Multiple Regression sec VIIIb Example apnea risk calculator

Multiple Regression (sec VIIIb)

Example: apnea risk calculator How many hours of sleep do you get: a. 5 or less (15 pts), b. 6 -8 (0 pts) , c. 9+ (5 pts) What best describes your sleeping pattern: a. sound sleep all night (0 pts) b. wake up once and go back to sleep (5 pts) c. wake up multiple times & go back to sleep (10 pts) d. wake up and cannot get back to sleep (15 pts) Do you snore: a. No (0 pts) b Yes (10 pts)

Multiple Regression - Overview Multiple Regression in statistics is the science and art of creating an equation that relates an outcome Y, to one or more predictors, X 1, X 2, X 3, . . Xk. Linear Regression Y = a + b 1 X 1 + b 2 X 2 + b 3 X 3 +. . . + bk Xk + e =Ŷ+e where the “b”s are the weights and "e" is the residual error between the observed Y and the prediction (Ŷ). In linear regression, bi is the average change in Y for a single unit change in Xi. “Performance” stats: R 2, SDe
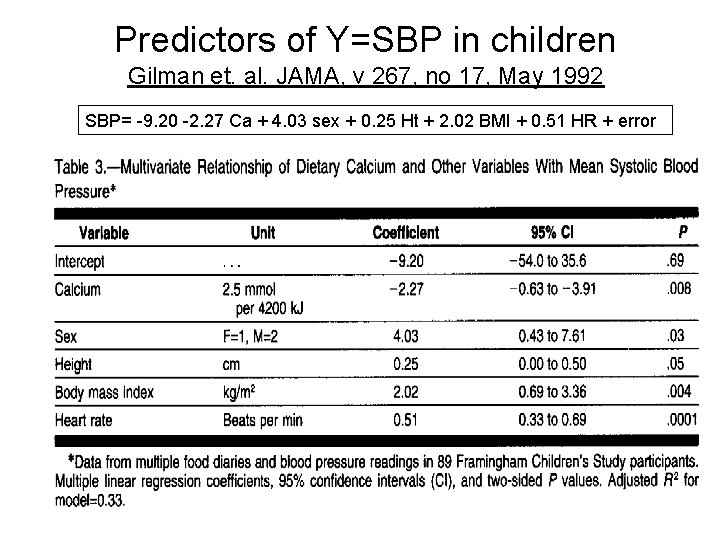
Predictors of Y=SBP in children Gilman et. al. JAMA, v 267, no 17, May 1992 SBP= -9. 20 -2. 27 Ca + 4. 03 sex + 0. 25 Ht + 2. 02 BMI + 0. 51 HR + error

Logistic Regression In multiple logistic regression, Y is 0 or 1 with mean P, the “risk”. The logit of P, not P, is assumed a linear function of the Xs Logit(P) = ln(P/(1 -P)) = a + b 1 X 1 + b 2 X 2 + b 3 X 3 +. . . + bk Xk (“Logit” = log of the odds since P/(1 -P) is the odds) odds=exp(a + b 1 X 1 + b 2 X 2 + b 3 X 3 +. . . + bk Xk) risk = P = odds/(odds+1) “Performance” stats: Sensitivity, Specificity, Accuracy, Concordance (C), mean deviance Poisson Regression When Y is a positive integer (0, 1, 2, 3…), we model the log of Y so Y can never be negative. This is the multiple Poisson regression model. ln(mean Y) = a + b 1 X 1 + b 2 X 2 + b 3 X 3 +. . . + bk Xk mean Y = exp(a + b 1 X 1 + b 2 X 2 + b 3 X 3 +. . . + bk Xk) mean Y cannot be negative. “Performance” stats: R 2, mean deviance
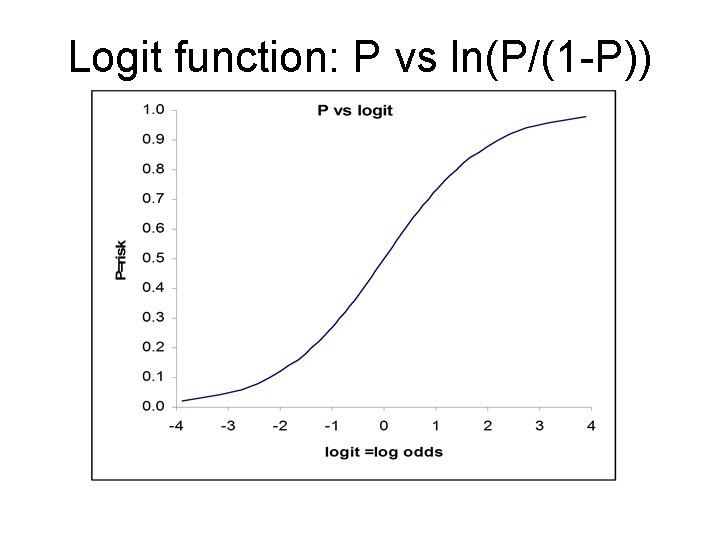
Logit function: P vs ln(P/(1 -P))
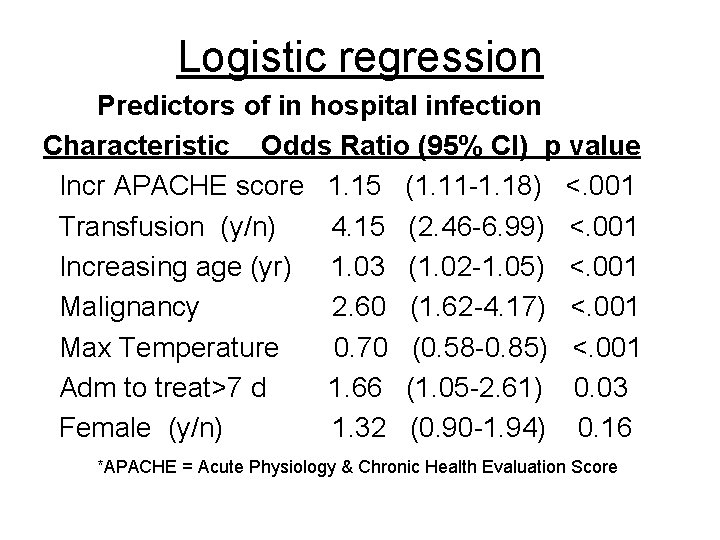
Logistic regression Predictors of in hospital infection Characteristic Odds Ratio (95% CI) p value Incr APACHE score 1. 15 (1. 11 -1. 18) <. 001 Transfusion (y/n) 4. 15 (2. 46 -6. 99) <. 001 Increasing age (yr) 1. 03 (1. 02 -1. 05) <. 001 Malignancy 2. 60 (1. 62 -4. 17) <. 001 Max Temperature 0. 70 (0. 58 -0. 85) <. 001 Adm to treat>7 d 1. 66 (1. 05 -2. 61) 0. 03 Female (y/n) 1. 32 (0. 90 -1. 94) 0. 16 *APACHE = Acute Physiology & Chronic Health Evaluation Score

