fraud to edit a multiple sequence alignment • Incorporate additional knowledge if possible • Alignment editors help to keep the data organised and help to prevent unwanted mistakes

Alignment Editors • e. g. GDE, Bioedit, Seaview, Jalview etc. • Some alignment editors have begun to function as sequence analysis platforms (e. g. tools on Bio. Edit, GDE) • Construct sub-sequences (GDE, Seaview) • Annotate sequences (Seaview)

Aligning weakly similar sequences

Sequence contains conserved regions • e. g. DIALIGN (Morgenstern, Dress, Werner) – re-aligns regions between conserved blocks http: //bibiserv. techfak. uni-bielefeld. de/ useful if sequences contains consistent conserved blocks • Block Maker – searches for conserved words that may be inconsistent http: //blocks. fhcrc. org/

Profile Alignment Gribskov et al. 1987 • Position specific scores • Allows addition of extra sequence(s) to an alignment • Allows alignment of alignments • Gaps introduced as whole columns in the separate alignments • Optimal alignment in time O(a 2 l 2) a = alphabet size, l = sequence length • Information about the degree of conservation of sequence positions is included

Good reasons to use profile alignments – Adding a new sequence to an existing multiple alignment that you want to keep fixed (align sequence to profile) – Searching a database for new members of your protein family (pfsearch) – Searching a database of profiles to find out which one your sequence belongs to (pfscan) – Combining two multiple sequence alignments (profile to profile)

Profile Alignment Using Clustal. X • • Profile Alignment Mode Align sequence to profile Align profile 1 to profile 2 Secondary structure parameters

Profile searching using PSIBLAST • Position Specific Iterative • Perform search – construct profile – perform search • Convergence (hopefully…) • Increased sensitivity for distantly related sequences • Available on-line (NCBI)

Databases of Aligned Sequences • Hovergen http: //pbil. univlyon 1. fr/databases/hovergen. html (vertebrate alignments) • Pfam http: //www. sanger. ac. uk/Software/Pfam/ (protein domain alignments and profile HMMs) • BLOCKS http: //blocks. fhcrc. org/ • Ribosomal Database Project http: //rdp. cme. msu. edu/html/ alignments and trees derived from r. RNA sequences • Interpro – combines information from other sources • Many more…

Probabilistic Models of Sequence Alignment • Hidden Markov Models – sequence of states and associated symbol probabilities • Produces a probabilistic model of a sequence alignment • Align a sequence to a Profile Hidden Markov Model – Algorithms exist to find the most efficient pathway through the model

Markov Chain: A chain of things. The probability of the next thing depends only on the current thing Hidden Markov Model: A sequence of states which form a Markov Chain. The states are not observable. The observable characters have “emission” probabilities which depend on the current state.

Some more recent developments • The need to align genomes – alignment tools required that can align very large regions of genomes – poses a computational challenge – programmes such as dialign can be run in parallel on multiprocessor machines

Some more recent developments • MUSCLE – Faster (uses a k-mer frequency to calculate first pairwise alignments) – Progressive (repeats the MSA using the more accurate kimura distance between aligned amino acid sequences) – Has a third optimisation stage that involves making profile alignments of sub-trees and accepting the new alignment if it improves the SP score.

• Mu. Si. C - multiple sequence alignment with constraints – web server that allows a user to enter a set of
- Slides: 48