Multimodal Integration and Intersubject Registration surfer nmr mgh

Multimodal Integration and Inter-subject Registration surfer. nmr. mgh. harvard. edu

Overview • Intra-subject registration • Affine transformations • Registration, Automatic and Manual • f. MRI Integration • f. MRI Analysis Intro • Registration • Viewing on Volume and Surface • ROI analyses • Surface-based group analysis • Inter-subject registration 2
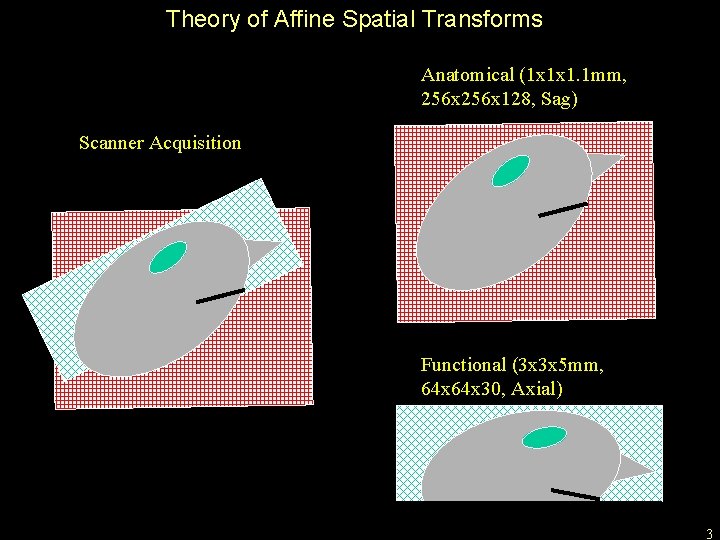
Theory of Affine Spatial Transforms Anatomical (1 x 1 x 1. 1 mm, 256 x 128, Sag) Scanner Acquisition Functional (3 x 3 x 5 mm, 64 x 30, Axial) 3
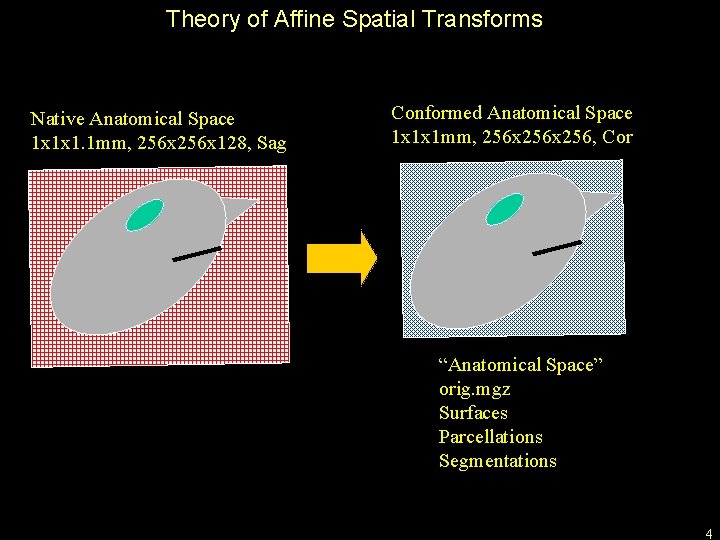
Theory of Affine Spatial Transforms Native Anatomical Space 1 x 1 x 1. 1 mm, 256 x 128, Sag Conformed Anatomical Space 1 x 1 x 1 mm, 256 x 256, Cor “Anatomical Space” orig. mgz Surfaces Parcellations Segmentations 4
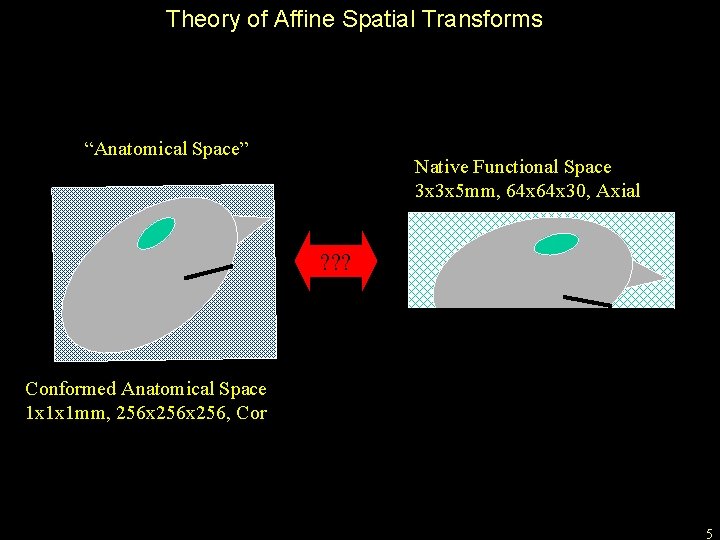
Theory of Affine Spatial Transforms “Anatomical Space” Native Functional Space 3 x 3 x 5 mm, 64 x 30, Axial ? ? ? Conformed Anatomical Space 1 x 1 x 1 mm, 256 x 256, Cor 5
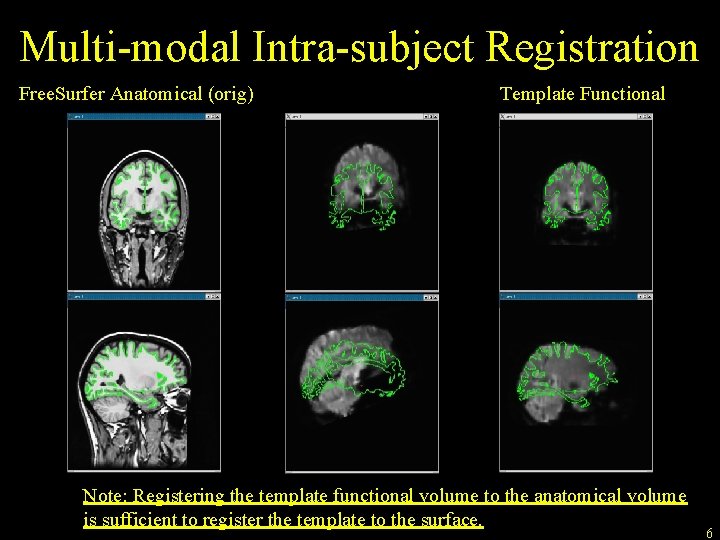
Multi-modal Intra-subject Registration Free. Surfer Anatomical (orig) Template Functional Note: Registering the template functional volume to the anatomical volume is sufficient to register the template to the surface. 6

