MORPHOLOGICAL PHYSIOLOGICAL BIOCHEMICAL STUDIES AND QUANTIFICATION GENE EXPRESSION


MORPHOLOGICAL, PHYSIOLOGICAL, BIOCHEMICAL STUDIES AND QUANTIFICATION GENE EXPRESSION UNDER DROUGHT STRESS IN UPLAND COTTON

Abdelmoghny, A. M. 1; Suman B. Singh 2, H. B. Santosh 2, K. P. Raghavendra 2, J. A. Sheeba 2 and K. R. Kranthi 2 Cotton Research Institute (CRI), Agricultural Research Center (ARC), 9 El-Gamma St, Giza, Egypt 1 ICAR, Central Institute for Cotton Research (CICR), Post Box No. 2, Shankar Nagar Post Office, Nagpur, Maharashtra 440 010, India 2

Pedigree of the studied six cotton genotypes Genotypes No. Name 891/ IC 1 357406 2 28 I (CNH 28 I) 3 Nagpur-9 4 Suraj 5 2389/IC 359637 6 LRA 5166 Pedigree JAN 579 -15 -5 -10 LRA 5166(CCH 526612 x HLS 329) 117 -F 2 -CO 2 -P Laxmix (Reba B 50 x AC 122)


The genotypes were sown in plastic pots measuring 38 cm x 40 cm, filled with loamy clay soil. All plants were regularly irrigated with 1 L tap water daily, Induced Drought Stress after 60 days from sown for 13 days

Each set (drought and control) having six genotypes arranged in randomized complete block design (RCBD) with triplicate, each replicate consists of eighteen pots and each pot has one plant for each genotype.


28 I 891 2389

LRA 1566 Nagpur 9 Suraj
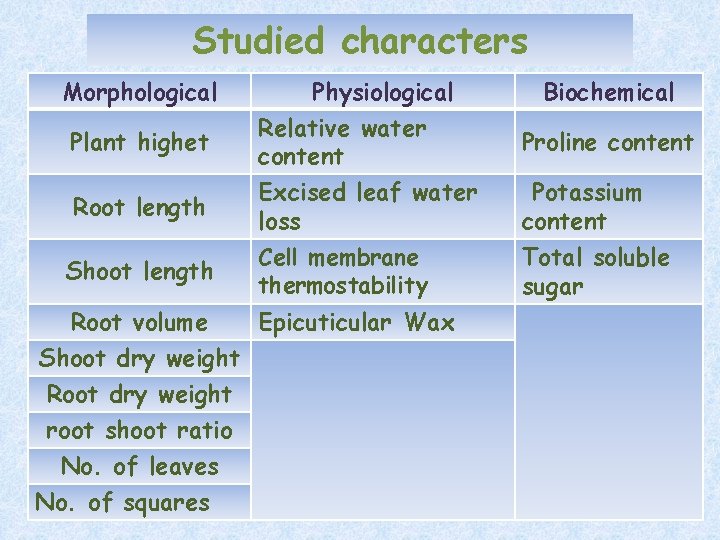
Studied characters Morphological Plant highet Root length Physiological Relative water content Excised leaf water loss Shoot length Cell membrane thermostability Root volume Epicuticular Wax Shoot dry weight Root dry weight root shoot ratio No. of leaves No. of squares Biochemical Proline content Potassium content Total soluble sugar


Drought- related traits S. No Trait Genotypes ( exhibited higher value) Morphological Parameters 1. Root length 891 (IC 357406) , Nagpur 9 and Suraj 2. Root Volume Suraj, 28 I and Nagpur 9 3. Root Shoot Ratio 891 (IC 357406), Nagpur 9 and Suraj Physiological / Biochemical Parameters 1. Epicuticular Wax content Suraj, 28 I , LRA 5166 2. Relative Water Content Nagpur 9, 891 (IC 357406), Suraj 3. Excised Leaf Water loss 891 (IC 357406), Nagpur 9, Suraj 4. Proline content 28 I, Suraj 5. Total Soluble Sugars Non Significant 6. Cell Membrane Stability Nagpur 9, 28 I , 2389 (IC 259637) 7. Potassium Content LRA 5166, 2389 (IC 259637), 28 I

