MRI preprocessing and segmentation Bias References Segmentation References

MRI preprocessing and segmentation

Bias References

Segmentation References
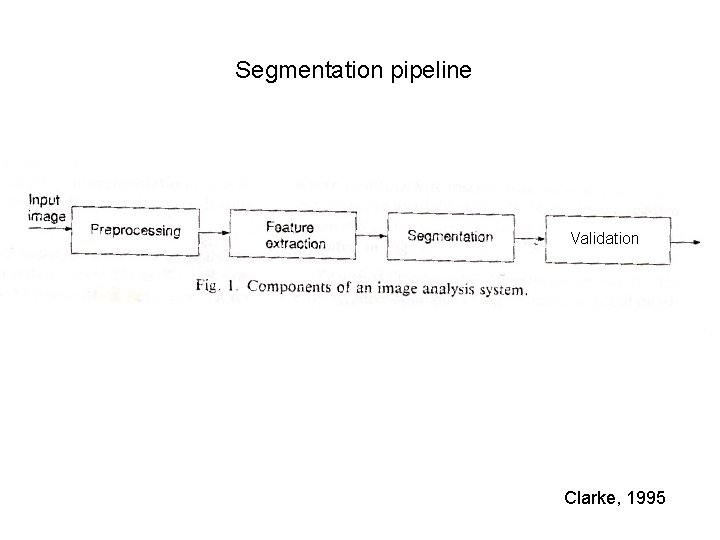
Segmentation pipeline Validation Clarke, 1995

1. Preprocessing 1. 1. Brain extraction 1. 2. Removal of field inhomogeneities (bias-field)
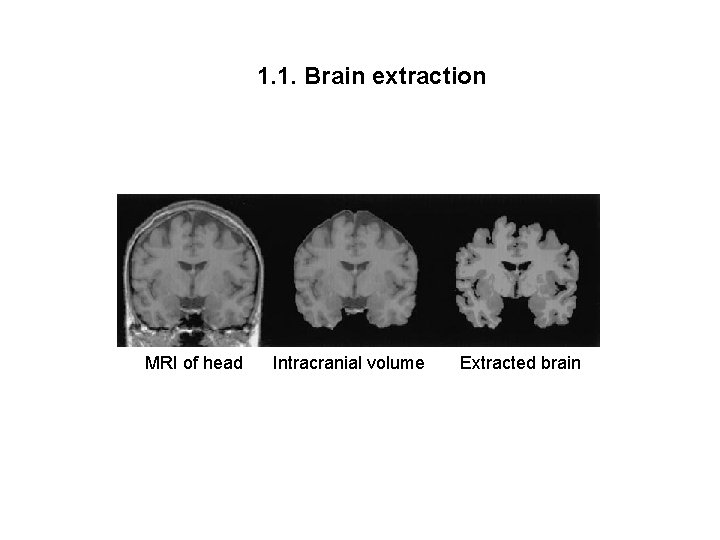
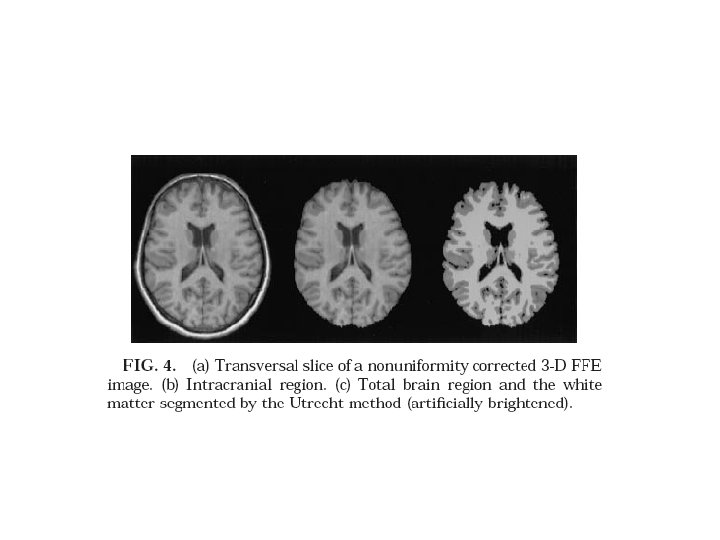
1. 1. Brain extraction MRI of head Intracranial volume Extracted brain
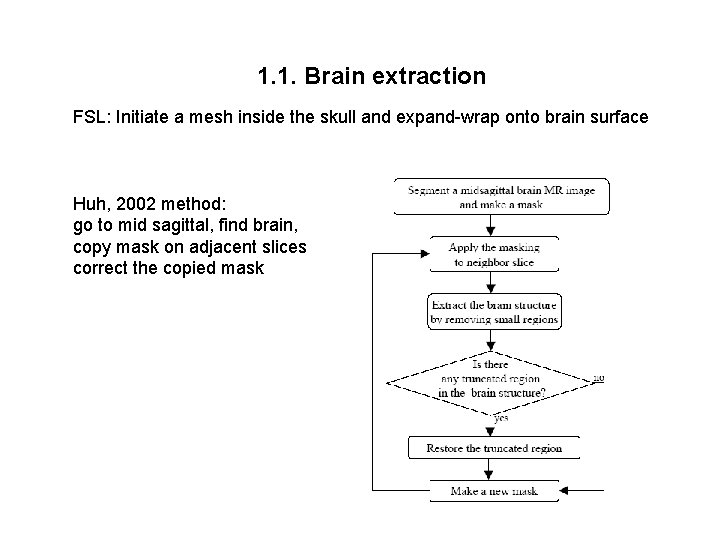
1. 1. Brain extraction FSL: Initiate a mesh inside the skull and expand-wrap onto brain surface Huh, 2002 method: go to mid sagittal, find brain, copy mask on adjacent slices correct the copied mask
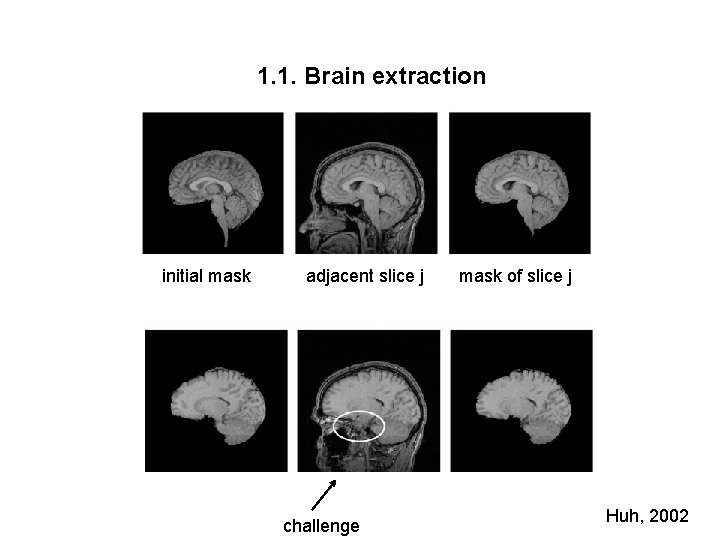
1. 1. Brain extraction initial mask adjacent slice j challenge mask of slice j Huh, 2002
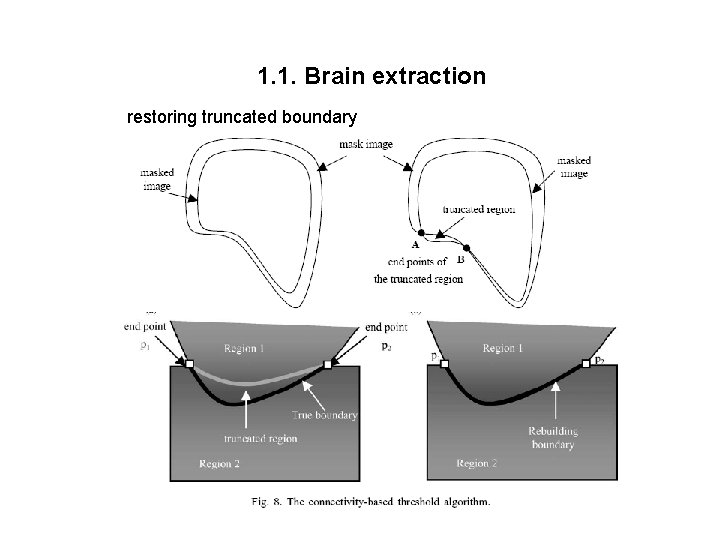
1. 1. Brain extraction restoring truncated boundary