Free. Surfer Registration and Template Volume Free. Surfer Subject-Specific • Volumes • Surfaces • Thickness • ROIs Template Volume Registration • f. MRI • DTI • ASL • PET • … Template Volume: • In voxel-for-voxel registration with parameter map • Best gray-white contrast 7

Free. Surfer Registration Matrix • 4 x 4 Matrix • As many as 12 DOF (usually 6) • Simple Text file • Coordinate system not easy to explain mgh-02407836 -v 2 3. 437500 5. 000000 0. 150000 9. 999985 e-01 -1. 428481 e-03 -8. 293565 e-04 5. 281680 e-01 4. 641568 e-04 -2. 388080 e-01 9. 710670 e-01 -4. 041043 e+01 1. 585159 e-03 9. 710652 e-01 2. 388064 e-01 -1. 376212 e+00 0 1 round 8

Free. Surfer Registration Matrix • 4 x 4 Matrix • As many as 12 DOF (usually 6) • Simple Text file • Coordinate system not easy to explain mgh-02407836 -v 2 Free. Surfer subject name 3. 437500 Functional In-plane resolution mm 5. 000000 Functional Between-plane resolution mm 0. 150000 Intensity (for visualization) 9. 999985 e-01 -1. 428481 e-03 -8. 293565 e-04 5. 281680 e-01 4. 641568 e-04 -2. 388080 e-01 9. 710670 e-01 -4. 041043 e+01 1. 585159 e-03 9. 710652 e-01 2. 388064 e-01 -1. 376212 e+00 0 1 Legacy round 9

Automatic Intra-subject Registration bbregister -–subject bert –-mov mmtemplate. nii --bold --init-fsl –-reg register. dat Command name Free. Surfer subject name Multimodal template volume Multimodal contrast Initialize with FSL-FLIRT Output registration file • BB = Boundary-based, about 5 min. • Registers template to conformed anatomical of given subject (bert) • Registration is initialized with FSL-FLIRT • 6 DOF • Initialization also with --init-spm and --init-header • About 5 min 10

Manual Intra-subject Registration tkregister 2 –-mov mmtemplate. nii –-reg register. dat --surfs Command name Multimodal template volume registration file Display white surf • Note similarity to bbregister command • Subject not needed (alread in register. dat file) tkregister 2 --help 11
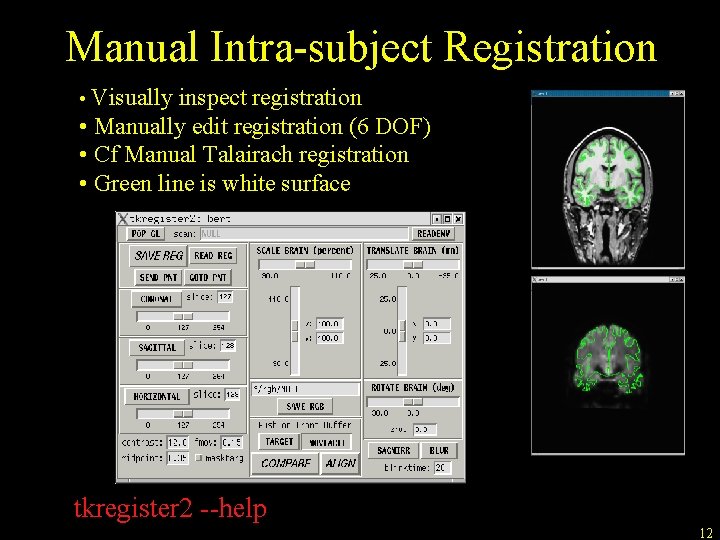
Manual Intra-subject Registration • Visually inspect registration • Manually edit registration (6 DOF) • Cf Manual Talairach registration • Green line is white surface tkregister 2 --help 12
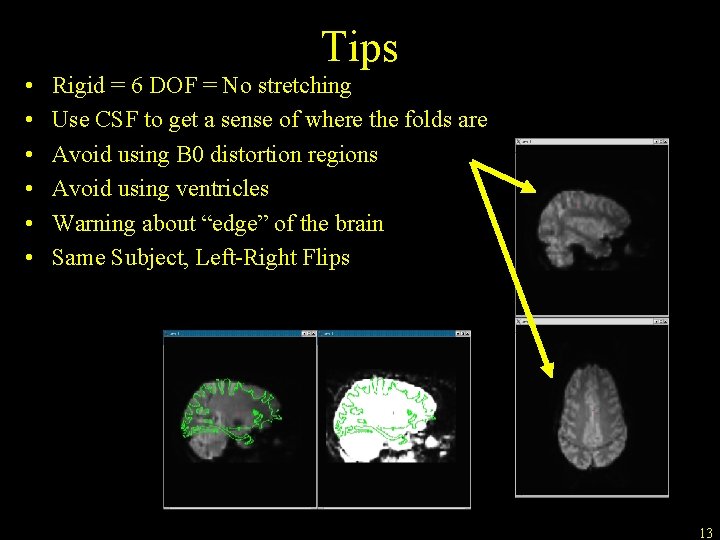
Tips • • • Rigid = 6 DOF = No stretching Use CSF to get a sense of where the folds are Avoid using B 0 distortion regions Avoid using ventricles Warning about “edge” of the brain Same Subject, Left-Right Flips 13