Multiple Proportional Hazards Regr (Cox model) For time dependent outcomes (ie time to death), we model the hazard rate, h , the event rate per unit time (for death, it is the mortality rate). Since h > 0, we model the log of the hazard as a linear function of the Xs so h can never be zero (similar to Poisson regression). ln(h) so h = a + b 1 X 1 + b 2 X 2 + b 3 X 3 +. . . + bk Xk = exp(a + b 1 X 1 + b 2 X 2 + b 3 X 3 +. . . + bk Xk) >0 If h 0=exp(a) is the ‘baseline’ hazard, (that is, a=log(h 0)) the hazard ratio is HR = h/h 0 = exp(b 1 X 1 + b 2 X 2 + b 3 X 3 +. . . + bk Xk) no intercept ‘a’. If S 0(t) is the ‘baseline’ survival curve corresponding to the baseline hazard, then the survival curve for a given combination of X 1, X 2, … Xk is given by S(t) = S 0(t)HR where HR is computed with the equation above. exp(bi) is the hazard rate ratio for a one unit change in Xi. “Performance” stats: Harrell’s Concordance (C) (0. 5 < C < 1. 0)
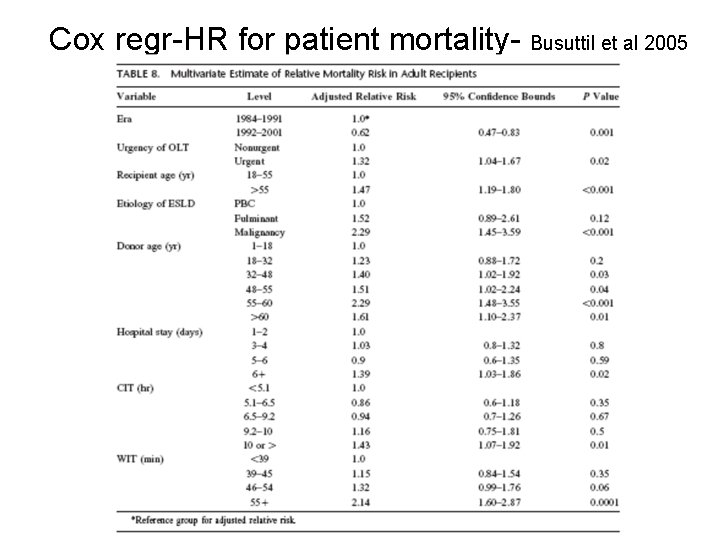
Cox regr-HR for patient mortality- Busuttil et al 2005
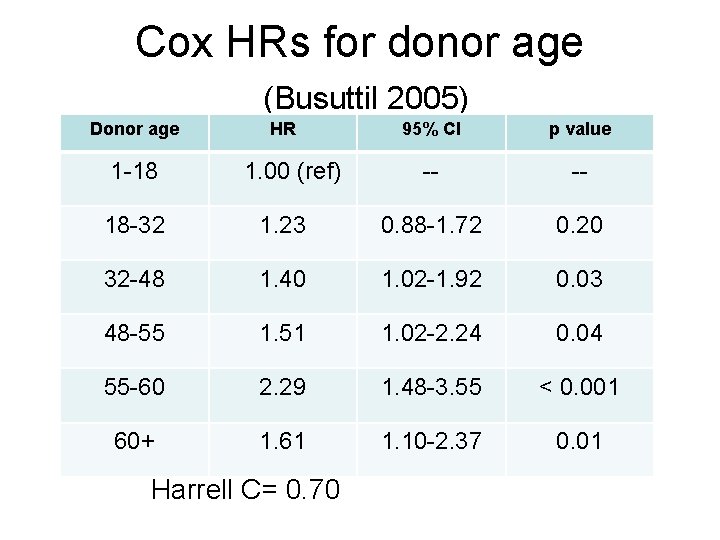
Cox HRs for donor age (Busuttil 2005) Donor age 1 -18 HR 1. 00 (ref) 95% CI p value -- -- 18 -32 1. 23 0. 88 -1. 72 0. 20 32 -48 1. 40 1. 02 -1. 92 0. 03 48 -55 1. 51 1. 02 -2. 24 0. 04 55 -60 2. 29 1. 48 -3. 55 < 0. 001 60+ 1. 61 1. 10 -2. 37 0. 01 Harrell C= 0. 70

Regression coeff interpretation Outcome (Y) Regression interpretation continuous Linear b is the average change in Y per one unit increase in X, the rate of change Binary Logistic exp(b)=eb=odds ratio (OR) for a one unit increase in X Low Positive integers (0, 1, 2, 3. . ) Poisson exp(b)= mean ratio (MR) for a one unit increase in X Hazard rate Cox exp(b)=hazard rate ratio (HR) for a one unit increase in X (P=proportion) (h=events/time) S(t) = S 0(t)HR