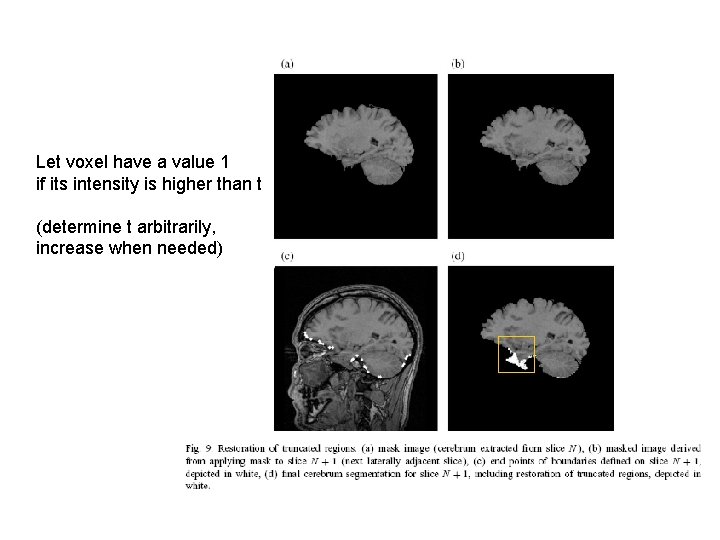
Let voxel have a value 1 if its intensity is higher than t (determine t arbitrarily, increase when needed)
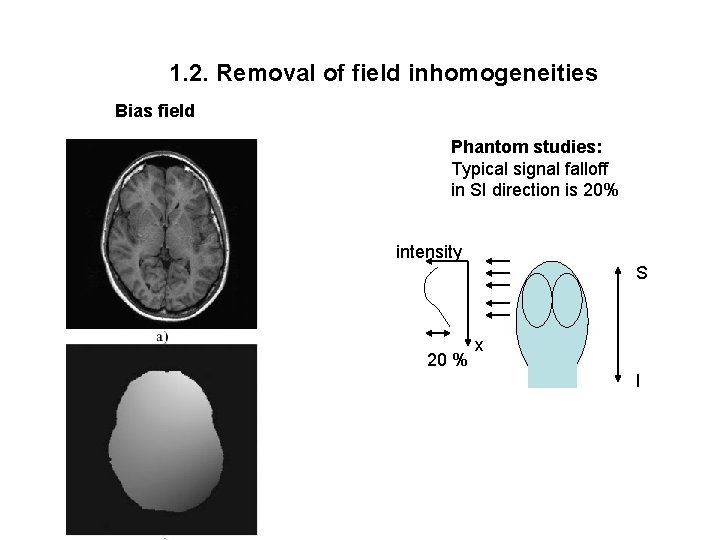
1. 2. Removal of field inhomogeneities Bias field Phantom studies: Typical signal falloff in SI direction is 20% intensity S 20 % x I

1. 2. Removal of field inhomogeneities Statistical methods: probabilistic, gaussian and mixture models of bias-field Polynomial methods: smooth polynomial fit to bias-field

1. 2. Removal of field inhomogeneities Polynomial method example: Milchenko, 2006

Milchenko, 2006
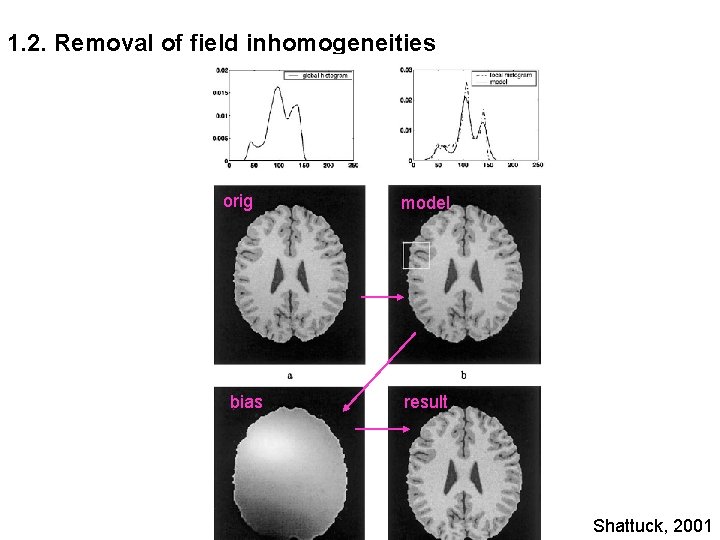
1. 2. Removal of field inhomogeneities orig bias model result Shattuck, 2001

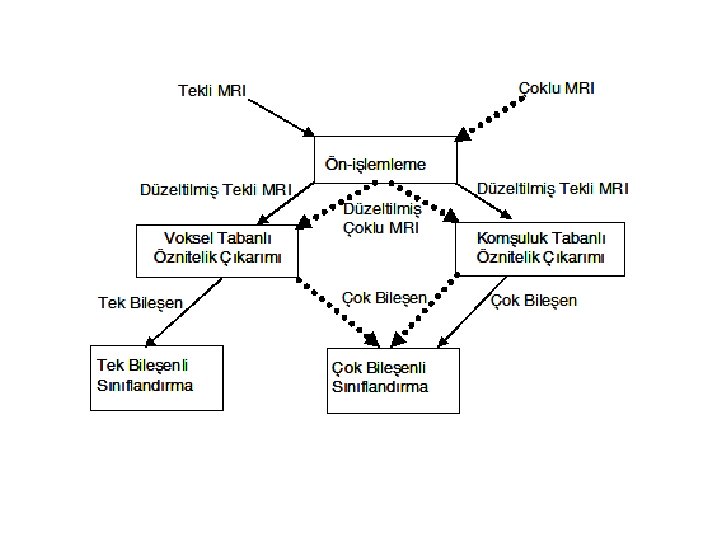
2. Feature extraction Features: - Intensities in a single MRI: univariate classification - Feature vector from a single MRI: multi-variate class. ex: [I(x, y, z) f(N(x, y, z)) g(N(x, y, z))] where N : neighbourhood around (x, y, z) f: distribution of I in neighborhood (entropy g: average I in neighborhood or f, g specify edge or boundary information - Intensities in multiple MRIs with different contrast: multi-variate (multi-spectral)
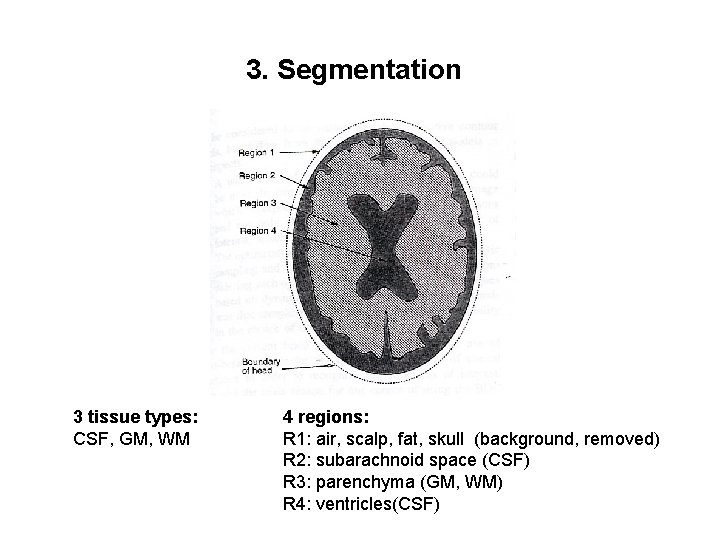
3. Segmentation 3 tissue types: CSF, GM, WM 4 regions: R 1: air, scalp, fat, skull (background, removed) R 2: subarachnoid space (CSF) R 3: parenchyma (GM, WM) R 4: ventricles(CSF)
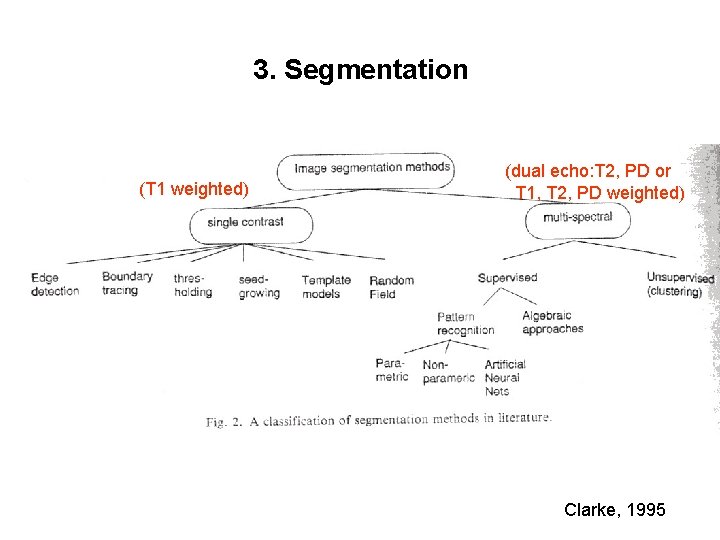
3. Segmentation (T 1 weighted) (dual echo: T 2, PD or T 1, T 2, PD weighted) Clarke, 1995
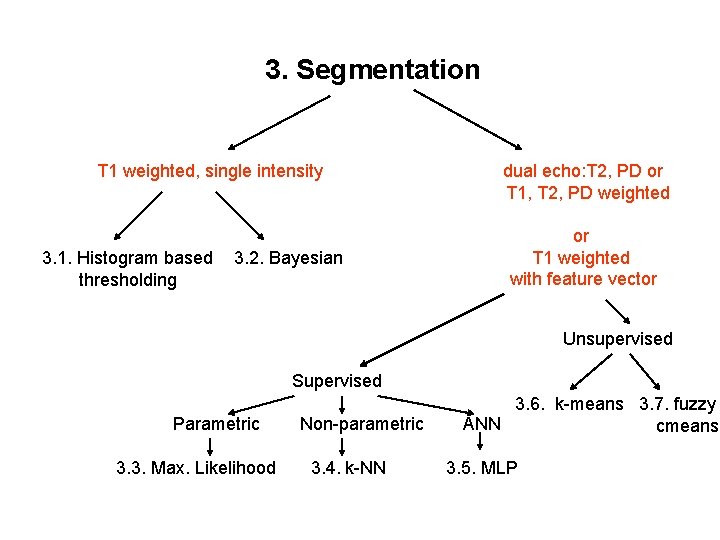
3. Segmentation T 1 weighted, single intensity 3. 1. Histogram based thresholding 3. 2. Bayesian dual echo: T 2, PD or T 1, T 2, PD weighted or T 1 weighted with feature vector Unsupervised Supervised Parametric 3. 3. Max. Likelihood Non-parametric 3. 4. k-NN 3. 6. k-means 3. 7. fuzzy ANN cmeans 3. 5. MLP