Command-line Tools Automatic Registration: • bbregister --help • fslregister –help • spmregister –help • reg-feat 2 anat –help } Manual Registration: • tkregister 2 --help Transformations: • mri_vol 2 surf --help • mri_vol 2 vol --help • mri_label 2 vol --help • mri_surf 2 vol --help Free. Surfer Scripts

f. MRI Integration • Visualize individual f. MRI results on • surface • volume • ROI Volume Study: Count number of voxels above threshold in an anatomical ROI • ROI Intensity Study: Average HRF inside of an ROI • Surface-based f. MRI group analysis 15

Hemodynamic Response (BOLD) Time-to-Peak (~6 sec) Dispersion TR (~2 sec) Equilibrium (~16 -32 sec) Undershoot Delay (~1 -2 sec) 16
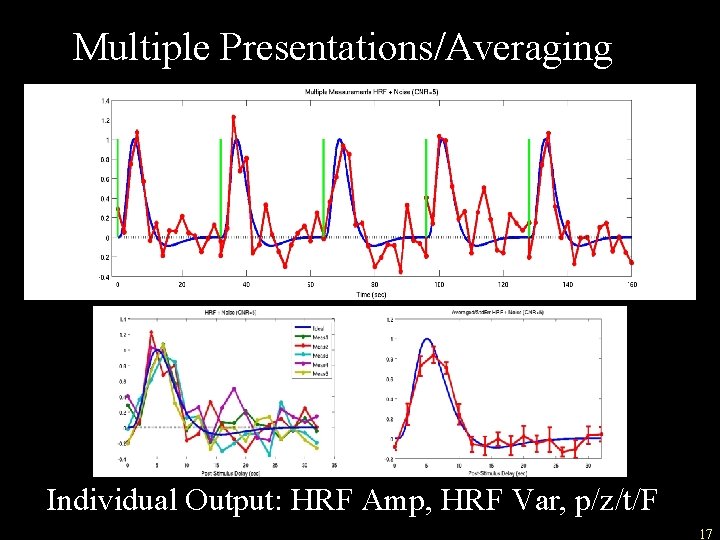
Multiple Presentations/Averaging Individual Output: HRF Amp, HRF Var, p/z/t/F 17

f. MRI Preprocessing Overview • Motion Correction (MC Template) • Use for registration template • bbregister --mov mctemplate. nii --s subject --init-fsl --reg register. dat • tkregister 2 --mov mctemplate. nii --reg register. dat --surf • Do not use nonlinear resampling to talairach/MNI space • Do not spatially smooth (3 d) 18

f. MRI Analysis Overview • First-Level (Individual) Analysis • HRF Amplitude (or Contrast of Amplitudes) • cope (FSL), • CON (SPM), • ces (FSFAST) • Variance of Amplitude • varcope (FSL), ? ? ? (SPM), cesvar (FSFAST) • Activation/Significance Maps: • z • t • F • sig (-log 10(p)) • All in alignment with MC Template!!!! 19

Template and Map Functional Template+ Map 20

Volume Viewing tkmedit subject orig. mgz –aux brain. mgz –aparc+aseg -overlay sig. nii –reg register. dat -fthresh 2 –fmax 4 sig. nii – significance map in native functional space. could have been z, t, or F map as well. register. dat – freesurfer registration file fthresh – lower threshold (value depends on map). You can change this in the interface. fmax – saturation threshold. (value depends on map). You can change this in the interface. aparc+aseg – display aparc+aseg. mgz. Can load this from the interface too. 21
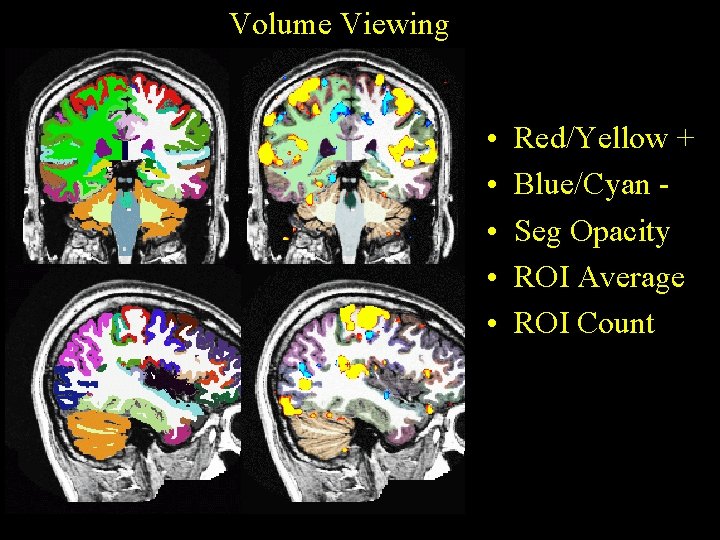
Volume Viewing • • • Red/Yellow + Blue/Cyan Seg Opacity ROI Average ROI Count

