Multiple Sequence Alignment CS 273 a Lecture 910










- Slides: 28

Multiple Sequence Alignment CS 273 a Lecture 9/10, Aut 10, Batzoglou

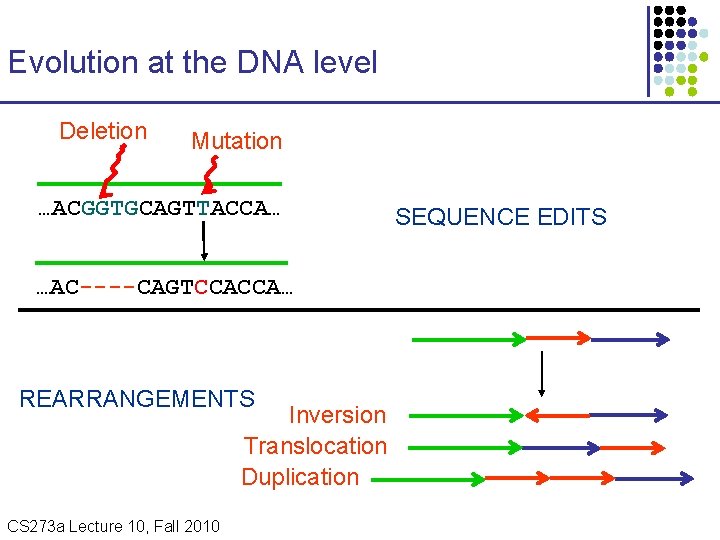
Evolution at the DNA level Deletion Mutation …ACGGTGCAGTTACCA… …AC----CAGTCCACCA… REARRANGEMENTS Inversion Translocation Duplication CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou SEQUENCE EDITS

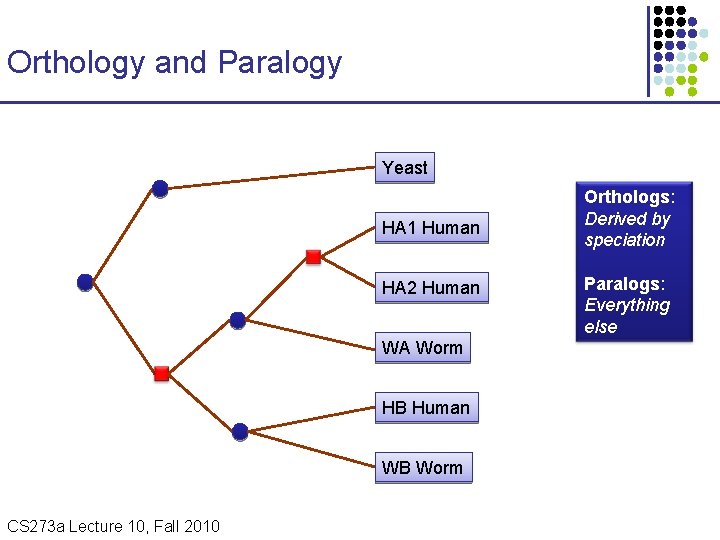
Orthology and Paralogy Yeast HA 1 Human HA 2 Human WA Worm HB Human WB Worm CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou Orthologs: Derived by speciation Paralogs: Everything else

Orthology, Paralogy, Inparalogs, Outparalogs CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

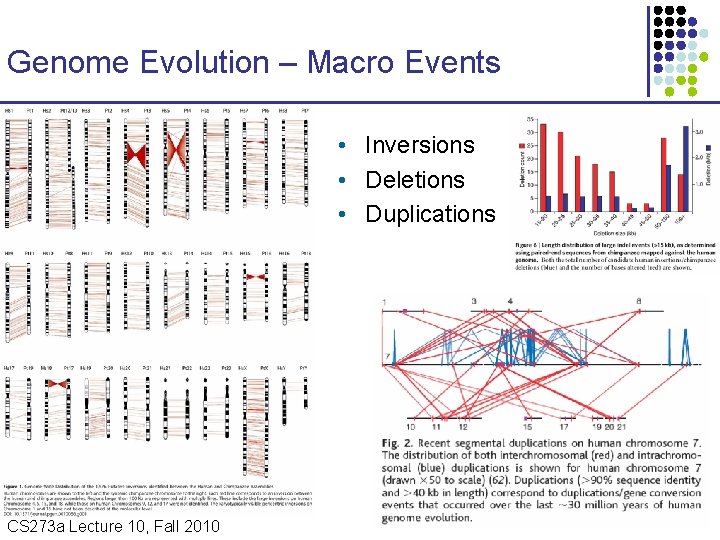
Genome Evolution – Macro Events • Inversions • Deletions • Duplications CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