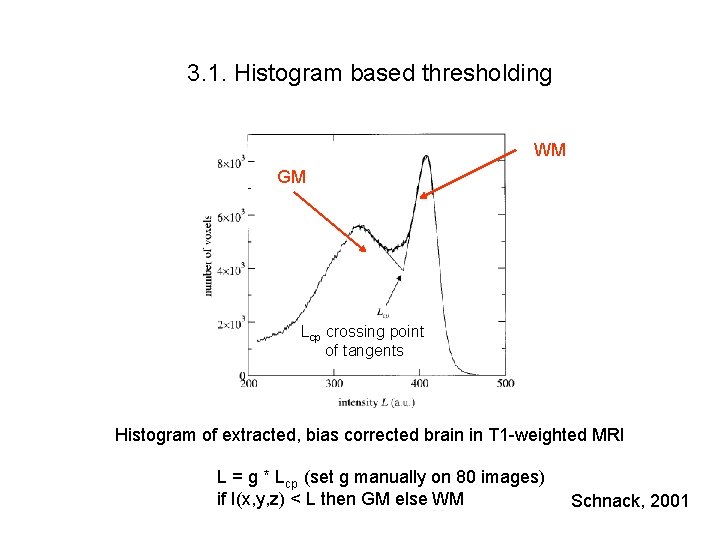
3. 1. Histogram based thresholding WM GM Lcp crossing point of tangents Histogram of extracted, bias corrected brain in T 1 -weighted MRI L = g * Lcp (set g manually on 80 images) if I(x, y, z) < L then GM else WM Schnack, 2001
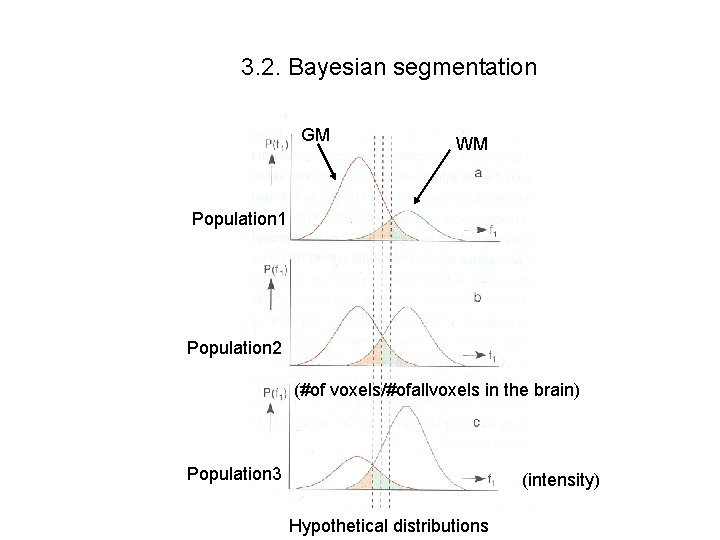
3. 2. Bayesian segmentation GM WM Population 1 Population 2 (#of voxels/#ofallvoxels in the brain) Population 3 (intensity) Hypothetical distributions
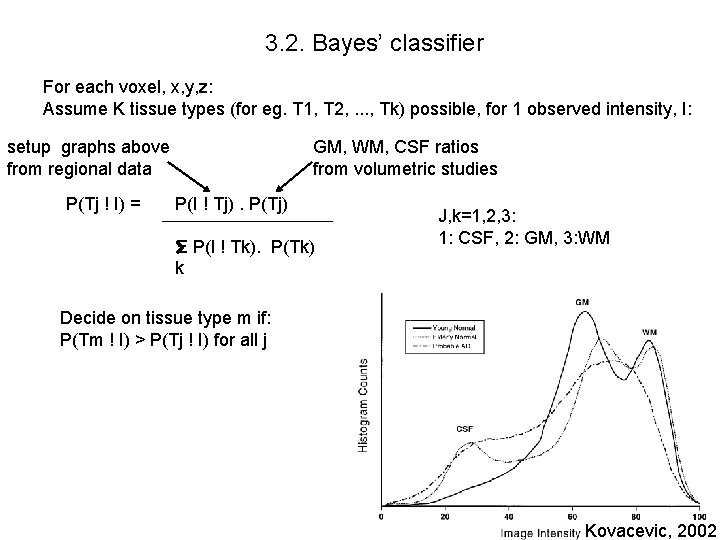
3. 2. Bayes’ classifier For each voxel, x, y, z: Assume K tissue types (for eg. T 1, T 2, . . . , Tk) possible, for 1 observed intensity, I: setup graphs above from regional data P(Tj ! I) = GM, WM, CSF ratios from volumetric studies P(I ! Tj). P(Tj) Ξ P(I ! Tk). P(Tk) k J, k=1, 2, 3: 1: CSF, 2: GM, 3: WM Decide on tissue type m if: P(Tm ! I) > P(Tj ! I) for all j Kovacevic, 2002