Sampling onto the Surface Pial White/Gray
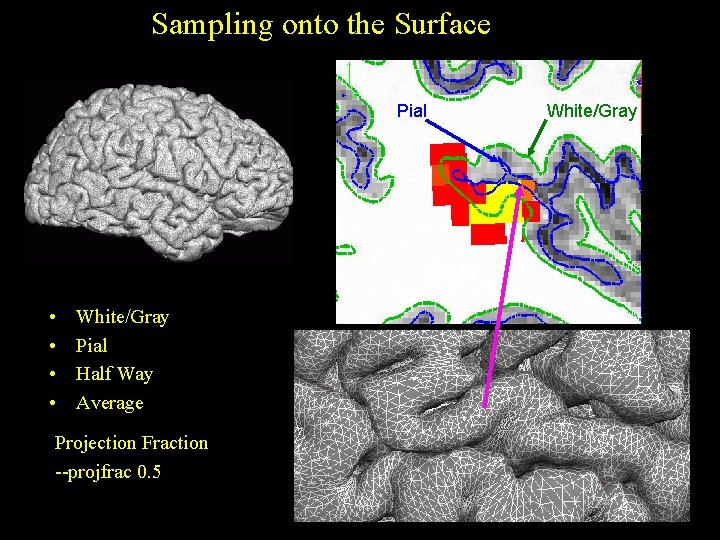
Sampling onto the Surface Pial • • White/Gray Pial Half Way Average Projection Fraction --projfrac 0. 5 White/Gray
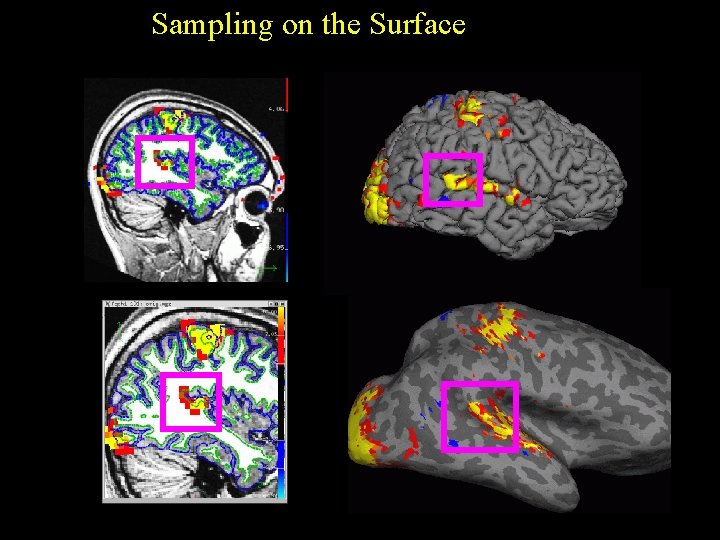
Sampling on the Surface
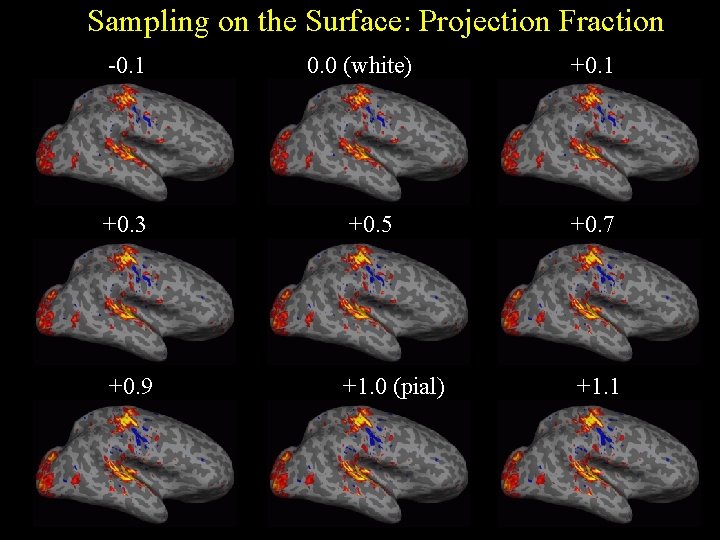
Sampling on the Surface: Projection Fraction -0. 1 0. 0 (white) +0. 1 +0. 3 +0. 5 +0. 7 +0. 9 +1. 0 (pial) +1. 1

Surface Viewing Resample HRF Contrast Significance to left hemisphere mri_vol 2 surf --mov sig. nii --reg register. dat --hemi lh --projfrac 0. 5 --o lh. sig. mgh map in native functional space freesurfer registration file hemisphere projection fraction (half) output (Nvertices-x-1 mgh format) Note similarity to bbregister and tkregister 2 commands. Load HRF Contrast Significance as overlay tksurfer subject lh inflated -aparc –overlay lh. sig. mgh 27
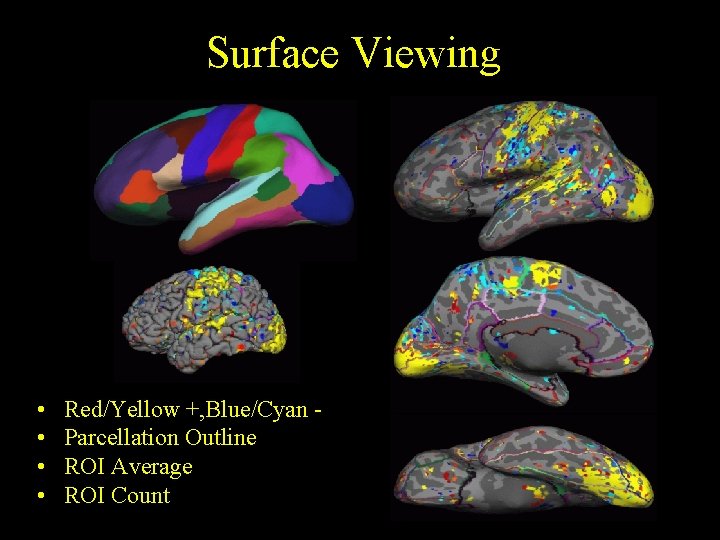
Surface Viewing • • Red/Yellow +, Blue/Cyan Parcellation Outline ROI Average ROI Count

ROI f. MRI Analyses: • Intensity – Full Anatomical ROI – Functionally Constrained ROI • Volume
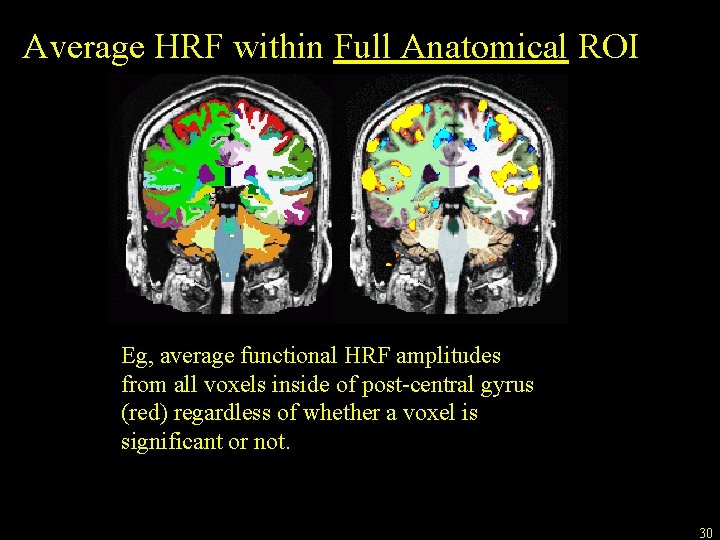
Average HRF within Full Anatomical ROI Eg, average functional HRF amplitudes from all voxels inside of post-central gyrus (red) regardless of whether a voxel is significant or not. 30