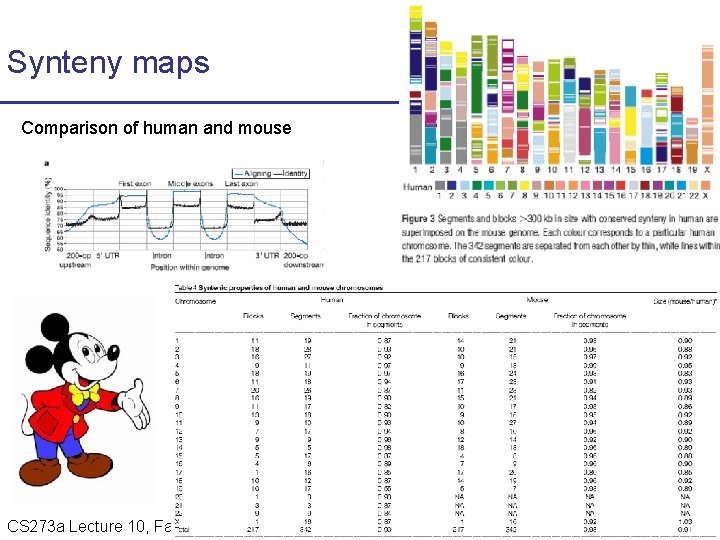
Synteny maps Comparison of human and mouse CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

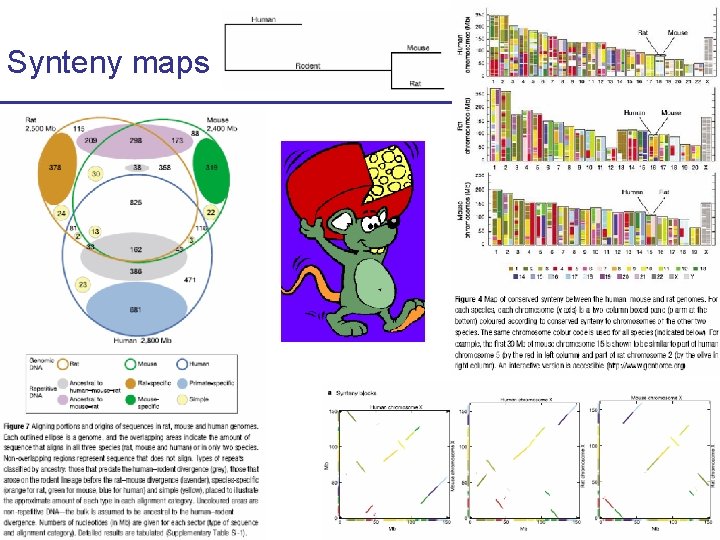
Synteny maps CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

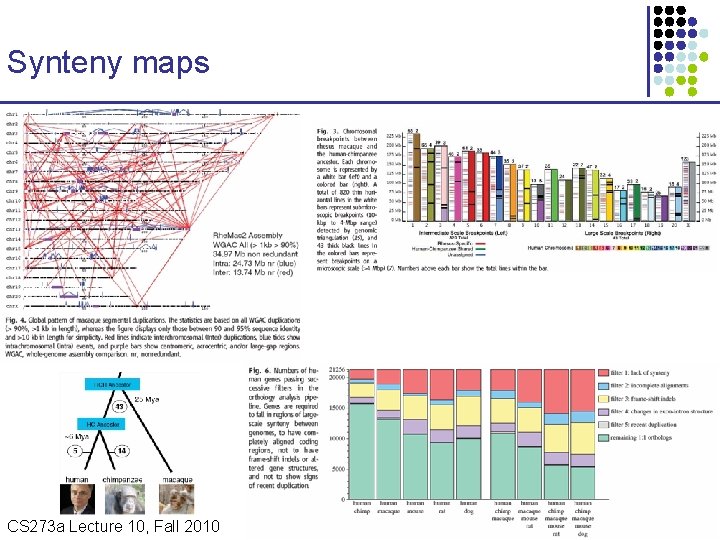
Synteny maps CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