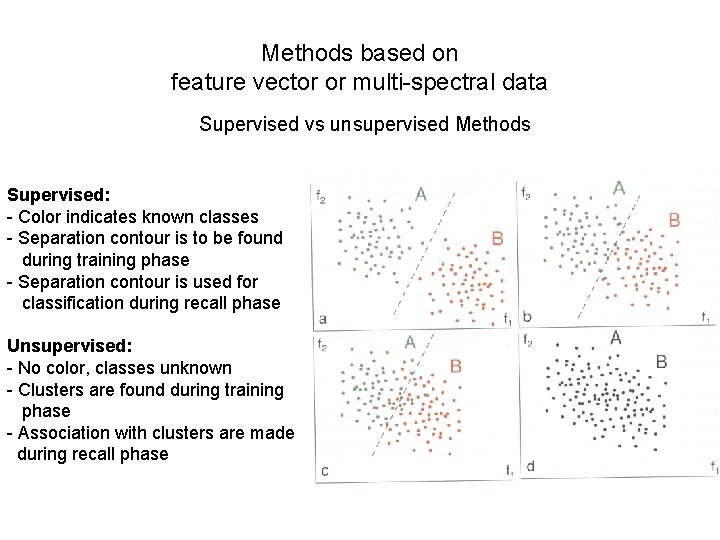
Methods based on feature vector or multi-spectral data Supervised vs unsupervised Methods Supervised: - Color indicates known classes - Separation contour is to be found during training phase - Separation contour is used for classification during recall phase Unsupervised: - No color, classes unknown - Clusters are found during training phase - Association with clusters are made during recall phase
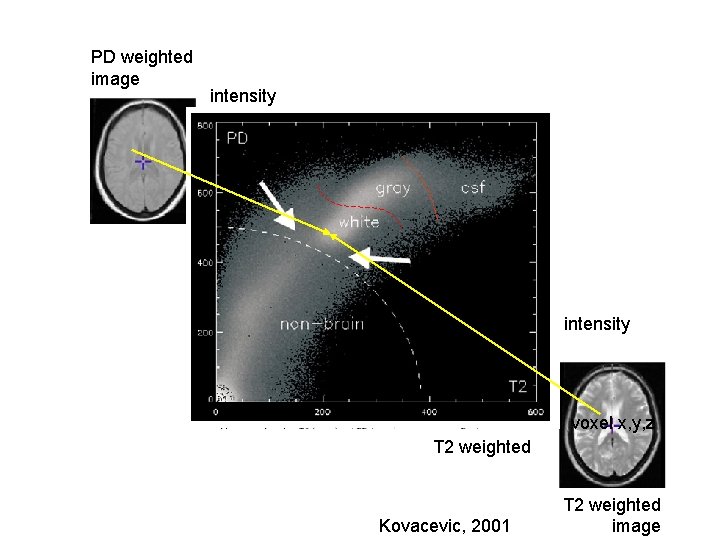
PD weighted image intensity voxel x, y, z T 2 weighted Kovacevic, 2001 T 2 weighted image
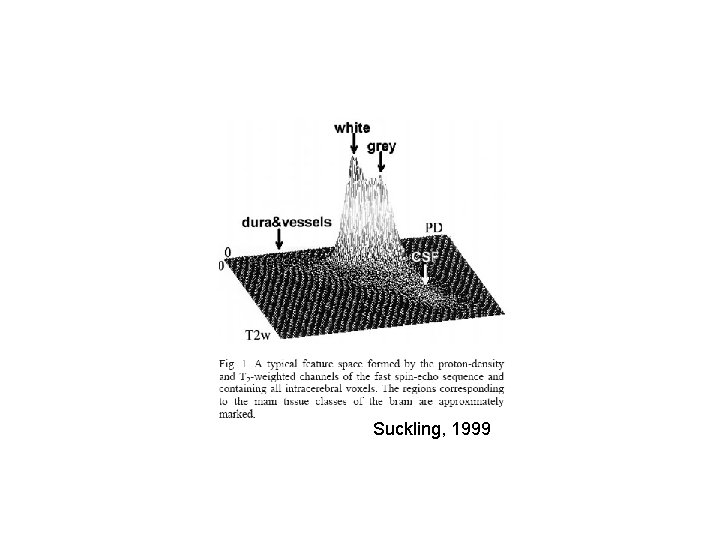
Suckling, 1999

3. 3. Maximum likelihood classifier - Assume the distribution P(I ! Tj) in Bayes can be obtained by a mixture of Gaussian or Normal distribution - Estimate means and co-variance matrix - For better results use Hidden Markov fields within neighborhoods 15 classes Zavaljevski, 2000