Average HRF within Full Anatomical ROI Resample HRF Contrast to anatomical space mri_vol 2 vol --mov ces. nii --reg register. dat --interp nearest --fstarg --o ces. anat. mgh Command name HRF map in functional space Free. Surfer Registration File Nearest neighbor interpolation Specify anatomical output space Output file in anatomical space Note similarity to bbregister, tkregister 2, and mri_vol 2 surf commands. Average HRF Contrast within ROIs mri_segstats --seg $SUBJECTS_DIR/subject/mri/aseg. mgz --ctab $FREESURFER_HOME/Free. Surfer. Color. LUT. txt --i ces. anat. mgh --sum ces. aseg. stats 31

Average HRF within Full Anatomical ROI Average HRF Contrast within ROIs mri_segstats --seg $SUBJECTS_DIR/subject/mri/aseg. mgz --ctab $FREESURFER_HOME/Free. Surfer. Color. LUT. txt --i ces. anat. mgh --sum ces. aseg. stats Notes: --seg is the segmentation (eg, aseg. mgz, aparc+aseg. mgz, etc) --ctab is matching color lookup table Output File: ces. aseg. stats • simple text file with same format aseg. stats • multiple subjects can be combined with asegstats 2 table 32
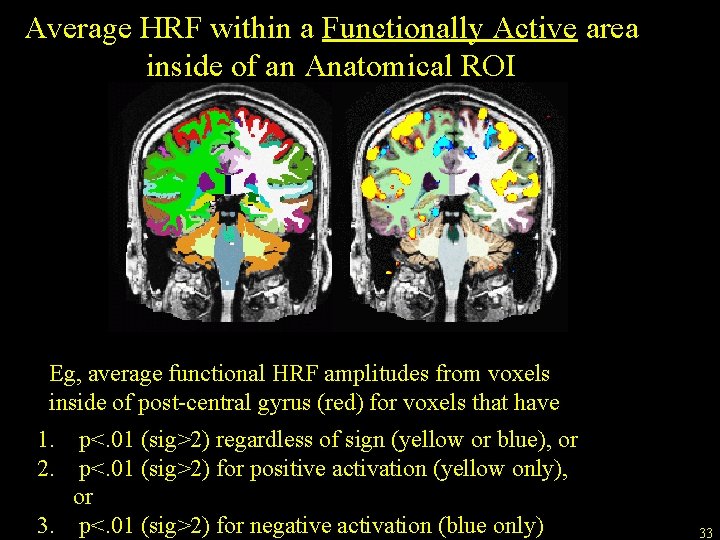
Average HRF within a Functionally Active area inside of an Anatomical ROI Eg, average functional HRF amplitudes from voxels inside of post-central gyrus (red) for voxels that have 1. p<. 01 (sig>2) regardless of sign (yellow or blue), or 2. p<. 01 (sig>2) for positive activation (yellow only), or 3. p<. 01 (sig>2) for negative activation (blue only) 33

Average HRF only within the Functionally Active Area of an Anatomical ROI, Count Voxels above threshold in each ROI Resample HRF Contrast Significance to anatomical space mri_vol 2 vol --mov sig. nii --reg register. dat --interp nearest --fstarg --o sig. anat. mgh Average HRF Contrast within functionally constrained ROIs mri_segstats --seg $SUBJECTS_DIR/subject/mri/aseg. mgz --ctab $FREESURFER_HOME/Free. Surfer. Color. LUT. txt --i ces. anat. mgh --sum ces. aseg. mask. stats --mask sig. anat. mgh --mask-thresh 2 --mask-sign abs 34

Average HRF only within the Functionally Active Area of an Anatomical ROI, Count Voxels above threshold in each ROI mri_segstats --seg $SUBJECTS_DIR/subject/mri/aseg. mgz --ctab $FREESURFER_HOME/Free. Surfer. Color. LUT. txt --i ces. anat. mgh --sum ces. aseg. mask. stats --mask sig. anat. mgh --mask-thresh 2 --mask-sign abs • Volume in stats file is volume above threshold (may be 0) • Sign is important for Average! • abs, pos, or neg • pos will always result in positive HRF average • neg will always result in negative HRF average • abs ? ? • Careful to avoid circularity • Can use a different contrast to mask 35

Surface-based Group Analysis mris_preproc --hemi lh --o lh. fsaverage. ces. mgh --iv subject 1/ces. nii subject 1 func/register. dat --iv subject 2/ces. nii subject 2 func/register. dat --iv subject 3/ces. nii subject 3 func/register. dat . . . After that, everything else is the same as a thickness study … mris_fwhm --i lh. fsaverage. ces. mgh --fwhm 10 --o lh. fsaverage. ces. sm 10. mgh --cortex mri_glmfit --surf fsaverage lh –cortex --y lh. fsaverage. ces. sm 10. mgh. . . 36

Tutorial 1. Registration – manual and automatic registration 2. f. MRI Integration (Sensorimotor Paradigm) 1. Individual 1. Volume view sig 2. Surface view sig 3. ROI analysis with and without functional constraint 2. Group 1. mris_preproc 2. ROI analysis (asegstats 2 table) 37