Synteny maps CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

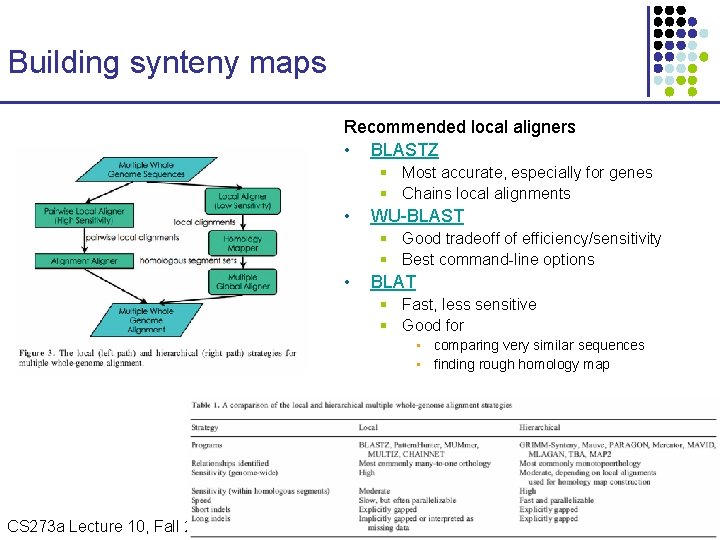
Building synteny maps Recommended local aligners • BLASTZ § Most accurate, especially for genes § Chains local alignments • WU-BLAST § Good tradeoff of efficiency/sensitivity § Best command-line options • BLAT § Fast, less sensitive § Good for • comparing very similar sequences • finding rough homology map CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

Index-based local alignment …… Dictionary: All words of length k (~10) Alignment initiated between words of alignment score T (typically T = k) Alignment: Ungapped extensions until score below statistical threshold Output: All local alignments with score > statistical threshold CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou query …… scan DB query Question: Using an idea from overlap detection, better way to find all local alignments between two genomes?

Local Alignments CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

After chaining CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

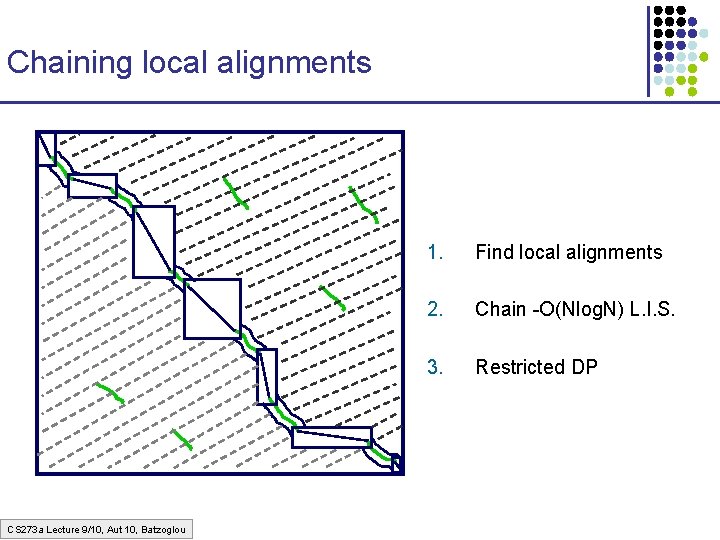
Chaining local alignments CS 273 a Lecture 9/10, Aut 10, Batzoglou 1. Find local alignments 2. Chain -O(Nlog. N) L. I. S. 3. Restricted DP

Progressive Alignment x y Example z Profile: (A, C, G, T, -) px = (0. 8, 0. 2, 0, 0, 0) w py = (0. 6, 0, 0. 4) • When evolutionary tree is known: s(px, py) = 0. 8*0. 6*s(A, A) + 0. 2*0. 6*s(C, A) + 0. 8*0. 4*s(A, -) + 0. 2*0. 4*s(C, -) § Align closest first, in the order of the tree § In each step, align two sequences. Result: x, y, or profiles px, py 0. 1, , to generate a new pxy = (0. 7, 0, 0, 0. 2) alignment with associated profile presult s(p , -) = 0. 8*1. 0*s(A, -) + 0. 2*1. 0*s(C, -) x Weighted version: § Tree edges have weights, proportional to the divergence in that edge Result: p = (0. 4, 0. 1, 0, 0, 0. 5) § New profile is a weighted average of two old x-profiles CS 273 a Lecture 9/10, Aut 10, Batzoglou