Multiple Linear Regression Example Consider predictors of Y=Bilirubin (mg/dl) in liver transplant candidates. Two predictors are X 1=Prothombin time (PT) in seconds X 2=ALT (alanine aminotransferase in U/L). A multiple regression equation (on the log scale) is Ŷ = (predicted) log Bilirubin = -3. 96 + 3. 47 log PT + 0. 21 log ALT
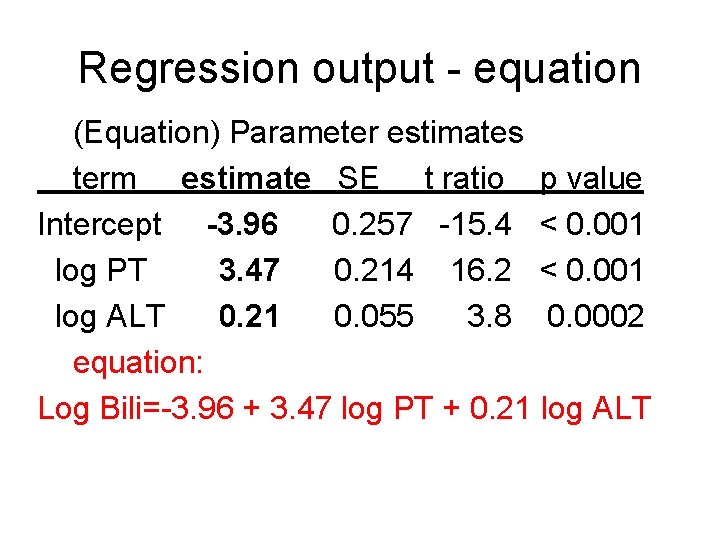
Regression output - equation (Equation) Parameter estimates term estimate SE t ratio p value Intercept -3. 96 0. 257 -15. 4 < 0. 001 log PT 3. 47 0. 214 16. 2 < 0. 001 log ALT 0. 21 0. 055 3. 8 0. 0002 equation: Log Bili=-3. 96 + 3. 47 log PT + 0. 21 log ALT
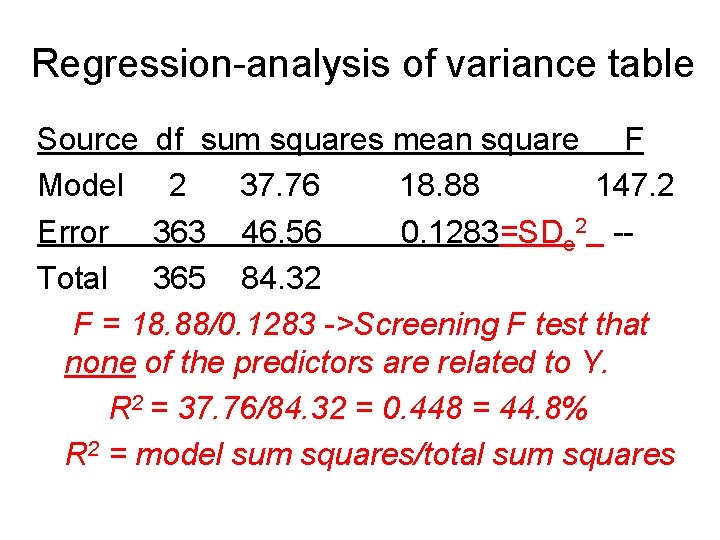
Regression-analysis of variance table Source df sum squares mean square F Model 2 37. 76 18. 88 147. 2 Error 363 46. 56 0. 1283=SDe 2 -Total 365 84. 32 F = 18. 88/0. 1283 ->Screening F test that none of the predictors are related to Y. R 2 = 37. 76/84. 32 = 0. 448 = 44. 8% R 2 = model sum squares/total sum squares
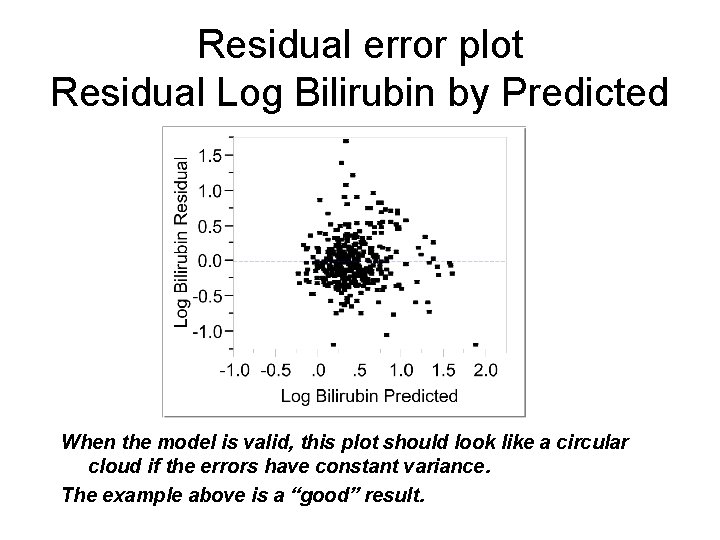
Residual error plot Residual Log Bilirubin by Predicted When the model is valid, this plot should look like a circular cloud if the errors have constant variance. The example above is a “good” result.
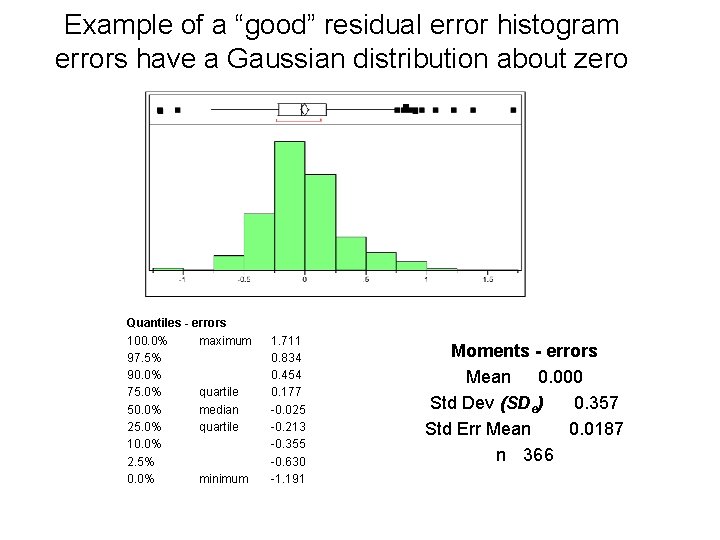
Example of a “good” residual error histogram errors have a Gaussian distribution about zero Quantiles - errors 100. 0% maximum 97. 5% 90. 0% 75. 0% quartile 50. 0% median 25. 0% quartile 10. 0% 2. 5% 0. 0% minimum 1. 711 0. 834 0. 454 0. 177 -0. 025 -0. 213 -0. 355 -0. 630 -1. 191 Moments - errors Mean 0. 000 Std Dev (SDe) 0. 357 Std Err Mean 0. 0187 n 366
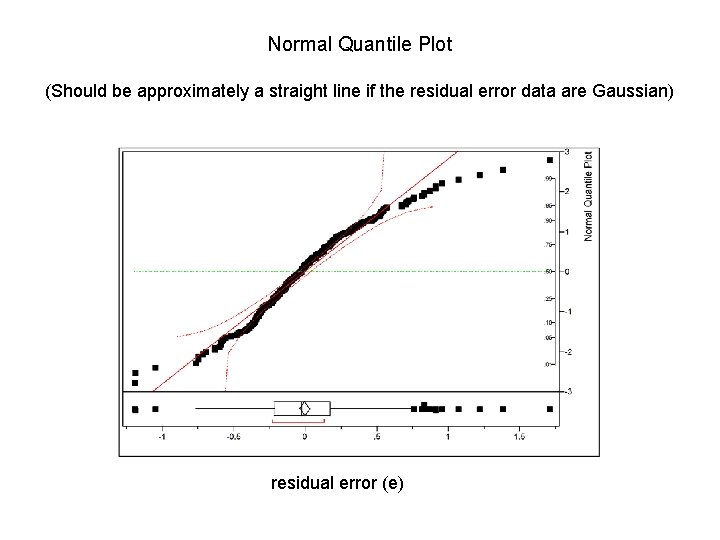
Normal Quantile Plot (Should be approximately a straight line if the residual error data are Gaussian) residual error (e)

