Mr BUMP Molecular Replacement with Bulk Model Preparation

Mr. BUMP – Molecular Replacement with Bulk Model Preparation Ronan Keegan, Martyn Winn CCP 4 group, Daresbury Laboratory Como May 23 rd 2006 1

The aim of Mr Bump • An automation framework for Molecular Replacement. • Particular emphasis on generating a variety of search models. • Can be used to generate models only. • Wraps Phaser and/or Molrep. • Also uses a variety of helper applications (e. g. Chainsaw) and bioinformatics tools (e. g. Fasta, Mafft) • Uses on-line databases (e. g. PDB, Scop) • In favourable cases, gives “one-button” solution • In unfavourable cases, will suggest likely search models for manual investigation (lead generation) 2
Target MTZ & Sequence Target ` Details • Currently: – Number of residues and molecular weight – Matthews Coefficient. – Estimated number of molecules in the a. s. u. 3
Target MTZ & Sequence Target ` Details Model ` Search Generate a list of structures that are possible templates for search models 4

Search for homologous proteins • FASTA search of PDB – Sequence based search using sequence of target structure. – Can be run locally if user has fasta 34 program installed or remotely using the OCA web-based service hosted by the EBI. – Local search is done against the complete list of PDB sequences derived from ATOM records in the PDB structure files. – All of the resulting PDB id codes are added to a list – Not interested in the alignment to target at this stage. 5

Search for similar structures • Secondary Structure based search (optional) – Top hit from the FASTA search is used as the template structure for a secondary structure based search. – Uses the SSM webservice provided by the EBI. – Any new structures found that aren’t included in the list of matches from the FASTA search are added to the list. – Provides structural variation, not based on direct sequence similarity to target • Manual addition – Can additional PDB id codes to the list, e. g. from FFAS or psi. BLAST searches 6

Multiple Alignment • After the set of PDB ids are collected in the FASTA and SSM searches, their coordinate-based sequences are collected and put through a multiple alignment with the target sequence • Aims: – Score template structures in a consistent manner, in order to prioritise them for subsequent steps – Extract pairwise alignment between template and target for use in Chainsaw step. Multiple alignment should give a better set of alignments than the original pair-wise FASTA alignments 7
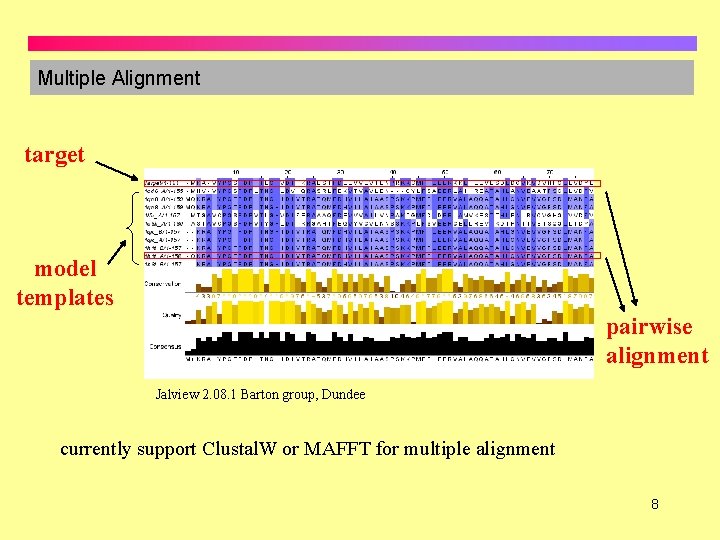
Multiple Alignment target model templates pairwise alignment Jalview 2. 08. 1 Barton group, Dundee currently support Clustal. W or MAFFT for multiple alignment 8

Template Model Scoring • Alignment Scoring: score = sequence identity X alignment quality • Sequence identity: • Alignment quality: – Ungapped sequence identity i. e. sequence identity of aligned target residues – Dependent on the alignment length, the number of gaps created in the template alignment and the extent of each of these gaps. – The penalties given for gaps and the size of the gaps is biased so that alignments that preserve domains of the structure rather than spreading the aligned residues out score higher. The top scoring models are then used for further processing 9
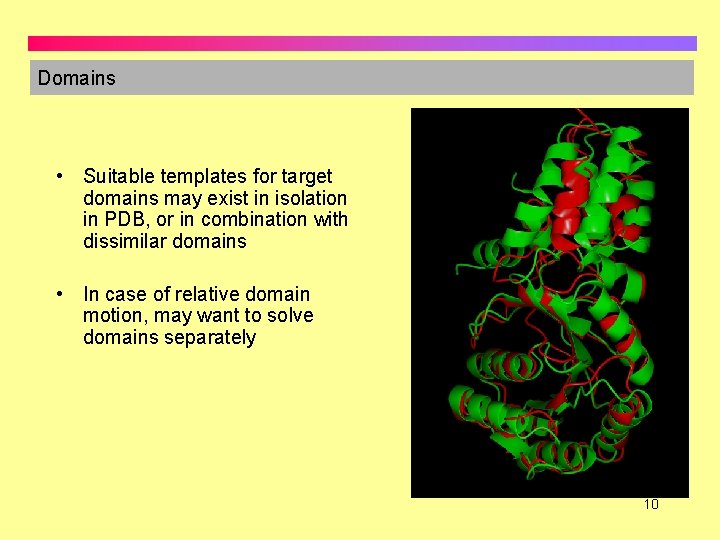
Domains • Suitable templates for target domains may exist in isolation in PDB, or in combination with dissimilar domains • In case of relative domain motion, may want to solve domains separately 10