Molecular Level q. RT-PCR

Gene expression analysis through q. PCR No. Genotypes Name 1 891/ IC 357406 2 28 I (CNH 28 I) 3 Nagpur-9 4 Suraj 5 2389/IC 359637 6 LRA 5166 Total RNA isolation: • Control and Drought stress plants • 3 biological replicates from each genotypes • Spectrum Plant RNA isolation kit Quality and quantity check: Gel electrophoresis and qubit flourometer c. DNA synthesis (Reverse transcription): Affyscript c. DNA synthesis kit q. PCR- Brilliant QPCR SYBR green Data Analysis: Mx. Pro software
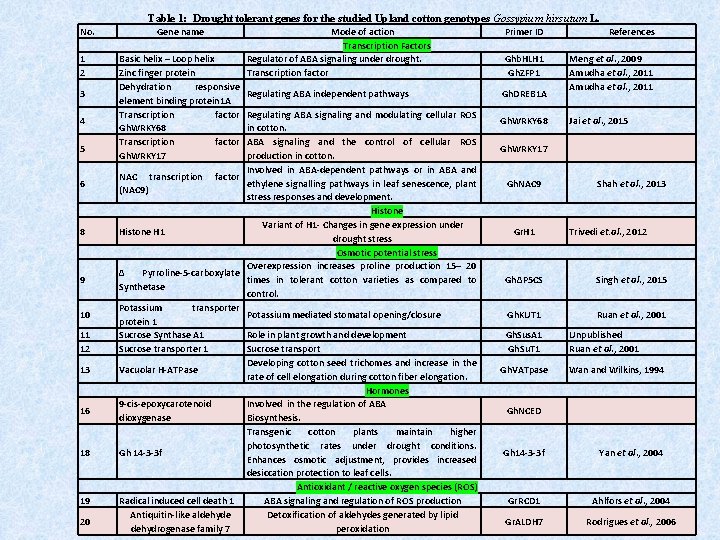
Table 1: Drought tolerant genes for the studied Upland cotton genotypes Gossypium hirsutum L. No. 1 2 3 4 5 6 8 9 10 11 12 13 16 18 19 20 Gene name Basic helix – Loop helix Zinc finger protein Dehydration responsive element binding protein 1 A Transcription factor Gh. WRKY 68 Transcription factor Gh. WRKY 17 Mode of action Transcription Factors Regulator of ABA signaling under drought. Transcription factor Regulating ABA independent pathways Regulating ABA signaling and modulating cellular ROS in cotton. ABA signaling and the control of cellular ROS production in cotton. Involved in ABA-dependent pathways or in ABA and NAC transcription factor ethylene signalling pathways in leaf senescence, plant (NAC 9) stress responses and development. Histone Variant of H 1 - Changes in gene expression under Histone H 1 drought stress Osmotic potential stress Overexpression increases proline production 15– 20 Δ Pyrroline-5 -carboxylate times in tolerant cotton varieties as compared to Synthetase control. Potassium transporter Potassium mediated stomatal opening/closure protein 1 Sucrose Synthase A 1 Role in plant growth and development Sucrose transporter 1 Sucrose transport Developing cotton seed trichomes and increase in the Vacuolar H-ATPase rate of cell elongation during cotton fiber elongation. Hormones 9 -cis-epoxycarotenoid Involved in the regulation of ABA dioxygenase Biosynthesis. Transgenic cotton plants maintain higher photosynthetic rates under drought conditions. Gh 14 -3 -3 f Enhances osmotic adjustment, provides increased desiccation protection to leaf cells. Antioxidant / reactive oxygen species (ROS) Radical induced cell death 1 ABA signaling and regulation of ROS production Antiquitin-like aldehyde Detoxification of aldehydes generated by lipid dehydrogenase family 7 peroxidation Primer ID References Ghb. HLH 1 Gh. ZFP 1 Meng et al. , 2009 Amudha et al. , 2011 Gh. DREB 1 A Gh. WRKY 68 Jai et al. , 2015 Gh. WRKY 17 Gh. NAC 9 Gr. H 1 Shah et al. , 2013 Trivedi et. al. , 2012 GhΔP 5 CS Singh et al. , 2015 Gh. KUT 1 Ruan et al. , 2001 Gh. Sus. A 1 Gh. Su. T 1 Gh. VATpase Unpublished Ruan et al. , 2001 Wan and Wilkins, 1994 Gh. NCED Gh 14 -3 -3 f Yan et al. , 2004 Gr. RCD 1 Ahlfors et al. , 2004 Gr. ALDH 7 Rodrigues et al. , 2006

Standardization of PCR condition: L 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 Fifteen primers working at annealing temperature 60 C with out primer dimer was chosen for subsequent analysis L

QPCR analysis of gene expression Methodology: Comparative qunatification Chemistry: SYBR green Reference/normaliser gene: Actin Calibrator: c. DNA from control samples Unknown: c. DNA from drought stress sample Genes under study