Interpreting multiple regression coefficients The multiple regression coefficients will not in general be the same as the individual regression coefficient for each variable one at a time, even though the same Y is being modeled. variable Log PT Log ALT Simple regression (one Y, one X) 3. 560 0. 310 Multiple (simultaneous) (b 1 X 1+b 2 X 2) 3. 470 0. 211 Log Bilirubin = - 3. 70 + 3. 56 log PT, R 2 = 0. 425 Log Bilirubin = -. 105 + 0. 310 log ALT, R 2 = 0. 049 Log Bilirubin = -3. 96 + 3. 47 log PT + 0. 211 log ALT , Log PT coefficients don’t match R 2 = 0. 448

Orthogonally (vs collinearity) In general, regression coefficients from simple and multiple regression are not the same ↔ controlling for covariates does not give the same answer as ignoring covariates. Only when all the X variables have correlation zero with each other will the simple and multiple regression coefficients be the same orthogonally (Collinearity is when Xs are strongly correlated. It is the “opposite” of orthogonality).
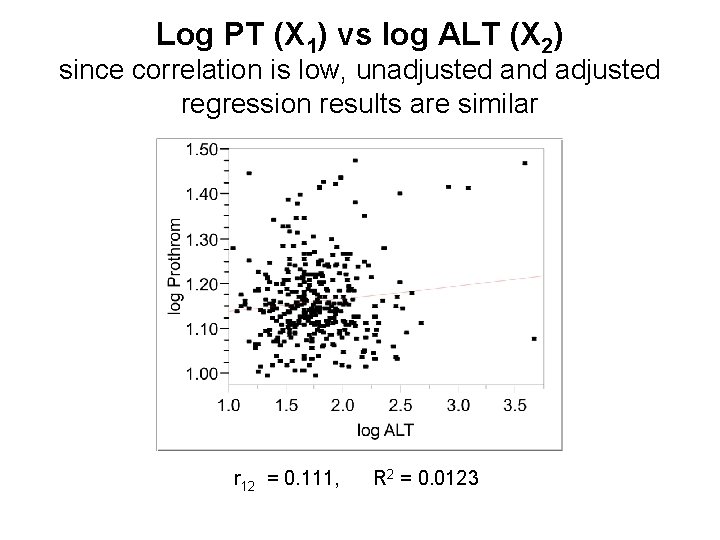
Log PT (X 1) vs log ALT (X 2) since correlation is low, unadjusted and adjusted regression results are similar r 12 = 0. 111, R 2 = 0. 0123

Non orthogonality and variables selection
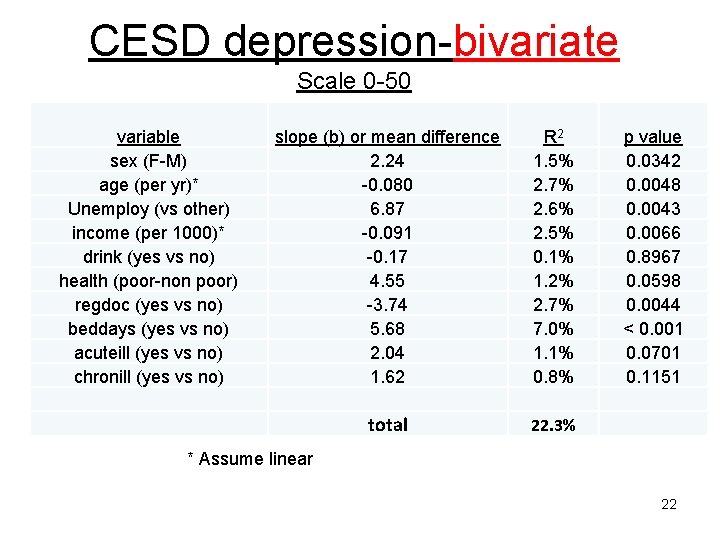
CESD depression-bivariate Scale 0 -50 variable sex (F-M) age (per yr)* Unemploy (vs other) income (per 1000)* drink (yes vs no) health (poor-non poor) regdoc (yes vs no) beddays (yes vs no) acuteill (yes vs no) chronill (yes vs no) slope (b) or mean difference 2. 24 -0. 080 6. 87 -0. 091 -0. 17 4. 55 -3. 74 5. 68 2. 04 1. 62 R 2 1. 5% 2. 7% 2. 6% 2. 5% 0. 1% 1. 2% 2. 7% 7. 0% 1. 1% 0. 8% total 22. 3% p value 0. 0342 0. 0048 0. 0043 0. 0066 0. 8967 0. 0598 0. 0044 < 0. 001 0. 0701 0. 1151 * Assume linear 22
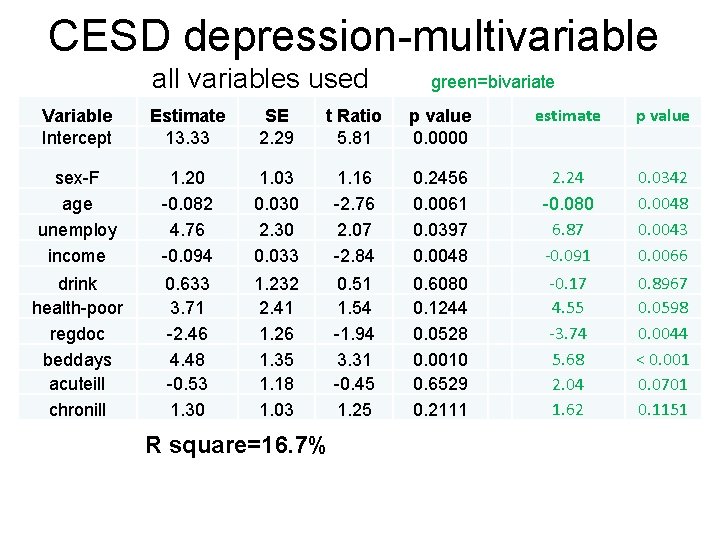
CESD depression-multivariable all variables used green=bivariate Variable Intercept Estimate 13. 33 SE 2. 29 t Ratio 5. 81 p value 0. 0000 estimate p value sex-F age unemploy income 1. 20 -0. 082 4. 76 -0. 094 1. 03 0. 030 2. 30 0. 033 1. 16 -2. 76 2. 07 -2. 84 0. 2456 0. 0061 0. 0397 0. 0048 2. 24 -0. 080 6. 87 -0. 091 0. 0342 0. 0048 0. 0043 0. 0066 drink health-poor regdoc beddays acuteill chronill 0. 633 3. 71 -2. 46 4. 48 -0. 53 1. 30 1. 232 2. 41 1. 26 1. 35 1. 18 1. 03 0. 51 1. 54 -1. 94 3. 31 -0. 45 1. 25 0. 6080 0. 1244 0. 0528 0. 0010 0. 6529 0. 2111 -0. 17 4. 55 -3. 74 5. 68 2. 04 1. 62 0. 8967 0. 0598 0. 0044 < 0. 001 0. 0701 0. 1151 R square=16. 7%
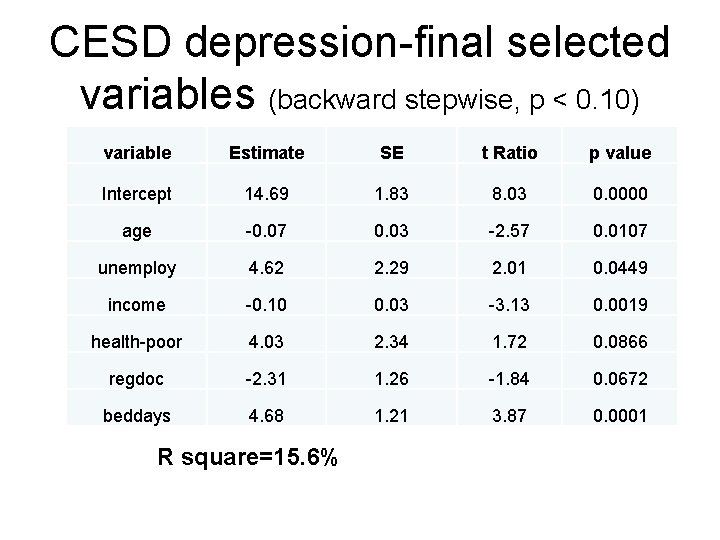
CESD depression-final selected variables (backward stepwise, p < 0. 10) variable Estimate SE t Ratio p value Intercept 14. 69 1. 83 8. 03 0. 0000 age -0. 07 0. 03 -2. 57 0. 0107 unemploy 4. 62 2. 29 2. 01 0. 0449 income -0. 10 0. 03 -3. 13 0. 0019 health-poor 4. 03 2. 34 1. 72 0. 0866 regdoc -2. 31 1. 26 -1. 84 0. 0672 beddays 4. 68 1. 21 3. 87 0. 0001 R square=15. 6%