Domains • Domains search: – Top scoring templates from multiple alignment are tested to see if they contain any domains. – Uses the SCOP database. This only lists domains that appear more than once in the PDB. – The database is scanned to to see if domains exist for each of the PDBs in the list of templates – Domains are then extracted from the parent PDB structure file and added to the list of template models as additional search models for MR. 11

Multimers • Multimer search: – Search for quaternary structures that may be used as search models. – Better signal-to-noise ratio than monomer, if assembly is correct for the target. – Multimeric structures based on top templates are retrieved using the PQS service at the EBI, and added to the list of search models – PQS will soon be replaced by the use of the PISA service at the EBI (Eugene Krissinel) 1 n 5 a 1 n 5 b 1 n 5 c 1 n 5 d SPLIT-ASU into 4 Oligomeric files of type TRIMERIC SPLIT-ASU into 2 Oligomeric files of type DIMERIC SYMMETRY-COMPLEX Oligomeric file of type DIMERIC 12
Target MTZ & Sequence Target ` Details Model ` Search Model ` Preparation 13

Search Model Preparation Search models prepared in four ways: 1. PDBclip – original PDB with waters removed, hydrogens removed, most probable conformations for side chains selected and chain ID’s added if missing. 2. Molrep – Molrep contains a model preparation function which will align the template sequence with the target sequence and prune the nonconserved side chains accordingly. 3. Chainsaw – Can be given any alignment between the target and template sequences. – Non-conserved residues are pruned back to the gamma atom. 4. Polyalanine – Created by excluding all of the side chain atoms beyond the CB atom using the Pdbset program 14

Search Model Preparation Ensemble for Phaser: • Top scoring search models are “superposed” to create a ensemble model. • This may provide a better search model than any of the individual models on their own. • Currently the default is to use the top 5 scoring search models but plan to create dynamically based on MW and RMSDs of constituent search models 15
Target MTZ & Sequence Target ` Details Model ` Search Model ` Preparation Molecular Replacement & Refinement ` 16

Molecular Replacement and Refinement • The search models can be processed with Molrep or Phaser or both. • The resulting models from molecular replacement are passed to Refmac for restrained refinement. • The change in the Rfree value during refinement is used to determine how good the resulting model is. • If the final value for Rfree is less than 0. 35 or it is less than 0. 5 and has fallen by more than 20 % from the initial Rfree, a solution is deemed to have been found. • Models that produce an Rfree below 0. 5 and the value looks to be falling will be highlighted as “marginal solutions” that are worthy of further investigation if no solution is found using the other search models. 17
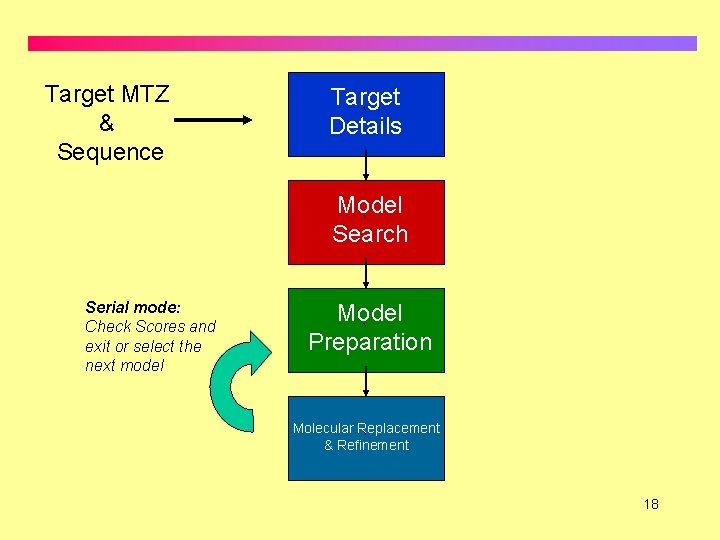
Target MTZ & Sequence Target ` Details Model ` Search Serial mode: Check Scores and exit or select the next model Model ` Preparation Molecular Replacement & Refinement ` 18
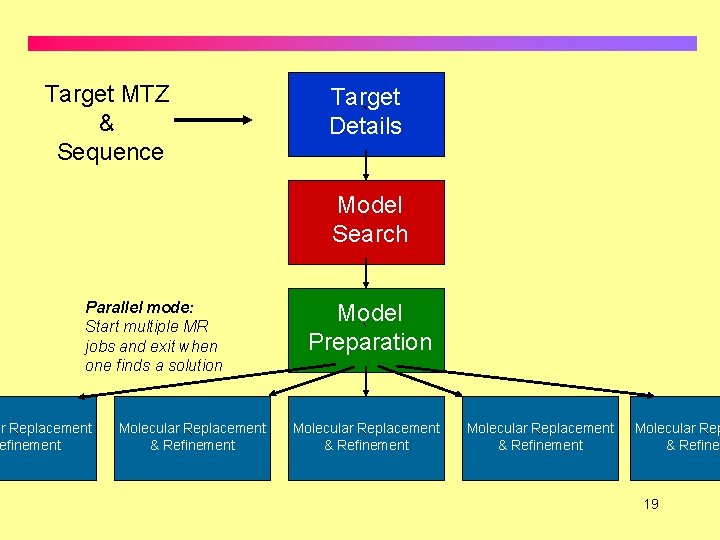
Target MTZ & Sequence Target ` Details Model ` Search Parallel mode: Start multiple MR jobs and exit when one finds a solution ar Replacement efinement ` Molecular Replacement & Refinement ` Model ` Preparation Molecular Replacement & Refinement ` Molecular Rep & Refinem ` 19