Analysis Selection/Setup Control (Calibrtor)Drought stress(Unknown Gene under study: b. HLH and DREB 1 A Normaliser/reference gene: Actin 4 Reference dye: ROX dye
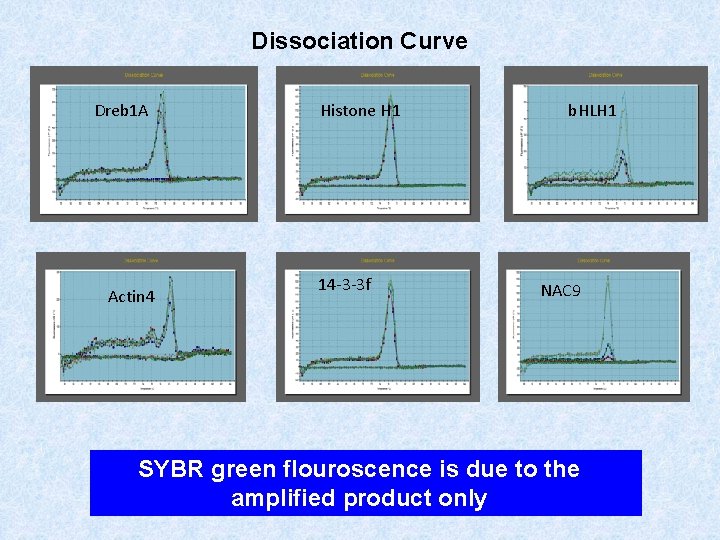
Dissociation Curve Dreb 1 A Actin 4 Histone H 1 14 -3 -3 f b. HLH 1 NAC 9 SYBR green flouroscence is due to the amplified product only