Another type of problem: intersubject uni-modal registration • Surface-based (2 D) registration does an excellent job of aligning cortical folds, but does not apply to non-cortical structures (e. g. basal ganglia). • Volumetric (3 D) registration applies to the entire brain but does not, in general, align folding patterns. • Goal: combined their strength 38
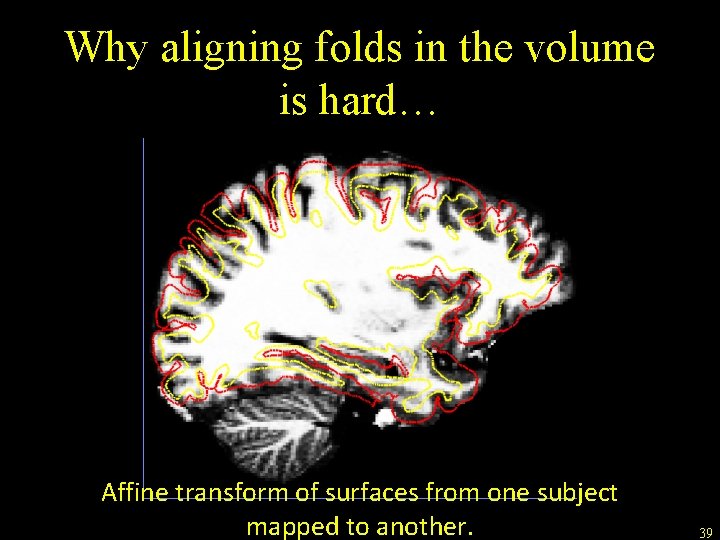
Why aligning folds in the volume is hard… Affine transform of surfaces from one subject mapped to another. 39
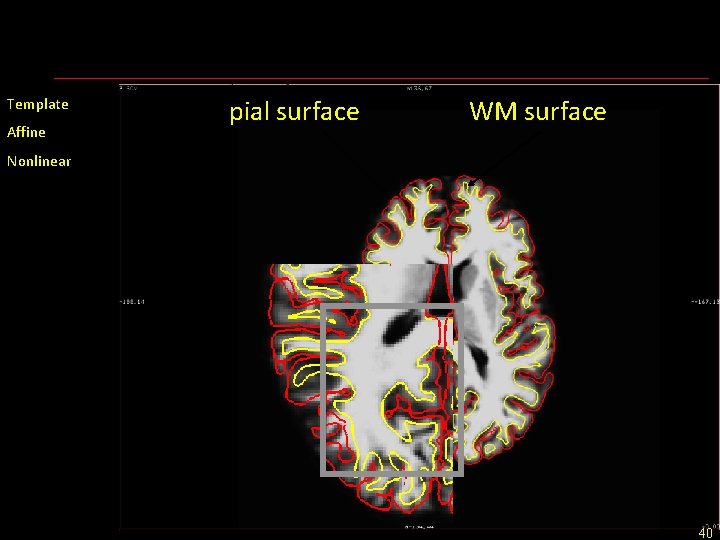
Template Affine pial surface WM surface Nonlinear 40
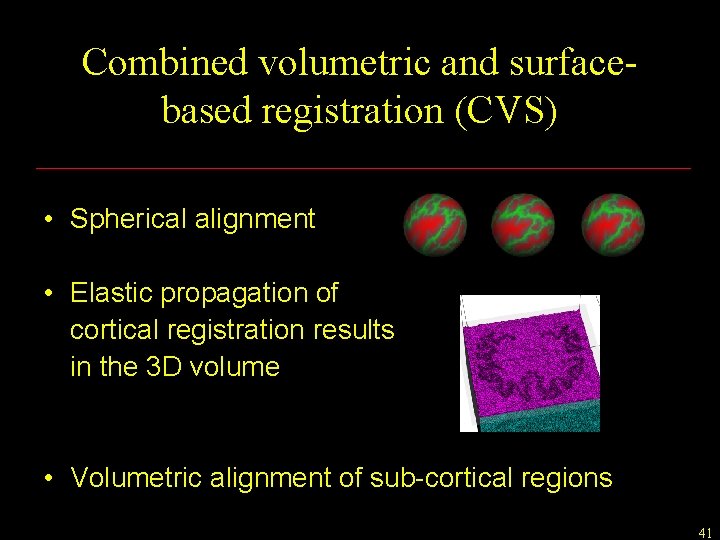
Combined volumetric and surfacebased registration (CVS) • Spherical alignment • Elastic propagation of cortical registration results in the 3 D volume • Volumetric alignment of sub-cortical regions 41
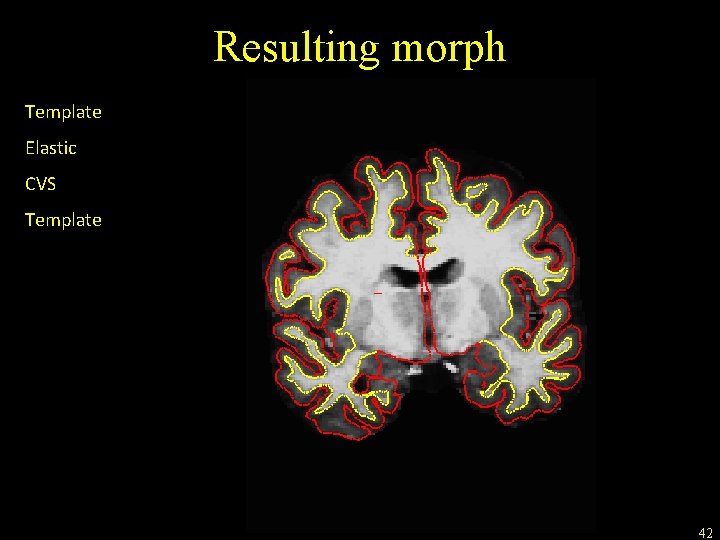
Resulting morph Template Elastic CVS Template 42
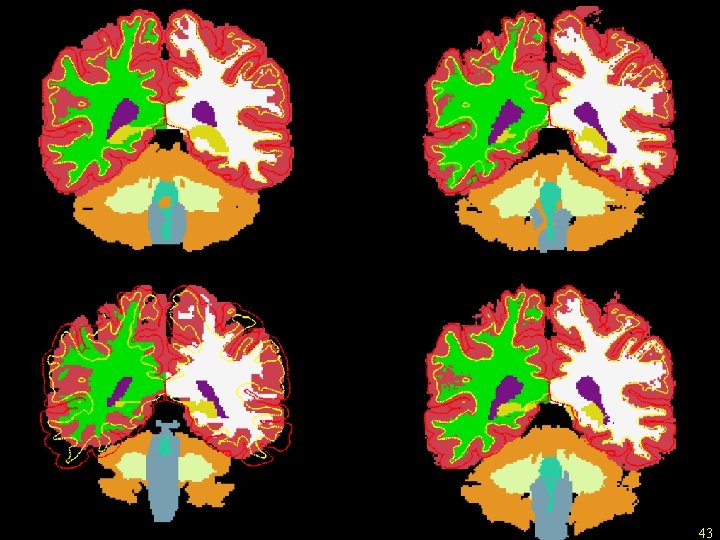
Template HAMMER FLIRT CVS 43
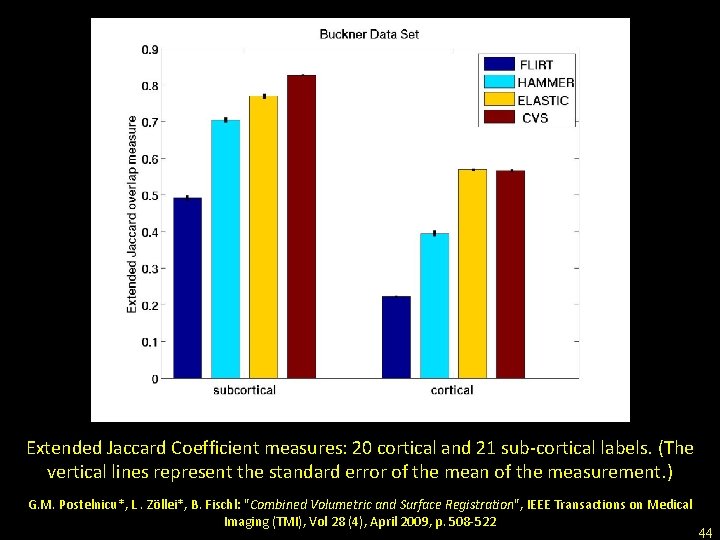
Extended Jaccard Coefficient measures: 20 cortical and 21 sub-cortical labels. (The vertical lines represent the standard error of the mean of the measurement. ) G. M. Postelnicu*, L. Zöllei*, B. Fischl: "Combined Volumetric and Surface Registration", IEEE Transactions on Medical Imaging (TMI), Vol 28 (4), April 2009, p. 508 -522 44