Mr. BUMP on compute clusters • Mr. BUMP can take advantage of a compute cluster to farm out the Molecular Replacement jobs. • Currently Sun Grid Engine enabled clusters are supported but support will be added for LSF and condor and any other types of queuing system if there is enough demand. 20
Pre-release version of Mr. BUMP • Pre-release made available in Jan 06 • Simple installation • Currently runs on Linux and OSX. • Windows version almost ready. • Comes with CCP 4 GUI. • Can also be run from the command line with keyword input • Good deal of interest and some successes • Regular updates (currently version 0. 3) http: //www. ccp 4. ac. uk/Mr. BUMP 21
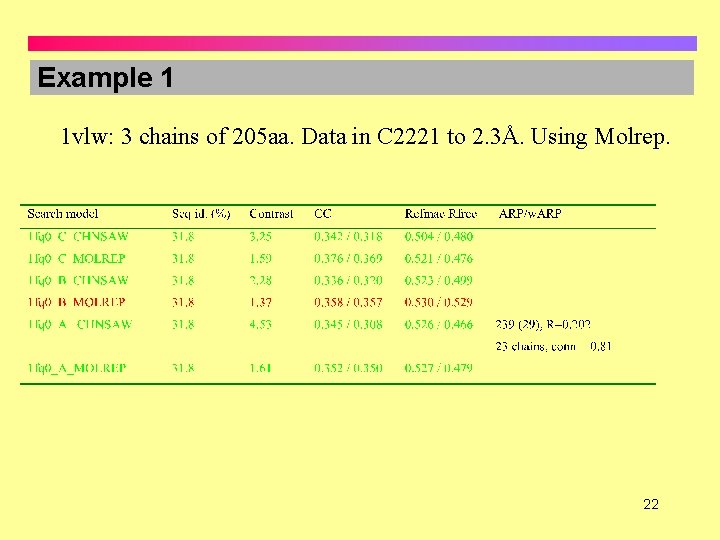
Example 1 1 vlw: 3 chains of 205 aa. Data in C 2221 to 2. 3Å. Using Molrep. 22

Example 2 Anon: Mr. BUMP marginal solution used yes = arp/warp builds and docks entire molecule no = arp/warp fails = wrong MR solution 23

A few observations. . . • In difficult cases, success in Mr. BUMP may depend on particular template, chain and model preparation method • Nevertheless, may get several putative solutions • Ease of subsequent model re-building, model completion may depend on choice of solution • First solution or check everything? • Expectation that quick solution required - in fact, most users seem happy to let Mr. BUMP run for long time (hours, days) • Worth checking “failed” solutions! 24

Future developments • Windows support (almost done) • Complexes (in progress) – Processing of multiple target sequences • Improved alignment: • • Multiple alignment against larger sequence database Alignment from profile-based search User-supplied alignment Incorporate PISA multimer determining service (in progress) • Model generation: • • Identification of flexible loops Normal mode generated conformations • Develop web-service version to allow CCP 4 i users to run jobs on CCP 4 cluster 25

Acknowledgements • Ronan Keegan, CCP 4 @ Daresbury • Thanks to authors of all underlying programs and services • Other suggestions from: • Dave Meredith, Graeme Winter, Daresbury Laboratory. • Eugene Krissinel, EBI, Cambridge. • Eleanor Dobson, YSBL, York University • Geoff Barton, Charlie Bond, University of Dundee • Randy Read, Airlie Mc. Coy, Cambridge • Funding: • BBSRC (e-HTPX, CCP 4) http: //www. ccp 4. ac. uk/Mr. BUMP 26
- Slides: 26