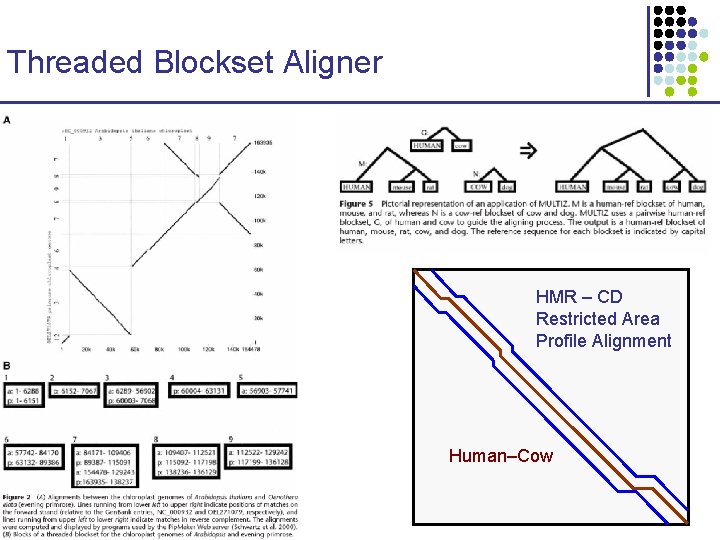
Threaded Blockset Aligner HMR – CD Restricted Area Profile Alignment Human–Cow CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

Reconstructing the Ancestral Mammalian Genome C Human: C Baboon: C G Dog: G C or G Cat: C CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

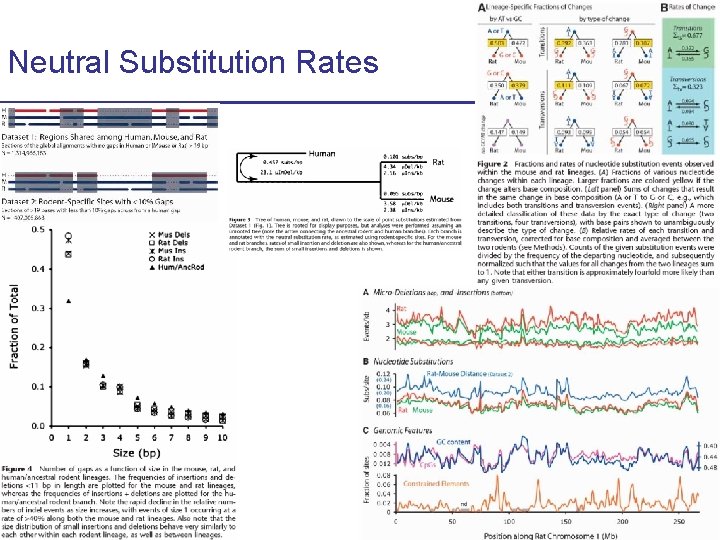
Neutral Substitution Rates CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

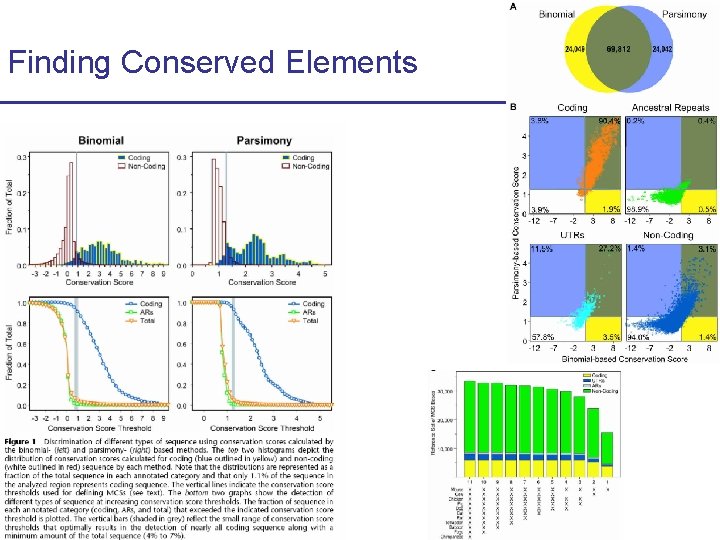
Finding Conserved Elements (1) • Binomial method § 25 -bp window in the human genome § Binomial distribution of k matches in N bases given the neutral probability of substitution CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