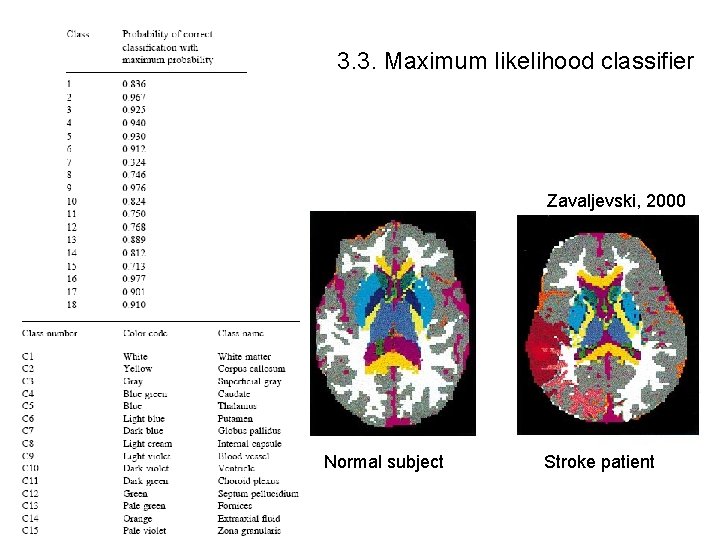
3. 3. Maximum likelihood classifier Zavaljevski, 2000 Normal subject Stroke patient
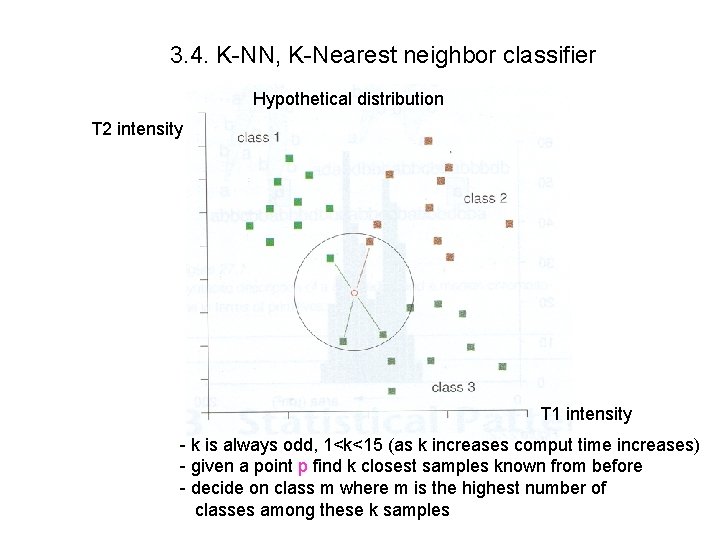
3. 4. K-NN, K-Nearest neighbor classifier Hypothetical distribution T 2 intensity T 1 intensity - k is always odd, 1<k<15 (as k increases comput time increases) - given a point p find k closest samples known from before - decide on class m where m is the highest number of classes among these k samples
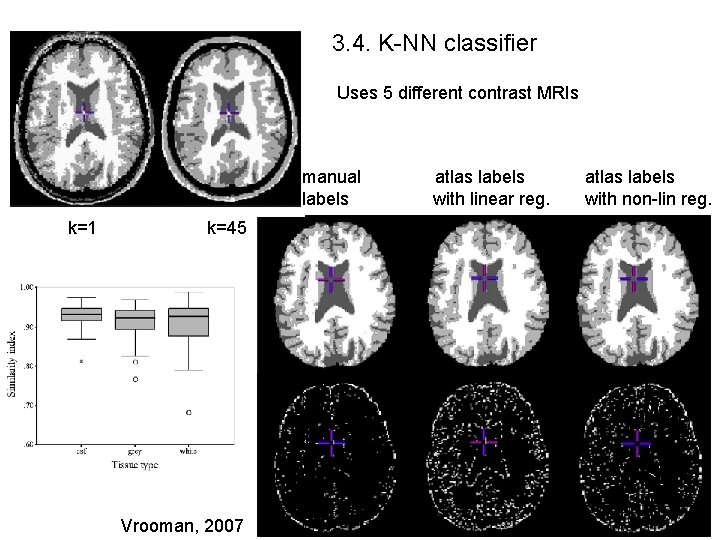
3. 4. K-NN classifier Uses 5 different contrast MRIs manual labels k=1 k=45 Vrooman, 2007 atlas labels with linear reg. atlas labels with non-lin reg.
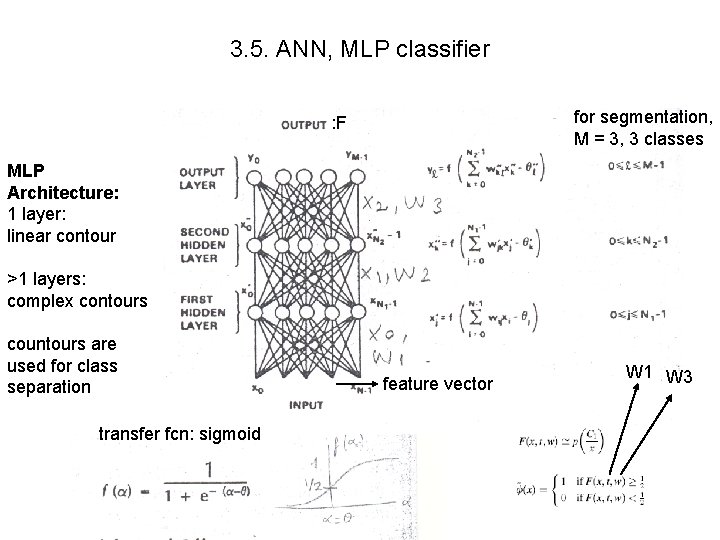
3. 5. ANN, MLP classifier for segmentation, M = 3, 3 classes : F MLP Architecture: 1 layer: linear contour >1 layers: complex contours countours are used for class separation transfer fcn: sigmoid feature vector W 1 W 3