Interaction Effects & subgroups The model Y = 0 + 1 X 1 + 2 X 2 + implies that change in Y due to X 1 (= 1) is the same (constant) for all values of X 2. An ADDITIVE model. In the model Y = 0 + 1 X 1 + 2 X 2 + 3 X 1 X 2 + the 3 term is an interaction term. Change in Y for a unit change in X 1 is ( 1+ 3 X 2) and is therefore not constant. Positive 3 is often termed a “synergism” Negative 3 is often termed an “antagonism” Additive only if β 3=0 How to implement? Make new variable W = X 1 X 2.

Interaction example Response: Y= log HOMA IR (MESA study) R 2=0. 280, Root Mean Square Error=SDe=0. 623 Mean Response= 0. 395, n= 6782 Parameter Estimates Term Estimate Std Error t Ratio Intercept -1. 388 0. 049 -28. 16 Gender -0. 669 0. 085 -7. 83 BMI 0. 061 0. 00168 36. 45 gender*BMI 0. 028 0. 00299 9. 38 p value <. 0001 Predicted log HOMA IR = -1. 39 – 0. 669 gender + 0. 061 BMI + 0. 028 gender * BMI (gender is coded 0 for female and 1 for male)
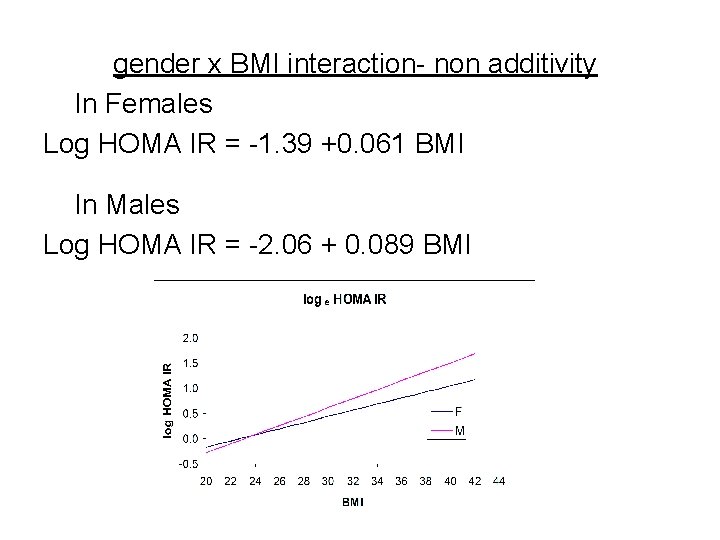
gender x BMI interaction- non additivity In Females Log HOMA IR = -1. 39 +0. 061 BMI In Males Log HOMA IR = -2. 06 + 0. 089 BMI
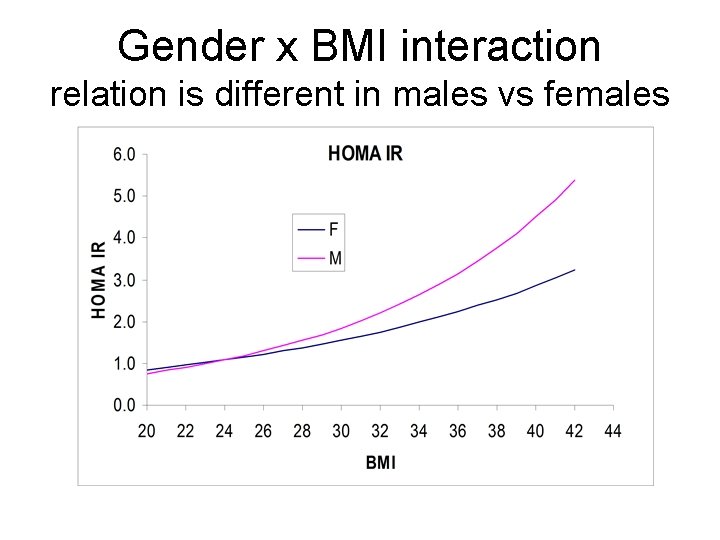
Gender x BMI interaction relation is different in males vs females

Hierarchically well formulated (WWF) regression models HWF Rule – To correctly evaluate the X 1*X 2 interaction, must also have X 1 and X 2 in the model. In general, one must include the lower order terms in order to correctly evaluate the higher order terms.