mri_cvs_register --mov subjid • registering the subject to, by default, the CVS atlas space • make sure that the SUBJECTS_DIR for subjid is correctly set Optional Arguments -- template subjid : subjid for template subject -- templatedir : recon directory for template (default is SUBJECTS_DIR) --outdir : output directory for all the results (default is SUBJECTS_DIR/subjid/cvs) … and many more: use --help 45

mri_cvs_register Optional Arguments (cont) --step 1 Only do step 1 (spherical registration). --step 2 Only do step 2 (elastic registration). --step 3 Only do step 3 (volumetric registration). --noaseg Do not use aseg volumes in the volumetric registration pipeline (default is 0). Setting this option could shorten significantly the time of registration, however, might also take away from the accuracy of the final results. 46

mri_cvs_register Optional Arguments (cont) --nocleanup Do not delete temporary files (default is 0). --keepelreg Do not delete elastic registration (default is 0) outcome. --cleanall Recompute all CVS-related morphs that might have been computed prior to the current CVS run (def = 0). --cleansurfreg Recompute CVS-related surface registration morphs that might have been computed prior to the current CVS run (def = 0). --cleanelreg Overwrite /recompute the CVS-related elastic registration morph that might have been computed prior to the current CVS run (default is 0). --cleanvolreg Overwrite / recompute CVS-related volumetric morphs that might have been computed prior to the current CVS run (default is 0). 47
CVS atlas 48

related commands • mri_cvs_check – checking whether all files needed for a successful CVS registration are present • mri_cvs_data_copy – copying the CVS-relevant recon directories over to a new location • mri_vol 2 vol – applying the CVS registration morph to files corresponding to the moving subject 49

Applying CVS morphs mri_vol 2 vol 1. applying CVS morph to aseg file mri_vol 2. --targ templateid --m 3 z morph. m 3 z --no. Def. M 3 z. Path --mov asegvol --o asegvol 2 CVS --interp nearest --no-save-reg applying morph to corresponding diffusion file mri_vol 2 vol --targ templateid --m 3 z morph. m 3 z --no. Def. M 3 z. Path --reg 2 anat. register. dat --mov diffvol --o diffvol 2 CVS --no-save-reg 50

Applying CVS morphs apply. Morph: use with older style CVS morphs (. tm 3 d) 1. applying CVS morph to aseg file apply. Morph 2. --templateid --transform morph. tm 3 d vol asegvol 2 CVS nearest applying morph to corresponding diffusion file apply. Morph --templateid –transform morph. tm 3 d vol diffvol 2 anat diffvol 2 CVS linear 51