q. PCR analysis ΔCT = (CT gene of interest - CT internal control) Δ Δ CT = (Δ CT, q - 2 Δ CT, b) Log Fold change = -2 Δ Δ Ct
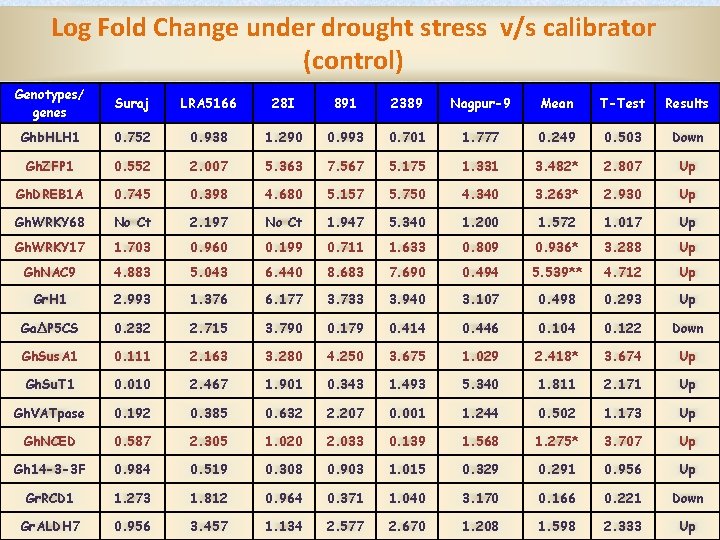
Log Fold Change under drought stress v/s calibrator (control) Genotypes/ genes Suraj LRA 5166 28 I 891 2389 Nagpur-9 Mean T-Test Results Ghb. HLH 1 0. 752 0. 938 1. 290 0. 993 0. 701 1. 777 0. 249 0. 503 Down Gh. ZFP 1 0. 552 2. 007 5. 363 7. 567 5. 175 1. 331 3. 482* 2. 807 Up Gh. DREB 1 A 0. 745 0. 398 4. 680 5. 157 5. 750 4. 340 3. 263* 2. 930 Up Gh. WRKY 68 No Ct 2. 197 No Ct 1. 947 5. 340 1. 200 1. 572 1. 017 Up Gh. WRKY 17 1. 703 0. 960 0. 199 0. 711 1. 633 0. 809 0. 936* 3. 288 Up Gh. NAC 9 4. 883 5. 043 6. 440 8. 683 7. 690 0. 494 5. 539** 4. 712 Up Gr. H 1 2. 993 1. 376 6. 177 3. 733 3. 940 3. 107 0. 498 0. 293 Up GaΔP 5 CS 0. 232 2. 715 3. 790 0. 179 0. 414 0. 446 0. 104 0. 122 Down Gh. Sus. A 1 0. 111 2. 163 3. 280 4. 250 3. 675 1. 029 2. 418* 3. 674 Up Gh. Su. T 1 0. 010 2. 467 1. 901 0. 343 1. 493 5. 340 1. 811 2. 171 Up Gh. VATpase 0. 192 0. 385 0. 632 2. 207 0. 001 1. 244 0. 502 1. 173 Up Gh. NCED 0. 587 2. 305 1. 020 2. 033 0. 139 1. 568 1. 275* 3. 707 Up Gh 14 -3 -3 F 0. 984 0. 519 0. 308 0. 903 1. 015 0. 329 0. 291 0. 956 Up Gr. RCD 1 1. 273 1. 812 0. 964 0. 371 1. 040 3. 170 0. 166 0. 221 Down Gr. ALDH 7 0. 956 3. 457 1. 134 2. 577 2. 670 1. 208 1. 598 2. 333 Up
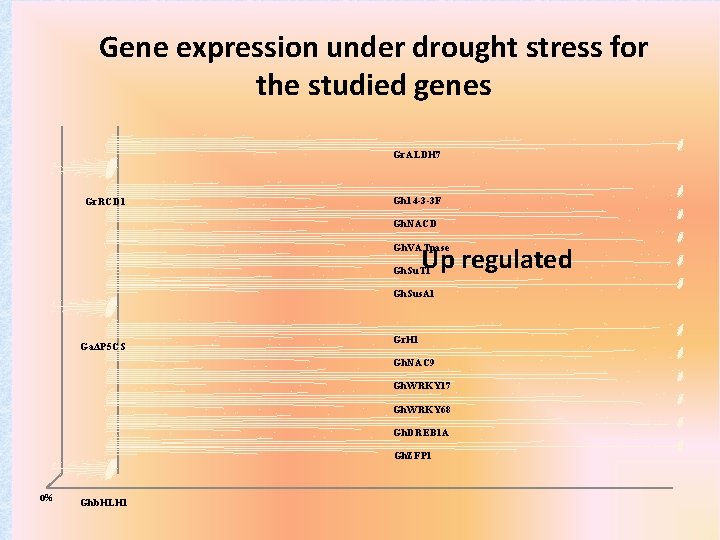
Gene expression under drought stress for the studied genes Gr. ALDH 7 Gr. RCD 1 Gh 14 -3 -3 F Gh. NACD Up regulated Gh. VATpase Gh. Su. T 1 Gh. Sus. A 1 GaΔP 5 CS Gr. H 1 Gh. NAC 9 Gh. WRKY 17 Gh. WRKY 68 Gh. DREB 1 A Gh. ZFP 1 0% Ghb. HLH 1
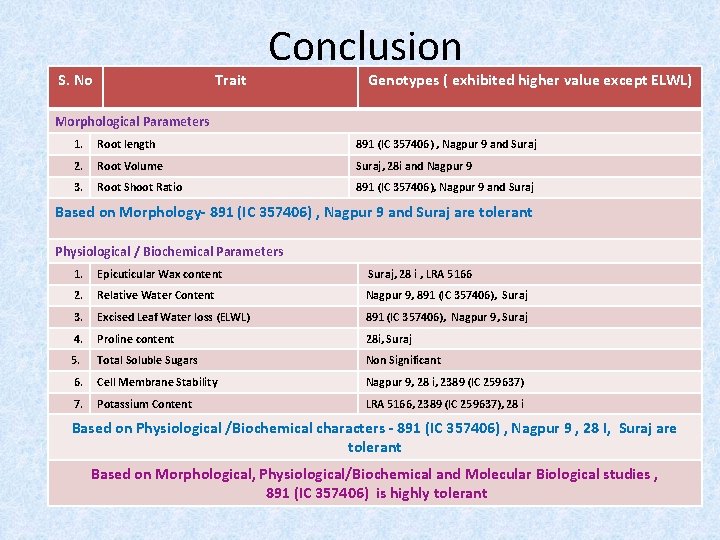
S. No Trait Conclusion Genotypes ( exhibited higher value except ELWL) Morphological Parameters 1. Root length 891 (IC 357406) , Nagpur 9 and Suraj 2. Root Volume Suraj, 28 i and Nagpur 9 3. Root Shoot Ratio 891 (IC 357406), Nagpur 9 and Suraj Based on Morphology- 891 (IC 357406) , Nagpur 9 and Suraj are tolerant Physiological / Biochemical Parameters 1. Epicuticular Wax content Suraj, 28 i , LRA 5166 2. Relative Water Content Nagpur 9, 891 (IC 357406), Suraj 3. Excised Leaf Water loss (ELWL) 891 (IC 357406), Nagpur 9, Suraj 4. Proline content 28 i, Suraj 5. Total Soluble Sugars Non Significant 6. Cell Membrane Stability Nagpur 9, 28 i, 2389 (IC 259637) 7. Potassium Content LRA 5166, 2389 (IC 259637), 28 i Based on Physiological /Biochemical characters - 891 (IC 357406) , Nagpur 9 , 28 I, Suraj are tolerant Based on Morphological, Physiological/Biochemical and Molecular Biological studies , 891 (IC 357406) is highly tolerant

ACKNOWLEDGEMENT RESEARCH TRAINING FELLOWSHIP FOR DEVELOPING COUNTRY SCIENTITSS (RTF-DCS) 2014 -2015 Centre for Science & Technology of the Non-Aligned and Other Developing Countries (NAM S&T Centre)

- Slides: 26