HWF example- NON HWF Model: chol = a 0 + a 1 smoke x age = SMOKEAGE 0, 1 (dummy) coding: smoke=0 or 1, Variable DF INTERCEPT 1 SMOKEAGE 1 Estimate 156. 863 0. 361 smokeage = smoke x age std error 3. 993 0. 182 t 39. 286 1. 988 p value 0. 0001 0. 0567 -------------------------------------- -1, 1 (effect) coding: smoke=-1 or 1, smokeage =smoke x age Variable DF INTERCEPT 1 SMOKEAGE 1 Estimate 162. 278 0. 055 std error 3. 103 0. 100 t 52. 320 0. 548 p value 0. 0001 0. 5881 p value changes just because coding changes!

HWF example (cont)- HWF: Model: chol = b 0 + b 1 smoke + b 2 age + b 3 smoke x age For HWF models, significance is the same regardless of coding 0, 1 (dummy) coding: smoke=0 or 1, smokeage = smoke x age Variable DF INTERCEPT 1 SMOKE 1 AGE 1 SMOKEAGE 1 Estimate 100. 221 3. 812 2. 010 -0. 009 std error 1. 110 1. 570 0. 036 0. 050 t 90. 304 2. 429 56. 297 -0. 178 p value 0. 0001* 0. 0224 0. 0001* 0. 8599 -1, 1 (effect) coding: smoke=-1 or 1, smokeage=smoke x age Variable DF INTERCEPT SMOKE AGE SMOKEAGE 1 1 Estimate std error 102. 127 0. 785 1. 906 0. 785 2. 005 0. 025 -0. 004 0. 025 * Testing non smokers, ** testing overall t p value 130. 138 0. 0001** 2. 429 0. 0224 79. 437 0. 0001** -0. 178 0. 8599

Regression assumptions Regression can simultaneously evaluate all factors and thus reduce confounding. But must check two critical assumptions. 1. If X is continuous/interval, check if relation of X & Y is linear on some scale. (otherwise polychotomize X) 2. Check if effects of X 1, X 2, X 3 … are additive by adding interaction terms. (Ex: X 4 = X 1 x X 2). Not additive if interactions are significant.

3 Also, in linear regression, prefer residual errors (e) to have a Gaussian distribution with a constant variance that is independent of Y. But additivity and linearity are more important since lack of additivity and linearity lead to bias and is more misleading.

Residual diagnostics & “model criticism” Assumptions of linear regression: 1. Linear relation between Y and each X except for random “noise” (but can transform X). 2. Effect of each X is additive (but can make interaction terms) 3. Errors (e) have constant variance and come from a Gaussian distribution 4. All observations from the same population 5. All observations independent (usually ok) A plot of Ŷ versus e, called a residual error (diagnostic) plot, can help verify if these assumptions are met.
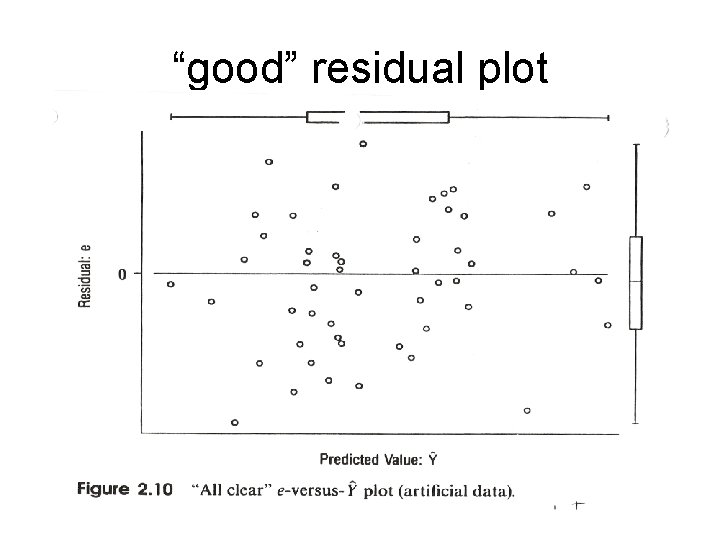
“good” residual plot
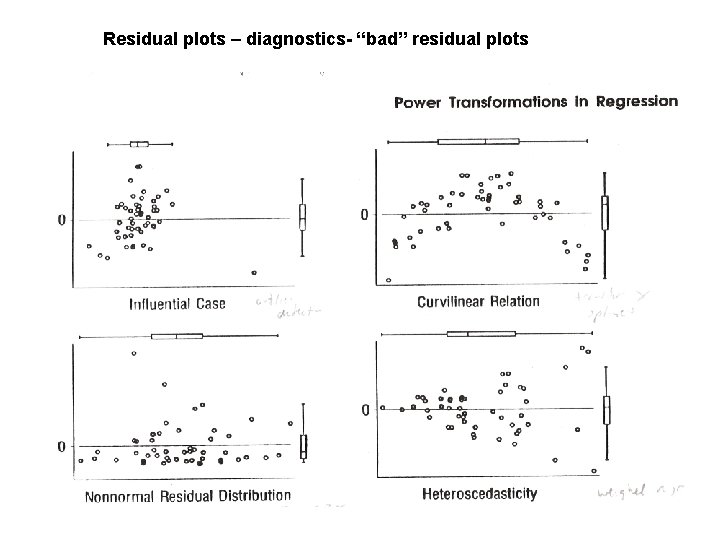
Residual plots – diagnostics- “bad” residual plots

Regression diagnostics Problem-outliers Solution – Find on residual plot & eliminate Problem-Curvilinear (non linear) Solution- try nonlinear trans formation (x 2, 1/x, log(x), ex) Problem-Errors not Gaussian Solution-Robust regression (future class) Problem-Non constant SDe or Var(e) Solution-Weights (future class)