Application of CVS to tractography • Goal: fiber bundle alignment • Study: compare CVS to methods directly aligning DWI-derived scalar volumes • Conclusion: high accuracy cross-subject registration based on structural MRI images can provide improved alignment • Zöllei, Stevens, Huber, Kakunoori, Fischl: “Improved Tractography Alignment Using Combined Volumetric and Surface Registration”, Neuro. Image 51 (2010), 206 -213 52
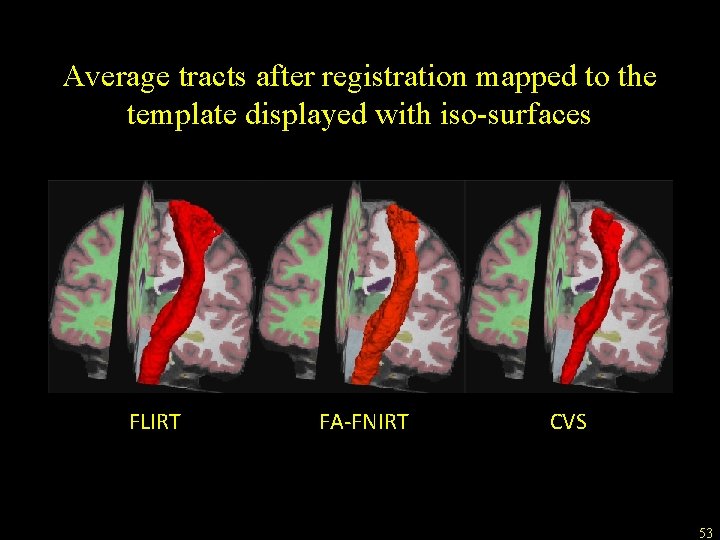
Average tracts after registration mapped to the template displayed with iso-surfaces FLIRT FA-FNIRT CVS 53
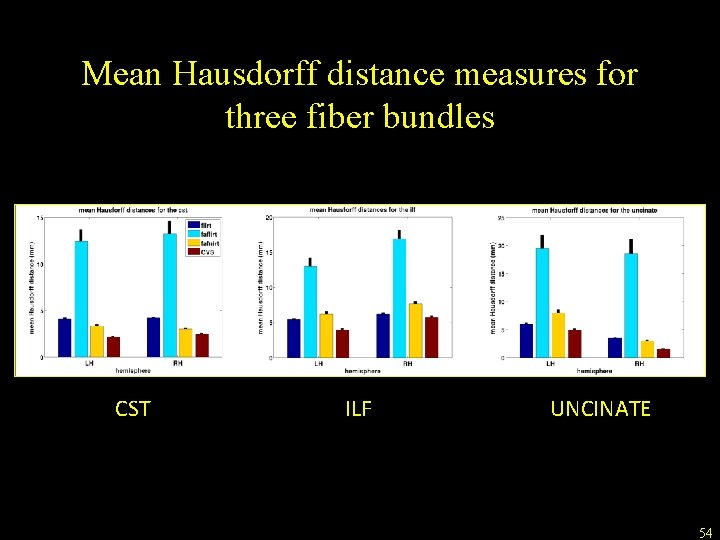
Mean Hausdorff distance measures for three fiber bundles CST ILF UNCINATE 54
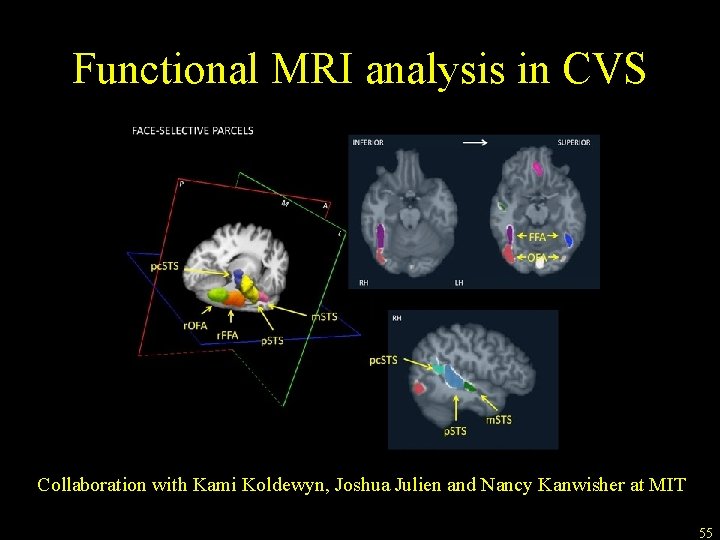
Functional MRI analysis in CVS space Collaboration with Kami Koldewyn, Joshua Julien and Nancy Kanwisher at MIT 55

Ongoing development • Improve CVS capability to register ex-vivo to in-vivo acquisitions • Implemented MI-based volumetric registration (for CVS step 3) to accommodate intensity profile differences • Qualitative preliminary results on 4 subjects • L. Zöllei, Allison Stevens, Bruce Fischl: Non-linear Registration of Intra-subject Ex-vivo and In-vivo Brain Acquisitions, Human Brain Mapping, June 2010 • L. Zöllei, B. Fischl, Automatic segmentation of ex-vivo MRI images using CVS in Free. Surfer, HBM 2011 56
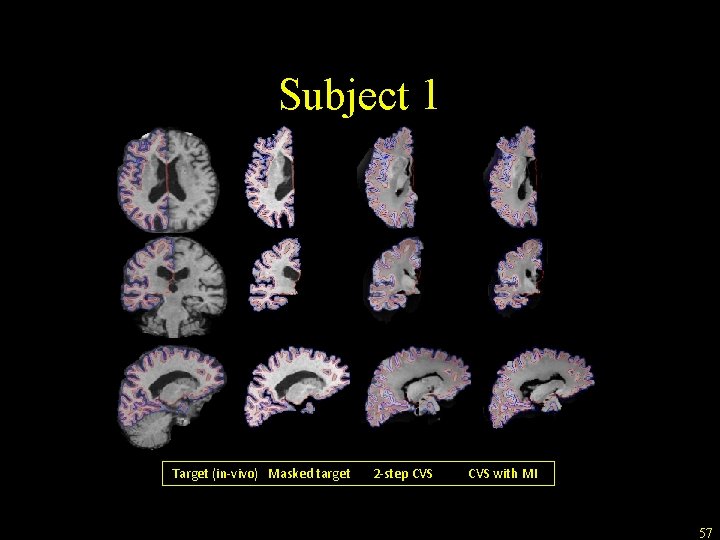
Subject 1 Target (in-vivo) Masked target 2 -step CVS with MI 57

registration tool summary • mris_register • fslregister: bet + flirt • bbregister • mri_robust_register • mri_cvs_register – mri_nl_align 58

registration morph summary • . dat, . lta, . xfm, . fslmat: encode rigid and affine transformations – mri_vol 2 vol • sphere. reg: encodes spherical morph – mris_resample • . m 3 z, . tm 3 d: encode nonlinear volumetric morphs – mri_vol 2 vol, apply. Morph 59

Summary • Multi/Cross-modal map (HRF Amplitude, FA) • Multimodal Integration requires a Template • A Template is: • Same size as multimodal map • In Voxel-to-voxel alignment with map • Has better anatomical contrast • Baseline functional • Low-B DTI • Usually a MC template • Volume and Intensity ROI Analyses • Functionally-constrained ROI 60
- Slides: 60