3. 5. ANN, MLP classifier Results This page is empty on purpose

3. 6. k-means classifier This classifier is not used much in segmentation, but explained here as an introduction to fuzzy c-means Algorithm: - k is equal to number of classes - choose k arbitrary initial seed points (*) - assume seed points are class centroids 1 for each sample point j, find distance to all k centroids Let j belong to class m if j is closest to centroid m 2 for each class k, recalculate centroids repeat steps 1 and 2 above until no change in centroids Note how class assignments change at each iteration Minimized measure:
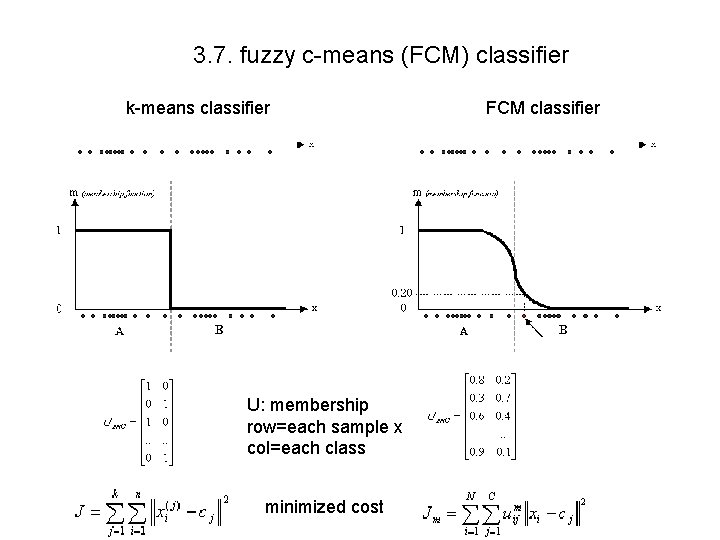
3. 7. fuzzy c-means (FCM) classifier k-means classifier U: membership row=each sample x col=each class minimized cost FCM classifier