Finding Conserved Elements (2) A • Parsimony Method § Count minimum # of mutations explaining each column § Assign a probability to this parsimony score given neutral model § Multiply probabilities across 25 -bp window of human genome CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou C A A G

Finding Conserved Elements CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

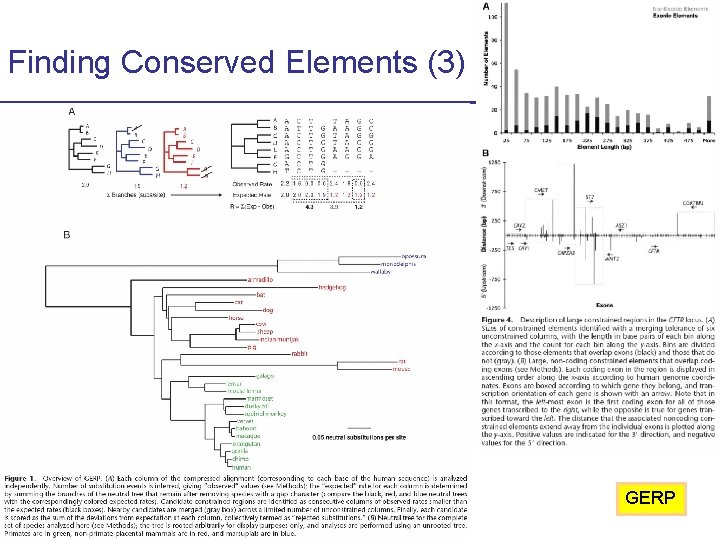
Finding Conserved Elements (3) GERP CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

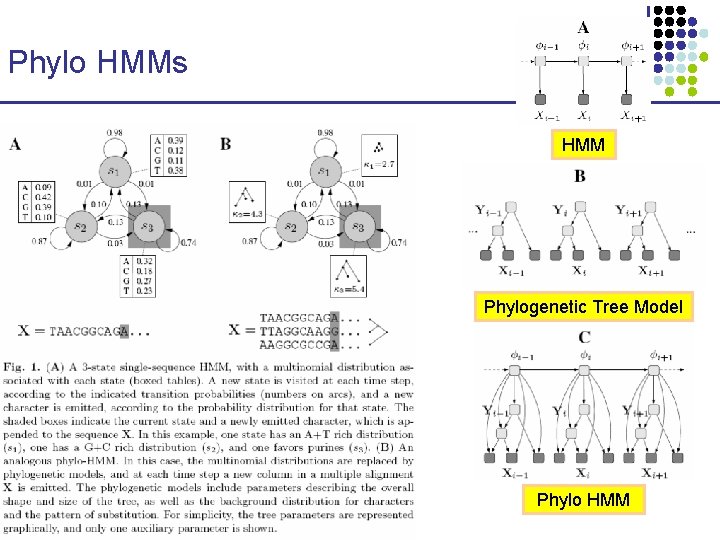
Phylo HMMs HMM Phylogenetic Tree Model Phylo HMM CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

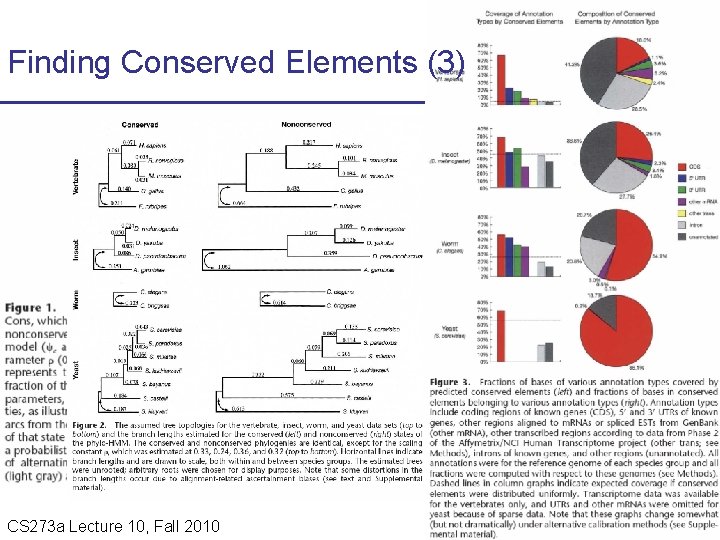
Finding Conserved Elements (3) CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