Nonlinear Regression Log(Bilirubin)= -3. 96 + 3. 47 log(PT) + 0. 211 log(ALT) is a nonlinear model in terms of PT and ALT but is a linear model in terms of log PT, log ALT and the regression coefficients b 0=-3. 96, b 1=3. 47 and b 2=0. 211. Consider model of the form: Ŷ= Drug concentration = b 1 10 b 2 X This is nonlinear in b 2 but can be made linear with a transformation: log 10(concentration)=log 10(b 1) + b 2 x What about: Drug concentration = b 0+ b 1 10 b 2 X This model is nonlinear in b 2 and cannot be transformed. Nonlinear regression software is needed to estimate b 0, b 1, & b 2.
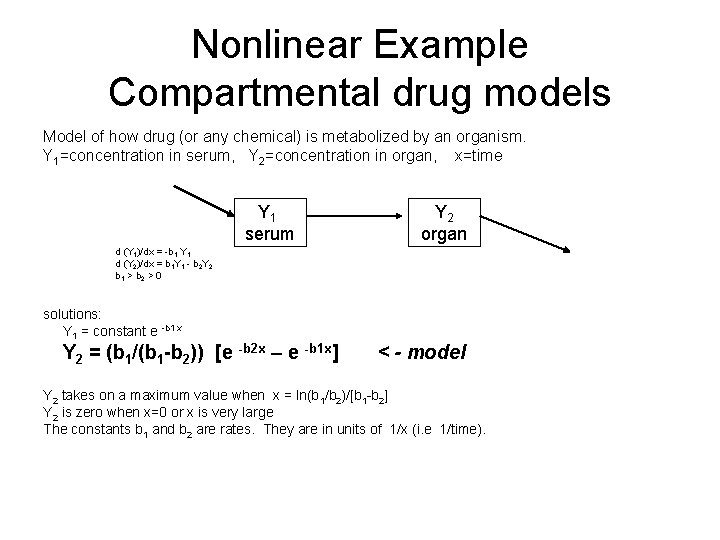
Nonlinear Example Compartmental drug models Model of how drug (or any chemical) is metabolized by an organism. Y 1=concentration in serum, Y 2=concentration in organ, x=time Y 1 serum Y 2 organ d (Y 1)/dx = -b 1 Y 1 d (Y 2)/dx = b 1 Y 1 - b 2 Y 2 b 1 > b 2 > 0 solutions: Y 1 = constant e -b 1 x Y 2 = (b 1/(b 1 -b 2)) [e -b 2 x – e -b 1 x] < - model Y 2 takes on a maximum value when x = ln(b 1/b 2)/[b 1 -b 2] Y 2 is zero when x=0 or x is very large The constants b 1 and b 2 are rates. They are in units of 1/x (i. e 1/time).
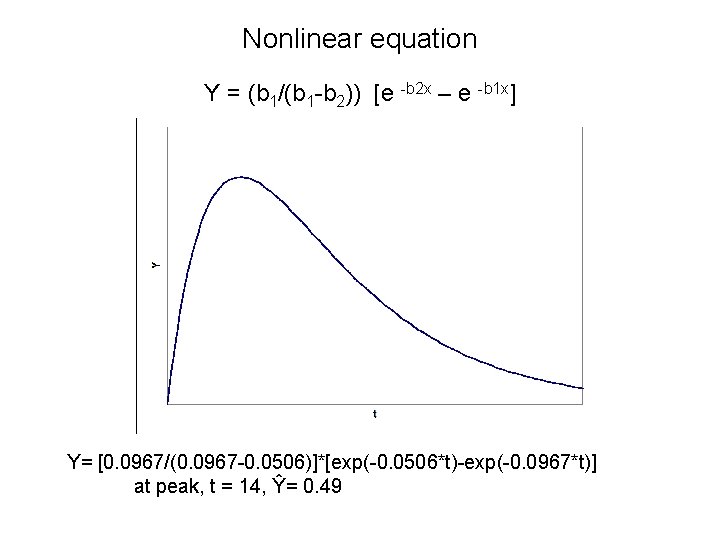
Nonlinear equation Y = (b 1/(b 1 -b 2)) [e -b 2 x – e -b 1 x] Y= [0. 0967/(0. 0967 -0. 0506)]*[exp(-0. 0506*t)-exp(-0. 0967*t)] at peak, t = 14, Ŷ= 0. 49
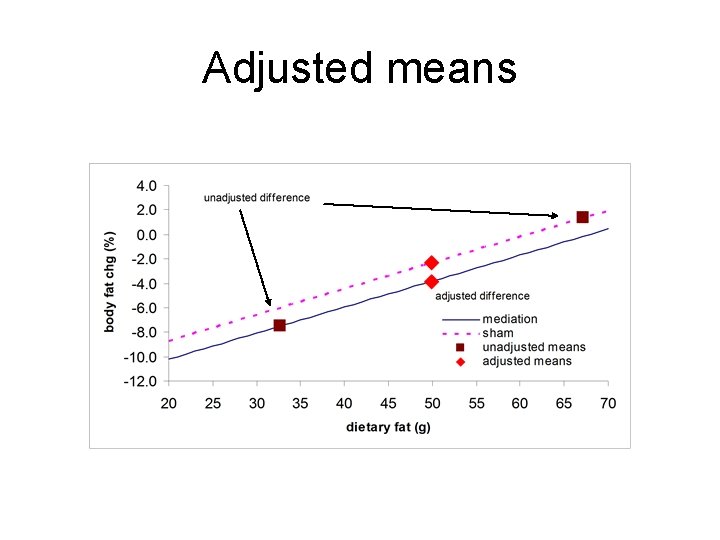
Adjusted means

Ex: Meditation & change in pct body fat Overweight persons chose a meditation program or a “sham” (lectures) as part of a weight loss effort. They were NOT randomized. Change in percent body fat by treatment group (mediation or sham) over three months Unadjusted Means Level n Mean pct body SEM Mean dietary fat (gm) fat change (before study start) 1 -meditate 439 -7. 51% 0. 47% 32. 7 g 2 -sham 704 1. 34% 0. 35% 67. 1 g Unadjusted Mean difference (sham - meditation) = 8. 85% SE of the difference = SEd = √ 0. 47222 + 0. 35322 = 0. 59% t = mean diff/SEd = 8. 85% / 0. 59% = 15. 1, p < 0. 0001 Overall unweighted dietary fat = 49. 9 g

Result via “regression” Y= change in body fat, X = 1 if sham, 0 if meditation Y = a + b X + error = a + b sham + error term estimate SE t p value a -7. 51 0. 46 -16. 4 < 0. 001 b 8. 85 0. 59 15. 2 < 0. 001 Ŷ = -7. 51 + 8. 85 sham