![3. 7. fuzzy c-means (FCM) classifier initial Initialize U=[uij] matrix, U(0) At k-step: calculate 3. 7. fuzzy c-means (FCM) classifier initial Initialize U=[uij] matrix, U(0) At k-step: calculate](http://slidetodoc.com/presentation_image_h2/3b2d7dfb7125704a22a959d2dd4c8145/image-36.jpg)
3. 7. fuzzy c-means (FCM) classifier initial Initialize U=[uij] matrix, U(0) At k-step: calculate the centers vectors C (k)=[cj] with U(k) Update U(k) , U(k+1) If || U(k+1) - U(k)||< iteration 8 then STOP; otherwise return to step 2. iteration 37
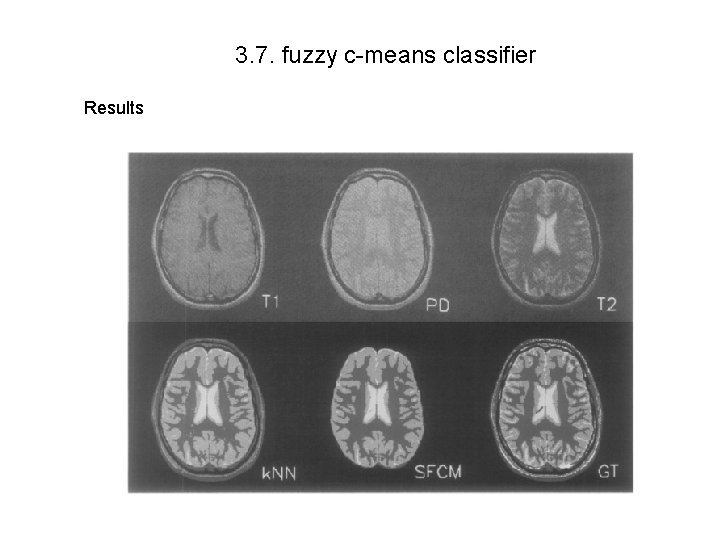
3. 7. fuzzy c-means classifier Results

4. Validation Important issues: - Partial volume effect, visualization - Validation in manually segmented image - Performance comparison with other methods on simulated image: Ex: Brainweb from Mcgill
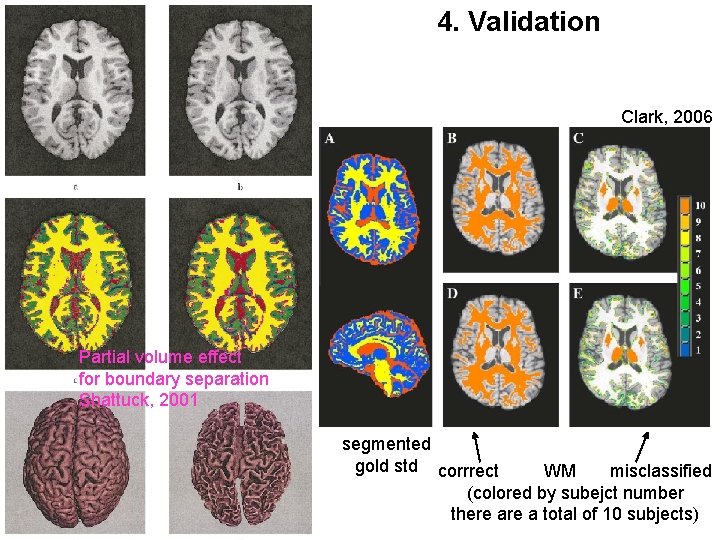
4. Validation Clark, 2006 Partial volume effect for boundary separation Shattuck, 2001 segmented gold std corrrect WM misclassified (colored by subejct number there a total of 10 subjects)
- Slides: 39