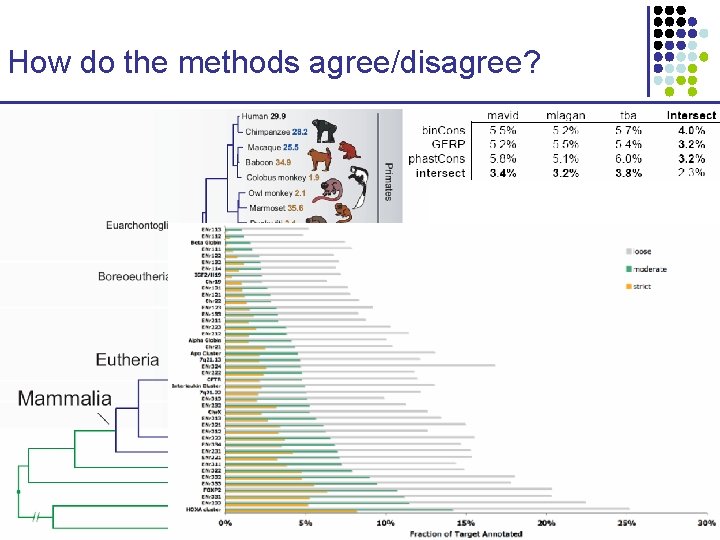
How do the methods agree/disagree? CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

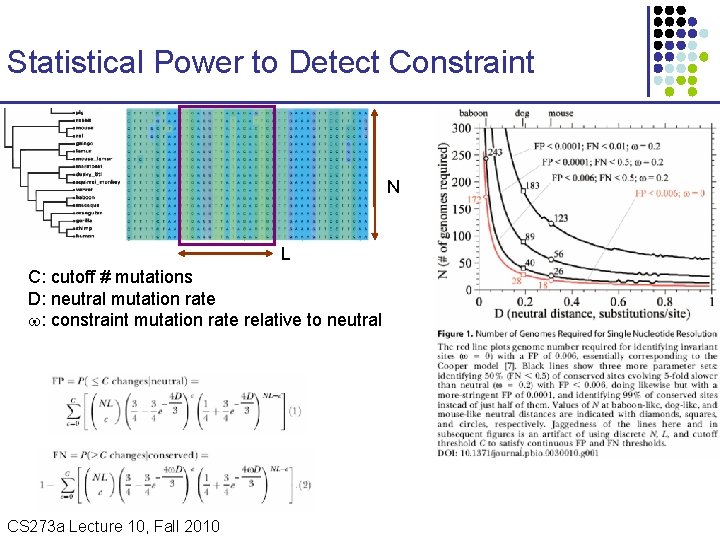
Statistical Power to Detect Constraint N L C: cutoff # mutations D: neutral mutation rate : constraint mutation rate relative to neutral CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou

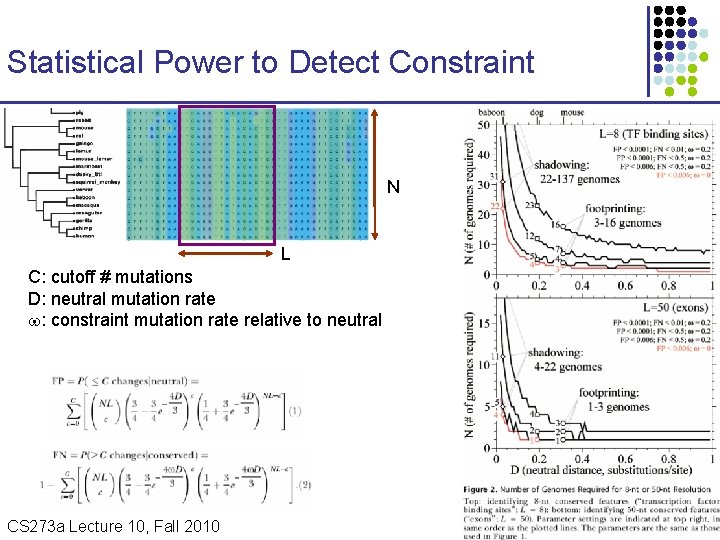
Statistical Power to Detect Constraint N L C: cutoff # mutations D: neutral mutation rate : constraint mutation rate relative to neutral CS 273 a Lecture 10, Fall 2010 CS 273 a Lecture 9/10, Aut 10, Batzoglou