Regr-control for dietary fat pct body fat= Y = a + b 1 X 1 + b 2 X 2 + error X 1= 1 if sham, 0 if meditation X 2 = dietary fat in g term estimate SE t p value a -14. 5 0. 54 -26. 6 < 0. 001 b 1 1. 51 0. 56 2. 7 0. 007 b 2 0. 21 0. 011 18. 9 <0. 001 body fat chg=-14. 5 +1. 51 sham +0. 21 diet fat
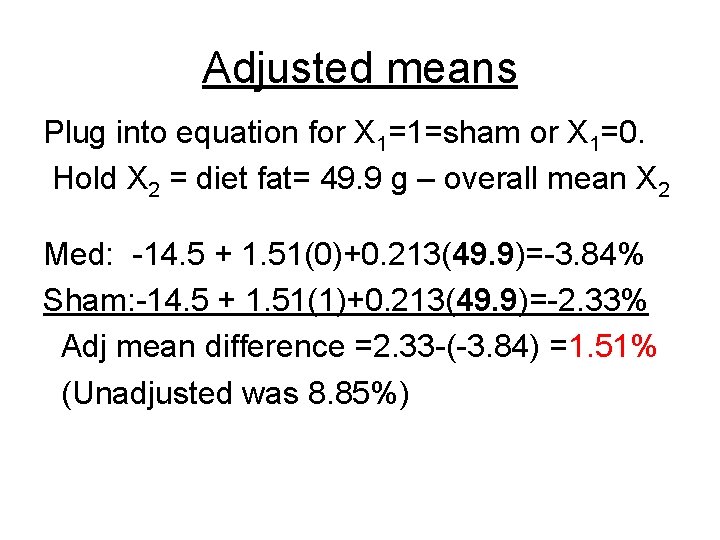
Adjusted means Plug into equation for X 1=1=sham or X 1=0. Hold X 2 = diet fat= 49. 9 g – overall mean X 2 Med: -14. 5 + 1. 51(0)+0. 213(49. 9)=-3. 84% Sham: -14. 5 + 1. 51(1)+0. 213(49. 9)=-2. 33% Adj mean difference =2. 33 -(-3. 84) =1. 51% (Unadjusted was 8. 85%)

General procedure Adjusted means for X 1, controlling for (confounders) X 2, X 3, X 4 … 1. Estimate model regression coefficients 2. Plug in different values for X 1, and values for all other Xs held constant at their overall means. Gives adjusted means and their SEs. Assumes ? ?

Test for linearity Assume “f(βs, X)” is an unknown non linear function that is part of the relationship of X to Y. The spline is an estimate of f(βs, X). We fit the model: Y = β 0 + β 1 X + f(βs, X) + error where f(βs, X)=β 2 g 2(X) + β 3 g 3(X) + β 4 g 4(X)+ … +βkgk(X) so Y – β 0 – β 1 X = f(βs, X) + error If all βs=0, then we have “proven” that the relation between Y and X is linear. We examine the spline plot to see if a parametric form for f(X) is suggested (ie f(X)=X 2, f(X)=log(X), …). (Note: fractional polynomial alternative not available in JMP) 47

Example: 2+2=4 knot spline (s=2, 3) Y = β 0 + β 1 X + β 2 g 2(X) + β 3 g 3(X) + error where g 2 and g 3 are unknown functions each estimated by a cubic spline. The X axis is broken into 4 segments. If β 2=0 and β 3=0, then Y = β 0 + β 1 X + error and we have proven that, except for error, the relation between Y and X is linear with slope β 1.

VARIABLE SELECTION in regression 49

Model performance criteria A possible criterion for variable selection is to find the variables that maximize R square = 1 - (SDe)/SDy (or estimated R square = 1 – SSerror/SStotal) where SDe = residual error variance. So, maximizing R square same as minimizing SDe. But SDe decreases (at least a bit) when ANY variable is added, even if not significant = overfitting. 50
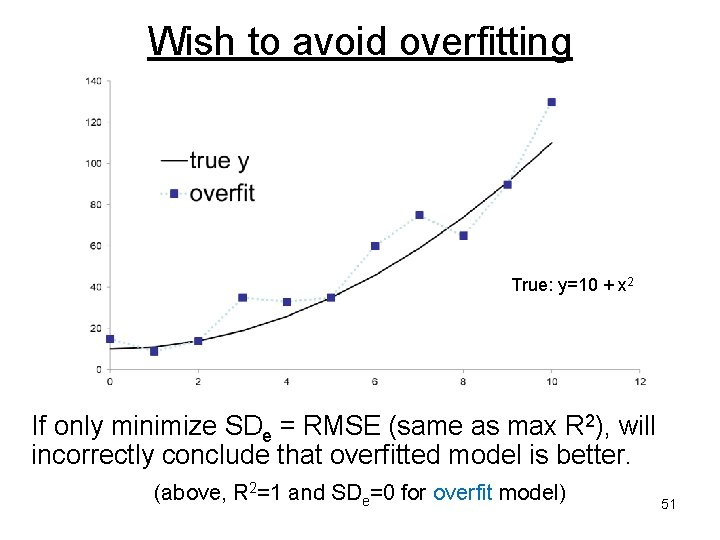
Wish to avoid overfitting True: y=10 + x 2 If only minimize SDe = RMSE (same as max R 2), will incorrectly conclude that overfitted model is better. (above, R 2=1 and SDe=0 for overfit model) 51

Finding the “best” modelalternatives to p values Akaike’s Information Criterion AIC = n log(SDe 2) + 2 K Schwarz’s Bayesian Information Criterion BIC = n log(SDe 2) + K log n Minimize AIC or BIC Similar to min SDe 2<-> max R 2 but with a penalty for adding more variables. SDe 2 = MSE= MS error, K=num variables, n=sample size 52

Variable searching Have pool of many potential predictors X 1, X 2, … XK-1, some of which may be interactions and/or non linear transformations. Question- Are they all simultaneously significant? Which are “important”? Would like a parsimonious model. Criteria: Omitting variables should not greatly lower R 2 or greatly increase RMSE=SDe Omitted variables should not be statistically significant. 53

Which predictors to include? Scientific criteria: Known established predictors should be included (forced in) regardless of the p value or R 2 (unless there are proxies for it). Example: age is a predictor of pregnancy Practical criteria: potential predictor variable should vary. potential predictor should not be missing for most subjects. Remove “outliers” Harrell: Regression modelling strategies, Springer, 2001 54

Strategy: First do bivariate assessment (Y, one X) – Helps find outliers, mistakes, variables that do not vary. But do NOT use this as a final screen. X can be non significant in bivariate and significant in multivariate or the reverse. Then do multivariate – may wish to include at least the important two way interactions, particularly if one of the variables is the “primary” predictor of interest. Check for 55 linearity if X is continuous.

Variable selection methods (partial list) 1. Forward stepping – Include most significant, reevaluate, include next most significant, re-evaluate, … 2. Backward stepping – Start with all, remove least significant, re-evalaute, remove next least significant, re-evaluate, … (see Harrell) #1 & #2 can be executed with a “reverse look” at each step. Can use AIC, BIC instead of p values 3. All possible subsets–Look at every possible combination and sort by AIC, BIC. 4. “Shrinkage” methods-LASSO/Elastic Net 5. “Boosting” (not in JMP yet, only R) 6. Particle Swarm (not in JMP or R) 7. Information gain ranking (not in JMP) 8. “Genetic” Algorithms (not in JMP) 56
- Slides: 56